Abstract
Background
Chronic rhinosinusitis (CRS) is a heterogeneous condition characterized by differing inflammatory endotypes. The identification of suitable biomarkers could enable personalized approaches to treatment selection.
Objective
This study aimed to identify and summarize clinical studies of biomarkers in adults with CRS in order to inform future research into CRS endotypes.
Methods
We conducted systematic searches of MEDLINE and Web of Science from inception to January 30, 2022 and included all clinical studies of adult CRS patients and healthy controls measuring biomarkers using enzyme-linked immunosorbent assays or Luminex immunoassays. Outcomes included the name and tissue type of identified biomarkers and expression patterns within CRS phenotypes. Study quality was assessed using the National Institutes of Health quality assessment tool for observational cohort and cross-sectional studies. A narrative synthesis was performed.
Results
We identified 78 relevant studies involving up to 9394 patients, predominantly with CRS with nasal polyposis. Studies identified 80 biomarkers from nasal tissue, 25 from nasal secretions, 14 from nasal lavage fluid, 24 from serum, and one from urine. The majority of biomarkers found to distinguish CRS phenotypes were identified in nasal tissue, especially in nasal polyps. Serum biomarkers were more commonly found to differentiate CRS from controls. The most frequently measured biomarker was IL-5, followed by IL-13 and IL-4. Serum IgE, IL-17, pentraxin-3 and nasal phospho-janus kinase 2, IL-5, IL-6, IL-17A, granulocyte-colony stimulating factor, and interferon gamma were identified as correlated with disease severity.
Conclusion
We have identified numerous potential biomarkers to differentiate a range of CRS phenotypes. Future studies should focus on the prognostic role of nasal tissue biomarkers or expand on the more limited studies of nasal secretions and nasal lavage fluid.
We registered this study in PROSPERO (CRD42022302787).
Keywords
Introduction
Chronic rhinosinusitis (CRS) is an inflammatory disorder of the nose and paranasal sinuses which persists for more than 12 weeks without resolution. 1 CRS is estimated to affect 11% of the global population1,2 and an estimated one in three patients have poorly controlled symptoms in secondary care.3,4 Existing measures of disease activity, such as nasoendoscopy, disease specific quality of life scores, or computed tomography scans1,5 can be retrospective in nature. Changes in these measures are often apparent only after disease control has regressed significantly, so better modalities to predict treatment response are needed.
Biomarkers are characteristics which can be objectively measured as an indicator of biological processes or responses to a therapeutic intervention. 6 Since CRS is a heterogeneous condition, the identification of suitable biomarkers for determining CRS endotypes and predicting treatment effectiveness will enable personalized approaches to treatment selection.7,8
CRS was traditionally classified into CRS with nasal polyps (CRSwNP) or CRS without nasal polyps (CRSsNP) however this approach does not capture the complexity of CRS phenotypes. The most recent European Position Paper on Sinusitis (EPOS2020) has set out type 2 and non-type 2 inflammation as two important endotypes of primary CRS. 1 The type 2 endotype is characterized by phenotypes such as CRSwNP and eosinophilic CRS with nasal polyps (ECRSwNP), defined as a tissue eosinophil count of ≥10 per high powered field or blood eosinophils ≥250 cells per microliter.
Biologic agents for treating asthma such as omalizumab are increasingly being used for CRSwNP, which has established a need for improved subtyping of the disease to enhance treatment efficacy. 9 Emerging evidence exists that CRS biomarkers are not mutually exclusive and can be used to determine CRS prognosis independently of phenotype. 10 Therefore, the aim of this review was to identify and summarize clinical studies of biomarkers in adults with CRS in order to inform future research into CRS endotypes.
Materials and Methods
Search Protocol and Selection Criteria
The protocol for this review was designed according to the Preferred Reporting Items for Systematic Review and Meta-Analysis guidelines 11 and was registered in PROSPERO (CRD42022302787). Systematic searches were initially performed of MEDLINE via Ovid SP and Web of Science from January 1, 2006 to October 1, 2018, and subsequently updated from inception to January 30, 2022 with no limits. The full search strategy is outlined in Supplemental file 1. Text-based synonyms and medical subject headings terms for the key search themes outlined in Table 1 were combined with Boolean operators. Additional studies were identified from the references of relevant studies and systematic reviews.
Search Strategy.
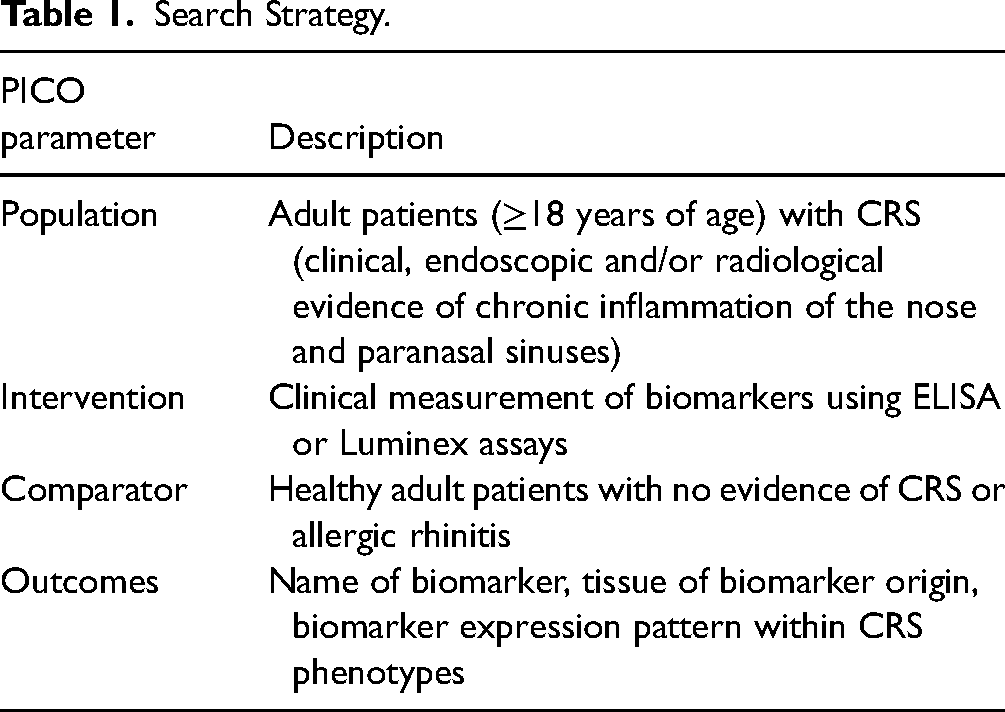
All clinical studies of adult patients (≥18 years of age) with CRS and healthy controls which reported the measurement of biomarkers through enzyme-linked immunosorbent assay (ELISA) or Luminex were included. CRS was defined as clinical, endoscopic, and/or radiological evidence of chronic inflammation of the nose and paranasal sinuses. Studies of patients with all subtypes of CRS including primary and secondary CRS were eligible for inclusion. Healthy control patients were defined as adults (≥18 years of age) with no evidence of CRS or allergic rhinitis. Eligible study designs included randomized controlled trials and observational studies (including cohort and cross-sectional studies) so that all relevant data could be considered. Animal studies, in vitro studies, reviews, editorials, letters, and conference abstracts were excluded. Studies were excluded if they used ELISA or Luminex in conjunction with other techniques and did not report separate results specifically for ELISA or Luminex alone.
Quality Assessment and Data Extraction
Two authors (SG and AE or ACP and JM) independently screened titles and abstracts to determine relevance for full text review. Two authors (SG and AE or LL and ACP) independently assessed full texts against the above inclusion and exclusion criteria and extracted data from included studies. Two authors (SG and LL) assessed the quality of included studies using The National Institutes of Health quality assessment tool for observational cohort and cross-sectional studies. 12 Disagreements at any stage were resolved by discussion with a third author (CP).
Data was extracted on study demographics, CRS phenotypes, number of patients, tissue site of analysis, name and expression pattern of individual biomarkers.
Outcomes and Data Synthesis
The primary outcome was the expression pattern of each biomarker in individual CRS phenotypes. A narrative synthesis of included studies was performed. Meta-analysis was not possible due to heterogeneity in assay techniques and study populations. Biomarkers were grouped by tissue site and CRS phenotype. Studies with conflicting results were identified separately.
