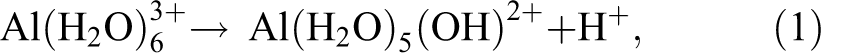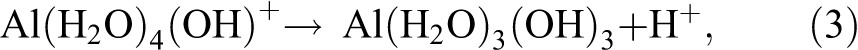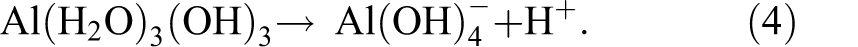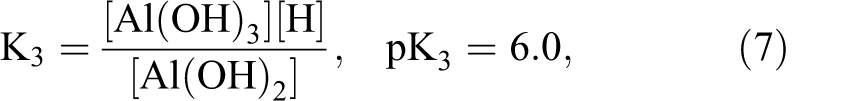Abstract
The crystal structure of the newly synthesized ligand pyridoxal amino methyl phosphonic acid (PYRAMPA) from green materials in deionized water and methanol is presented. It took only two steps to prepare this new ligand. The ligands’ stability constants with Al3+ are also reported in aqueous solutions at 25 °C ± 0.1 °C. The ligand was introduced as part of our concerted efforts to identify and synthesize an orally effective aluminum sequestering agent due to the known toxic effect of aluminum in living organisms particularly humans. PYRAMPA is a multidentate ligand suitable to sequester aluminum with very high stability.
Introduction
Occurrence of Aluminum in Nature and Its Coordination Chemistry
Aluminum is the most abundant metal ion in the earth’s crust and the third-most abundant element after oxygen and silicon. The earth’s crust contains 8.1% by weight of aluminum compared to 5.0% by weight of iron. 1 In nature, aluminum exists only in the oxidation state Al3+. The primary ore is bauxite, which is mainly the metal oxide. The ion is a major component of large number of minerals, for example, mica, feldspar, and clays. In contrast to its abundance in the crust, the ocean concentration is below 1 μM. Until recently, most natural waters contained insignificant amounts of aluminum, except for those in some volcanic regions and in alum springs. At neutral pH, aluminum is extremely insoluble, and the concentration of free Al3+ is very low in both surface and subsoil waters. However, the solubility increases at low pH levels. Because of the effects of acid rain and the use of acidifying fertilizers, the concentration of the free Al3+ has increased in the soil, in lakes and in rivers. Therefore, aluminum has become more accessible to living organisms. 1,2
Aluminum is one of the most hydrolytic metal ions known in aqueous media.
3–5
Equilibria among the free Al3+ ion and four hydrolyzed species are described in four reactions as shown in eqs 1–4.
By dropping the attached water molecules and the charges on the complexes for clarity, eqs 1–4 yield the following equilibrium constants, as shown in eqs 5–8. The negative log (−log) of the equilibrium constant is defined as the pKa of the hydrolysis process. The four pKa’s for the aluminum ion are very close and are squeezed in less than one pH unit due to the cooperativity in Al3+ hydrolysis reactions.
5
Many researches propose that aluminum biochemistry is very complex and not fully understood. 1–8 A report by the World Health Organization accessed on January 24, 2013, indicates that oral ingestion of aluminum additives in the diet constitutes the main source of aluminum exposure for humans. 7 Many papers associate and relate the accumulation of aluminum with Alzheimer’s disease; the following seven references are not the whole list, but just simple examples. 6,8–13 We have published some reports that dealt with the aluminum issue whether it is the speciation of aluminum among serum ligands or introducing new ligands that can treat aluminum overload. 14–17
In this article, we are presenting the crystal structure and the stability constants of the newly synthesized ligand pyridoxal amino methyl phosphonic acid (PYRAMPA) with aluminum in aqueous solution under ambient conditions. The conditions set forth for the stability constants measurements were 25 °C ± 0.1 °C, under inert atmosphere and ionic strength of I = 0.1 M KNO3. It appeared that PYRAMPA forms very stable aluminum complexes.
