Abstract
MicroRNAs (miRNAs) are small, noncoding RNAs that influence diverse biological outcomes through the repression of target genes during normal development and pathological responses. In particular, the alteration of miRNA expression has dramatic consequences for the progression of tumorigenesis. miRNAs undergo two processing steps that transform a long primary transcript into the mature miRNA. Although the general miRNA biogenesis pathway is well established, it is clear that not all miRNAs are created equally. Recent studies show that miRNA expression is controlled by diverse mechanisms in response to cellular stimuli. In this review, we discuss the mechanisms that govern the regulation of miRNA biogenesis with particular focus on how these mechanisms are perturbed in cancer.
Introduction
MicroRNAs (miRNAs) are a newly described class of short (21-24 nucleotide [nt]), noncoding RNAs that influence diverse biological functions through the repression of target genes during normal development and pathological responses. miRNAs are generated through two processing steps that convert a long, RNA polymerase II (RNA pol II)–derived primary transcript into a mature 21- to 24-nt miRNA (Fig. 1). 1 The mature miRNA then promotes the association of a large protein complex, termed the RNA-induced silencing complex (RISC), with specific regions in the 3′ untranslated region (3′UTR) of target genes (Fig. 1). 2 Selection of miRNA targets is mediated by imperfect base pairing between the miRNA and miRNA binding site present in the 3′UTR of the target mRNA. The imperfect nature of the miRNA:mRNA interaction means that a single miRNA can target tens to hundreds of mRNAs. 3-5 Association of the miRNA-RISC with a target mRNA results in the repression of the target gene by promoting mRNA degradation and/or translational inhibition. 6-10 Through the repression of targets, miRNAs elicit critical changes in gene expression programs, which have been reported to underlie diverse aspects of biology, including developmental timing, differentiation, proliferation, cell death, and metabolism. 11-13
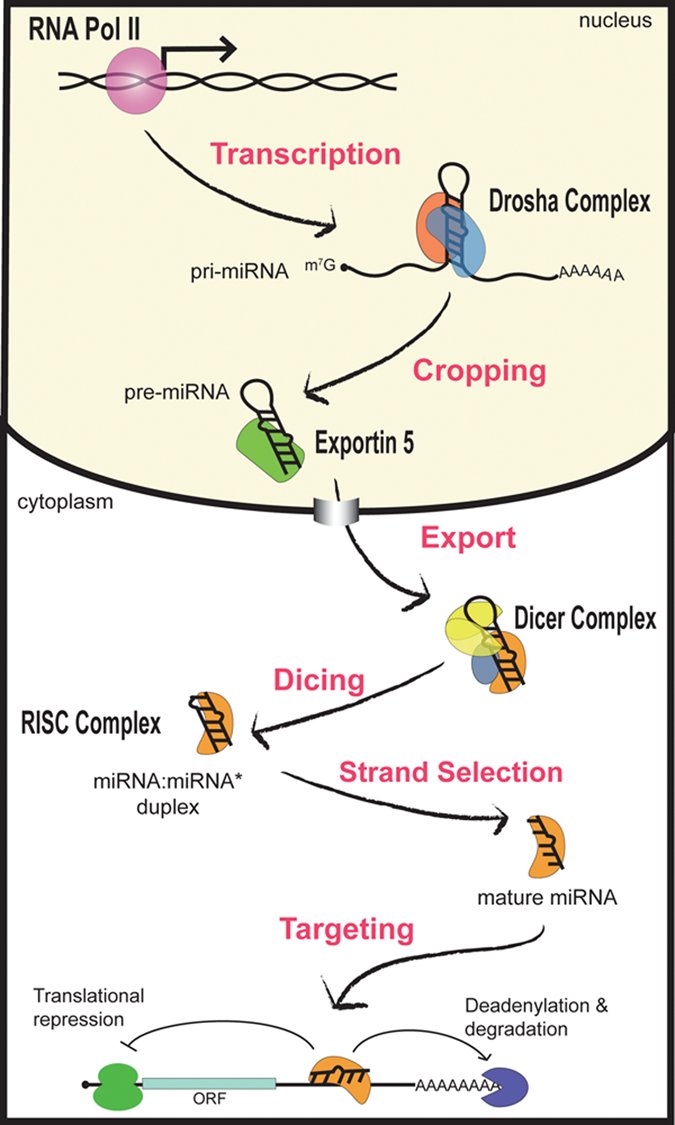
General mechanism of miRNA biogenesis. miRNAs are initially transcribed as a long, capped, and polyadenylated pri-miRNA. The Drosha complex crops the pri-miRNA into a hairpin-shaped pre-miRNA. Next, Exportin-5 promotes the nuclear translocation of the pre-miRNA, which is further processed by the Dicer complex. Following Dicing, the resulting miRNA:miRNA* is dissociated, and the mature miRNA is incorporated into the RISC, where it functions to mediate gene silencing either by translational inhibition or by promoting the degradation of target mRNAs.
