Abstract
Notochordal cells (NCs), abundantly found in the developing nucleus pulposus (NP), show potential for intervertebral disc regeneration because of their unique instructive and healthy matrix-producing capacity. However, NCs are lost early in life, and attempts at in vitro expansion have failed because they lose their specific phenotype. Therefore, much effort is focused on the generation of cells resembling the properties of healthy matrix-producing NP-like cells from human induced pluripotent stem cells (hiPSCs). They are considered a promising alternative for employing native NCs. Given the ongoing challenges in the field to fine-tune the differentiation protocol and obtain a high yield of mature matrix-producing cells, this study aims to build on the epigenetic memory and instructive capacity of healthy NP tissue. For this, we employed the epigenetic memory of tissue-specific hiPSCs derived from TIE2+ NP progenitor cells (NPPCs) and microenvironmental cues of decellularized porcine NC-derived matrix (dNCM), consisting of matrix components and bioactive factors to differentiate hiPSC into mature, healthy matrix-producing cells for NP repair. As a comparison, donor-matched minimally invasive peripheral blood mononuclear cell-derived hiPSCs were used. The results show that employing NPPC-derived hiPSCs instructed by natural cues provided by dNCM resulted in an increased expression of healthy phenotypic and matrisome-related NP markers. Furthermore, within this in vitro environment, differentiation of blood-derived hiPSC lines led to augmented differentiation into the hematopoietic and neural cell lineage. In conclusion, we demonstrate that hiPSCs derived from NPPCs achieve enhanced differentiation outcomes in the presence of dNCM, highlighting the potential impact of the epigenetic memory.
Impact Statement
This study highlights the influence of the epigenetic memory in hiPSCs, which primarily affects early passages. This is of clinical relevance as early but stable hiPSC passages are preferred to minimize the risk of passage-induced mutations. This work showcases possible benefits of the tissue-specific epigenetic memory and environmental cues for the particular application to the intervertebral disc. This concept may also provide avenues for other fields exploring hiPSC therapeutic strategies, as well as the modeling of genetic diseases with hiPSC models.
Keywords
Introduction
Chronic lower back pain affects the quality of life of millions of people worldwide. 1 It often starts within the core of the intervertebral disc (IVD), the nucleus pulposus (NP), 2 which is composed of a particularly proteoglycan-rich extracellular matrix (ECM) essential for maintaining a high-water content required for absorbing mechanical stresses during daily life activities. Degeneration of the avascular and nutrient-restricted NP contributes significantly to chronic back pain and consequent disability.2,3 Therefore, there is a pressing need for advanced regenerative approaches capable of restoring the NP tissue quality and functionality. 4
The onset of NP degeneration is linked to the disappearance of notochordal cells (NCs), which are crucial for NP development and maintenance (reviewed by Séguin et al. 5 ). NCs possess a vacuolated morphology and distinct functional characteristics, secreting essential ECM components including proteoglycans and collagen type II in a high ratio, and instructive stimuli promoting NP homeostasis (reviewed by Bach et al. 4 ). However, the use of NCs for therapeutic applications is limited by their absence in the adult NP, accompanied by considerable ethical challenges6,7 and loss of their specific phenotype during isolation and expansion (reviewed by Williams et al. 8 ). Human induced pluripotent stem cells (hiPSCs) have emerged as a promising alternative in regenerative medicine due to their unique ability to replicate and differentiate into various cell types, including NC-like cells. 9
In this relatively new field, the lack of specific markers that can exclusively identify human NP cells at any stage of development, from embryonic to mature NCs to non-vacuolated NP cells still capable of producing the appropriate ECM, hampers the exact determination of the hiPSC differentiation efficiency. In the end, obtaining a hiPSC-derived cell population that is able to thrive and secrete the NP-specific proteoglycan-rich ECM within the hostile degenerative disc environment 10 is most crucial from a functional perspective. Striving to improve differentiation efficacy, this field may benefit from hiPSCs potentially retaining residual epigenetic marks of their somatic cell lineage. 11 The epigenetic signatures, predominantly observed in early passages, 1 not only reflect the donor cell identity but could also influence the hiPSCs differentiation potential and lineage bias when exposed to differentiation cues.12–18
In this context, the microenvironment that the iPSCs have been derived from, that is, the tissue source, influences hiPSC differentiation capacity, as does the microenvironment that the hiPSCs encounter during differentiation. ECM-embedded cues are essential in directing differentiation and may even induce tissue-specific specification of uncommitted fetal stage precursor cells.19,20 Interestingly, ECM derived from healthy NC-rich NP tissue has already shown potential for IVD regeneration (reviewed by Bach et al. 4 ) and thus seems a promising candidate to guide hiPSC differentiation.