Results
Characteristics of Included Studies
We identified 6152 unique records, of which 143 were selected for full text review (Figure 1). Seventy-eight studies met the inclusion criteria and were included in the final synthesis. Included studies were published between 2003 13 and 2022.14,15 Twenty-seven studies were conducted in China, 12 in South Korea, 10 in the USA, five in Belgium and Turkey, three in Japan and Germany, two in Taiwan, India, and Austria, and one in Russia, Australia, the Netherlands, Luxembourg, Switzerland, Romania, Brazil, Egypt, Lithuania, Hungary, Slovakia, and the UK (Figure 2). The number of participants per study ranged from 20 16 to 573. 17 In total, up to 9394 participants were included across all 78 studies, of which up to 5572 patients had CRSwNP (Figure 3).
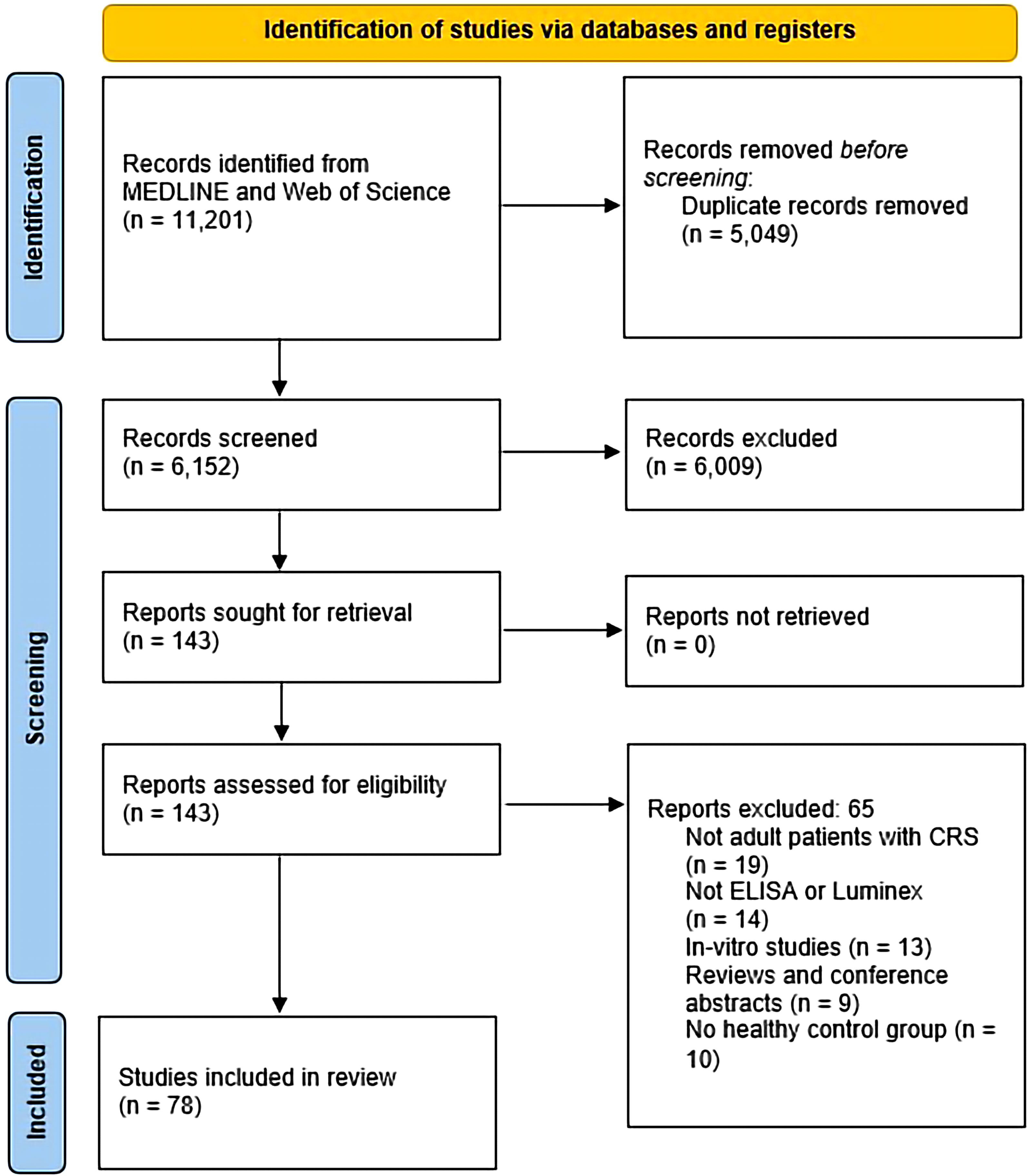
PRISMA 2020 flow diagram. 18
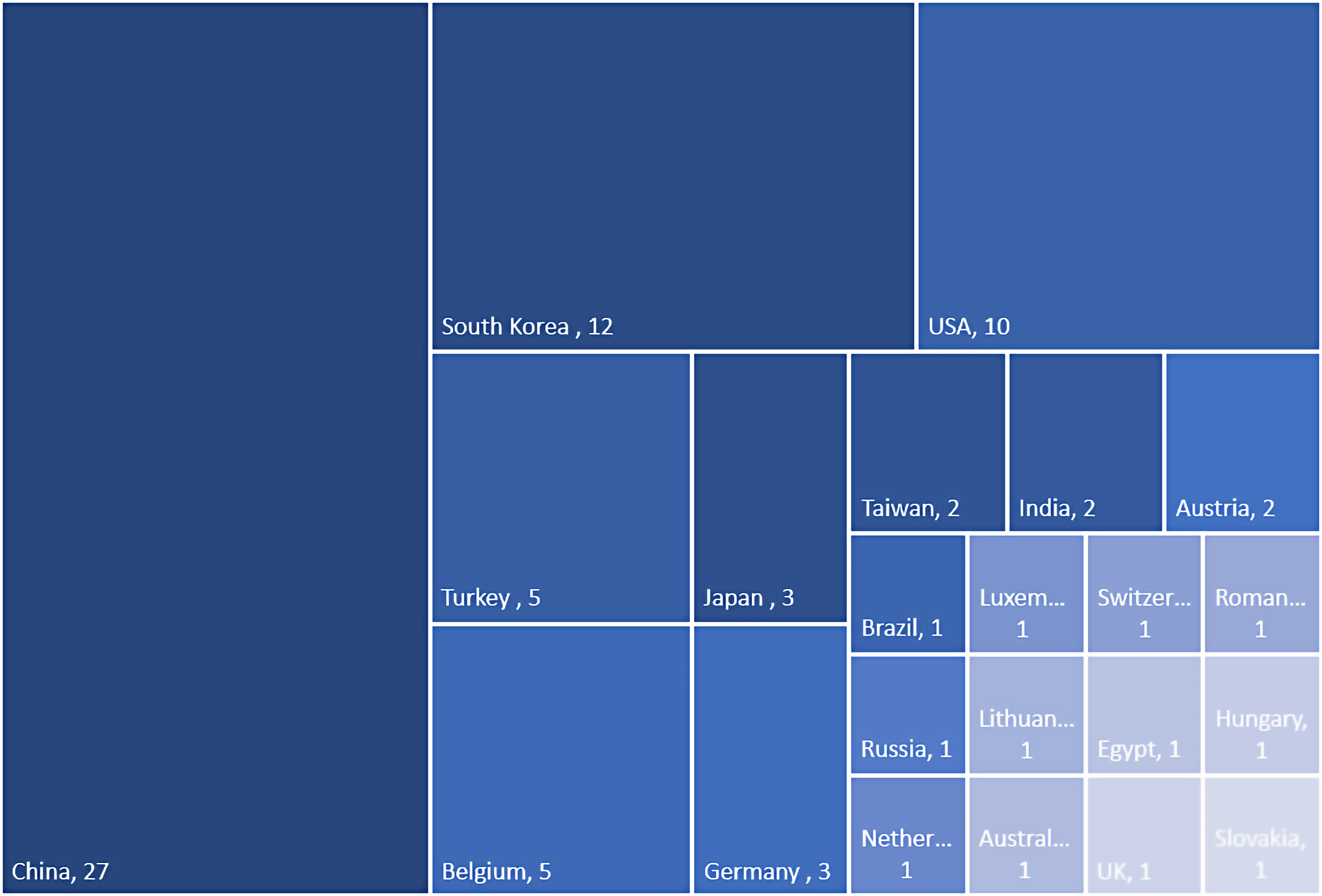

Geographical setting of included studies.