Results and Discussion
Herein, we are reporting the synthesis of the novel PYRAMPA, its crystal structure, and its 3-D crystal packing. The stability constants for this ligand with Al3+ in aqueous solutions under ambient conditions were Log β112 = 24.60 ± 0.09, Log β111 = 21.35 ± 0.15, Log β110 = 15.26 ± 0.43, and Log β11-1 = 7.69 ± 0.27. See the exact definition of the Log β complexes in the experimental section below.
The PYRAMA ligand was made possible from the parent ligand aminomethylphosphonic acid (AMPA). The parent ligand AMPA was tested to chelate aluminum in a previous study; 15 it was found to form insoluble complexes with aluminum at neutral pH value. For this reason, we prepared PYRAMPA that formed soluble aluminum complexes at neutral pH value. Attempts have been made to improve the binding abilities of aminoalkylphosphonic acids such as AMPA presented in reference 15. One of these attempts to improve the binding abilities of AMPA is to include the phenolate functionality to the aminoalkylphosphonic acid ligand lead to the synthesis of 2- Hydroxybenzylaminomethyl phosphonic acid (HBAMPA) presented in reference 14 (Harris and Hamada). Another attempt was to make this added phenolate functionality more effective in binding the aluminum ion and at the same time to increase the ligand solubility at neutral pH value. We met the second goal of increasing the effectiveness of the phenolate group to chelate metal ions and lower the ligand solubility by synthesizing PYRAMPA. The aqueous coordination chemistry of PYARMPA involving the Al3+ ion is described in this article.
Potentiometric Titrations of PYRAMPA Ligand with Al3+
Free PYRAMPA was defined as an H4L ligand in which there are four dissociable protons. The four protons that dissociated are as follows: the pyridine, the second phosphonic acid, the phenolate, and the amine functional groups. It appeared that the pyridine group has a pKa of 2.88 ± 0.08, the second phosphonic acid group has a pKa of 5.28 ± 014, the phenolate OH group has a pKa of 8.02 ± 0.01, and the amine group has a pKa of 11.12 ± 0.16. All functional groups possess pKa values that are in the correct ranges of these functionalities. 18–20 As it was the case with AMPA and HBAMPA, the first phosphonic acid OH proton was too small or too acidic to have a measurable pKa value. The potentiometric titrations of the free PYRAMPA showed an initial pH of 3.3 and three distinct inflections. Supplementary Figure 1 shows the family of titration curves of the free PYRAMPA, the 1:1 and the 1:2 Al3+:PYRAMPA systems. It is worth mentioning that with the 1:1 and the 1:2 titrations there was no precipitation at any given pH value.

Projection view of the PYRAMPA molecule with 50% thermal ellipsoids.
With the 1:1 titrations there was high equilibration time above pH 8.0 or above 5 equivs of KOH. In Supplementary Figure 1, the darkened data points in the 1:1 titration curve is indicating the high equilibration time above pH 8. The initial inflection appeared at 3.0 equivs, which is accounted for by the formation of the monoprotonated 111 species complex. The species notation 111 indicates that the first index stands for the number of Al3+ ions in that species, the second is the number of PYRAMPA, and the last index stands for the number of the hydrogen ions. The model used to fit these 1:1 titration data points was that which contained the simple 1:1 complexes namely
Protonation Constants and Stability Constants in the Form of Log βs for the Different Complexes Observed in the Al3+–PYRAMPA Titration Systems, 25 °C, I = 0.10 M KNO3.

