Abstract
Background
Previous research has shown diminished nasal immune function following nasal saline irrigation (NSI), returning to baseline at 6 hours. The aim of this study was to examine the immune nasal proteome before and after 14 days of nasal irrigation.
Methods
Seventeen healthy volunteers received either isotonic (IsoSal) or low salt (LowNa) NSI. Nasal secretions were collected before and 30 min after NSI at baseline and again after 14 days. Specimens were analyzed using mass spectrometry to detect proteins of relevance to nasal immune function.
Results
One thousand eight hundred and sixty-five proteins were identified with significant changes in 71 proteins, of which 23 were identified as part of the innate immune system. Baseline analysis demonstrated an increase of 9 innate proteins after NSI, most after IsoSal. After 14 days, a greater increase in innate peptides was present, with most now in the LowNa group. When NSI solutions were compared, a significant increase in 4 innate proteins, including a 211% in lysozyme, was detected in the LowNa group.
Conclusion
LowNa NSI demonstrates evidence of improving the innate immune secretions, especially lysozyme, in healthy volunteers.
Keywords
Introduction
Chronic rhinosinusitis (CRS) is a common chronic inflammatory condition with a prevalence of up to 9% of the Australian population and has a significant impact on patient quality of life (QOL) and morbidity.1,2 Evidence-based treatments for CRS include the use of large volume nasal saline irrigation (NSI) 3 to remove thickened secretions to alleviate symptoms and improve QOL.2,4 NSI is hypothesized to function by thinning mucous, improving mucociliary clearance, reducing edema and decreasing antigen load in the sinonasal cavity. 5
The innate immunity of the sinonasal airway is multifaceted 6 comprising of both a physical barrier of mucous producing epithelial cells 7 and a mucous layer with airway surface liquid (ASL) 8 containing antimicrobial polypeptides, produced by the submucosal glands.7,9 The hypotonic ASL is bilaminar, comprising an outer viscous layer and deeper aqueous periciliary layer.10–12 These secretions contain numerous cationic antimicrobial peptides including lysozyme, lactoferrin, antitrypsin, defensins, s100 proteins, and surfactant,10,13,14 with lysozyme and lactoferrin being the most abundant. 9
Antimicrobial peptides are cationic and hydrophobic at physiological pH. 9 ASL tonicity is a product of active electrolyte transport within airway epithelium, which directly impacts the salt sensitive bactericidal activity of the cationic antimicrobial peptides.12,15 Previous studies have identified that ASL composition becomes more isotonic in patients with chronic airway irritation or cystic fibrosis, impeding bactericidal capability of cationic antimicrobial proteins, which further perpetuates the disease process.12,16 Commercially available NSI formulations are classified as isotonic (IsoSal), low salt (LowNa), or hypertonic. In previous studies, these solutions demonstrated a multi-dimensional effect, altering nasal protein concentration in addition to modifying protein activity. 17
Previous work comparing the effect of NSI on innate proteins demonstrated that, with single nasal irrigation, lysozyme and lactoferrin levels were depleted, although levels returned to baseline at 6 hours. 17 Interestingly, at 24 hours post-irrigation, LowNa NSI resulted in a significant increase in lysozyme and lactoferrin concentration compared to IsoSal NSI 17 although this study only looked at 2 individual proteins. While some work has been done on the whole nasal proteome with older mass spectrometry methodology,18–21 this pilot study (prior to a planned more focused study on patients with CRS) aimed to utilize contemporary proteomic methodology including more sensitive instrumentation and quantitative data independent analysis (DIA) to test the hypothesis that short-term NSI will have positive benefits to the nasal immune proteome by comparing secretions both before and after 14 days of IsoSal or LowNa daily NSI.
Materials and Methods
Subjects and Tissue Samples
The project was approved by the Southern Adelaide Local Health Network Human Research Ethics Committee (reference number 278.19). Seventeen healthy volunteers were recruited via research advertisement. Individuals were screened with the 22-item Sino-Nasal Outcome Test (SNOT-22), with a score ≤7 representative of normal 22 and required for inclusion in the study. Nine participants were randomly allocated to isotonic saline, and 8 to hypotonic saline.