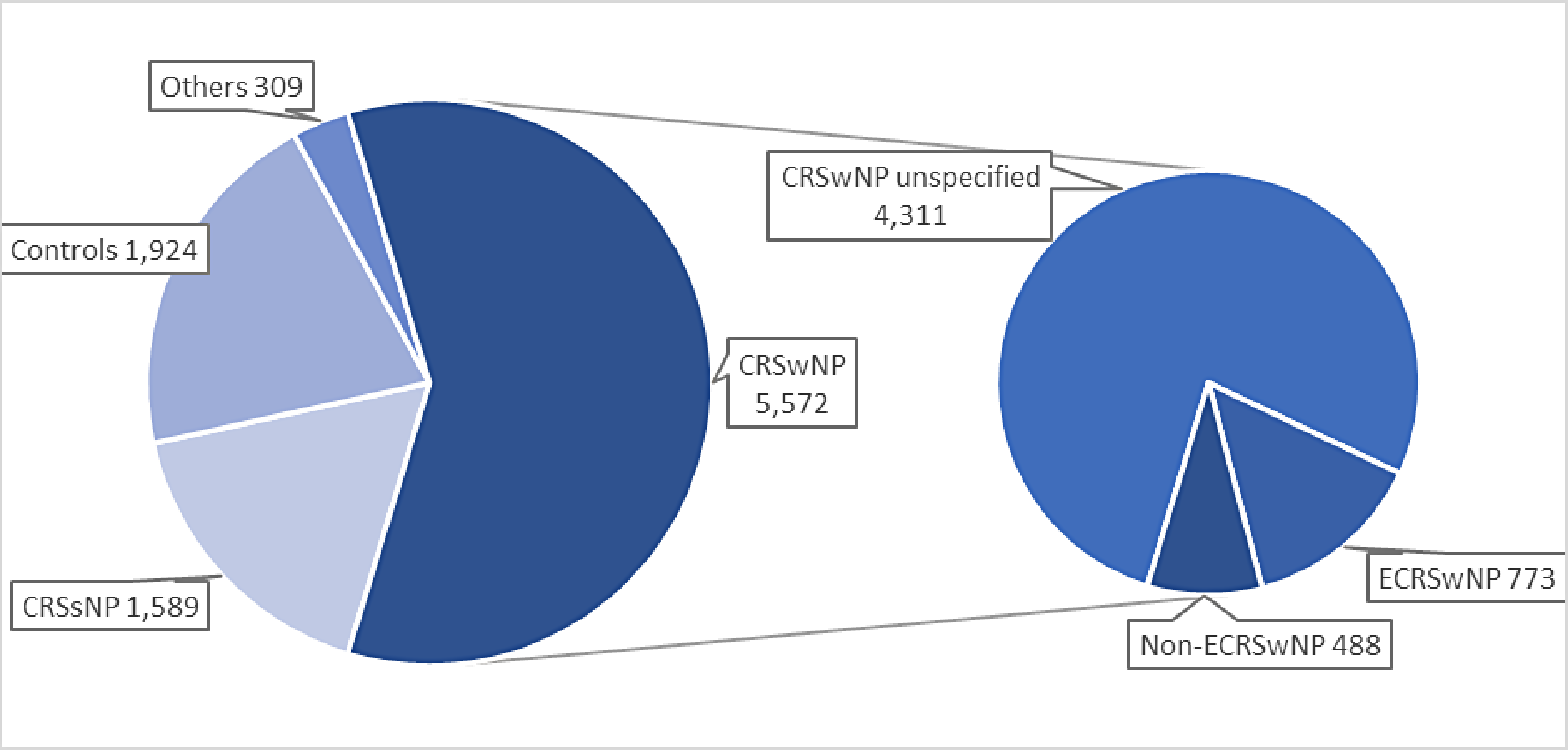
Patient phenotypes from included studies.
A summary of biomarkers by site of identification is outlined in Figure 4. The majority of studies analyzed biomarkers from nasal tissue samples, including nasal polyps (44 studies), uncinate process (24 studies), inferior turbinate (21 studies), ethmoid mucosa (10 studies), middle turbinate (five studies), osteomeatal complex mucosa (two studies), maxillary sinus mucosa (one study), and unspecified nasal tissue (six studies). Five studies analyzed nasal secretions and a further seven studies analyzed nasal lavage fluid. Nineteen studies analyzed serum and one study analyzed urine. 19 The most common phenotypic comparisons characterized by each biomarker are outlined in Table 2.
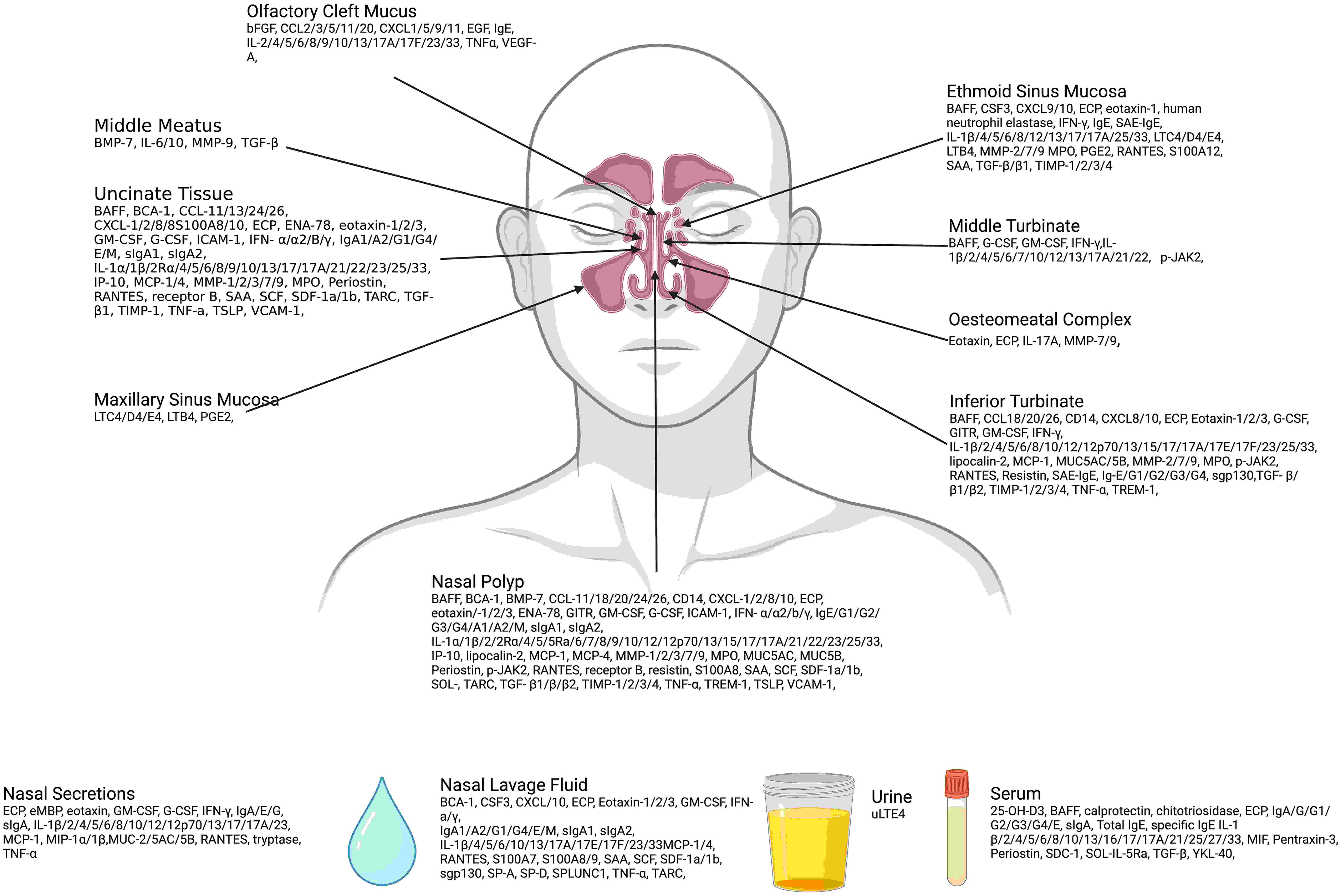
Site of identification of biomarkers. Created with BioRender.com.
Biomarkers Identified as Elevated or Decreased by CRS Phenotypic Comparison.