The model that fits the 1:2 titration system was the same model given to the 1:1 runs, which consists of 112, 111, 110, and 11-1. The major species was the mono-protonated complex
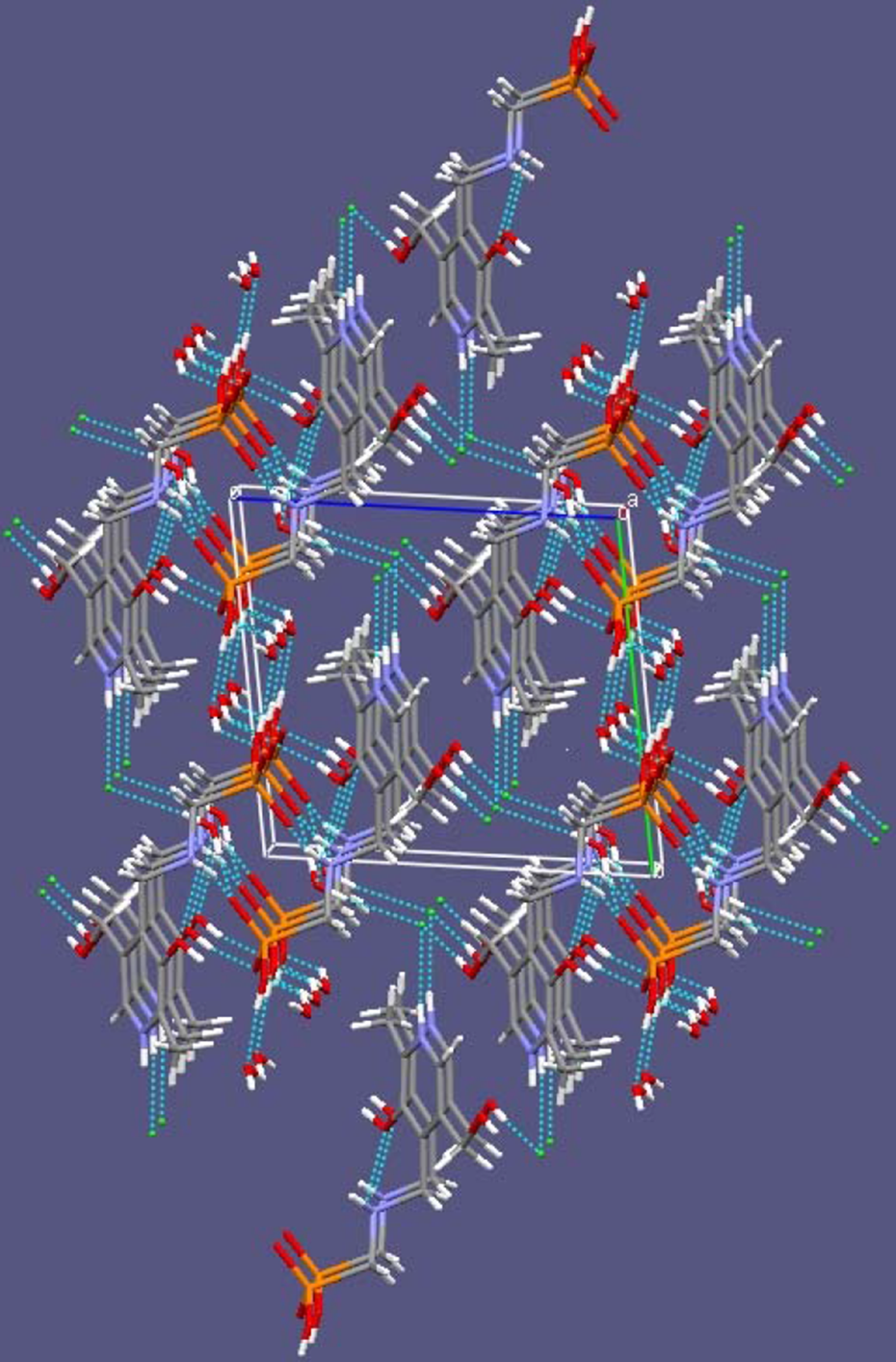
Crystal packing of PYRAMPA via hydrogen bonding and Cl− counter ion.

Synthetic routes used to prepare the new PYRAMPA from commercially available and green compounds using water and methanol as solvents.