Mucus Collection
After obtaining fully informed consent, participants were randomly allocated to receive NSI using either isotonic (IsoSal; FLO Sinus Care©, ENT Technologies, Melbourne, Australia) 23 or low salt (LowNa, FLO CRS©, ENT Technologies) 24 formulations. Minimally modified nasal secretions were collected by the placement of a sinus secretion collector (8 mm×3 mm×3 mm of polyurethane foam; Corning, New York, USA) between the right inferior turbinate and nasal septum for 10 min as utilized in previous studies.17,25
At baseline, nasal secretions were collected prior to (T1), and at 30 min after (T2) NSI. Daily NSI occurred for 14 days, with nasal secretions collected on day 14 prior to (T3) and 30 min after (T4) NSI (Figure 1). This provided a total of 4 samples per individual, totaling 8 comparative sample groups. T1/T2 (short-term benefits in the saline-naïve nose), T1/T3 (long-term benefits), and T3/T4 (short-term benefits in the saline-managed nose) were chosen as time points to compare between and within groups.
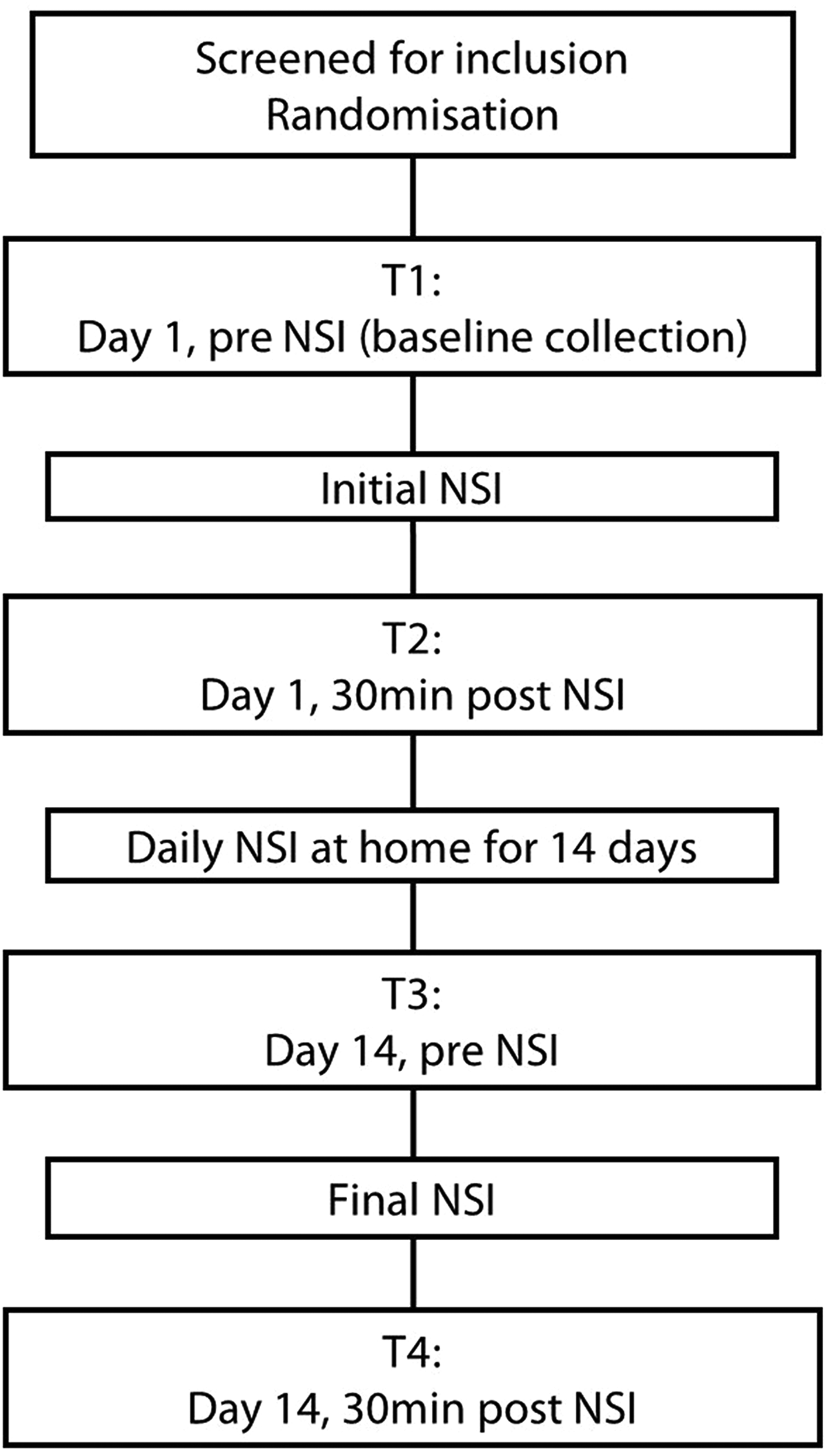
Project timeline.