We explored whether we could employ the epigenetic memory of NP progenitor cell (NPPC)-derived hiPSCs (iNPPCs) 21 instructed by microenvironmental cues derived from their tissue of origin to facilitate the differentiation toward mature matrix-producing cells for NP repair. NPPC numbers decrease with IVD degeneration and are an interesting target for regenerative strategies because of their multilineage differentiation capacity.21–24 To test our hypothesis, we compared the differentiation potential of iNPPCs to donor-matched peripheral blood mononuclear cell-derived hiPSCs (iPBMCs) instructed by transforming growth factor beta 1 (TGF-β1) based on previous studies25,26 or tissue-specific decellularized NC-derived matrix (dNCM).
Materials and Methods
Three donor-matched IVD and blood samples from patients undergoing trauma surgery were collected at Inselspital Hospital of the University of Bern, Switzerland; the three male donors were aged 20, 32, and 55 years after informed consent and ethical approval (Swiss ethics approval no. 2019-00097). IVD cells were isolated from freshly collected tissue within 2 h after surgery by mild two-step digestion procedure overnight in Pronase (10-165-921-001, Roche, Basel, Switzerland) and Collagenase II (4176, Worthington, Columbus, OH). Cells were then expanded in monolayer in normoxia and in low glucose-Dulbecco’s modified Eagle medium (DMEM; 31600, Thermo Fisher, Waltham, MA), containing 10% fetal calf serum and 5 ng/mL basic fibroblast growth factor (bFGF-2) (recombinant human basic fibroblast growth factor b-FGF2; Cat. No. 100-18B 100-18B; Peprotech, Inc., London, UK) until ∼80% confluent. Cells were then trypsinized, washed, and labeled with TIE2+ antibody Tie2-PE (#FAB3131A) at 1:50 (R&D Systems, Inc., Minneapolis, MN), gated and sorted as previously described21,27 using a BD FACS ARIA III (Becton & Dickinson, Inc., Heidelberg, Germany). About 2 × 5 mL blood samples were collected (in ethylenediaminetetraacetic acid [EDTA]-containing monovettes); all materials were sent by overnight courier from Bern, Switzerland, to the IRMB, University of Montpellier, INSERM, Montpellier, France, to ensure fast processing of these cells and not involving any freezing steps.