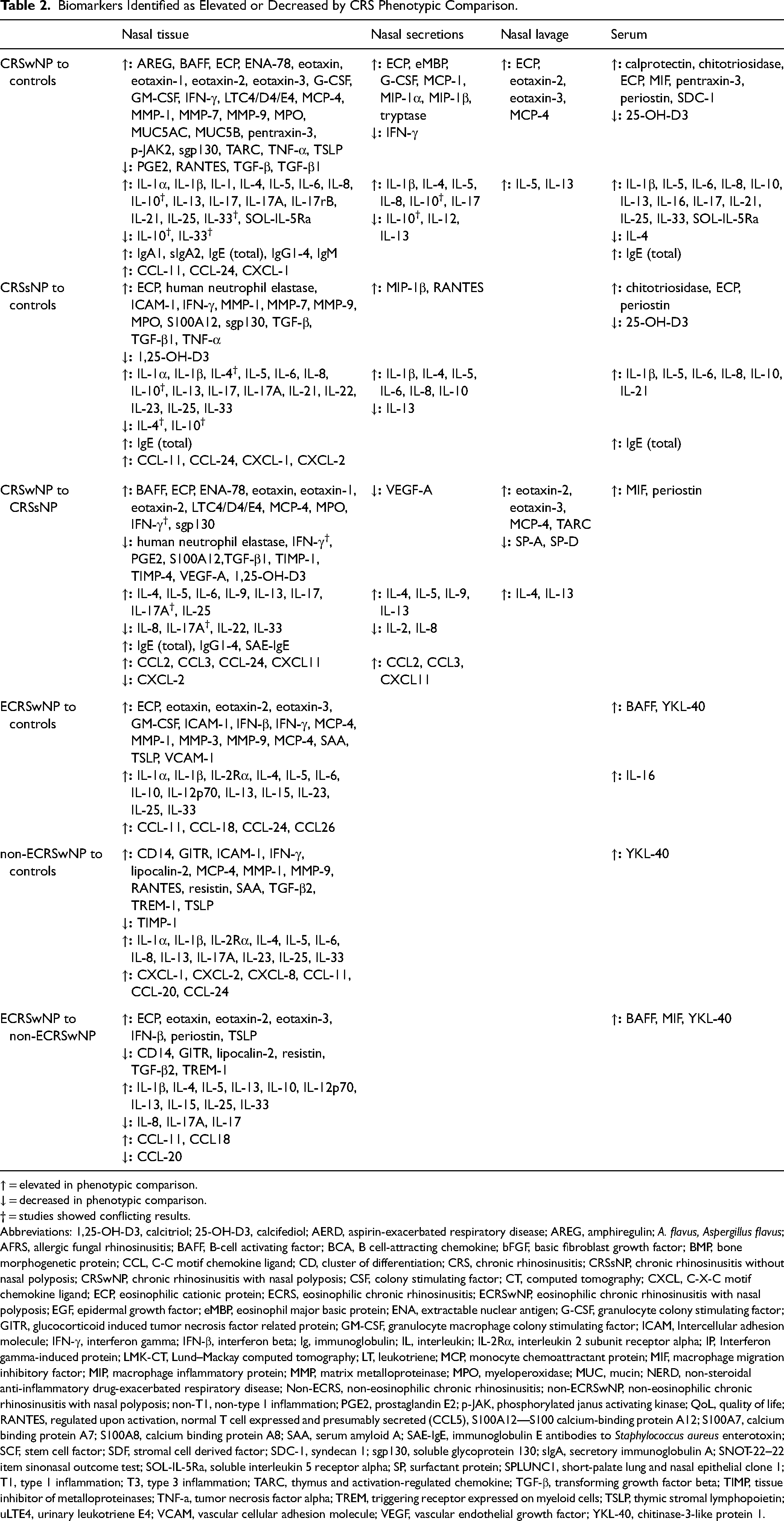
↑ = elevated in phenotypic comparison.
↓ = decreased in phenotypic comparison.
† = studies showed conflicting results.
Abbreviations: 1,25-OH-D3, calcitriol; 25-OH-D3, calcifediol; AERD, aspirin-exacerbated respiratory disease; AREG, amphiregulin; A. flavus, Aspergillus flavus; AFRS, allergic fungal rhinosinusitis; BAFF, B-cell activating factor; BCA, B cell-attracting chemokine; bFGF, basic fibroblast growth factor; BMP, bone morphogenetic protein; CCL, C-C motif chemokine ligand; CD, cluster of differentiation; CRS, chronic rhinosinusitis; CRSsNP, chronic rhinosinusitis without nasal polyposis; CRSwNP, chronic rhinosinusitis with nasal polyposis; CSF, colony stimulating factor; CT, computed tomography; CXCL, C-X-C motif chemokine ligand; ECP, eosinophilic cationic protein; ECRS, eosinophilic chronic rhinosinusitis; ECRSwNP, eosinophilic chronic rhinosinusitis with nasal polyposis; EGF, epidermal growth factor; eMBP, eosinophil major basic protein; ENA, extractable nuclear antigen; G-CSF, granulocyte colony stimulating factor; GITR, glucocorticoid induced tumor necrosis factor related protein; GM-CSF, granulocyte macrophage colony stimulating factor; ICAM, Intercellular adhesion molecule; IFN-γ, interferon gamma; IFN-β, interferon beta; Ig, immunoglobulin; IL, interleukin; IL-2Rα, interleukin 2 subunit receptor alpha; IP, Interferon gamma-induced protein; LMK-CT, Lund–Mackay computed tomography; LT, leukotriene; MCP, monocyte chemoattractant protein; MIF, macrophage migration inhibitory factor; MIP, macrophage inflammatory protein; MMP, matrix metalloproteinase; MPO, myeloperoxidase; MUC, mucin; NERD, non-steroidal anti-inflammatory drug-exacerbated respiratory disease; Non-ECRS, non-eosinophilic chronic rhinosinusitis; non-ECRSwNP, non-eosinophilic chronic rhinosinusitis with nasal polyposis; non-T1, non-type 1 inflammation; PGE2, prostaglandin E2; p-JAK, phosphorylated janus activating kinase; QoL, quality of life; RANTES, regulated upon activation, normal T cell expressed and presumably secreted (CCL5), S100A12—S100 calcium-binding protein A12; S100A7, calcium binding protein A7; S100A8, calcium binding protein A8; SAA, serum amyloid A; SAE-IgE, immunoglobulin E antibodies to Staphylococcus aureus enterotoxin; SCF, stem cell factor; SDF, stromal cell derived factor; SDC-1, syndecan 1; sgp130, soluble glycoprotein 130; sIgA, secretory immunoglobulin A; SNOT-22–22 item sinonasal outcome test; SOL-IL-5Ra, soluble interleukin 5 receptor alpha; SP, surfactant protein; SPLUNC1, short-palate lung and nasal epithelial clone 1; T1, type 1 inflammation; T3, type 3 inflammation; TARC, thymus and activation-regulated chemokine; TGF-β, transforming growth factor beta; TIMP, tissue inhibitor of metalloproteinases; TNF-a, tumor necrosis factor alpha; TREM, triggering receptor expressed on myeloid cells; TSLP, thymic stromal lymphopoietin; uLTE4, urinary leukotriene E4; VCAM, vascular cellular adhesion molecule; VEGF, vascular endothelial growth factor; YKL-40, chitinase-3-like protein 1.
Quality Assessment
The quality assessment scores for individual studies are outlined in Supplemental file 2. Twenty-five studies received an overall quality rating of “good,” 43 as “fair,” and 10 as “poor.” Most studies had clearly defined outcome measures but only eight studies commented on blinding of participants and only two provided a sample size justification. All included studies were of cross-sectional or cohort study design.