Mucus Analysis
Samples were microfuged for 10 min at 21 000 × G (4 °C), then transferred into 1.7 mL maximum recovery tubes (Corning, New York, USA) and stored at −20 °C.
Protein concentration estimation for each sample was performed using an EZQ™ Protein Quantitation Kit (Thermo Fischer Scientific). Lipids and other non-proteinaceous compounds present in mucous were isolated by sample clean-up and enzymatic digestion with Sera-Mag SpeedBead Carboxylate-Modified Magnetic beads (Cytiva). Briefly, 10 µg of each nasal fluid protein extract was reduced with 10 mM TCEP (tris(2-carboxyethyl)phosphine) followed by alkylation with 20 mM chloroacetamide. About 100 µg of magnetic SpeedBeads comprised of a 1:1 ratio of hydrophilic and hydrophobic beads and washed 3 × in ddH20, and were added to each of the reduced/alkylated nasal fluid samples. Ethanol was then added to a 50% concentration (v/v) to precipitate protein followed by three washes with 80% ethanol. Ethanol was removed and the beads were air dried briefly and resuspended in 100 mM ammonium bicarbonate with 500 ng Pierce™ Trypsin Protease, MS Grade (Thermo Fischer Scientific) and incubated at 37 °C overnight. Speedbeads were removed using a magnet and fresh 100 µg of Sera-Mag SpeedBeads added as described above to clean up the peptides. Acetonitrile was added to each sample to 95% (v/v) concentration to bind peptides onto the beads followed by 1 × 100% acetonitrile wash. Following clean-up, peptides were eluted off the magnetic beads with 2% DMSO and stored at −80 °C until mass spectrometry analysis was performed.
Mass Spectrometry Analysis of Nasal Proteins-Liquid Chromatography
Peptides were analyzed with a Dionex Ultimate 3000 uHPLC (ultra-high performance liquid chromatography) coupled to a Thermo Fusion Lumos mass spectrometer fitted with a FAIMS ion mobility interface. An inhouse pulled column created from 75 µm inner diameter fused silica capillary packed with 1.9 µm ReproSil-Pur C18 beads (Dr Maisch, Ammerbuch, Germany) to 25 cm, coupled with an Acclaim PepMap™ 100 trap column (75 µm × 2 cm, 3 µm C18, 100 Å, Thermo Fischer) were utilized. Solvent A was 0.1% formic acid in water and solvent B was 0.1% formic acid in 80% acetonitrile. For each injection, 1 µg of peptides were separated using a 60 min gradient from 3 to 31.2% B, followed by a 25 min washing and equilibration in Buffer A.
Spectral Library Generation
A nasal peptide spectral library of MS1/MS2 spectra was generated to assist identification of peptides contained in DIA spectra. Gas phase fractionation (GPF) was employed to increase the depth of coverage of the library. GPF targets only a fraction of the mass range in each injection to reduce complexity and increase depth of coverage. A pooled sample comprised of 1 µL of each nasal fluid protein digest was injected 11 times, each only analyzing a 60 m/z window of the 400–1070 m/z total mass range. 3 µL of the pooled sample was utilized for each GPF analysis. Briefly, precursor spectra including a wide spectrum (390–1100 m/z at 60 000 resolution) and a narrow spectrum matching 1 of the 11 60 m/z mass ranges were performed at each FAIMS compensation voltage (CV) (−45 V and −65 V). An AGC target of 4e5 and a maximum injection time of 60 ms were used for all MS scans. 4 m/z DIA spectra (4 m/z precursor isolation windows at 30 000 resolution, AGC target 5e4, maximum inject time 60 ms, 33% normalized collision energy in the HCD cell) were acquired utilizing a staggered window pattern over the same 60 m/z mass range as the narrow MS scan. DIA spectra were also acquired at both −45 and −65 V FAIMS CVs. Comprehensive details of the GPF method employed are enclosed in the online Supplemental Methods document.