In careful qualitative UV measurements, we were able to prove that the phenolate functionality is participating in the chelation of the Al3+ ion. Supplementary Figure 3 is the qualitative UV absorption spectra for 255 μM free PYRAMPA taken from pH 2.1 to 11.97. From these spectra, the ligand is in its totally protonated state, as H4L has one characteristic peak at 295 nm. The deprotonation of the pyridine group shifted this peak to the higher wavelength with the characteristic peak at 330 nm. Upon increasing the pH further, the phenolic group was deprotonated and the peak was decreased to lower wavelength value with the absorption peak at 310 nm. The peak at 310 was the fingerprint of deprotonated phenolate group. 14–19
Supplementary Figure 4 is the qualitative UV absorption spectra for 255 μM total Al3+ concentration of the Al3+:PYRAMPA reaction mixture in 1:1 ratio taken from pH 2.3 to 9.2. The characteristic absorption peak at 295 nm was observed for the phenolate chromophore at both pH 2.3 and pH 2.5. When the pH was further increased to 3.3, the absorption peak showed shift to 300 nm and 325 nm with a shoulder indicating the deprotonation of the pyridine group as it was the case for the free PYRAMPA. When the pH was further increased to 4.0, the characteristic peak at 310 due to the deprotonation of the phenolate group was observed. The observation of the fingerprint of the deprotonated phenolate at 310 nm at the acidic pH = 4.0 is the clear evidence that the phenolate group is participating in the coordination of the Al3+ ion.
Experimental
Synthesis of PYRAMPA
Scheme 1 shows the detailed routes of PYRAMPA synthesis from commercially available chemicals in deionized water and methanol as solvents. A solution of 500 mg (2.46 mmoles) of the hydrochloride salt of pyridoxal was added dropwise to a 273 mg (2.46 mmoles) methanol solution of AMPA. A net amount of 385 mg (6.9 mmoles) solid KOH was added to basify the methanol solution of AMPA. The characteristic deep reddish-yellow color of the Schiff base was instantaneously observed. The methanol was stripped off under vacuum; the microcrystalline material of the tripotassium salt of the Schiff base weighted 791 mg (2.1 mmoles) with the percentage yield of 85.88%. The reduction step was carried out in 25 mL dry methanol by adding 2 equivs of solid sodium borohydride 162 mg (4.2 mmoles). The distinct reddish-yellow color of the Schiff base disappeared indicating the transformation of the Schiff base functionality to the free amine functionality (step #2 in scheme #1). The reduced material was concentrated by pumping off most of the methanol. The concentrated free amine or PYRAMPA was acidified by the addition of minimum amount of 6.0 M HCl. A white precipitate appeared instantaneously that was filtered via filter paper. The filtrate was allowed to stand for 72 h for the crystallization of PYRAMPA. The PYRAMPA ligand with the empirical formula (C9 H20 Cl N2 O7 P) had a melting point of 264 ± 1 °C.
Crystallography Data
PYRAMPA was crystallized in the triclinic, P-1 type space group in which α, β, and γ ≠ 90°. All the crystal data and structure refinement for PYRAMPA are given in Table 2. Table 3 shows the atomic coordinates and equivalent isotropic displacement parameters; Table 4 shows the bond lengths; Table 5 shows the anisotropic displacement parameters; Table 6 shows the hydrogen coordinates and isotropic displacement parameters; and Table 7 shows the torsion angles in degrees.
Crystal Data and Structure Refinement for PYRAMPA.

Atomic Coordinates (× 104) and Equivalent Isotropic Displacement Parameters (Å 2 × 103) for PYRAMPA.

Note: U(eq) is defined as one-third of the trace of the orthogonalized Uij tensor.
Bond Lengths [Å] and Angles [°] of PYRAMPA.

Anisotropic Displacement Parameters (Å 2 × 103) for PYRAMPA.

Hydrogen Coordinates (× 104) and Isotropic Displacement Parameters (Å 2 × 103) for PYRAMPA.

Torsion Angles [°] for PYRAMPA.