Serum Biomarkers
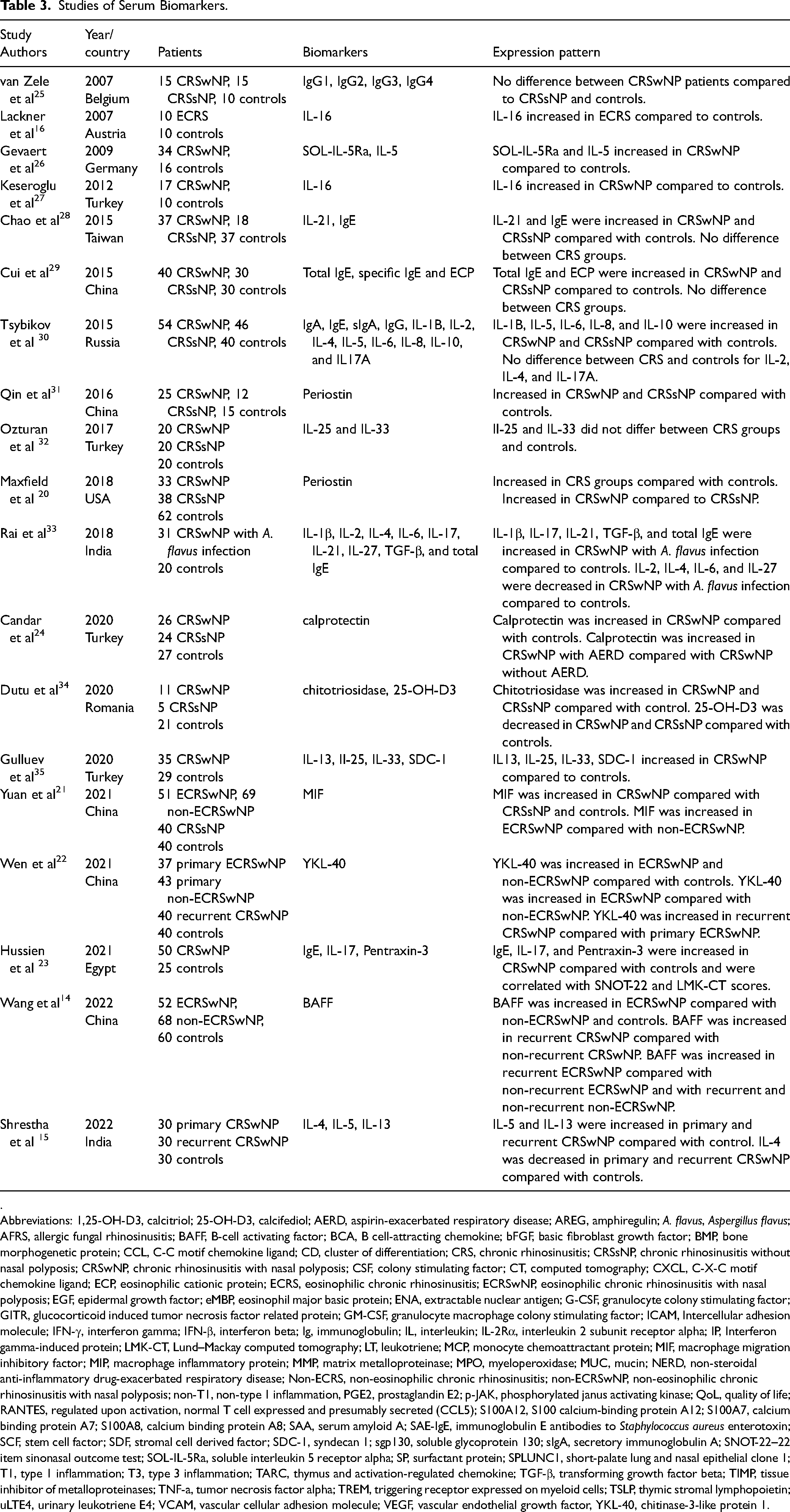
Twenty-four biomarkers were identified in serum samples of CRS patients from 19 studies (Table 3). The majority were found to distinguish patients with CRSwNP or CRSsNP from controls, and only four biomarkers were found to differentiate CRS phenotypes. Macrophage migration inhibitory factor and periostin were elevated in CRSwNP compared to CRSsNP.20,21 Chitinase-3-like protein 1 (YKL-40) and B-cell activating factor (BAFF) were found to distinguish patients with ECRSwNP and recurrent CRSwNP.14,22
Studies of Serum Biomarkers.
.
Abbreviations: 1,25-OH-D3, calcitriol; 25-OH-D3, calcifediol; AERD, aspirin-exacerbated respiratory disease; AREG, amphiregulin; A. flavus, Aspergillus flavus; AFRS, allergic fungal rhinosinusitis; BAFF, B-cell activating factor; BCA, B cell-attracting chemokine; bFGF, basic fibroblast growth factor; BMP, bone morphogenetic protein; CCL, C-C motif chemokine ligand; CD, cluster of differentiation; CRS, chronic rhinosinusitis; CRSsNP, chronic rhinosinusitis without nasal polyposis; CRSwNP, chronic rhinosinusitis with nasal polyposis; CSF, colony stimulating factor; CT, computed tomography; CXCL, C-X-C motif chemokine ligand; ECP, eosinophilic cationic protein; ECRS, eosinophilic chronic rhinosinusitis; ECRSwNP, eosinophilic chronic rhinosinusitis with nasal polyposis; EGF, epidermal growth factor; eMBP, eosinophil major basic protein; ENA, extractable nuclear antigen; G-CSF, granulocyte colony stimulating factor; GITR, glucocorticoid induced tumor necrosis factor related protein; GM-CSF, granulocyte macrophage colony stimulating factor; ICAM, Intercellular adhesion molecule; IFN-γ, interferon gamma; IFN-β, interferon beta; Ig, immunoglobulin; IL, interleukin; IL-2Rα, interleukin 2 subunit receptor alpha; IP, Interferon gamma-induced protein; LMK-CT, Lund–Mackay computed tomography; LT, leukotriene; MCP, monocyte chemoattractant protein; MIF, macrophage migration inhibitory factor; MIP, macrophage inflammatory protein; MMP, matrix metalloproteinase; MPO, myeloperoxidase; MUC, mucin; NERD, non-steroidal anti-inflammatory drug-exacerbated respiratory disease; Non-ECRS, non-eosinophilic chronic rhinosinusitis; non-ECRSwNP, non-eosinophilic chronic rhinosinusitis with nasal polyposis; non-T1, non-type 1 inflammation, PGE2, prostaglandin E2; p-JAK, phosphorylated janus activating kinase; QoL, quality of life; RANTES, regulated upon activation, normal T cell expressed and presumably secreted (CCL5); S100A12, S100 calcium-binding protein A12; S100A7, calcium binding protein A7; S100A8, calcium binding protein A8; SAA, serum amyloid A; SAE-IgE, immunoglobulin E antibodies to Staphylococcus aureus enterotoxin; SCF, stem cell factor; SDF, stromal cell derived factor; SDC-1, syndecan 1; sgp130, soluble glycoprotein 130; sIgA, secretory immunoglobulin A; SNOT-22–22 item sinonasal outcome test; SOL-IL-5Ra, soluble interleukin 5 receptor alpha; SP, surfactant protein; SPLUNC1, short-palate lung and nasal epithelial clone 1; T1, type 1 inflammation; T3, type 3 inflammation; TARC, thymus and activation-regulated chemokine; TGF-β, transforming growth factor beta; TIMP, tissue inhibitor of metalloproteinases; TNF-a, tumor necrosis factor alpha; TREM, triggering receptor expressed on myeloid cells; TSLP, thymic stromal lymphopoietin; uLTE4, urinary leukotriene E4; VCAM, vascular cellular adhesion molecule; VEGF, vascular endothelial growth factor, YKL-40, chitinase-3-like protein 1.
Studies of serum biomarkers for CRS disease activity were limited. However, a study of 50 patients with CRSwNP by Hussein et al found that serum immunoglobulin E (IgE), interleukin-17 (IL-17), and pentraxin-3 were correlated with SNOT-22 and LMK-CT scores. 23 Study quality was fair due to the use of age and sex matched controls but no specific CRS inclusion criteria. Candar et al studied 26 patients with CRSwNP and found that serum calprotectin was elevated in those with aspirin-exacerbated respiratory disease (AERD) compared to those without AERD. 24 Study quality was limited by a small sample size of six AERD patients.