Quantitative Shotgun Data Independent Acquisition (DIA) Mass Spectrometry
For single-injection runs, the Thermo Fusion Lumos with FAIMS interface installed, was configured to acquire 23 20 m/z precursor isolation windows (400–800 m/z), followed by 4 100 m/z windows (800–1200 m/z) using a staggered window pattern. This schema was performed twice, once with a FAIMS CV of −45 and again with a CV of −65. An ms2 resolution of 30 000, AGC target 5e4, maximum inject time of 54 ms, and normalized HDC collision energy of 30% was employed for all DIA scans. Precursor spectra over a 350–1200 m/z mass range were acquired at each FAIMS CV prior to DIA scans. A resolution of 30 000, AGC target 1e6, maximum inject time 60 ms were used for all full scan MS spectra.
DIA Data Analysis
Spectronaut™ (version 15.0.210615.50606, Biognosis) was used for both spectral library generation and data analysis. Default setting were used for all Spectronaut analysis steps, with the p-values corrected to a q-value representing final statistical difference. A q-value <0.05 was regarded as significant. In all analyses, differentially expressed proteins are reported as the percentage change in one group relative to the other.
Results
A total of 17 participants with normal SNOT-22 questionnaire (mean score 3.13 ± 2.94) were recruited with one participant withdrawal due to nasal hypersensitivity following the initial NSI. The demographics of the 16 individuals who completed the study are presented in Table 1.
Demographics.
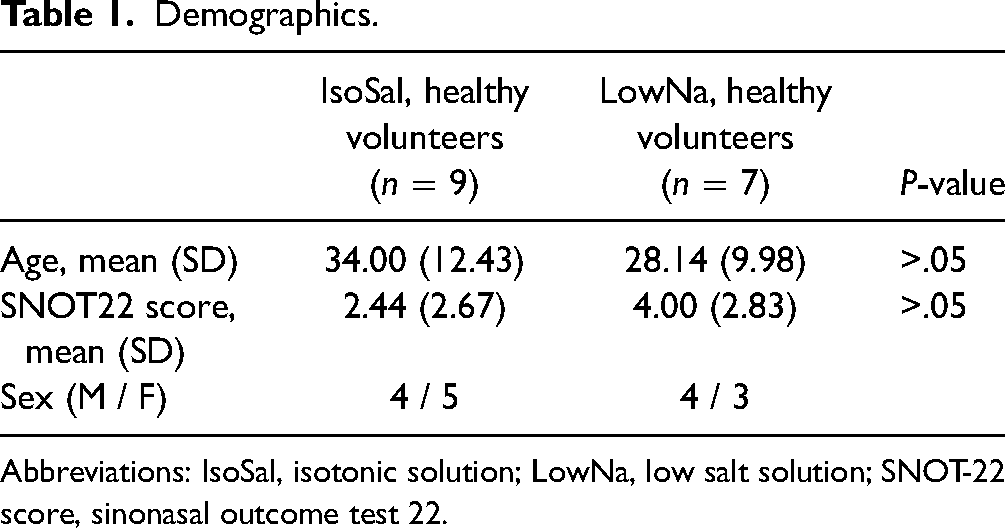
Abbreviations: IsoSal, isotonic solution; LowNa, low salt solution; SNOT-22 score, sinonasal outcome test 22.
Spectronaut™ analysis quantified 1865 unique proteins in the library. From all samples (range 1379–1828 per sample) 71 (3.8%) proteins were identified as having statistically significant differential abundance between comparison groups. Utilizing the UniProt© proteomic descriptive database 24 (32.4%) of these proteins were identified as being an integral part of the innate immune system (Table 2).
Innate Proteins Demonstrating Significant Change in Abundance Following Nasal Saline Irrigation.

