Abstract
Background
Immune-related diseases can interact with each other, and growing evidence suggests that these diseases are associated with allergic rhinitis (AR). However, it is unclear whether previously observed associations reflect causal relationships.
Objective
This study estimated the genetic association between various immune-related diseases and AR using two-sample Mendelian randomization (MR).
Methods
Eight immune-related diseases were selected as exposure factors, and AR was selected as the outcome. The 8 immune-related disease categories included atopic dermatitis (AD), Graves’ disease (GD), asthma, Crohn's disease (CD), multiple sclerosis (MS), rheumatoid arthritis (RA), systemic lupus erythematosus (SLE), and ulcerative colitis (UC). Data from GWAS (Genome-Wide Association Studies) were selected to construct instrumental variables (IVs) for each disease, and multiple single-nucleotide polymorphisms (SNPs) were selected as IVs. Corresponding data were retrieved according to the selected SNPs, and all data were summarized and analyzed.
Results
A total of 416 SNPs were screened as IVs, and the results of IVW support a causal relationship between AR risk and AD (OR: 1.026, 95% CI: 1.014-1.038, P = 9.59 × 10−6), asthma (OR: 1.057, 95% CI: 1.029-1.086, P = .0001), and CD (OR: 1.006, 95% CI: 1.002-1.011, P = .0085). Furthermore, GD (OR: 0.995, 95% CI: 0.991-0.999, P = .0213) and SLE (OR: 0.997, 95% CI: 0.995-1.000, P = .025) may be protective factors.
Conclusion
This MR study found that AD, asthma and CD increase the risk of AR in populations of European ancestry, GD and SLE may be protective factors. These results suggest that confounding factors may have influenced associations previously reported in observational studies.
Introduction
Allergic rhinitis (AR) is a hypersensitivity reaction driven by type 2 helper T cells (Th2) as a result of an inhaled allergen-induced immunoglobulin E (IgE)-mediated inflammatory response in the nasal mucosa.1,2 Common symptoms of AR include rhinorrhea, sneezing, nasal pruritus and nasal obstruction and are often accompanied by atopic dermatitis (AD), asthma, conjunctivitis and atopic comorbidities. The estimated prevalence of AR in the United States and Europe is 15% to 30%. 3 In recent years, worsening environmental pollution and climate change have had a serious impact on airway diseases. A longitudinal study involving 18 cities across China showed that AR prevalence is increasing by approximately 1% per year, resulting in an enormous social and economic burden that cannot be ignored. 4
In particular, many studies show that the primary characteristic of many immune-related diseases is a high T-cell count, as well as a significant imbalance of T-cell subsets. 5 Autoimmune diseases (AIDs) refer to the loss of autoimmune immune tolerance and excessive autoantibody production, which leads to chronic systemic disease with immune abnormalities and has a negative impact on physical health and quality of life. 6 The pathogenesis of AID has not been fully elucidated. Common AIDs include autoimmune hyperthyroidism, Crohn's disease, multiple sclerosis, rheumatoid arthritis, systemic lupus erythematosus, ulcerative colitis, etc. Both AID and atopic allergies result from a dysregulated immune system. 7 Some studies have suggested that type 1 helper T cells (Th1) upregulation could favor AIDs and that Th2 upregulation could favor atopic allergies. Concerning the relationship between these immune-related diseases, past studies have produced discordant results, with the magnitude of reported associations varying considerably across studies. Therefore, a causal relationship between immune-related diseases and AR has not been established.
In traditional observational epidemiology, the association between exposure and outcome may be influenced by confounding factors and reverse causal associations, thus limiting the inference of causality between outcome and exposure.8,9 Many studies have attempted to evaluate the causal relationship between AR and AID, with results varying greatly. Mendelian randomization (MR) is based on the principle of the random distribution of chromosomes during meiosis. Since chromosome allocation is a completely random process, the distribution of genetic variation in a population can be used as a measure to verify causality between exposure factors and response outcomes.
MR uses single-nucleotide polymorphisms (SNPs) as instrumental variables (IVs) to infer causal associations between exposure and outcome, overcoming the influence of confounding factors and reverse causal associations on causal inference. Therefore, we performed a two-sample MR study to explore the causal relationship between immune-related disease and AR using the largest and most recently available GWAS dataset.