Nasal Secretion Biomarkers
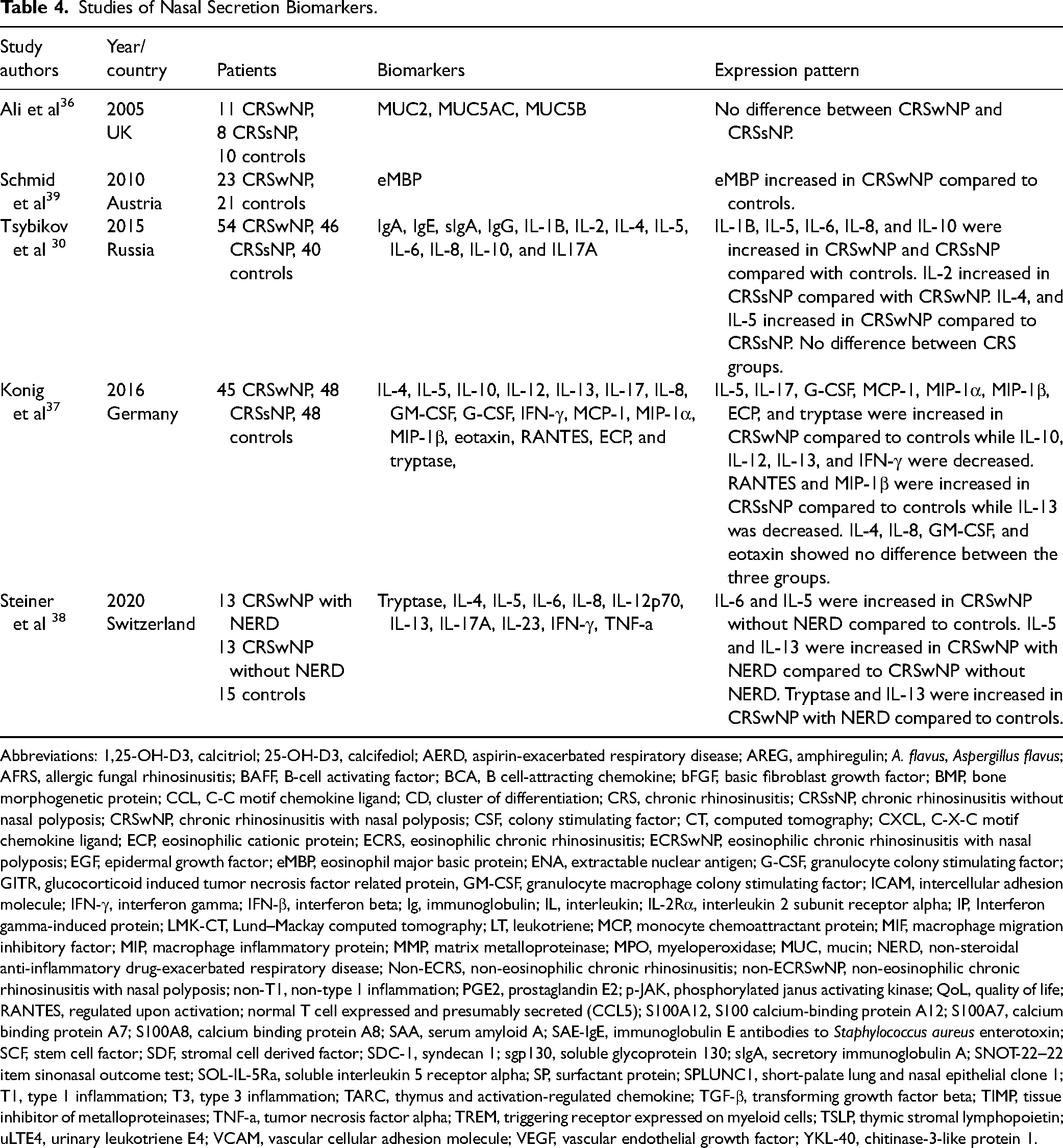
Twenty-five biomarkers were identified in nasal secretions across five studies (Table 4). Nasal secretion harvesting techniques differed in each study. Secretions were sampled using cotton swabs applied to the inferior turbinate, 36 cotton swabs with phosphate-buffered solution in the anterior nares, 30 absorbent cotton wool left for 20 min in the middle meatus, 37 neurosurgical patties left above the inferior turbinate for 10 min, 38 or using a “Sinus Secretion Collector” in the middle meatus. 39
Studies of Nasal Secretion Biomarkers.
Abbreviations: 1,25-OH-D3, calcitriol; 25-OH-D3, calcifediol; AERD, aspirin-exacerbated respiratory disease; AREG, amphiregulin; A. flavus, Aspergillus flavus; AFRS, allergic fungal rhinosinusitis; BAFF, B-cell activating factor; BCA, B cell-attracting chemokine; bFGF, basic fibroblast growth factor; BMP, bone morphogenetic protein; CCL, C-C motif chemokine ligand; CD, cluster of differentiation; CRS, chronic rhinosinusitis; CRSsNP, chronic rhinosinusitis without nasal polyposis; CRSwNP, chronic rhinosinusitis with nasal polyposis; CSF, colony stimulating factor; CT, computed tomography; CXCL, C-X-C motif chemokine ligand; ECP, eosinophilic cationic protein; ECRS, eosinophilic chronic rhinosinusitis; ECRSwNP, eosinophilic chronic rhinosinusitis with nasal polyposis; EGF, epidermal growth factor; eMBP, eosinophil major basic protein; ENA, extractable nuclear antigen; G-CSF, granulocyte colony stimulating factor; GITR, glucocorticoid induced tumor necrosis factor related protein, GM-CSF, granulocyte macrophage colony stimulating factor; ICAM, intercellular adhesion molecule; IFN-γ, interferon gamma; IFN-β, interferon beta; Ig, immunoglobulin; IL, interleukin; IL-2Rα, interleukin 2 subunit receptor alpha; IP, Interferon gamma-induced protein; LMK-CT, Lund–Mackay computed tomography; LT, leukotriene; MCP, monocyte chemoattractant protein; MIF, macrophage migration inhibitory factor; MIP, macrophage inflammatory protein; MMP, matrix metalloproteinase; MPO, myeloperoxidase; MUC, mucin; NERD, non-steroidal anti-inflammatory drug-exacerbated respiratory disease; Non-ECRS, non-eosinophilic chronic rhinosinusitis; non-ECRSwNP, non-eosinophilic chronic rhinosinusitis with nasal polyposis; non-T1, non-type 1 inflammation; PGE2, prostaglandin E2; p-JAK, phosphorylated janus activating kinase; QoL, quality of life; RANTES, regulated upon activation; normal T cell expressed and presumably secreted (CCL5); S100A12, S100 calcium-binding protein A12; S100A7, calcium binding protein A7; S100A8, calcium binding protein A8; SAA, serum amyloid A; SAE-IgE, immunoglobulin E antibodies to Staphylococcus aureus enterotoxin; SCF, stem cell factor; SDF, stromal cell derived factor; SDC-1, syndecan 1; sgp130, soluble glycoprotein 130; sIgA, secretory immunoglobulin A; SNOT-22–22 item sinonasal outcome test; SOL-IL-5Ra, soluble interleukin 5 receptor alpha; SP, surfactant protein; SPLUNC1, short-palate lung and nasal epithelial clone 1; T1, type 1 inflammation; T3, type 3 inflammation; TARC, thymus and activation-regulated chemokine; TGF-β, transforming growth factor beta; TIMP, tissue inhibitor of metalloproteinases; TNF-a, tumor necrosis factor alpha; TREM, triggering receptor expressed on myeloid cells; TSLP, thymic stromal lymphopoietin; uLTE4, urinary leukotriene E4; VCAM, vascular cellular adhesion molecule; VEGF, vascular endothelial growth factor; YKL-40, chitinase-3-like protein 1.
Biomarkers found to differentiate CRS phenotypes included IL-4, IL-5, IL-9, IL-13, C-C motif chemokine ligand 2 (CCL2), CCL3, C-X-C motif chemokine ligand 11 (CXCL11), and IgE which were increased in CRSwNP compared to CRSsNP, while the reverse was true for IL-2, IL-8, and vascular endothelial growth factor A (VEGF-A). Immunoglobulins were not associated with CRS phenotypes in nasal secretions.
Steiner et al studied 13 patients with CRSwNP and non-steroidal anti-inflammatory drug-exacerbated respiratory disease (NERD) and 13 patients with CRSwNP without NERD. They found that the former group had increased levels of IL-5 and IL-13. 38 Study quality was limited by a small sample size of 13 NERD patients and no specific CRSwNP inclusion criteria.
Nasal Lavage Fluid Biomarkers
Fourteen biomarkers were identified in nasal lavage fluid across seven studies (Table 5). IL-4, IL-13, eotaxin-2, eotaxin-3, monocyte chemoattractant protein-4, and thymus and activation-regulated chemokine were increased in CRSwNP compared to CRSsNP. No studies of CRSwNP subtypes were performed and immunoglobulins were not found to distinguish CRS phenotypes. The largest study was by Klingler et al of 126 patients with CRSsNP which identified that CXCL9 and CXCL10 were elevated in patients characterized by the T1 endotype compared to non-T1 CRSsNP and controls. 40 Study quality was fair with detailed CRS inclusion criteria but nasal lavage fluid samples were highly diluted.
Studies of Nasal Lavage Biomarkers.