Abbreviations: A-1-acid, Alpha-1 acid glycoprotein; A-1B-gly, Alpha 1B glycoprotein; Act2, actin-related protein 2; Annex, Annexin; Casp14, caspase 14; CortBH, corticosteroid binding globulin; Hapt, haptoglobin; IBM-A3 HCVR, IBM-A3 heavy chain variable region; IgHV3-20, immunoglobulin heavy variable 3-20; IGL1714, IGL c1714_light_IGKV6-21_IGKJ4; IgHeavyA2, immunoglobulin heavy constant alpha 2; IgHeavyA1, immunoglobulin heavy constant alpha 1; IgHeavyMu, immunoglobulin heavy constant mu; IgJ, immunoglobulin J chain; LC1, Lipocalin 1; LeukA4, Leukotriene A(4) hydrolase; LTF, Lactotransferrin; Lysoz, Lysozyme; Myelop, myeloperoxidase; PIR, polymeric immunoglobulin receptor; ProS100, Protein S100; ProS100-A7, Protein S100-A7; ProS100-A8, Protein S100 A8; RAB7.
Analysis findings of the NSI comparison groups are summarized in Table 3.
Nasal Mucous Proteins Demonstrating Significant Change in Abundance/Concentration Following Initial and 14 Days of Nasal Saline Irrigation.
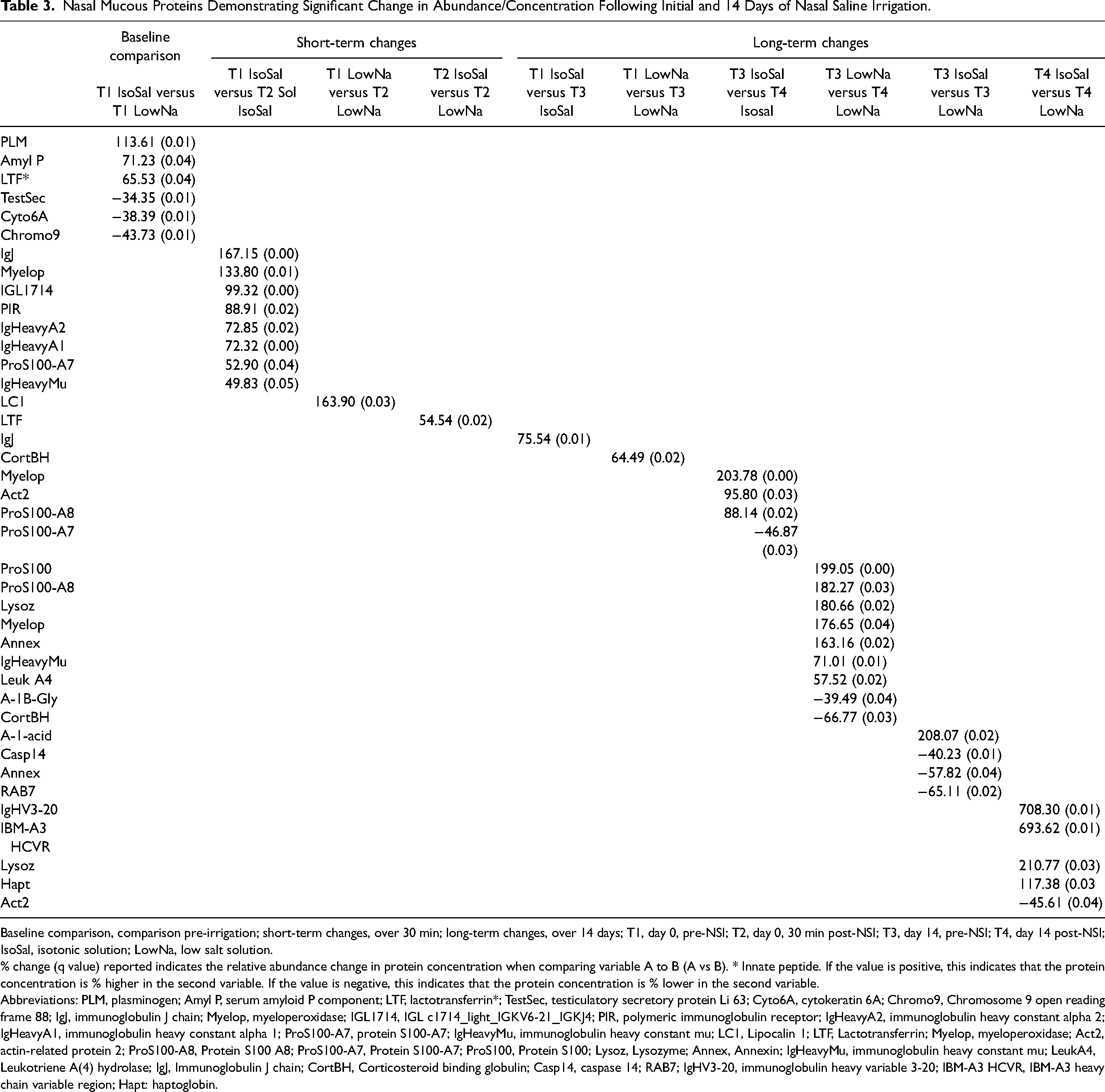
Baseline comparison, comparison pre-irrigation; short-term changes, over 30 min; long-term changes, over 14 days; T1, day 0, pre-NSI; T2, day 0, 30 min post-NSI; T3, day 14, pre-NSI; T4, day 14 post-NSI; IsoSal, isotonic solution; LowNa, low salt solution.
% change (q value) reported indicates the relative abundance change in protein concentration when comparing variable A to B (A vs B). * Innate peptide. If the value is positive, this indicates that the protein concentration is % higher in the second variable. If the value is negative, this indicates that the protein concentration is % lower in the second variable.