Methods
Mendelian Randomization
The MR analysis was performed based on the following three assumptions: (a) genetic variants were strongly associated with exposure; (b) genetic variants were only associated with the outcome through exposure; and (c) the association was independent of any potential confounders (Figure 1). In general, the use of GWAS aggregate data satisfies the first hypothesis well, while the last 2 hypotheses sometimes do not. SNPs can directly affect exposure. Since genetic variation is not influenced by potential confounding factors or outcome, outcome can only be affected through exposure, which is affected by genetic variation. Therefore, the association observed from MR should be causal.
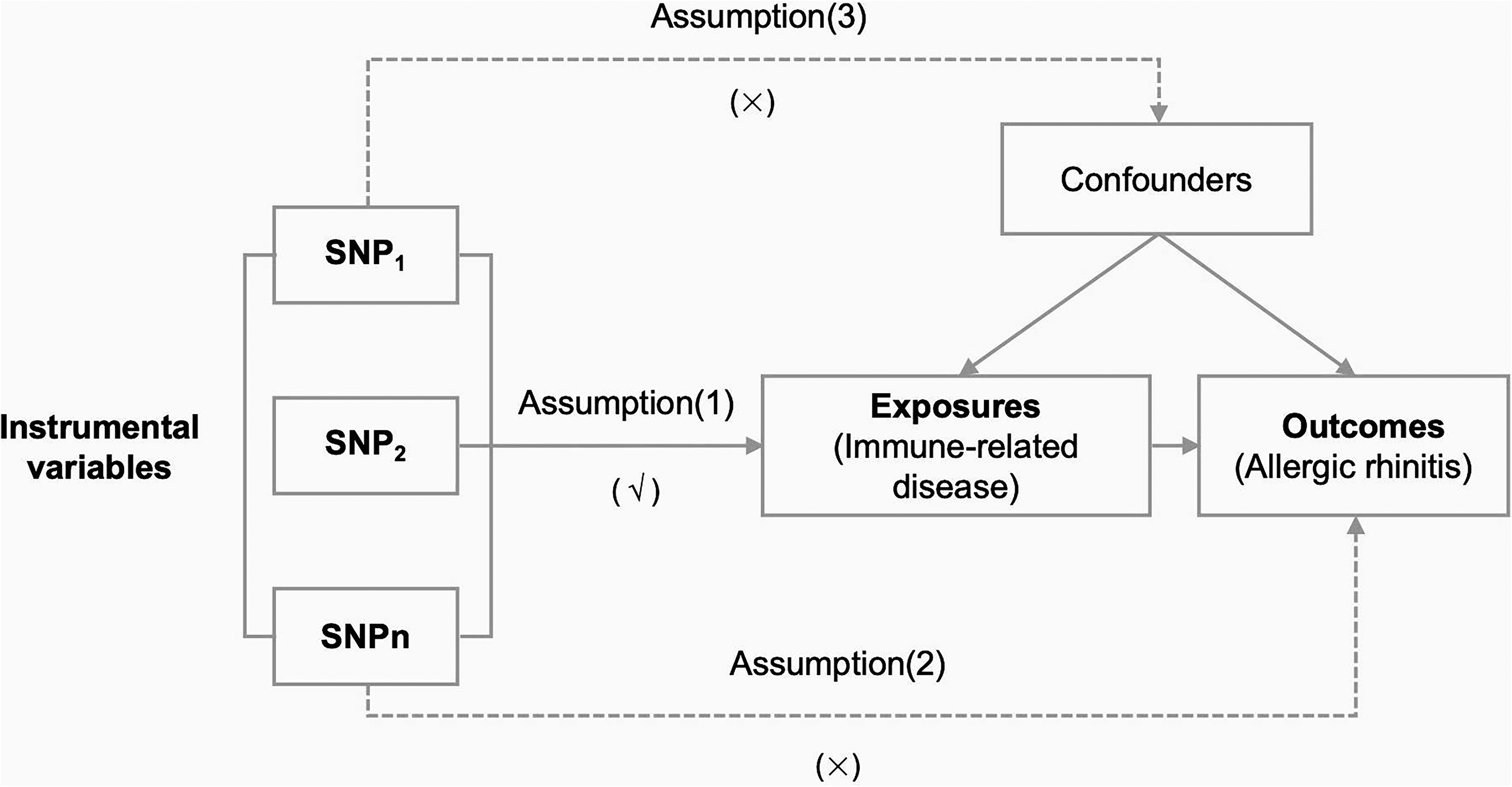
Overview of Mendelian randomization.