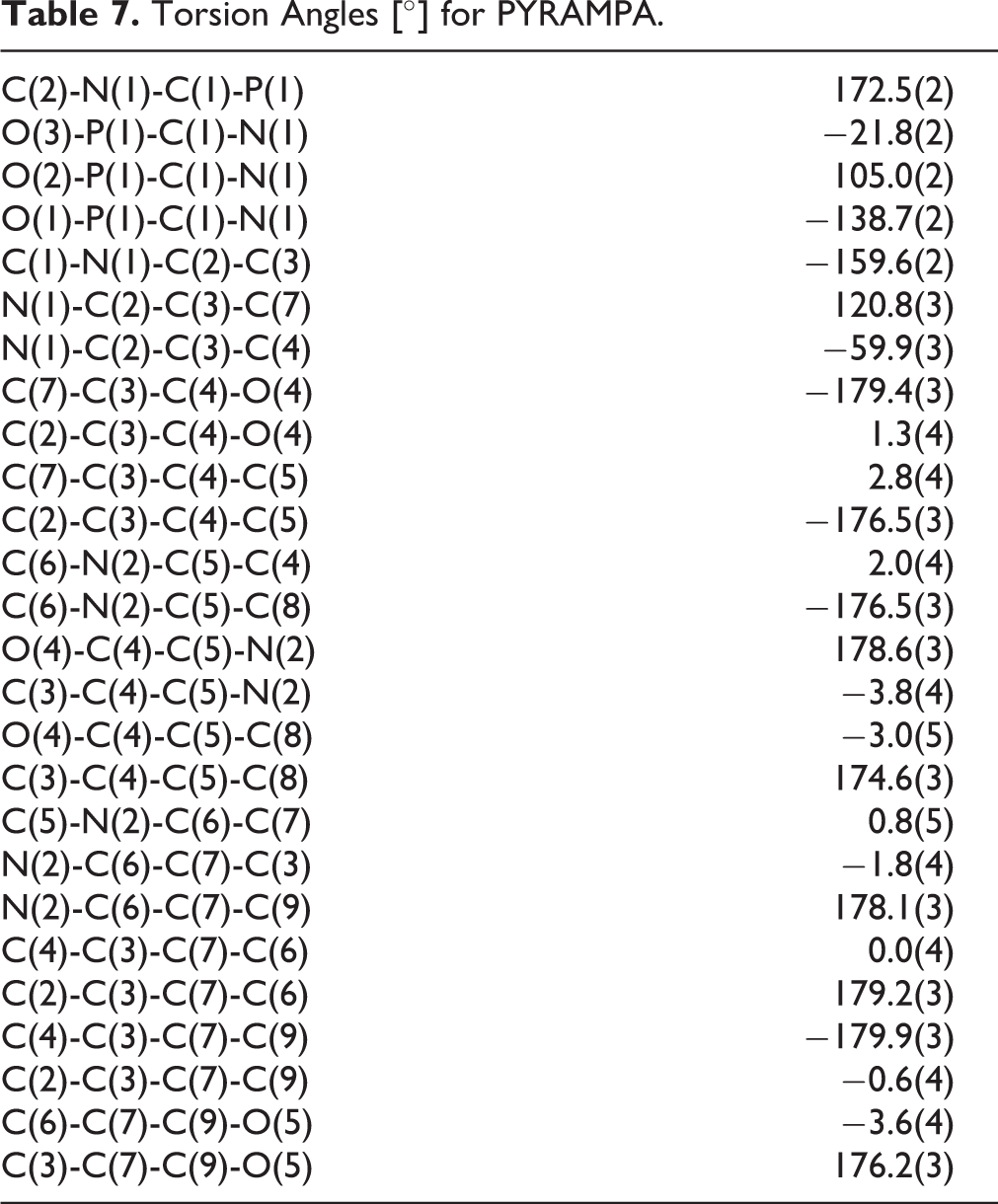
Potentiometric Titration
Potentiometric titrations for the free ligands PYRAMPA, the 1:1 and the 1:2 Al3+:PYRAMPA systems were performed using a thermostated all-glass titration cell that was maintained at 25.0 ± 0.1 °C by an external circulating water bath. The 0.1 M KOH titrant solutions were prepared from DILUT-IT ampoules purchased from J.T. Baker Chemical Co. and kept under an ascarite CO2 scrubber in the reservoir of a Metroham model 655 autoburette. The absence of carbonate was confirmed for each KOH solution by Gran’s plots that were used elsewhere. 14–18 Stock aluminum solutions were prepared by dissolving reagent grade aluminum chloride in a 100 mM hydrochloric acid solution to prevent aluminum from the hydrolysis process illustrated in eqs 1–4.