Abbreviations: PLM, plasminogen; Amyl P, serum amyloid P component; LTF, lactotransferrin*; TestSec, testiculatory secretory protein Li 63; Cyto6A, cytokeratin 6A; Chromo9, Chromosome 9 open reading frame 88; IgJ, immunoglobulin J chain; Myelop, myeloperoxidase; IGL1714, IGL c1714_light_IGKV6-21_IGKJ4; PIR, polymeric immunoglobulin receptor; IgHeavyA2, immunoglobulin heavy constant alpha 2; IgHeavyA1, immunoglobulin heavy constant alpha 1; ProS100-A7, protein S100-A7; IgHeavyMu, immunoglobulin heavy constant mu; LC1, Lipocalin 1; LTF, Lactotransferrin; Myelop, myeloperoxidase; Act2, actin-related protein 2; ProS100-A8, Protein S100 A8; ProS100-A7, Protein S100-A7; ProS100, Protein S100; Lysoz, Lysozyme; Annex, Annexin; IgHeavyMu, immunoglobulin heavy constant mu; LeukA4, Leukotriene A(4) hydrolase; IgJ, Immunoglobulin J chain; CortBH, Corticosteroid binding globulin; Casp14, caspase 14; RAB7; IgHV3-20, immunoglobulin heavy variable 3-20; IBM-A3 HCVR, IBM-A3 heavy chain variable region; Hapt: haptoglobin.
Following the initial irrigation (T2) there were 8 innate proteins with higher levels compared to T1 in the IsoSal group, whereas only one innate protein was higher than baseline in the LowNa group. Comparison of IsoSal v LowNa groups at T2 showed lactotransferrin was increased in the LowNa group.
Following 13 days of irrigation and immediately prior to the last dose (T3), there was a single (but different) innate protein that increased when compared to baseline (T1) in each of the IsoSal and LowNa groups. Comparison of IsoSal v LowNa groups at T3 demonstrated a slight decrease in 3 proteins and a large increase in alpha 1 acid glycoprotein 2 in the LowNa group.
Comparing pre- and post-irrigation on day 14 (T3 vs T4), there was an increase in 3 proteins in the IsoSal group, and an increase of 7 proteins in the LowNa group, including a 180% increase in lysozyme. There was also noted to be a reduction in one protein in the IsoSal group and 2 proteins in the LowNa group. Comparison of IsoSal v LowNa groups at T4 demonstrated a significant increase in 4 innate proteins, including a 211% increase in lysozyme in the LowNa group.