Data Source Description
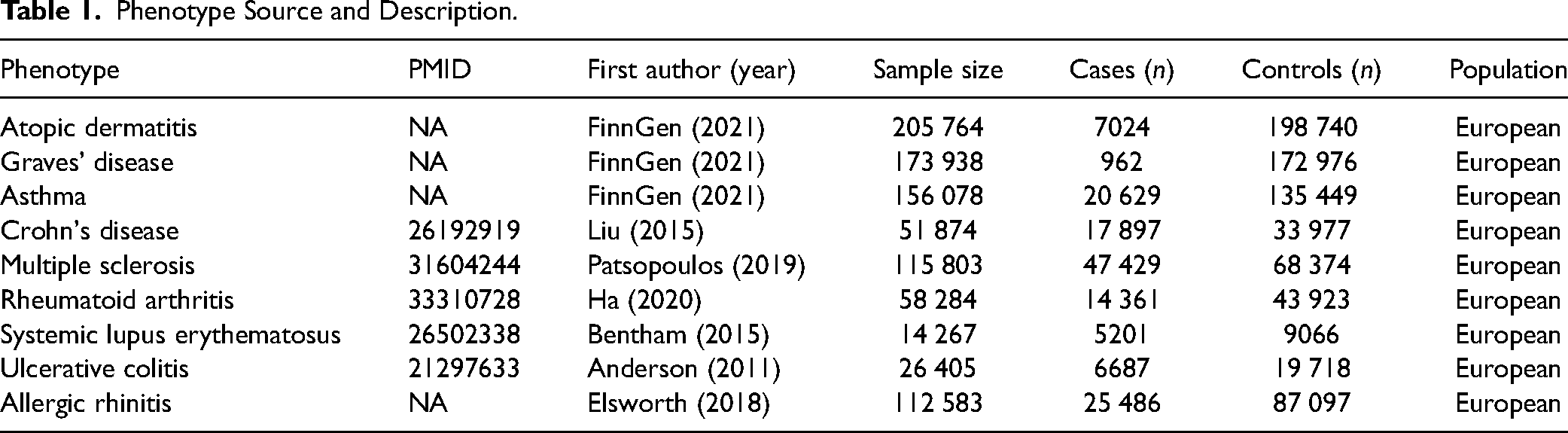
This MR study used summary-level data from a published GWAS dataset on AR and immune-related disease, which obtained appropriate patient consent and ethical approval. Eight immuno-related diseases were selected as exposure variables, and AR was selected as the outcome variable. The summary data of AD (finn-b-L12_ATOPIC), autoimmune hyperthyroidism (finn-b-AUTOIMMUNE_HYPERTHYROIDISM), asthma (finn-b-J10_ASTHMA), Crohn's disease (ieu-a-12), multiple sclerosis (ieu-b-18), rheumatoid arthritis (ebi-a-GCST90013534), systemic lupus erythematosus (ebi-a-GCST003156), ulcerative colitis (ebi-a-GCST000964) and AR (ukb−b−7178) were obtained from the official website of GWAS (https://gwas.mrcieu.ac.uk). The above GWAS databases all represent European populations (Table 1).
Phenotype Source and Description.
Statistical Analysis
All statistical analyses were performed using R software (version 4.0.2).
To verify the causal association between immune-related disease and AR, we selected SNPs in immune-related disease as measurement evidence and screened out SNPs significantly associated with disease in the whole genome (P < 5 × 10−8). To avoid confounding factors, a computational evaluation of linkage disequilibrium (LD) was performed to ensure that the selected SNPs were not associated with other loci. We removed the SNPs that had a linkage imbalance with each other (r2 < 0.01), and independent SNPs were identified in 8 immune-related diseases to evaluate causality.
In this study, inverse variance weighting (IVW) of random effects was used as the primary analysis method, which provides the most accurate estimate but is sensitive to pleiotropy. Therefore, the weighted median method and MR‒Egger regression were adopted for sensitivity analysis. The weighted median method produces consistent estimates of causation, assuming that 50% of the weight comes from valid SNPs. Therefore, MR‒Egger regression can detect possible pleiotropic effects and provide estimates after adjusting for pleiotropic effects.
We calculated Cochrane's Q value to assess heterogeneity. If P > .05, it was determined that there was no heterogeneity among SNPs. If heterogeneity existed, we used the random effects model for recalculation. The leave-one-out method was used for sensitivity analysis, and the causal effect of the remaining SNPs on outcome treatment was analyzed after SNPs were removed one-by-one to evaluate the influence of a single SNP on the outcome.