miRNAs Act as Both Oncogenes and Tumor Suppressors
A number of large-scale expression profiling studies have shown that the expression of miRNAs is deregulated under different pathological conditions, such as cancer (Table 1). Several reports have identified miRNA signatures that distinguish between normal and tumor-derived samples. 14-16 These studies indicate that the expression of miRNAs may serve as a better indicator of tumor origin and prognosis than conventional protein coding gene arrays. 15 Furthermore, a miRNA profiling study of pancreatic tissues indicates that the expression of as few as 25 miRNAs could distinguish among pancreatic adenocarcinoma, chronic pancreatitis, and normal pancreatic tissue. 17,18 Additional studies have shown that the pattern of miRNA expression serves as a highly reliable tool to predict therapeutic outcomes in leukemia and colon cancer. 19,20
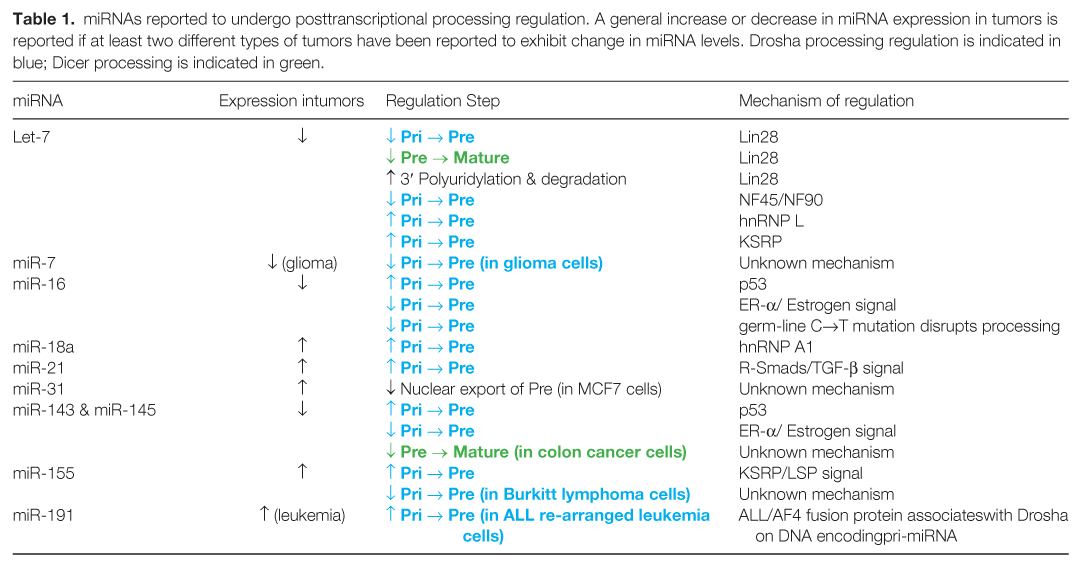
miRNAs reported to undergo posttranscriptional processing regulation. A general increase or decrease in miRNA expression in tumors is reported if at least two different types of tumors have been reported to exhibit change in miRNA levels. Drosha processing regulation is indicated in blue; Dicer processing is indicated in green.

miRNAs have been suggested to function as oncogenes or tumor suppressors based on their inhibition of tumor-suppressive and oncogenic target mRNAs, respectively. For example, miR-21 is frequently found to be overexpressed in different types of cancer and targets multiple tumor suppressor genes, including Bcl2, PTEN, programmed cell death 4 (PDCD4), sprouty, Fas ligand, tissue inhibitor of metalloproteinase 3 (TIMP3), and tropomyosin 1 (TPM1). 21-28 Increased expression of miR-21 results in the downregulation of multiple targets of miR-21 and contributes to increased proliferation and/or decreased apoptosis. Although several tumor suppressors have been identified as miR-21 targets, the combinatorial effect of these targets and the cell-type specificity of miR-21 targeting are poorly understood.
Similarly to miR-21, miR-155 is found to be highly expressed in different types of tumors and promotes oncogenesis. miR-155 is processed from an exon of the noncoding RNA B cell integration cluster (BIC), which was originally described as a viral DNA integration site in virally induced lymphomas.
29
Indeed, overexpression of miR-155 is sufficient to promote high-grade lymphoma
Although some miRNAs are demonstrated to promote oncogenesis, several large-scale miRNA expression studies have indicated that global miRNA levels are reduced in tumor samples in comparison with normal tissues. 36 This observation suggests a role for miRNA in the maintenance of homeostasis, possibly by maintaining differentiated cell phenotypes. Indeed, miRNAs have been proposed to stabilize gene expression programs and may be particularly important for the maintenance of homeostasis in the face of aberrant stimuli. 37
Several miRNAs have been reported to play tumor-suppressive roles. In particular, let-7 expression is frequently reduced in different cancers.
38
Expression of exogenous let-7 dramatically inhibits tumor growth in a lung cancer model
MicroRNA Biogenesis: The Basics
The mechanism of miRNA biosynthesis is evolutionarily conserved and involves sequential endonucleolytic cleavages mediated by two RNase III enzymes, Drosha and Dicer (Fig. 1). 44 Following transcription by RNA pol II, Drosha processes the primary miRNA transcript (pri-miRNA) into an approximately 60- to 100-nt hairpin structure termed the precursor-miRNA (pre-miRNA) in the nucleus (Fig. 1). 45-48 Following cleavage by Drosha, the pre-miRNA is transported out of the nucleus through the interaction with Exportin-5 and Ran-GTP. The pre-miRNA then undergoes further processing catalyzed by Dicer (Fig. 1). 49,50 This cleavage event gives rise to an approximately 22-nt double-stranded (ds) RNA product containing the mature miRNA guide strand and the miRNA* passenger strand (Fig. 1). The mature miRNA is then loaded into the RNA-induced silencing complex (RISC), while the passenger strand is degraded (Fig. 1). 51 Although major progress has been made in understanding the basic mechanism of miRNA biogenesis, less is known about the mechanisms that regulate miRNA biogenesis and how these systems might be deregulated during oncogenesis. In the following sections, we discuss various mechanisms of the regulation of miRNA biosynthesis and their potential link to carcinogenesis.