All titrations were carried out under argon atmosphere; the ionic strength of all solutions was adjusted to 0.10 M by the addition of 1.0 M KNO3 solution. The pH was measured directly with an accumet model 25 pH meter equipped with an accu-pHast combination electrode both were purchased from Fisher Scientific. The titrations were conducted using a computer-controlled autotitrator. The autotitrator added the titrant aliquots, stirred the component of the titration cell, and monitored the pH versus time. When the pH drift fell below 0.001 pH unit per minute, the final pH was recorded and the burette was prompted for the following addition of the titrant. In a typical experimental run, the data points were collected throughout the pH range of 2.50 to 10.50. The equilibration time was preset and long enough for complete equilibration. A default preset timing was chosen to be 1000 s. Any data point that exceeded the preset time was discarded from the calculations. The concentrations of the reactants were in the order of 0.001 M or 0.002M. The titrations were conducted in which the metal:ligand ratios were 1:1 and 1:2. The stability constants were calculated using the least squares data fit that was described in detail in previous reports. 14 , 15 In all stability constants calculations for Al3+, four different hydrolysis constants for the free metal ions have been accounted for from Mesmer and Baes.3
Stability Constants Measurements
In brief, the typical titration system contains these basic components, the metal ion, M (Al3+ in that case) ligand L (PYRAMPA in that case), and hydrogen ion. Equation 9 shows the general formula of a complexation reaction. The overall stability constants for the formed complexes (βs) were expressed as shown in eq 10.

Where M
i
L
j
H
k
represents the different combination of the formed complexes, M represents the free unhydrolyzed hexaaquo aluminum ion, L represents the uncomplexed, totally deprotonated form of the ligand PYRAMPA, and H represents the free hydrogen ion concentration which is given by the pH value. Each complex has one particular (β) value. The pH data points were fit with different models where a model is a unique combination of different complexes that can form simultaneously using the least squares data regression. The value of goodness of fit (GOF) as given in eq 11 is the criterion that governs the success of the proposed model. Under the potentiometric experimental conditions, a value of .01 or less is considered as an acceptable value.
where pHobs and pHcalc are the observed and calculated pH’s, respectively, N obs is the number of observed data points, and Nv is the number of variables in the proposed model.
The ultraviolet–visible (UV–vis) spectra were collected using the computer-controlled cary-14 spectrophotometer at 25.0 ± 0.1 °C, within the concentration range of 250 μM total ligand in 0.1 M KNO3 solutions.
Concluding Remarks
By examining the values of the stability constants of the newly synthesized ligand PYRAMPA with Al3+ in aqueous solutions at room temperature, one can state that PYRAMPA is an effective aluminum binder. A series of the simple one-to-onecomplexes were formed with relatively high stabilities. For example, by comparing the Log β value for the 110 Al3+–PYRAMPA complex of 15.26 to that of Al3+–ethylenediaminetetraacetatic acid (EDTA) of 16.40 from the literature, 20 one can make the argument that the tridentate PYRAMPA is more effective than the hexadentate EDTA ligand. 20 Figure 1 is the projection view of the PYRAMPA molecule with 50% thermal ellipsoids. Figure 2 is the crystal packing of PYRAMPA via hydrogen bonding and Cl− counter ion in a beautiful nanostructure. That will have nanocharacter/structure applications that need to be assessed in a future research work. It is worth mentioning here that PYRAMPA was such an effective Al3+ binder that it hindered the formation of many aluminum hydrolytic species shown in eqs 1–4.
Footnotes
Acknowledgments
The authors would like to thank the Research Board of the University of Missouri for financial support of this work (under Grant No. S-3-40454) and NSF (MRI grant # CHE-0420497).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Research Board of the University of Missouri (under Grant No. S-3-40454) and NSF (MRI grant # CHE-0420497).
Supplemental Material
The online supplemental material is available with this article.