Results
Selection of Instrumental Variables
A total of 416 SNPs significantly correlated with immune-related disease were screened out on the basis of the three assumptions of MR and screening IVs. The number of SNPs used as IVs were as follows: 15 SNPs, AD; 4 SNPs, GD; 11 SNPs, Asthma; 120 SNPs, Crohn's disease; 61 SNPs, MS; 77 SNPs, RA; 40 SNPs, SLE; 88 SNPs, UC. All SNPs were significantly correlated with the corresponding AID (P < 5 × 10-8), as shown in Table 2 and Supplemental Table S1.
MR Analysis of Immune-Related Disease and Allergic Rhinitis.
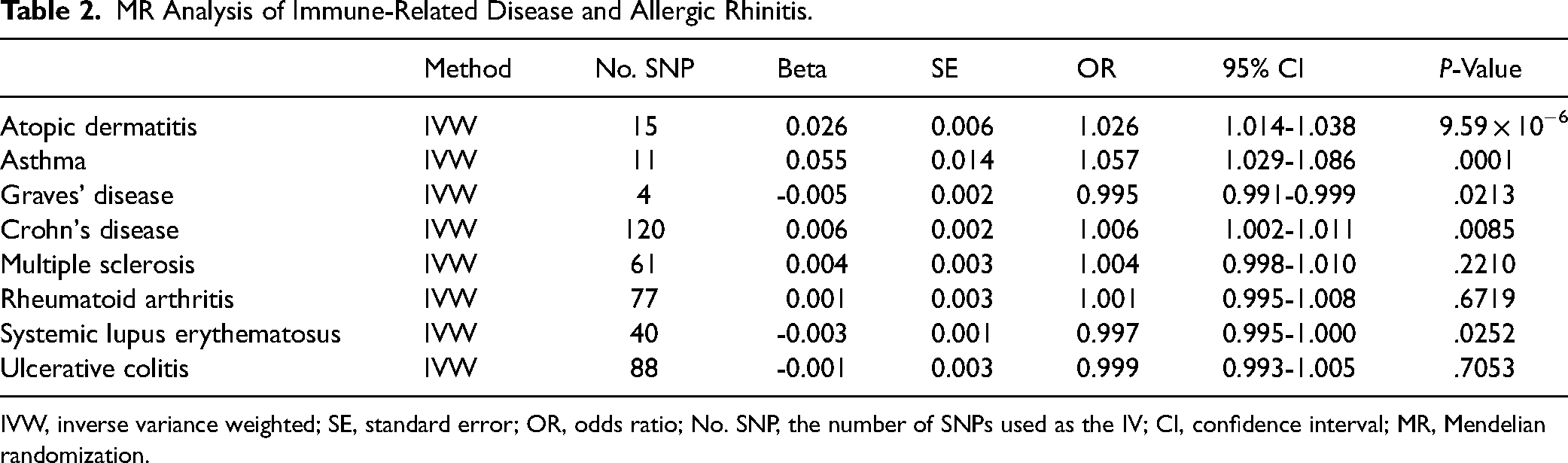
IVW, inverse variance weighted; SE, standard error; OR, odds ratio; No. SNP, the number of SNPs used as the IV; CI, confidence interval; MR, Mendelian randomization.
Causal Effects of Immune-Related Disease on AR
In this study, MR was used to analyze the causal association between 8 immune-related diseases and AR (Figure 2, Table 2). In the discovery stage, the genetic prediction of AD, GD, asthma, CD, and SLE was associated with the risk of AR. Specifically, genetically predicted AD [odds ratio (OR): 1.026, 95% confidence interval (CI): 1.014-1.038, P = 9.59 × 10−6], asthma (OR: 1.057, 95% CI: 1.029-1.086, P = .0001) and CD (OR: 1.006, 95% CI: 1.002-1.011, P = .0085) were associated with a higher risk of AR. In contrast, higher genetically predicted GD (OR: 0.995, 95% CI: 0.991-0.999, P = .0213) and SLE (OR: 0.997, 95% CI: 0.995-1.000, P = .0252) were associated with a lower risk of AR. Furthermore, findings calculated using the IVW method showed that UC had a protective effect on AR, while MS and RA were risk factors for AR, there was no significant difference in the results. Therefore, a causal relationship between UC, MS, or RA with AR could not be established (Supplemental Figures S1 to S8).