Genomic Organization and Transcription of MicroRNAs
miRNAs are encoded in diverse regions of the genome. Approximately 50% of miRNAs are derived from non-protein coding RNA transcripts, while an additional approximately 40% are located within the introns of protein coding genes. 52,53 Many miRNAs are encoded as clusters that range from 2 to 19 miRNA hairpins encoded in close proximity. Detection of polycistronic transcripts containing multiple miRNAs by Northern blot suggests that clustered miRNAs can be derived from a single transcript. 54 Interestingly, many mi-RNAs are mapped at or near fragile sites, common breakpoints, and other sites of genomic rearrangements associated with human cancer. 55 For example, the miR-15a/miR-16 cluster was found to be located in the 13q14.3 region, which is frequently deleted in chronic lymphocytic leukemia (CLL). 56 Furthermore, an important target of miR-15a/miR-16-1, B cell lymphoma protein 2 (BCL-2), is highly expressed in CLL, and the levels of miR-15a/miR-16-1 are inversely correlated to BCL-2 expression. 57 Ectopic expression of miR-16 reduces cell growth and induces apoptosis in a leukemic xenograft model. 58 These results indicate that loss of miRNA expression due to chromosomal rearrangements might be an underlying cause of oncogenetic programs; therefore, it is plausible that restoration of normal miRNA expression in tumor cells could have a therapeutic potential.
miRNA Transcription
Transcription is one of the major regulatory steps for the biosynthesis of mi-RNAs. Although RNA Pol III has been demonstrated to generate the transcripts of a subset of miRNAs, 59,60 the majority of miRNAs are transcribed by RNA Pol II and bear the characteristic 7-methyl-guanylate cap at the 5′ end and poly (A) tail at the 3′ end. 45,61 Large-scale mapping of the promoters of 175 human miRNA genes through nucleosome positioning and chromatin immunoprecipitation–on–genomic DNA microarray chip (or ChIP-on-chip) analysis suggests that many characteristics of miRNA promoters, such as the relative frequencies of CpG islands, TATA box, TFIIB recognition, initiator elements, and histone modifications, are similar to the promoters of protein coding genes. 60,62 The DNA binding factors that regulate miRNA transcription largely overlap with those that control protein coding genes, including c-myc and p53, as well as cell type–specific transcription factors such as MEF2, PU.1, and REST. 63-66 Furthermore, transcription of primary miRNA transcripts can be dynamically regulated in response to growth factor stimulation, such as platelet-derived growth factor (PDGF), transforming growth factor-β (TGF-β), and bone-derived neurotrophic factor (BDNF). 67-70
miRNA Transcriptional Regulation by c-Myc
The proto-oncogene
Overexpression of both c-Myc and the miR-17 cluster is known to give rise to more aggressive tumors in a mouse xenograft model. 33 The miR-17-92 cluster contains 6 miRNAs, including mi-R-17-5p and miR-20, which are known to repress the cell cycle regulator E2F1. 73 Interestingly, c-Myc also promotes E2F1 transcription, suggesting that c-Myc is likely to fine tune cell cycle progression via the regulation of both miRNA and mRNA expression. 76 In addition to activating transcription, c-Myc also decreases the expression of several tumor suppressor miRNA genes, including the miR-15a, -29, -34, and let-7 families. 74 Exogenous expression of c-Myc–repressed miRNAs in lymphoma cells reduced cell growth, indicating that downregulation of a subset of miRNAs is an important mechanism of c-Myc–mediated tumorigenesis. 74 ChIP analysis revealed that this repression is mediated, at least in part, through direct binding of c-Myc to miRNA promoters. 74 Interestingly, c-Myc has also been shown to increase transcription of Lin-28B, which in turn mediates the posttranscriptional repression of let-7 family members (discussed further below). 77
Epigenetic Control of miRNAs
As the majority of miRNA genes are transcribed by RNA Pol II, many of the mechanisms of epigenetic control known to regulate the expression of RNA Pol II–transcribed protein coding genes are also likely to apply to miRNA genes. For example, in bladder cancer, the expression of miR-127 is highly reduced due to promoter hypermethylation.
78
Several additional miRNA loci, such as miR-9-1, -193a, -137, -342, -203, and -34b/c, are also found to be hypermethylated in multiple human cancers similar to known tumor suppressor gene loci.
79,80
miRNA promoters are also regulated by histone modifications during development and pathogenesis. For example, low expression of miR-1 in lung cancer cells can be rescued by treating with histone deacetylase (HDAC) inhibitors, suggesting that the promoter of miR-1 may be aberrantly acetylated in tumors. Restored miR-1 expression reduced cell growth, mobility, and tumor formation
Alterations of MicroRNA Processing
Although transcription of the primary miRNA transcript is the initiating step of microRNA biogenesis, several lines of evidence suggest that posttranscriptional regulation of miRNA processing also plays an important role in the regulation of miRNA expression. For example, while many miRNAs are encoded within introns or as clusters of miRNAs, the mature miRNA is often differentially expressed relative to the host transcript. Furthermore, analysis of mature and primary transcripts of miRNAs in 68 tumor and 22 normal samples indicated that the correlation between the primary transcript and mature miRNA was disrupted in the tumor samples, perhaps suggesting global alterations in miRNA processing.
82
Experimental knockdown of proteins that are essential for miRNA biogenesis, such as DiGeorge syndrome critical region 8 (DGCR8), Drosha, or Dicer, in mouse lung adenocarcinoma cells dramatically elevates properties of tumorigenesis, including proliferation, soft agar colony formation, and
miRNA Processing by the Drosha Complex
Following transcription, the pri-miRNA undergoes two sequential cleavages to become the mature miRNA (Fig. 1). The first step of miRNA processing is catalyzed in the nucleus by the RNase III enzyme Drosha.
46,48,84
Pri-miRNAs generated by the transcriptional machinery are generally several thousands of nucleotides long and contain a distinctive stem loop structure. Drosha cleaves at the base of the stem to generate an approximately 60- to 100-nt hairpin RNA with a characteristic 2-nt overhang at the 3′ end (Fig. 1).
46
Although Drosha contains a highly conserved RNase III domain, purified Drosha is unable to efficiently generate pre-miRNAs
Regulation of Drosha Processing
DEAD-Box RNA Helicase–Dependent Pri-miRNA Processing
The DEAD-box RNA helicases p68 (DDX5) and p72 (DDX17) were initially identified as components of the Drosha microprocessor complex by immunoprecipitation–mass spectrometry analysis and subsequently shown to also associate with DGCR8.
47,87
Knockout of either p68 or p72 in mouse results in embryonic lethality (E11.5 or P2, respectively), and animals with double knockout of p68 and p72 show earlier lethality without obvious specific degeneration of organogenesis.