Generation of the hiPSC lines resulting in iNPPCs and iPBMCs (Fig. 1A) was performed as published previously (https://doi.org/10.1016/j.scr.2025.103736) and shortly included in Supplementary Data and Supplementary Table S1. This study entails the use of isogenic iPSC lines to reduce variation, thus improving power. 28
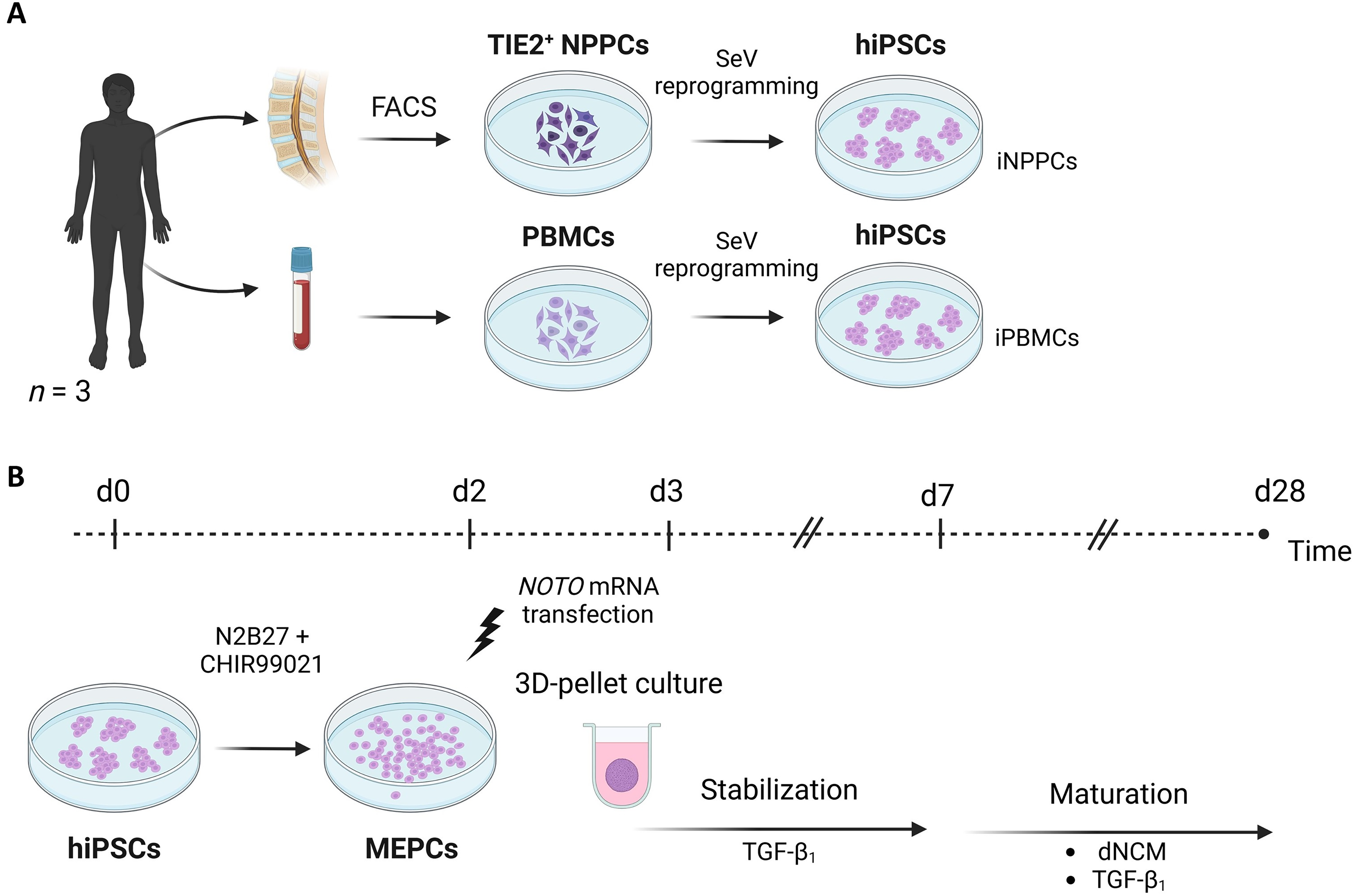
Graphical representation of the study design.
dNCM was generated according to Schmitz et al. 29 from 12-week-old piglet spines.
Experiment
Mesendoderm progenitor cells (MEPCs) of the iNPPC and iPBMC lines were generated with 48 h of CHIR stimulation following the protocol of Warin et al. 30 (Fig. 1B). Since 3D-cell culture systems better support cell phenotype and matrix production, 31 from this point, we deviated from this protocol to generate matrix-producing cells. MEPCs were resuspended in serum-free differentiation medium (high glucose-DMEM, 11965, Thermo Fisher), 1% ITS+ (354352, Corning, New York, NY), 40 µg/mL L-proline (P5607, Sigma-Aldrich), 1% NEAA, 50 µg/mL Ascorbic acid-2-Phosphate (A8960, Sigma-Aldrich, St. Louis, MO) supplemented with 10 ng/mL TGF-β1 (240-B-010, R&D Systems, Inc.) to promote 3D cell pellet formation inspired by previous maturation protocols.25,26 Furthermore, cells were reverse transfected with 6 μg/mL synthetic NOTO messenger RNA (NOTO-mRNA) (TriLink BioTechnologies, San Diego, CA) using 31.2 μL/mL of Lipofectamine RNAiMAX Transfection Reagent (13778075, Thermo Fisher) in Opti-MEM I Reduced Medium (31985062, Thermo Fisher). 30 Fifty microliters of cell suspension was added to 8 µL transfection mix per well in low-adherent 96 well plates (650970, Greiner Bio-One, Kremsmünster, Austria), resulting in 35,000 cells/well, and centrifuged at 500 g. For determining the transfection distribution within the 3D structure, cells were reverse transfected with green fluorescent protein (GFP) mRNA in separate control pellets using the same protocol.
After a 7-day stabilization period, for half of the pellets, TGF-β1 was replaced by 3 mg/mL dNCM, while the other half was continued in TGF-β1 as control maturation medium.25,26 The total differentiation period entailed 28 days (Fig. 1B), and media were replaced daily.