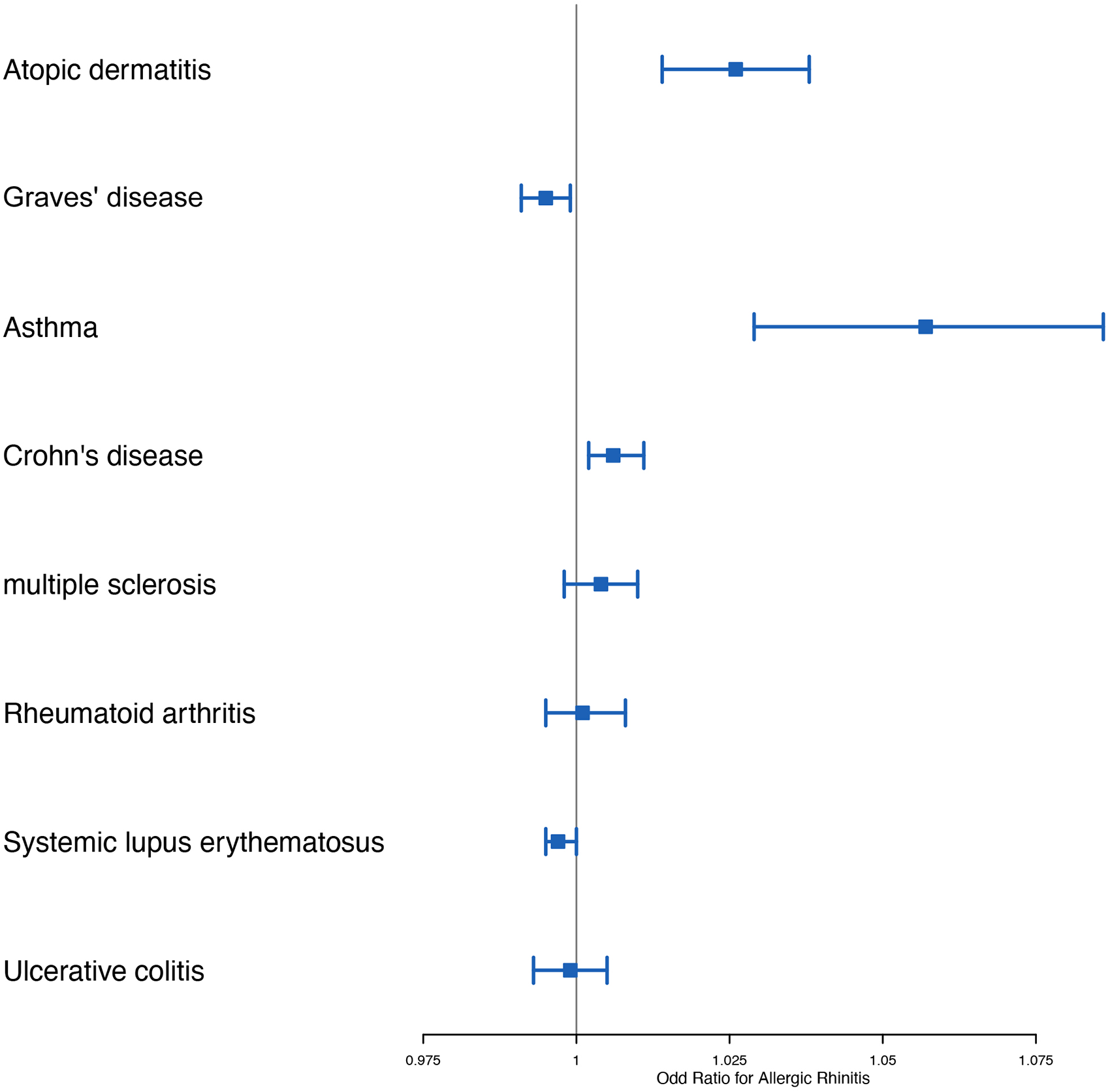
Results of Mendelian randomization.
Sensitivity Analyses
Heterogeneity was observed between the SNPs for immune-related disease, although none of the MR‒Egger regression intercepts deviated from null. Horizontal pleiotropy was observed between the SNPs of AD (P for intercept = .024), MS (P for intercept = .024), and SLE (P for intercept = .006), as shown in Supplemental Table S1.