Discussion
This pilot study, in healthy volunteers, has demonstrated it is possible to easily collect nasal secretions on multiple occasions and compare the effect of a therapeutic intervention on the nasal proteome using mass spectrometry. It is now our intention to use these results for a more targeted study on patients with CRS. This study found that NSI utilization over 14 days enhanced the production of several innate immune proteins which have potential to improve nasal health and the ability to protect against disease. This is particularly topical in the context of the current COVID-19 epidemic, where NSIs have been advocated as a method of improving nasal health and potentially reducing infection risk. 45
A recent scoping review identified 10 studies reporting on the proteomics of nasal secretions but the number of proteins identified was limited (range 12–366, perhaps due to older methods of mass spectrometry). 19 Using contemporary proteomic methodology and analysis techniques this study identified 1865 unique proteins from minimally manipulated nasal mucous of healthy people, to our knowledge, the largest number of proteins ever identified within nasal mucous. 46
Various NSI formulations are available commercially with some reports demonstrating effects on nasal immunity. Our findings suggest that LowNa nasal irrigations may provide additional benefit to the nasal innate proteome. However, this study did not assess protein activity, which is highly relevant because some innate immune proteins are disabled to some degree in ionic solutions greater than 34 Millimoles. 47 Previous work from our group demonstrated that isotonic NSI inhibited the functional activity of cationic antimicrobial proteins and this effect occurred after the use of all commercially available isotonic NSI solutions in Australia. 17 Specifically, following irrigation, innate protein concentration and activity (measured using ELISA) initially reduced at 1 h but increased at 6 h, with the best results being observed following LowNa irrigation. 17
LowNa solutions have also been shown to provide clinical benefit over IsoSal in a randomized controlled trial following endoscopic sinus surgery 48 for the treatment of CRS. Participants were randomly allocated to a either a isotonic solution (with an ionic strength of 154 mM) 49 or an isotonic but low salt solution (with an ionic strength of 26 mM), 49 identical formulations as used in the current study. The increased abundance of innate peptides following 14 days of LowNa NSI in the current study suggests an additional mechanism for the described clinical benefit beyond maintaining functionality of the cationic antimicrobial proteins.
Short-term Irrigation Effects
Lysozyme and lactoferrin have been identified as the most important antimicrobial proteins in the innate nasal immune response. 49
Both IsoSal and LowNa irrigation resulted in an increase of a myriad of innate proteins after 30 min. Increased expression of immunoglobulin chains, polymeric immunoglobulin receptor, myeloperoxidase, and protein S100-A7 were noted for IsoSal, with increased lipocalin levels, believed to mediate airway inflammation with important antibacterial and antifungal functions 50 noted for LowNa. These findings support the hypothesis that nasal irrigation may help reset the innate protein environment, aiding in the optimization of nasal health.
Interestingly, no significant change in the levels of lysozyme in the short term was identified. This potentially contradicts the findings of our previous work where both lysozyme and lactoferrin were reduced at 1 h following irrigation, although a 30-min time point was not evaluated in this study. It is also possible that changes in these proteins may not have been large enough to be detected using mass spectrometry. Combining both mass spectrometry and ELISA on the same samples would potentially provide stronger data and it is our intention to use both in future work on nasal innate immune secretions.
Long-term Irrigation Effects
Following a 14-day period of NSI in healthy volunteers, several innate proteins were noted to significantly change. Innate proteins of interest included myeloperoxidase, protein S100-A8, actin-related protein, protein S100-A7, annexin, immunoglobulin chain, corticosteroid binding globulin, alpha 1-B glycoprotein, leukotriene A4 hydrolase, immunoglobulin heavy constant mu chain, annexin, and lysozyme.
Different responses were observed with the NSI formulations investigated at day 14. For IsoSal irrigation there was a change in the relative abundance of 4 innate proteins consisting of a relative increase in myeloperoxidase, protein S100-A8, actin-related protein, and a relative decrease in protein S100-A7. Those using LowNa showed increased myeloperoxidase, protein S100/S100-A8, lysozyme, annexin, and immunoglobulin chain, with decreased levels of corticosteroid binding hormone and alpha 1-B glycoprotein.