88
Microarray analysis indicates that the steady-state levels of approximately 35% (94 of 266 surveyed) of mature miRNAs in p72-null mouse embryonic fibroblasts (MEFs) are reduced versus the control, wild-type MEFs. Most of the miRNAs that are reduced in p72 knockout mice are similarly reduced in p68 knockout mice, while combined knockout of p68 and p72 does not further reduce mature miRNA levels. This observation suggests that the two proteins are largely functionally redundant or act as a heterodimer in the Drosha complex. Interestingly, while the level of pre-miRNA was also reduced in p72 or p68 knockout MEFs, the level of pri-miRNA was unchanged. Furthermore,
The role of p68/p72 in miRNA processing is further supported by the positive regulation of pri-miRNA processing mediated by multiple p68-interacting proteins, including the Smads, p53, and estrogen receptor α (ER-α) (Fig. 2).
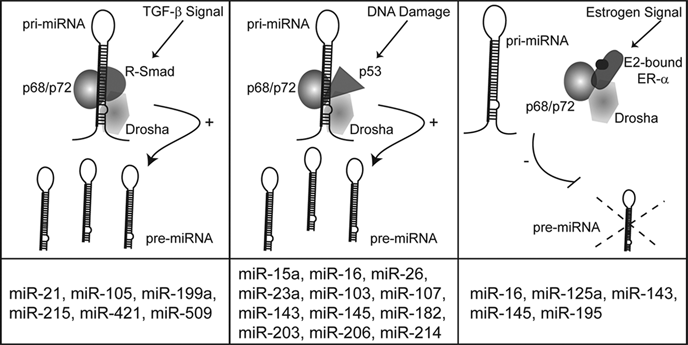
p68/p72-dependent miRNA processing pathways. The RNA helicases p68 and p72 play a critical role in the posttranscriptional regulation of numerous miRNAs in response to cellular signals, including TGF-β stimulation, DNA damage, and estrogen stimulation. The downstream mediators of TGF-β stimulation and DNA damage, the Smads and p53, act to promote miRNA processing. Conversely, when bound to E2, ER-α reduces the processing of a subset of miRNAs. The bottom column indicates miRNAs known to be posttranscriptionally regulated by each pathway. For clarity, DGCR8 and other potential members of the Drosha processing complex are not shown.
The Smads are the signal transducers of the transforming growth factor-β (TGF-β) family of signaling pathways. In the canonical pathway, ligand binding to the type I and type II TGF-β receptors promotes the nuclear accumulation of receptor-specific Smads (R-Smads) in complex with the common Smad (co-Smad), Smad4. The complex of R-Smad and co-Smad binds to specific DNA sequences within the promoter of target genes and regulates gene transcription either positively or negatively. TGF-β and its family member, bone morphogenetic protein 4 (BMP4), are particularly important for the differentiation of vascular smooth muscle cells (VSMCs). Treatment with either BMP4 or TGF-β increases expression of VSMC-specific genes; this process is due, at least in part, to the miR-21–mediated repression of programmed cell death protein-4 (PDCD4). miR-21 is rapidly induced by BMP4 or TGF-β in VSMC, which results in a subsequent decrease in PDCD4 and increased VSMC gene expression. 28 Interestingly, although knockdown of the R-Smads prevents upregulation of mature and pre-miR-21 in response to BMP4, no alteration in pri-miR-21 transcription is detected, suggesting that R-Smads posttranscriptionally affect the level of miR-21. Furthermore, BMP4 could increase the expression of pre- and mature miR-21 derived from an expression construct driven by a cytomegarovirus (CMV) promoter, suggesting that miR-21 is posttranscriptionally regulated at the Drosha processing step. The identification of R-Smads as binding partners of p68 by the yeast 2-hybrid system suggests that R-Smads could associate with the Drosha complex. 91 Consistently, coimmunoprecipitation and RNA-IP studies indicate that R-Smad is present in a complex with Drosha and p68 on the pri-miR-21 hairpin following BMP4 or TGF-β stimulation. 28 An increase in Drosha binding to pri-miR-21 is also observed after growth factor treatment, suggesting that R-Smads may promote the association of Drosha with miRNA hairpins. 28 These results indicate that in addition to the transcriptional regulation mediated by the R-Smads, TGF-β can also regulate gene expression through miRNA processing. In addition to miR-21, several other miRNAs are posttranscriptionally induced by BMP and TGF-β, suggesting that rapid modulation of miRNA levels plays an important role in cellular response to cytokine signaling (Fig. 2). 28,92
The tumor suppressor protein p53 was recently identified to modulate miRNA processing through association with p68 and Drosha, similarly to the R-Smads.
93
Under conditions of DNA damage, several miRNAs, such as miR-143 and miR-16, are posttranscriptionally induced (Fig. 2).
93
p53 is essential for this process as p53-null HCT116 cells do not induce miRNAs in response to DNA damage.
93
Coimmunoprecipitation studies indicate that p53 is present in a complex with both Drosha and p68, and the addition of p53 to
Recently, it was reported that estrogen receptor-α (ERα) also associates with p72 and Drosha upon estradiol (E2) stimulation. Unlike the Smads or p53, however, association of E2-bound ERα with the Drosha complex inhibits the association of Drosha with a subset of pri-miRNAs (Fig. 2). As a result, the association of E2-bound ERα leads to a reduction of some miRNAs. As estrogen dependency is a critical determinant of breast cancer status, it will be interesting to determine the influence of E2/ERα-mediated miRNA processing in the context of breast cancer.
Altogether, these results indicate that the association of p68/Drosha with transcription factors, such as p53, R-Smads, and ERα, is important for the rapid regulation of miRNA expression in response to extracellular stimuli. It is intriguing to speculate that other known p68-interacting transcription factors, such as MyoD, Runx2, androgen receptor, and β-catenin, 94-97 might also play a role in the Drosha microprocessor complex. Furthermore, although each of the described mechanisms requires the RNA helicases p68 or p72, the miRNAs that are regulated are not completely overlapping (Fig. 2). We have recently shown that R-Smads select miRNAs for regulation based on the presence of a conserved dsRNA sequence within the pri-miRNA stem 92 ; it will be interesting to determine if p53 and/or ERα similarly selects pri-miRNAs based on a specific sequence or structure.