Additionally, the leave-one-out analysis showed no marked difference in causal estimations of AID on AR, suggesting that none of the identified causal associations were driven by any single IV (Supplemental Figures S9 to S16).
Discussion
SNP is a key tool used in GWAS to study genetic susceptibility, pleiotropy, and polymorphism. SNP is a single-base difference in the DNA sequence that exists widely in the human genome and can be used to find genetic variations associated with specific traits, diseases, or physiological characteristics. 10 Genetic susceptibility refers to an individual's tendency or risk for a certain disease, disorder, or physiological characteristic at the genetic level. This susceptibility can be caused by genetic variations in one or multiple genes and may also interact with environmental factors.10,11 Gene polymorphism refers to the presence of different alleles of the same gene in a population. SNPs can lead to the existence of different alleles, and the polymorphism of SNPs allows different individuals to have different genetic variations on the same gene. 12 Pleiotropy refers to the phenomenon that a gene or a genetic variation affects multiple traits or functions. This means that an SNP or a gene can produce multiple effects in different biological processes. 13 Therefore, through the SNP data analysis of GAWS, it help us understand why some people are more likely to suffer from certain diseases or show specific physiological characteristics, which reflects genetic susceptibility, gene pleiotropy, and gene polymorphism.
In this study, we performed two-sample MR analyses, using the tool of SNPs, to investigate the causal association between 8 common immune-related diseases (AD, GD, asthma, CD, SLE, RA, MS, and UC) and AR. AR is a systemic inflammatory disorder involving pathophysiological mechanisms, numerous cells and inflammatory mediators that are similar to autoimmune disorders. 14 Combining evidence from MR, we identified that AD, asthma, and CD were causally associated with the risk of AR, that GD and SLE may be protective factors, and that there is no causal relationship between MS, RA, UC, and AR.
The risk factors associated with developing atopic diseases are complex, and the time pattern described in atopic progression may not be simple. The development of atopic diseases is strongly influenced by both genetic and environmental factors. Excluding environmental factors, underlying atopy is thought to be the key link between AD, AR, and asthma.15,16
Allergic diseases are pathogenically linked through a common genetic predisposition and frequently occur together with a temporal mechanistic sequence, ie, although not all patients follow a set pattern in disease progression, AD is often considered the first step in the development of atopic diseases. 17 In general, patients with AD may develop a typical sequence of AD, AR, and asthma at certain ages. Several studies show that children with AD have an increased odds ratio of developing AR compared to children without AD. Additionally, AD patients that present with specific IgE antibodies for common environmental allergens by 2 to 4 years of age are at higher risk for developing AR than those with AD without IgE sensitization.18,19 Therefore, the role of AD as a risk factor for AR has been established. The results of this study are consistent with the conclusions of previous studies. Epidemiologic studies illustrate strong associations between rhinitis and asthma. The findings of several studies support AR as a risk factor for developing asthma, as it can precede asthma in the atopic march 16 ; however, asthma as a risk factor for AR has not been well assessed.
Previous studies have shown that atopic manifestations are more common in patients with CD than in the general population. 20 Epidemiological evidence also suggests that children with CD have an increased risk of AR. 21 Epidemiological evidence shows that CD is a risk factor for AR, but there are few studies that establish the pathological basis for this association. 22 Additional research is needed to further clarify the mechanism of causation. During the immune response of CD, Th0 cells differentiate into Th2 cells under the influence of IL-4, then, Th2 cells further produces IL-13, increasing intestinal permeability, inducing epithelial cell apoptosis, and thereby damage the epithelial barriers of the intestine. 23 This mechanism is similar to AR, where IL-4 and IL-13 are key cytokines in the immune response and cause destruction of the nasal mucosal epithelial barrier. Therefore, CD and AR may have a causal relationship through the synergistic effect of Th2 cells, which may be one of the reasons why CD is a risk factor for AR.
At present, it is generally believed that asthma and AR belong to the same airway disease, “same airway, same disease,” and both immune mechanisms are similar, both associated with Th 2 cells, the main cytokines are IL-5 and IL-4 and IL-13. 24 Some monoclonal antibodies, such as omalizumab (IgE), mepolizumab, reslizumab, benralizumab (IL-5) and dupilumab (IL-4 / IL-13), can not only treat asthma, but also can relieve the symptom of AR.25,26 This study found that the increased risk of AR in asthma patients, which may be related to the continuity of nasal and respiratory mucosa and similar immune mechanisms.