To check whether the hiPSC lines successfully differentiated toward cells with MEPC identity, on day 2, the typical morphology of cells leaving the colonies was evaluated, and gene expression of pluripotency, Wnt and Nodal signaling, and mesendodermal markers was determined (Supplementary Table S2). Day 2 monolayers were exposed to RNeasy® Lysis Buffer (79216, Qiagen, Hilden, Germany) supplemented with 1% beta mercaptoethanol (31350, Thermo Fisher). RNA was extracted using the RNeasy Mini kit (74106, Qiagen), and complementary DNA (cDNA) was prepared (the iScript™ cDNA Synthesis Kit (1708891, Bio-Rad, Hercules, CA) according to the manufacturer’s instructions. Quantitative PCR (qPCR) was performed using the IQ™ Syber Green Supermix (1708887, Bio-Rad) and the CFX384 Touch™ Real-Time PCR Detection System (53146, Bio-Rad) (Supplementary Table S2).
The cycle threshold (Ct) value of each target gene was normalized to the mean Ct value of the reference genes (GAPDH and YWHAZ 32 ) (i.e., ΔCt = Cttarget − Ctmean ref). The E−ΔCt-value was calculated with E indicating the amplification efficiency of the target gene.
hiPSCs stimulated by CHIR in eight-well chamber slides (80826, Ibidi, Gräfelfing, Germany) were used to determine protein expression of FOXA2 and TBXT via immunofluorescent (IF) staining (Supplementary Table S3) and imaged by confocal microscopy (Leica TCS SP8 X, Vienna, Austria).
For evaluating the reverse transfection using synthetic NOTO-mRNA, transfected and non-transfected cell pellets were collected after 24 h. RNA was extracted (RNeasy Micro kit [74004, Qiagen]), and cDNA synthesis and qPCR were performed as described above to assess endogenous and total NOTO-mRNA expression (Supplementary Table S2). Also, NOTO-mRNA- and GFP-mRNA-transfected cell pellets were collected, fixed, embedded in agarose, and, after processing, embedded in paraffin. Four micrometer sections mounted on Microscope KP+ slides (KP-3056, Klinipath, Duiven, The Netherlands) were used for IF staining for GFP (Supplementary Table S3).
On day 28, cell pellets were assessed for cell fate, identity, and differences in transcriptome. The diameter of pellets was assessed on stereoscopic microscopic images (Olympus SZ61). Assuming that the pellets were spheroid, the volume was estimated by (4/3)πr3. To further determine the proliferation and matrix-producing capacity, DNA, glycosaminoglycan, and collagen content were measured on days 7 and 28. 3D cell pellets were washed with phosphate-buffered saline and digested using papain digestion buffer containing 23.99 g/L Na2HPO4 (71640, Sigma-Aldrich), 3.722 g/L EDTA.2H2O (Titriplex; 108418, Merck, Darmstadt, Germany), 1.57 mg/mL Cysteine HCL (C7880, Sigma-Aldrich), and 250 μg/mL Papain from papaya (P3125-100, Sigma-Aldrich). To each cell pellet, 75 μL of the digestion solution was added, followed by overnight incubation at 60°C, whereafter, the digested samples were vortexed and incubated for an additional hour at 60°C.
In the digested samples, the double-stranded (ds) DNA content was measured using the Qubit® dsDNA High Sensitivity Assay Kit (Q32851, Thermo Fisher) according to the manufacturer’s instructions to estimate the number of living cells. The sulfated glycosaminoglycan content was quantified by 1,9-dimethyl methylene blue 33 binding assay at pH 3.0 using chondroitin sulfate from shark cartilage (C4348) as standard and extracting the measured absorbance at 595 nm from 540 nm. To evaluate the collagen content, hydroxyproline (HYP) content was measured with a chloramine-T assay 34 using trans-4-hydroxyproline (H54409, Sigma-Aldrich) as standard and reading absorbance at 550 nm.
For histological evaluation, day-28 pellets were processed as described for the samples collected for GFP immunofluorescence. To determine cell morphology and ECM deposition, hematoxylin and eosin (H&E) and Safranin O/Fast Green staining were performed on deparaffinized sections (Supplementary Data).