DEAD-Box RNA Helicase–Independent Processing
In addition to the mechanisms described above, several processing mechanisms independent of the DEAD-box RNA helicases have been studied. For example, hnRNP A1 is required for the processing of a member of the miR-17-92 cluster, miR-18a. The addition of hnRNP A1 to
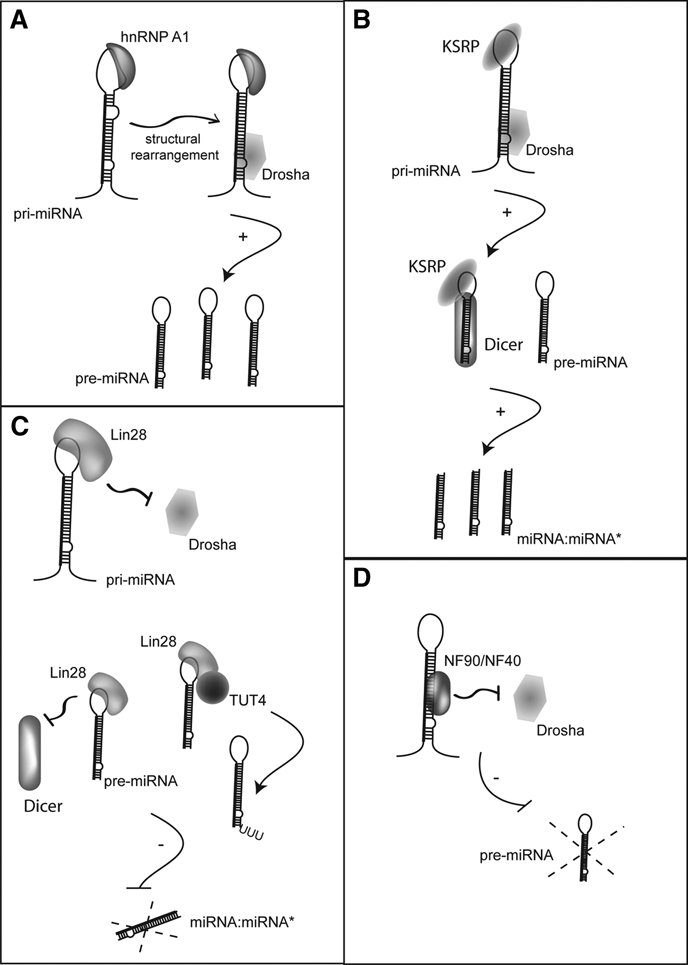
RNA helicase-independent mechanisms of miRNA processing regulation. (
The importance of the loop region in the regulation of pri-miRNA processing is further supported by recent studies, which indicate the KH-type splicing regulatory protein (KSRP) directly interacts with G-rich regions present within the loop of a subset of pri-miRNAs to promote both Drosha- and Dicer-mediated miRNA processing (Fig. 3B). 100 Transient knockdown of KSRP in HeLa cells strongly reduced the expression of a group of mature miRNAs, including let-7a and miR-206. Furthermore, knockdown of KSRP altered cellular responses mediated by these miRNAs, including proliferation and skeletal muscle differentiation. KSRP was found to be present in both the Drosha and Dicer complexes, and knockdown of KSRP reduced the cropping activity of Drosha and Dicer. 100 The interaction of KSRP with sequential pri-miRNA and pre-miNRA cleavage steps may serve to promote the synergistic activation of processing for a subset of miRNAs (Fig. 3B). Alternatively, KSRP association with either the Drosha or Dicer complexes may be sufficient to alter the miRNA processing in response to cellular stimuli. For example, lipopolysaccharide (LPS) treatment of macrophages dramatically increases mature miR-155 without significantly altering the expression of the pri-miR-155. 101 Mobility shift assays indicate KSRP interacts with pri-miR-155, and knockdown of KSRP prevents LPS-mediated elevation of miR-155. 101 Altogether, these results suggest a role of KSRP in promoting the posttranscriptional regulation of miR-155 in response to LPS. Interestingly, miR-155 is derived from the noncoding BIC RNA, which has long been associated with induction of lymphomas. 29 BIC (pri-miR-155) is strongly induced by B cell receptor signaling in a variety of cell types. In the Burkitt lymphoma–derived Ramos cell line, induction of BIC does not give rise to an increase in mature miR-155. 102 This effect seems to be unique to the Ramos cell line as expression of pri-miR-155 in HEK293 or Hodgkin lymphoma cell lines gives rise to very high levels of miR-155. 102 The mechanism of this cell type–specific effect is unclear; however, in light of the results above, it may be interesting to examine if KSRP levels or activity are altered in some lymphomas.
In addition to the positive regulation of miRNA processing mediated by hnRNPs and KSRP discussed above, negative regulation of the Drosha processing of some pri-miRNAs has also been reported. The let-7 family of mi-RNAs is encoded as multiple copies in the genome; for example, the human genome contains 9 mature let-7 sequences derived from 12 precursors.
103
While most let-7 primary transcripts are highly expressed throughout development, the mature let-7 is detectable only in highly differentiated cells, suggesting a mechanism of posttranscriptional regulation during differentiation.
103
Analysis of the pri-miRNA sequences of the human let-7 family, as well as let-7 from different species, indicates the presence of conserved nucleotide sequences in the hairpin loop. This observation suggests that this region might be critical for the posttranscriptional regulation of the let-7 family of miRNAs.