The causal relationship between GD and AR is still unclear. Although some studies have suggested that GD may be a risk factor for AR, the conclusions of these studies are inconsistent and not fully confirmed. 27 Although the mechanisms underlying these two diseases are different, they are both related to the immune system, and thus may have some overlapping effects. Epidemiological evidence suggests an association between GD and AR. 28 Studies have reported that the prevalence of seasonal AR is significantly higher in patients with GD, and it has been noted that recurrence of GD can be induced by an allergy to pollen. 29 Meanwhile, GD and AR share a common inflammatory pathological basis, which can be used as a possible mechanism to explain the correlation between GD and AR. Hidaka et al found elevated eosinophilic neurotoxin (EDN), a product of activation and stimulation of eosinophilic degranulation in allergic diseases, in untreated patients with Graves’ disease. In addition, EDN levels significantly correlated with the activity of serum thyrotropin receptor antibodies against GD, the major pathogenic antibody. 30 These data suggest that the allergic condition is closely related to GD and that GD can increase the risk of AR. However, some studies suggest otherwise. Several studies suggest that Th2 cell-mediated abnormal humoral immunity is important in GD, although which cell type is predominantly involved in the pathogenesis of GD remains controversial. In this study, several published GWAS data sets were used and analyzed by MR. The results indicated by IVW (OR: 0.995, 95% CI: 0.991-0.999, P = .0213) supported a causal association between GD and a decreased risk of AR.
Similar findings are observed in studies on the relationship between SLE and AR, and many have shown a clear link between AR and SLE. 31 As early as 1976, Goldman et al found that SLE patients were more likely to be affected by AR than the general population. Recent studies by Taiwanese scholars have shown that AR incidence in SLE patients is approximately 2 times higher than that in the non-SLE control group. 28 While there appears to be a strong relationship between the two disorders, previous studies largely involve case‒control or cross-sectional designs; thus, the direction of causality remains uncertain.
MS is a chronic AID that affects the central nervous system. Its cause is not completely understood, and the immune mechanism is thought to be related to CD4+ T cells, Th1 and Th17 cells. 32 Considering that the immune mechanism of AR is mainly Th2-related, which may be one of the reasons why MS and AR have no causal relationship with each other. Although UC is also considered a Th2-driven disease, its main cytokines involved are TNF, IL-23, IL-19, and IL-36. While the main cytokines involved of AR is IL-4, IL-5, IL-13, IL-25, which are quite different from UC. Moreover, the Th2 mechanism in UC mainly functions in the gut homing of T cells. 33 Therefore, this may be one of the reasons why UC is not causally related to AR.
RA is a disease with a variety of cytokines and autoantibodies involved in its pathogenesis, if untreated, eventually often lead to cartilage erosion and bone destruction. There are some clinical evidences indicated that patients with RA have a lower prevalence of atopic disease, 34 and the results of the a meta-analysis were consistent. 35 We analyzed the causes of this phenomenon as follows: First, there are obvious differences between RA and AR in pathogenesis and location. The lesions of AR are mainly confined to type I allergic disease of respiratory mucosa, while RA is type III allergic disease, the pathological process is the deposition of immune complex in synovial membrane of joint and causes inflammation. 36 In terms of immune mechanism, the levels of cytokines secreted by Th1 and Th2 in RA and AR are obviously unbalanced, in which RA is dominated by Th1 cells, while AR is dominated by Th2 cells. In addition, Millauer et al simultaneously measured the level of IgE–anti-IgE complex in RA patients and found that it was significantly increased, indicating that anti-IgE antibodies exist in RA patients, which may bind to IgE to form complexes, thereby preventing basophil and mast cell sensitization, and thereby reducing the incidence of allergic diseases. 37 Therefore, AR patients are less likely to suffer from RA.
In our study, we found that some AIDs may be protective of AR, while others may increase the risk of AR, which we believe reflects complex immune system interactions that may be associated, but not necessarily converge, with the pathological basis of inflammatory or immune diseases. Overall, the pathological basis of inflammatory or immune diseases may be similar in some ways because they all involve the abnormal activity of the immune system, but there are also important differences that may be related to the pathogenesis of specific diseases and the types of immune responses, such as the activation of different types of immune cells, different curing inflammatory factors, and different types of immune responses.