When comparing protein levels between the NSI formulations after 14 days most innate protein levels were higher with LowNa use than IsoSal, except for corticosteroid-binding globulin and alpha-1B-glycoprotein which were lower. Of particular interest are the lysozyme levels which were 211% greater in the LowNa group, along with approx. 700% increase in various immunoglobulins, suggesting the nasal mucosa may have been primed to produce a greater short-term increase in some innate proteins from repeated daily NSI over 14 days.
We were interested to determine if daily NSI influenced baseline levels of proteins in the nasal mucosa. Despite short-term changes in protein levels following irrigation, and a greater number of proteins increasing levels after 14 days, the pre-irrigation sample showed little difference to the initial baseline sample. This data indicates ongoing daily NSI does not increase baseline levels of the nasal mucous proteome but may prime the mucosa ready for peptide release upon NSI, and therefore suggests any clinical effect may be relatively short-lived, supporting the recommendation for repeated NSI throughout the day.
A reduction in levels of innate proteins can alter microbial colonization. Previous studies have demonstrated reduced lysozyme and lactoferrin levels in patients with nasal polyps relative to healthy controls, altering microbial colonization. 51 Pathologic deficiencies in the antimicrobial properties of airway secretions can lead to distortion of the nasal microbiome and may contribute to the pathophysiology of CRS. 52
We regard this study as a pilot for further exploration of innate nasal proteins. Recently a down-regulation of mucosal immunity was demonstrated in patients with CRS, 53 but the use of NSI was not controlled in either patient group. Other work has produced alternative findings by demonstrating an increase in the innate immune system in patients with allergic rhinitis 18 and this is an interesting area of nasal physiology which, in our opinion, has been poorly studied thus far.
Having demonstrated short-term and long-term changes in a myriad of innate proteins, further work could potentially explore the effects of disease and therapeutic interventions on individual proteins in greater detail. This study has several limitations. Sample preparation included precipitation and clean up on magnetic beads. Some proteins may be lost through this process. In addition, very low molecular weight proteins such as defensins (2-5 kDa) may not be detected by the mass spectrometer. For example, beta-defensin 1 only generates 2 tryptic peptides of mass that is detectable by MS. Larger proteins can generate 10-fold or more peptides which increases the ability of the mass spectrometer to detect the protein. Additionally, in this study, changes in innate protein levels over time were measured, with no attempt to assess their antimicrobial functionality. Previous studies have noted that antimicrobial protein levels in CRS patients were equivocal to healthy volunteers, although their functionality reduced. 17 As the use of proteomic analysis to investigate nasal secretions becomes more widespread, we believe the methodology is likely to add to the academic rhinologist's armamentarium in the search for the holy grail as to “what causes and how to treat CRS?” LowNa formulation of NSI demonstrates evidence of improving the innate immune secretions in healthy volunteers. This pilot study has identified several innate proteins for future research in both health and disease.
Supplemental Material
sj-docx-1-ajr-10.1177_19458924231159176 - Supplemental material for The Nasal Innate Immune Proteome After Saline Irrigation: A Pilot Study in Healthy Individuals
Supplemental material, sj-docx-1-ajr-10.1177_19458924231159176 for The Nasal Innate Immune Proteome After Saline Irrigation: A Pilot Study in Healthy Individuals by Emily A. Harcourt-Smith, MD, Emerson T. Krstic, MD, Belinda J. Soekov-Pearce, MD, Alex D. Colella, PhD, Nusha Chegeni, BScHons, Timothy K. Chataway, PhD, Charmaine M. Woods, PhD, Kamelya Aliakbari, M, and A. Simon Carney, DM in American Journal of Rhinology & Allergy
Footnotes
Author Contributions
EAH-S: literature review, ethics approval, patient recruitment, sample collection, and manuscript writing; ETK: patient recruitment, sample collection, and manuscript writing; BJS-P: patient recruitment, sample collection, and manuscript writing; ADC: data analysis, manuscript writing; NC: secretion analysis, data analysis, manuscript writing; TC: study inception, secretion analysis, data analysis, manuscript writing; KA: secretion analysis; ASC: study inception, patient recruitment, sample collection, and manuscript writing.
Authors’ Note
This content was presented at The Australian Society of Otolaryngology Head & Neck Surgery (ASOHNS) Annual Scientific Meeting June 12, 2022.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by a stipend from the Flinders University School of Medicine Advanced Studies Research Program.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