104,105
While pri-let-7 is processed efficiently
The precise mechanism of Lin-28 inhibition of Drosha processing is still unclear. As Lin-28 has a strong affinity for let-7, it is plausible that Lin-28 prevents interaction with Drosha/DGCR8 by mutual exclusion (Fig. 3C). Alternatively, similarly to hnRNP A1 discussed above, the interaction of Lin-28 with the loop region could rearrange the secondary structure of the hairpin and inhibit Drosha processing. Furthermore, as discussed further below, recent reports suggest that Lin-28 inhibits the biogenesis of let-7 through multiple mechanisms (Fig. 3C). 107-109 It is interesting to note that Lin-28, KSRP, and hnRNPs all interact with conserved nucleotides present within the loop of pri-miRNAs. The balanced interaction of proteins with the pri-miRNA loop may allow for the fine tuning of miRNA levels during development or in response to extracellular cues, such as growth factors. For example, although KSRP interacts strongly with pri-let-7g in differentiated cells, high levels of Lin28 block KSRP association in embryonic carcinoma p19 cells. 100 Interestingly, in some cell types, low expression of let-7 is not mediated by Lin-28, but through the negative regulation of Drosha processing of let-7 by hnRNP-A1. Thus the balanced expression of diverse proteins allows for the tight regulation of miRNA biogenesis. 110
Repression of Drosha processing of let-7 family members is also mediated by the nuclear factor 45 (NF45) and nuclear factor 90 (NF90) proteins.
111
Cancer-Specific Drosha Processing Alterations
Altered Drosha processing has been described for the tumor suppressor miR-7 in glioblastoma. 113 While pri-miR-7 is expressed at equal levels in normal and tumor samples, both pre-miRNA and mature miRNA are dramatically reduced in glioblastoma tumor samples and cell lines. Restoration of miR-7 levels in primary cell lines reduced tumor cell viability and invasiveness by targeting the epidermal growth factor receptor (EGFR) and inhibiting the downstream Akt signaling cascade. 113 As the levels of several other miRNAs are unaffected, the deregulation of processing is specific to miR-7. 113 Although the mechanism is unknown, reduced pre-miR-7 in the tumor could arise from either 1) increased expression of a processing inhibitor or 2) decreased expression of a processing activator. It will be interesting to examine if miR-7 is regulated by one of the mechanisms described above or if a yet unidentified mechanism is responsible for the altered regulation of miR-7 in cancer cells.
Chromosomal abnormalities associated with cancer may also give rise to altered pri-miRNA processing activities. For example, the acute lymphoblastic leukemia-1 (ALL-1) gene (also termed myeloid/lymphoid or mixed lineage leukemia [MLL]) is located at chromosome band 11q23 and is involved in chromosomal abnormalities associated with ALL and acute myeloblastic leukemia (AML). Chromosomal translocation results in the fusion of 5′ fragments of the ALL-1 gene with a part of 1 of 50 partner genes. Fusion of the ALL-1 to ALL1-fused gene from chromosome 4 (AF4) is particularly common and is associated with poor prognosis. Interestingly, a subset of miRNAs was found to be upregulated in cells bearing an ALL1/AF4 fusion protein. 114 Further analysis indicates that this upregulation is due, at least in part, to increased Drosha processing of these miRNAs. 114 Coimmunoprecipitation studies indicate that the ALL1 fusion proteins directly interact with Drosha in a DNA-dependent manner. Furthermore, ChIP studies indicate that the fusion protein could promote the recruitment of Drosha to the DNA regions encoding the affected pri-miRNAs genes. 114 Although the mechanism of ALL1/AF4-mediated processing is unclear, recent studies have shown that, in some cases, pri-miRNA processing is coordinated with transcription. 86 Thus, it is possible that Drosha could be recruited to the promoter of a miRNA gene, which in turn promotes processing of the primary transcripts. It will be interesting to determine if other fusion proteins generated by chromosomal translocations similarly give rise to altered pri-miRNA processing activities.
Pre-miRNA Export by Exportin-5
Following the first processing step by Drosha, the pre-miRNA hairpin is translocated from the nucleus to the cytoplasm by exportin-5 (Fig. 1). 115 Like other nuclear export receptors, exportin-5 cooperates with the small GTPase Ran to mediate directional transport of its cargo. Pre-miRNAs interact with exportin-5 and the GTP-bound form of Ran in the nucleus. Following export to the cytoplasm, GTP is hydrolyzed, and the pre-miRNA is released from exportin-5 (Fig. 1). 49,116
The export of pre-miRNA from the nucleus to the cytoplasm may be differentially regulated under certain physiological conditions. For example, the hairpins of pre-miR-105, -128, and -31 are detected in many cells at relatively high abundance. However, under some conditions, the mature miRNA is undetectable.
36
For example, pre-miR-31 is expressed at comparable levels in the pancreatic cancer cell line HS766T and the MCF7 breast cancer cell line. However, while HS766T cells show high levels of mature miR-31, no expression is detected in the MCF7 cells.
Dicer Cleavage and the RISC
Following translocation into the cytoplasm, the pre-miRNA is cleaved near the terminal loop by the RNase type III protein Dicer to generate an approximately 22-nt double-stranded mature miRNA (Fig. 1).
117,118
Dicer is highly conserved throughout evolution and present in nearly all eukaryotic organisms. Similarly to Drosha, several Dicer-associated proteins have been identified, including TAR RNA binding protein (TRBP) and protein kinase R–activating protein (PACT).
119,120
Association of TRBP and PACT with Dicer enhances Dicer stability and processing activity.
119,120
Furthermore, although recombinant Dicer protein can catalyze the efficient and specific cleavage of pre-miRNA
Regulation of Dicer Processing
In general, pre-miRNAs are expressed at a low level relative to pri-miRNA or mature miRNA, suggesting that Dicer is generally very efficient at processing pre-miRNA. 36,124 Unlike Drosha, which often requires accessory factors to promote processing, regulation of miRNAs at the Dicer step often involves inhibition of Dicer activity. Early evidence for posttranscriptional control of miRNA at the level of Dicer processing comes from studies of miRNA expression in colorectal neoplasia. Mature miR-143 and miR-145 are expressed at much lower levels in tumor samples than normal tissue. 125 However, Northern blots indicate that the level of pre-miR-143 or pre-miR-145 is not significantly different in the tumor samples, suggesting altered Dicer processing in the colorectal tumor samples. 125 These miRNAs act as tumor suppressors through the repression of the mitogen-activated protein kinase ERK5, suggesting that inhibition of their processing contributes to tumorigenesis. 126
miRNA processing by Dicer is also differentially regulated during normal development and tissue specification. For example, while pre-miR-138 is ubiquitously expressed in all tissues and HeLa cells, the mature miR-138 is only detected in adult mouse brain and fetal liver.