For contradictory results, a possible explanation for the present findings could be that the difference may be due to the fact that the differences in the design of different types of studies. Firstly, the randomized control trial (RCT) is experimental study designs that control for interventions between treatment and control groups, with the aim of evaluating whether an intervention is effective. 38 RCTs can eliminate the effects of population heterogeneity and confounding factors by random assignment, thereby making the results more reliable. 39 In contrast, MR studies involve random assignment at the individual level, with the aim of evaluating the effects of genetic variation on intermediate phenotypes and outcomes. 40 Additionally, so far, most GWAS studies have been conducted in middle-aged and elderly individuals, and the lack of association between SNPs and intermediate phenotypes or disease outcomes may be due to the cumulative effects of older age and other gene-environment or gene-gene interactions. 41 In conclusion, these factors may be the reason why results from MR studies are inconsistent with those from RCTs and cross-sectional studies. To obtain more accurate results, we need to use different types of study designs in combination and consider the effects of complex factors. It should be noticed that, epidemiological surveys and cohort studies may be affected by confounding factors, such as time, environment, and may bias the results of observational studies. These factors may be better controlled in MR. 42 MR focuses only on the role of genetic aspects on causality, and thus the conclusions drawn may differ from real-world studies. But MR explains causality from a genetic perspective, so it cannot fully explain the effects in the real world. The relationship between these complex chronic diseases still needs to be explored more in the future. The SNPs identified in our study are associated with disease risk and can be used as a new direction for biological research, but the role of these SNPs is unclear and more studies are needed. 43
Our study has several significant strengths. MR is less susceptible to confounding factors and reverse causality not observed in observational studies. Furthermore, multifactorial assessments of different models and sensitivity analyses increase the robustness of the results, thereby strengthening the conclusions drawn. Nevertheless, our study had several limitations. First, the sensitivity analysis indicated that heterogeneity could be observed among the SNPs of immune-related diseases; however, this did not affect the results of IVW. The random effects model was used to calculate effect size, which ensured that the conclusions drawn were reliable. Second, this study used aggregated GWAS data, which could not assess the nonlinear relationship between exposure and outcome. Moreover, due to the lack of individual data, stratified analyses by sex, age, or other factors could not be conducted. In addition, the results of this study are based on a European population and, therefore, the causal relationships established in this study may be racially biased. Confirmation in other genetically diverse populations is required. Besides, there are also some limitations in the relationship between exposure factors and outcomes inferred by MR, such as: failure to establish reliable genotype—intermediate phenotype or genotype—disease associations, confounding of genotype—intermediate phenotype—disease associations, pleiotropy and the multi-function of genes, canalization and developmental compensation and lack of suitable polymorphisms for studying modifiable exposures of interest. 43
Conclusion
This MR study found that AD, asthma and CD increase the risk of AR in populations of European ancestry, GD and SLE may be protective factors, and that there is no causal relationship between MS, RA, UC and AR risk. However, the effect of these influencing factors is relatively weak, which suggests that confounding factors may have influenced the associations previously reported in observational studies.
Supplemental Material
sj-docx-1-ajr-10.1177_19458924231207131 - Supplemental material for Association Between Immune-Related Disease and Allergic Rhinitis: A Two-Sample Mendelian Randomization Study
Supplemental material, sj-docx-1-ajr-10.1177_19458924231207131 for Association Between Immune-Related Disease and Allergic Rhinitis: A Two-Sample Mendelian Randomization Study by Jinming Zhao, Mengmeng Zhang and Zufei Li in American Journal of Rhinology & Allergy
Supplemental Material
sj-docx-2-ajr-10.1177_19458924231207131 - Supplemental material for Association Between Immune-Related Disease and Allergic Rhinitis: A Two-Sample Mendelian Randomization Study
Supplemental material, sj-docx-2-ajr-10.1177_19458924231207131 for Association Between Immune-Related Disease and Allergic Rhinitis: A Two-Sample Mendelian Randomization Study by Jinming Zhao, Mengmeng Zhang and Zufei Li in American Journal of Rhinology & Allergy
Footnotes
Authors’ Contributions
Drafting the manuscript: Jinming Zhao. Acquisition of data: Jinming Zhao, Zufei Li, and Mengmeng Zhang. Analysis of data: Jinming Zhao, Zufei Li, and Mengmeng Zhang. Revising the manuscript for important intellectual content: Jinming Zhao, Zufei Li, and Mengmeng Zhang. Conception and design of the study: Jinming Zhao, Zufei Li, and Mengmeng Zhang.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
All of the procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