After RNA extraction and purification, bulk RNA sequencing was performed using TruSeq RNA stranded polyA (Illumina) for paired-end sequencing on the Illumina NextSeq 2000. Differential expression analysis was performed with an in-house R-script using DESeq2 (v1.38.0), taking raw read counts as input (Supplementary Data; GEO accession no. GSE273124). To confirm observed transcriptome differences on the protein level, immunohistochemical (IHC) staining was performed for lineage-specific markers, including aquaporin-6 (AQP6), aggrecan (ACAN), caveolin-1 (CAV1), cluster of differentiation (CD)24, CD45, collagen type I (COL1A1), collagen type II (COL2A1), collagen type X (COL10A1), cytokeratin 8/18/19 (panKRT), SRY-box 2 (SOX2), SRY-box 9 (SOX9), brachyury (TBXT), tyrosine-protein kinase receptor Tie-2 (TIE2), and tubulin beta III (TUBB3) (Supplementary Data and Supplementary Table S4).
Statistical analysis
All statistical analysis was performed using GraphPad Prism version 9.1.0. Normal distribution was tested by the Shapiro–Wilk test. For normally distributed data, a two-way analysis of variance (ANOVA) test determined the fixed effects, that is, treatment (e.g., mTeSR1 vs. CHIR; TGF-β1 vs. dNCM) and cell source (NPPC- vs. PBMC-derived cell lines) with Tukey’s post hoc tests. Graphs show the mean ± standard deviation. When data were nonparametric, the Kruskal–Wallis test with Dunn’s multiple comparison test was performed in a one-way fashion, and graphs show median ± interquartile range. Statistical significance was assumed for p < 0.05. To assist interpretation, the mean difference (MD) with the 95% confidence interval (CI) is provided as MD [lower CI; upper CI] for ANOVA and MD for nonparametric tests derived from GraphPad Prism.
For the bulk RNA sequencing data, negative binomial generalized linear model fitting and a paired Wald’s test were used to assess the differentially expressed genes (DEGs) using R/Bioconductor package DESeq2 version 1.42, and p-values were adjusted for multiple testing errors with a 5% false discovery rate according to the Benjamini–Hochberg method. 35 DEGs were defined as those genes showing a | log2 fold change | ≥1 in expression at an adjusted p < 0.05.
Results
iNPPCs and iPBMCs successfully differentiate into MEPCs upon CHIR stimulation
To investigate the effect of the tissue source on the differentiation capacity of the hiPSCs, iNPPCs and donor-matched iPBMCs were generated and directed toward MEPCs 30 (Fig. 2A). CHIR stimulation of iNPPCs led to a 256-fold increase of the E−ΔCt-value of the Wnt target gene LEF1 (p < 0.0001; 8.1 [7; 9.1]) and 1163-fold increase in NODAL (p < 0.0001; −10.4 [9.2; 11.6] expression compared with mTeSR1-controls (Fig. 2B). In iPBMCs, CHIR stimulation increased LEF1 by 170-fold (p < 0.0001; 7.6 [6.6; 8.6]) and resulted in a 998-fold increase in NODAL (p < 0.0001; −10.1 [8.9; 11.2]) expression (Fig. 2B). Primitive streak (TBXT) and mesendodermal marker (FOXA2) were also upregulated. In iNPPCs, this resulted in a 75-fold increase in FOXA2 (p = 0.011; 12.2) and 4339-fold increase in TBXT (p = 0.0042; 13.8) expression and, for iPBMCs, a 115-fold (p = 0.051; −10.8) and 2672-fold (p = 0.0493; 10.2) increase, respectively (Fig. 2B). CHIR stimulation downregulated pluripotency markers in both iNPPC and iPBMC lines (Supplementary Fig. S1). Overall, no significant difference was shown between the iNPPC and iPBMC lines in the markers tested.