127
Potential alteration in nuclear export is ruled out as the pre-miR-138 is effectively exported into the cytoplasm; therefore, it is suggested that the regulation of Dicer processing of pre-miR-138 might be inhibited in many tissues. Furthermore, while pre-miR-138 is processed efficiently by Dicer
In addition to inhibition of the Drosha processing step discussed above, Lin-28 also inhibits the Dicer processing of let-7 family members (Fig. 3C). 107,128 Lin-28 shuttles between the nucleus and cytoplasm but is commonly enriched within the cytoplasm, suggesting that the cytoplasm may be the primary location of the Lin-28–let-7 interaction. The addition of purified Lin-28 decreases the association of let-7 with Dicer and results in reduced processing by Dicer, suggesting that Lin-28 can compete with Dicer for access to let-7 (Fig. 3C). 107 Furthermore, Lin-28 also promotes the 3′-polyuridylation of pre-let-7 by acting as a scaffold between let-7 and the terminal uridine transferase-4 (TUT4) (Fig. 3C). 108-110 Polyuridylation of pre-let-7 inhibits Dicer processing and promotes the degradation of pre-let-7. 109 Analysis of the Lin-28 binding site within the loop of let-7 identified a tetranucleotide sequence required for Lin-28/TUT4–mediated uridylation. Interestingly, this motif is located within the terminal loop of 15 additional pre-miRNAs, some of which have also been shown to undergo uridylation. 109 High-throughput sequencing analysis identifies numerous nontemplated uridine residues that are added to a variety of miRNAs, perhaps suggesting that this type of regulation extends beyond Lin-28/TUT4. 123
Regulation of Core Processing Machinery Levels
Regulation of Drosha/DGCR8 Expression
In addition to regulation by accessory factors, the total levels of Drosha and DGCR8 in the cell are tightly controlled and may play a role in the regulation of pri-miRNA processing. Increased Drosha expression is observed in late-stage cervical cancer samples and is associated with poor prognosis in esophageal cancer patients.
129,130
Interestingly, despite a 2- to 7-fold increase in Drosha expression in cervical cancer samples, only miR-31 is increased, while the other differentially expressed miRNAs were decreased.
130
DGCR8 was originally identified as a gene located within a common chromosomal deletion at ch22q11, which gives rise to a syndrome characterized by learning disabilities and heart defects termed DiGeorge syndrome.
131
Mouse models of a homologous chromosomal deletion exhibit a moderate decrease of only 59 miRNAs.
132
Given the critical role of Drosha and DGCR8 in mediating pri-miRNA processing, it is surprising that relatively minor changes are found in miRNA levels upon dramatic alteration of DGCR8/Drosha levels.
132
These findings may be explained, at least in part, by the observation that the total levels of Drosha and DGCR8 are coupled in an intricate feedback circuit. In addition to cleaving pre-miRNA hairpins, the Drosha microprocessor complex can also promote cleavage of hairpin structures within annotated protein coding genes, which are not further processed by Dicer.
133,134
This type of cleavage allows Drosha to modulate protein coding gene expression independent of miRNA production. Indeed, tiling microarray analysis in
In addition to the alteration of protein levels, the overall activity of the Drosha/DGCR8 complex may be altered under some circumstances. Cells grown to high confluency exhibit increased pre- and mature miRNA expression without alteration of Drosha or DGCR8 protein level.
135
Furthermore, extracts from high confluency cells promoted the more efficient conversion of pri- to pre-miRNA in
Regulation of Dicer Expression
The total levels of Dicer may serve as an important control point in miRNA biogenesis. A careful analysis of the Dicer 5′UTR indentified multiple alternatively spliced leader exons that are expressed in a tissue-specific manner.
137
Although differential splicing alters the translation efficiency of Dicer
The expression of Dicer may also be regulated by its cofactors. Dicer is associated with the dsRNA binding proteins TRBP and PACT. 119,120 Depletion of either of these cofactors decreases the steady-state levels of Dicer protein. 120,145 Furthermore, truncation mutations of TRBP are found in carcinoma and are associated with decreased miRNA levels and dramatic destabilization of Dicer. 146 Dicer protein levels can be rescued by full-length TRBP, suggesting that the interaction between Dicer and TRBP is a critical determinant of Dicer stability. 146 Furthermore, the expression or activity of Dicer can also be modulated by cellular signaling pathways. For example, MAPK/ERK signaling was found to promote the phosphorylation of TRBP. 147 Phosphorylated TRBP enhances miRNA production by increasing the stability of Dicer. Interestingly, increased abundance of Dicer is correlated with the increase in growth-promoting miRNAs and decrease of let-7, which has a tumor suppressor activity. 147 Understanding the molecular mechanisms that determine miRNA-specific alterations as a result of TRBP phosphorylation requires further investigation.
The Role of Ago Expression in miRNA Regulation
The Argonaute (Ago) proteins are the primary component of the RISC complex and the effectors of miRNA-mediated repression of target mRNAs.
148
The human genome contains 8 Ago family proteins: Ago1 to Ago4 and Piwi1 to Piwi4.
148
Although structurally similar to Agos, the Piwi family members interact with piRNAs and function in germline maintenance.
149
While all of the Ago proteins have the ability to interact with miRNAs and siRNAs, Ago2 is the only one with RNA cleavage activity and is thought to play a critical role in miRNA-mediated mRNA silencing.
148
Total levels of the Ago proteins within the cell also contribute to global miRNA regulation and biogenesis. Ectopic expression of Ago proteins results in a dramatic increase in mature miRNAs.