Abbreviations: 1,25-OH-D3, calcitriol, 25-OH-D3; calcifediol; AERD, aspirin-exacerbated respiratory disease; AREG, amphiregulin; A. flavus, Aspergillus flavus; AFRS, allergic fungal rhinosinusitis; BAFF, B-cell activating factor; BCA, B cell-attracting chemokine; bFGF, basic fibroblast growth factor; BMP, bone morphogenetic protein; CCL, C-C motif chemokine ligand; CD, cluster of differentiation; CRS, chronic rhinosinusitis; CRSsNP, chronic rhinosinusitis without nasal polyposis; CRSwNP, chronic rhinosinusitis with nasal polyposis; CSF, colony stimulating factor; CT, computed tomography; CXCL, C-X-C motif chemokine ligand; ECP, eosinophilic cationic protein; ECRS, eosinophilic chronic rhinosinusitis; ECRSwNP, eosinophilic chronic rhinosinusitis with nasal polyposis; EGF, epidermal growth factor; eMBP, eosinophil major basic protein; ENA, extractable nuclear antigen; G-CSF, granulocyte colony stimulating factor; GITR, glucocorticoid induced tumor necrosis factor related protein; GM-CSF, granulocyte macrophage colony stimulating factor; ICAM, Intercellular adhesion molecule; IFN-γ, interferon gamma; IFN-β, interferon beta; Ig, immunoglobulin; IL, interleukin; IL-2Rα, interleukin 2 subunit receptor alpha; IP, Interferon gamma-induced protein; LMK-CT, Lund–Mackay computed tomography; LT, leukotriene; MCP, monocyte chemoattractant protein; MIF, macrophage migration inhibitory factor; MIP, macrophage inflammatory protein; MMP, matrix metalloproteinase; MPO, myeloperoxidase; MUC, mucin; NERD, non-steroidal anti-inflammatory drug-exacerbated respiratory disease; Non-ECRS, non-eosinophilic chronic rhinosinusitis; non-ECRSwNP, non-eosinophilic chronic rhinosinusitis with nasal polyposis; non-T1, non-type 1 inflammation; PGE2, prostaglandin E2; p-JAK, phosphorylated janus activating kinase; QoL, quality of life; RANTES, regulated upon activation, normal T cell expressed and presumably secreted (CCL5); S100A12, S100 calcium-binding protein A12; S100A7, calcium binding protein A7; S100A8, calcium binding protein A8; SAA, serum amyloid A; SAE-IgE, immunoglobulin E antibodies to Staphylococcus aureus enterotoxin; SCF, stem cell factor; SDF, stromal cell derived factor; SDC-1, syndecan 1; sgp130, soluble glycoprotein 130; sIgA, secretory immunoglobulin A; SNOT-22, 22-item sinonasal outcome test; SOL-IL-5Ra, soluble interleukin 5 receptor alpha; SP, surfactant protein; SPLUNC1, short-palate lung and nasal epithelial clone 1; T1, type 1 inflammation; T3, type 3 inflammation; TARC, thymus and activation-regulated chemokine; TGF-β, transforming growth factor beta; TIMP, tissue inhibitor of metalloproteinases; TNF-a, tumor necrosis factor alpha; TREM, triggering receptor expressed on myeloid cells; TSLP, thymic stromal lymphopoietin; uLTE4, urinary leukotriene E4; VCAM, vascular cellular adhesion molecule; VEGF, vascular endothelial growth factor; YKL-40, chitinase-3-like protein 1.
Nasal Tissue Biomarkers
Eighty biomarkers across 55 studies (Table 6) were identified in nasal tissue, including 57 differentiating CRSwNP from controls, 35 differentiating CRSsNP from controls, 41 differentiating CRSwNP from CRSsNP, 30 differentiating ECRSwNP from non-ECRSwNP, and 12 differentiating refractory CRSwNP from controls or primary CRSwNP. The most widely measured biomarker among included studies was IL-5, followed by IL-13 and IL-4.
Studies of Nasal Tissue Biomarkers.

Abbreviations: 1,25-OH-D3, calcitriol; 25-OH-D3, calcifediol; AERD, aspirin-exacerbated respiratory disease; AREG, amphiregulin; A. flavus, Aspergillus flavus; AFRS, allergic fungal rhinosinusitis; BAFF, B-cell activating factor; BCA, B cell-attracting chemokine; bFGF, basic fibroblast growth factor; BMP, bone morphogenetic protein; CCL, C-C motif chemokine ligand; CD, cluster of differentiation; CRS, chronic rhinosinusitis; CRSsNP, chronic rhinosinusitis without nasal polyposis; CRSwNP, chronic rhinosinusitis with nasal polyposis; CSF, colony stimulating factor; CT, computed tomography; CXCL, C-X-C motif chemokine ligand; ECP, eosinophilic cationic protein; ECRS, eosinophilic chronic rhinosinusitis; ECRSwNP, eosinophilic chronic rhinosinusitis with nasal polyposis; EGF, epidermal growth factor; eMBP, eosinophil major basic protein; ENA, extractable nuclear antigen; G-CSF, granulocyte colony stimulating factor; GITR, glucocorticoid induced tumor necrosis factor related protein; GM-CSF, granulocyte macrophage colony stimulating factor; ICAM, Intercellular adhesion molecule; IFN-γ, interferon gamma; IFN-β, interferon beta; Ig, immunoglobulin; IL, interleukin; IL-2Rα, interleukin 2 subunit receptor alpha; IP, Interferon gamma-induced protein; LMK-CT, Lund–Mackay computed tomography; LT, leukotriene; MCP, monocyte chemoattractant protein; MIF, macrophage migration inhibitory factor; MIP, macrophage inflammatory protein; MMP, matrix metalloproteinase; MPO, myeloperoxidase; MUC, mucin; NERD, non-steroidal anti-inflammatory drug-exacerbated respiratory disease; Non-ECRS, non-eosinophilic chronic rhinosinusitis; non-ECRSwNP, non-eosinophilic chronic rhinosinusitis with nasal polyposis; non-T1, non-type 1 inflammation; PGE2, prostaglandin E2; p-JAK, phosphorylated janus activating kinase; QoL, quality of life; RANTES, regulated upon activation, normal T cell expressed and presumably secreted (CCL5); S100A12, S100 calcium-binding protein A12; S100A7, calcium binding protein A7; S100A8, calcium binding protein A8; SAA, serum amyloid A; SAE-IgE, immunoglobulin E antibodies to Staphylococcus aureus enterotoxin; SCF, stem cell factor; SDF, stromal cell derived factor; SDC-1, syndecan 1; sgp130 , soluble glycoprotein 130; sIgA , secretory immunoglobulin A; SNOT-22, 22-item sinonasal outcome test; SOL-IL-5Ra, soluble interleukin 5 receptor alpha; SP, surfactant protein; SPLUNC1, short-palate lung and nasal epithelial clone 1; T1, type 1 inflammation; T3, type 3 inflammation; TARC, thymus and activation-regulated chemokine; TGF-β, transforming growth factor beta; TIMP, tissue inhibitor of metalloproteinases; TNF-a, tumor necrosis factor alpha; TREM, triggering receptor expressed on myeloid cells; TSLP, thymic stromal lymphopoietin; uLTE4, urinary leukotriene E4; VCAM, vascular cellular adhesion molecule; VEGF, vascular endothelial growth factor; YKL-40, chitinase-3-like protein 1.
Control tissue was most often harvested from the uncinate process, although one study used olfactory cleft mucosa. 47 Biomarker expression was greatest in nasal polyp tissue in most cases when compared to other tissue sites, especially for IL-25 and eotaxin-1. 48
While Liu et al 49 and Stevens et al 44 found IL-10 to be increased in patients with CRSwNP compared to controls, Lucas et al found IL-10 was higher in controls. 50 Similarly, Kim et al 51 and Lin et al 52 found IL-33 to be increased in patients with CRSwNP compared to controls, while Ozturan et al found the opposite. 32 When comparing CRSwNP patients with CRSsNP, IL-17A and interferon gamma (IFN-γ) were found to be increased by Chen et al, 53 but the reverse was found by Kim et al. 54
A study of 309 patients by Ryu et al found that IL-5, CCL-11, CCL-24, IFN-γ, and periostin increased with ageing in CRS phenotypes when compared with controls, while IL-17A, CXCL-8, and IL-6 decreased with ageing. 55 Study quality was strengthened by a large sample size and adjustment of outcomes for confounding factors such as atopy status, smoking history, and disease duration. Similarly, Kim et al, in a study of 70 patients with CRSwNP, found IL-17A and IL-23 to be negatively correlated with age. 56 Wang et al conducted a multi-centre study of 573 patients across 6 countries in Europe, Asia, and Australia and found geographical variation in biomarkers. 17 IL-17 was increased in CRSwNP compared to CRSsNP and controls in Adelaide and Beijing, but increased in CRSsNP compared to CRSwNP and controls in Tochigi. IFN-γ was increased in CRSsNP compared to CRSwNP and controls in Beijing but did not differ in other regions. IL-8 was increased in CRSwNP compared to controls in all regions. Study quality was good due to the multi-center design, clear inclusion criteria, and standardized sampling procedures.