150
In contrast, mouse embryonic fibroblasts and hematopoietic cells deleted in
Ago2 mRNA and protein expression are also found to be increased in a subset of breast cancer cells. Epidermal growth factor (EGF) treatment enhances Ago2 stability, suggesting that Ago2 levels can be regulated at both the transcriptional and posttranscriptional level. 153 Furthermore, increased Ago2 expression was associated with a transformed phenotype in breast cancer cells. 153 Interestingly, Ago2 is recently found to be targeted for ubiquitination and degradation by Lin-41. 154 Lin-41 is expressed primarily in stem cells and undifferentiated cell types. Coimmunoprecipitation studies indicate that Lin-41 directly interacts with Ago2 and overexpression of Lin-41 reduced miRNA-mediated mRNA silencing. 154 Reduction of Ago2 by Lin-41 may serve to prevent expression of miRNAs and inhibit cellular differentiation, thus maintaining a stem cell–like phenotype. 154 Interestingly, Lin-41 is a target of let-7, which is highly regulated and important for cellular differentiation. 107 Similarly to Lin-28, the levels of Lin-41 are inversely correlated with the level of let-7. 154 It is interesting to speculate that in conditions that are characterized by low levels of let-7, such as lung cancer, Lin-41 may be increased, which could in turn result in reduced expression of Ago2. Indeed, immunohistochemical analysis of 68 gastric and colorectal tumors indicates a reduction in Ago2 protein in 40% and 35% of gastric and colorectal cancers, respectively. 155 In the future, it will be interesting to determine if alterations in Ago2 protein are a universal characteristic of poorly differentiated tumors.
Single Nucleotide Polymorphisms and Mutations Can Alter miRNA Processing
In addition to changes in the processing enzymes, single nucleotide polymorphisms (SNPs) and point mutations within the pri-miRNA have the potential to alter the secondary structure and thus prevent processing of the pri-miRNA. An analysis of pri-miRNA sequences in CLL identified mutations in 5 of 42 pri-miRNAs examined. 19 While no pri-miRNA mutations were identified in 160 normal samples, 15% of 75 CLL samples exhibited at least one mutation. 19 In particular, a germline cytosine-to-thymine mutation 7-bp 3′ to the pre-miR-16-1 region is associated with reduced mature miRNA-16-1 levels, indicating that mutations in the pre-miRNA may influence mature miRNA levels. 19 In addition to mutations, SNPs may also alter miRNA processing. For example, a common guanine/cytosine polymorphism within the precursor of miR-146 is associated with increased risk of papillary thyroid cancer. 156 The cytosine allele exhibits reduced expression of mature miR-146 and results in an increase in miR-146 targets compared to individuals with the guanine allele. 156 The SNP is located within the passenger strand of miR-146 (miR-146*), and the cytosine allele alters the secondary structure of pri-miR-146 and inhibits Drosha processing. 156 Similarly, a polymorphism within the mature sequence of miR-125a disrupts Drosha processing by introducing a bulge within the hairpin. 157 Processing of miR-125a could be rescued by the introduction of a complementary mutation on the passenger strand, which restores the stem structure. 157 Together, these studies emphasize the importance of secondary structure in the control of miRNA biogenesis.
Alterations in microRNA Binding Sites Can Alter miRNA Activity
In addition to altering miRNA biogenesis, SNPs may play a role in miRNA-regulated gene expression when they are located within 3′UTR of target mRNAs. For example, SNPs may perturb miRNA function by introducing or disrupting miRNA recognition sequences. A method to determine differences in mRNA expression associated with a miRNA binding site present in one allele, but not in the other, termed allelic imbalance sequencing (AI-seq), suggests that single nucleotide polymorphic sites may be responsible for the differential regulation of a large number of mRNAs (7.5%-15%).
158
Importantly, this means that the response of 2 individuals to the same miRNA may differ based on their genetic composition. Several physiologically relevant examples of altered miRNA targeting due to SNPs have been identified. For example, a SNP within the let-7 binding site of the
In addition to genetic alterations of the 3′UTR sequence, such as SNPs or mutations, the ability of a miRNA to repress a target mRNA is limited by the presence of miRNA binding sites within the 3′UTR. Alternative usage of cleavage and polyadenylation sites may provide a simple mechanism for the tissue-specific regulation of miRNA-mediated repression. Analysis of alternative 3′UTR isoforms indicates a global reduction of 3′UTR length in proliferating cells, 161 while differentiated cells express mRNAs with longer 3′UTRs. 162 As shortened 3′UTRs contain fewer miRNA binding sites, mRNAs in proliferating cells are more resistant to miRNA-dependent repression. 161 Furthermore, analysis of mRNA 3′UTRs indicates that the precise 3′ end of some mRNAs could distinguish different tumor subtypes and is associated with altered prognosis. 163 It is speculated that the effects of both 1) reduced expression of miRNAs and 2) increased expression of mRNAs with shorter 3′UTRs may synergistically contribute to the pathogenesis of human disease, including cancer.
Conclusions
miRNAs represent critical regulators of cell type differentiation, proliferation, and survival. Alterations in the expression of miRNAs are clearly linked to the progression of numerous human disorders, in particular, cancer. Multiple mechanisms of miRNA regulation have been described, and it is now clear that although the general biogenesis of miRNAs is conserved, diverse pathways influence these processes for the generation of some miRNAs. In the future, it will be important to investigate how multiple mechanisms of miRNA regulation cooperate to give rise to tightly controlled miRNA expression. Furthermore, an understanding of how miRNAs are regulated under normal conditions may allow a more complete understanding of the processes that are deregulated during cancer.
Footnotes
Acknowledgements
The authors thank Drs. Gelman and Sudol for giving us the opportunity to contribute to the special Dr. Hanafusa’s memorial issue of
The author(s) declared no potential conflicts of interest with respect to the authorship and/or publication of this article.
This work was supported by the National Institute of Health [grant numbers HD042149 and HL082854 to A.H., T32 HL069770 to B.N.D.] and by the American Heart Association [grant number 0940095N to A.H.].