Correlation to Disease Severity
Lin et al studied 61 patients with CRSwNP and found that phospho-janus kinase 2 (p-JAK2), IL-5, IL-6, and granulocyte-colony stimulating factor (G-CSF) were correlated with LMK-CT and SNOT-22 scores. 57 Patients were categorized into “mild” and “severe” CRS based on Lund–Kennedy score alone, and data on patients with CRSsNP was limited. Similarly, Pulshiper et al studied 70 patients and found that levels of S100 calcium-binding protein A12 (S100A12) were increased in CRSsNP compared to CRSwNP and were correlated to LMK-CT scores but not to the rhinosinusitis disability index, a quality of life score. 58 Clear CRS inclusion criteria were not specified and healthy controls included patients with nasal obstruction with associated quality of life implications. Kim et al stratified 69 patients into mild, moderate, and severe ECRSwNP using the Japanese Epidemiological Survey of Refractory Eosinophilic Chronic Rhinosinusitis score, which considers clinical examination findings, CT results and eosinophil counts. 59 They found that IL-17A and IFN-γ were decreased in severe ECRSwNP compared with mild ECRSwNP. Inclusion criteria were clear and each subgroup had a minimum of 13 patients, but subgroups were not matched for sex or asthma status.
Urine Biomarkers
One urine biomarker was identified in a USA study of 115 patients with unspecified CRS and 38 controls by Santarelli et al. 19 Urinary leukotriene E4 (uLTE4) was increased in patients with CRS compared to controls. Elevated uLTE4 levels were correlated with the presence of comorbid asthma but not with atopy. Subgroup analyses for other CRS phenotypes were not performed.
Discussion
This review summarizes the expression pattern among CRS phenotypes of 143 biomarkers identified from studies of nasal tissue, nasal secretions, nasal lavage fluid, serum, or urine. Biomarker profiles are presented to distinguish patients with CRSwNP, CRSsNP, ECRSwNP, non-ECRSwNP, refractory CRS, and primary CRS from each other or from controls. Biomarkers such as serum IgE, IL-17, and pentraxin-3 or nasal p-JAK2, IL-5, IL-6, IL-17A, G-CSF, and IFN-γ are identified as correlated with disease severity. Studies by Ryu et al, 55 Kim et al, 82 and Wang et al 17 identify biomarker profiles to vary by patient age, site of nasal tissue sample, and geographical patient group, respectively.
Previous systematic reviews have concentrated on specific biomarkers alone, such as periostin 92 or matrix metalloproteinases 93 while this review integrates findings from all available biomarkers. Additionally, this review is not limited to one specific CRS phenotype but compares data across multiple phenotypes. Limitations include the lack of studies from developing economies, which could reduce the generalizability of results to these settings. Additionally, patients from South America and Africa were not well represented among the included studies. However, studies are presented from 22 countries across six continents, and the majority of studies were conducted in countries where English is not the national language. This review also only considered studies which used ELISA or Luminex for the analysis of samples. These methods are relatively easy to perform and cost-effective, but as a result studies using polymerase chain reaction assays, aptamer-based analyses or other techniques were not included. However, biomarkers which are not detectable across a range of modalities are less likely to translate into clinical practice. Finally, we did not choose to specify the use of a single CRS diagnostic criterion for inclusion of a study within our review. This is because the studies dated back to 2006, prior to the publication of more recent CRS diagnostic criteria, and we believe that using any one set of criteria may limit the geographical variability of included studies.
Studies from this review suggest that nasal tissue has the largest body of evidence for biomarker analysis. Most biomarkers found to distinguish CRS phenotypes such as ECRS and refractory CRS were found in nasal tissue, specifically nasal polyp. This has the advantage of being a consistent site of collection and can be harvested using local anaesthetic in some cases.
On the other hand, serum has the advantages of ease of collection and reproducible sampling. However, the range of serum biomarkers used to distinguish CRS phenotypes from each other was limited in this study. Serum biomarkers appear to have more of a role in differentiating CRS patients from controls, so further investigation into the role of serum biomarkers in disease identification could be considered.
Studies of nasal lavage fluid and nasal secretions were limited, and were hampered by inconsistencies in collection methods including site of sampling, amount of mucus sampled, and collection devices. This is significant as the proteome varies considerably throughout the nose. In particular, the role of immunoglobulins and eotaxin was limited in nasal secretions when compared to nasal tissue, and phenotypes such as ECRSwNP were not categorized within studies. Studies of urine and sputum were limited and warrant further investigation given the relative ease of collection of these samples, although biomarker profiles are likely to be less apparent when compared to nasal tissue.
Increasing evidence shows that earlier diagnosis and characterization of CRS is linked to better outcomes. 94 Biomarkers are indicative of pathways which are important targets for biologic therapies currently under investigation for treating CRS, such as mepolizumab and omalizumab. Identifying a biomarker which can predict treatment response to these expensive therapies will be crucial to their uptake into clinical practice. 1 Currently, specific protein biomarkers are not widely used in clinical practice and so further validation studies are required.
This review has identified multiple knowledge gaps in CRS biomarker research, such as the use of nasal lavage fluid and nasal secretions in distinguishing patients with ECRSwNP, the value of understudied nasal tissue collection sites such as olfactory cleft and maxillary sinus mucosa, and the prognostic role of nasal tissue biomarkers such as immunoglobulins. Specific biomarker targets for further research are also identified in those cytokines found to be associated with disease severity, such as p-JAK2, IL-5, IL-6, IL-17A, G-CSF, and IFN-γ. Additionally, future research should focus on exploring the conflicting results seen for nasal tissue biomarkers such as IFN-γ, IL-4, IL-10, IL-17A, and IL-33. This may have in part been due to geographical variation between studies or the small sample sizes of some included studies.
Future CRS biomarker studies can avoid the weaknesses of some of the studies in this review by considering measuring biomarkers at multiple time points and across multiple geographic regions, utilizing standardized outcome measures such as those proposed in EPOS2020, 1 limiting the use of underpowered samples for biomarkers known to have reduced variation among sampling sites, specifying CRS inclusion criteria, and ensuring consistent reporting of the amount of sample harvested between patients.
Rather than relying on a single biomarker in isolation, CRS endotypes can be categorized by patient clusters with specific biomarker profiles. 95 Future research should focus on exploring the interplay between biomarkers described in this review through prospective studies to identify correlations with treatment response. With monoclonal antibodies for CRS gaining increasing evidence of efficacy 96 better means of identifying patients that should receive these expensive drugs is of paramount importance. Ultimately, a combination of several biomarkers will likely be the most promising approach in understanding the immunological mechanisms underlying the different phenotypes of CRS. Further studies of existing biomarkers should aim to bring patients closer to a personalized approach to CRS treatment.
Supplemental Material
sj-docx-1-ajr-10.1177_19458924231190568 - Supplemental material for Systematic Review of Protein Biomarkers in Adult Patients With Chronic Rhinosinusitis
Supplemental material, sj-docx-1-ajr-10.1177_19458924231190568 for Systematic Review of Protein Biomarkers in Adult Patients With Chronic Rhinosinusitis by Shyam A. Gokani, Andreas Espehana and Ana C. Pratas, Louis Luke, Ekta Sharma, Jennifer Mattock, Jelena Gavrilovic, Allan Clark, Tom Wileman, Carl M. Philpott in American Journal of Rhinology & Allergy
Supplemental Material
sj-docx-2-ajr-10.1177_19458924231190568 - Supplemental material for Systematic Review of Protein Biomarkers in Adult Patients With Chronic Rhinosinusitis
Supplemental material, sj-docx-2-ajr-10.1177_19458924231190568 for Systematic Review of Protein Biomarkers in Adult Patients With Chronic Rhinosinusitis by Shyam A. Gokani, Andreas Espehana and Ana C. Pratas, Louis Luke, Ekta Sharma, Jennifer Mattock, Jelena Gavrilovic, Allan Clark, Tom Wileman, Carl M. Philpott in American Journal of Rhinology & Allergy
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: SG is funded by a National Institute for Health and Care Research (NIHR) Academic Clinical Fellowship. ACP received PhD funding from the Sir Jules Thorn Charitable Trust. JM was funded by a PhD studentship from the University of East Anglia. TW is supported by Biotechnology and Biological Sciences Research Council Institute Strategic Programme Gut Microbes and Health [BB/R012490/1 BBS/E/F/000PR10353, BBS/E/F/000PR10355].
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