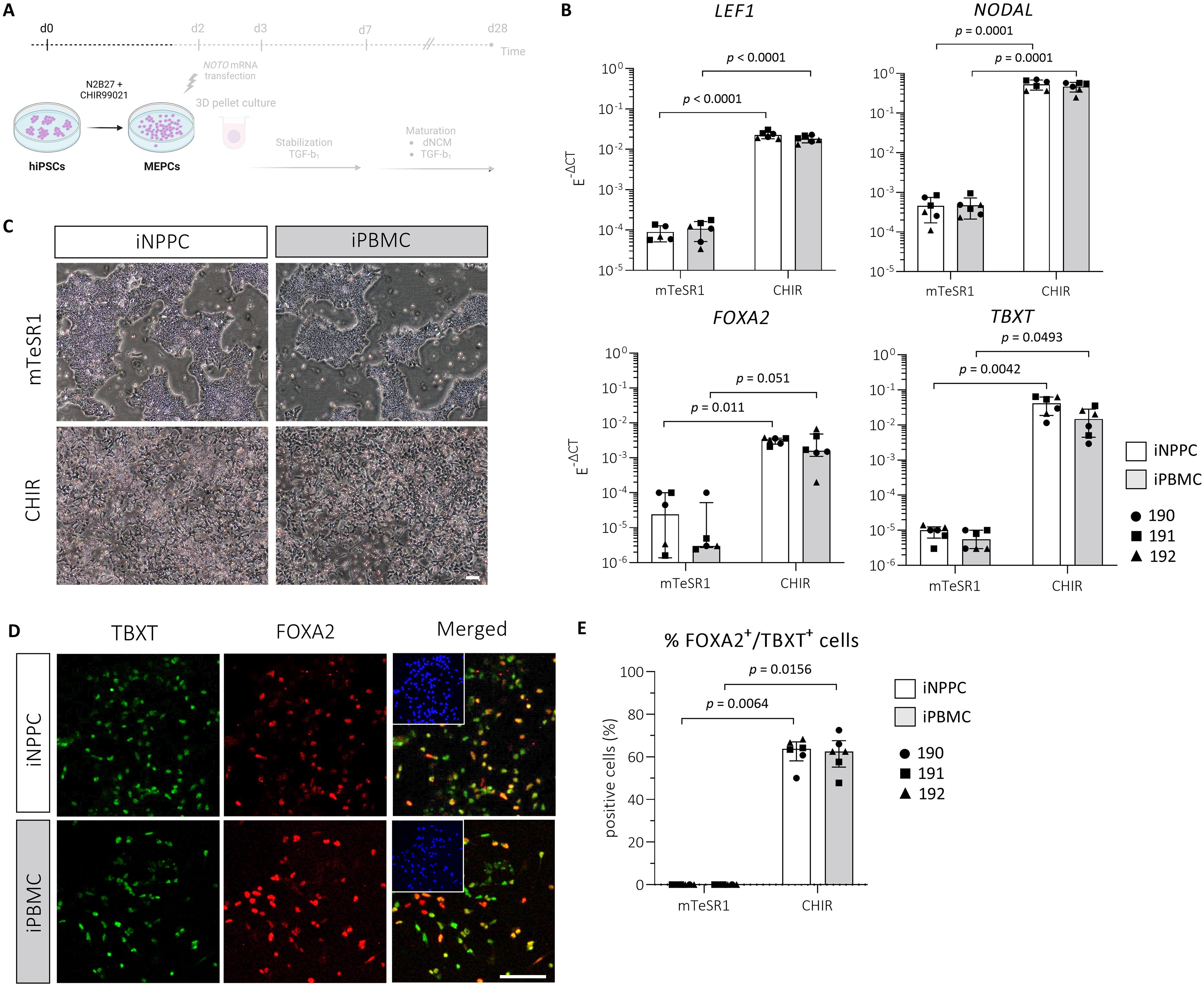
Three donor-matched iNPPC and iPBMC lines showed successful differentiation into MEPCs upon CHIR stimulation.
Accordingly, changes in cell morphology also indicated cell differentiation upon CHIR stimulation in iNPPC and iPBMC lines, shown by cells moving outside of the compact hiPSC colonies (Fig. 2C). Immunostains showed 62 ± 6% (p = 0.006; −12.5 [6.5; 19.0]) and 61 ± 8% (p = 0.016; −11.5 [6.5; 18.0]) TBXT+/FOXA2+ cells on day 2 in CHIR-treated iNPPC and iPBMC lines, respectively, compared with undetectable staining in mTeSR1-controls (Fig. 2D, E). Together, these results indicate that the hiPSC lines derived from the two tissue sources were able to obtain the MEPC identity to a similar extent.
MEPCs derived from iNPPCs and iPBMCs show successful transfection within the 3D-cell pellet structures
To drive the cells further toward the NC- lineage and support matrix production, MEPCs derived from iNPPCs and iPBMCs were reverse transfected with NOTO-mRNA on day 2 following CHIR stimulation and switched to a 3D-cell pellet culture (Fig. 3A). On day 3, total NOTO-mRNA expression increased compared with non-transfected controls, with a 6731-fold (p = 0.020; −11.8) and 2722-fold (p = 0.015; −12.2) increase in both iNPPC- and iPBMC-pellets, respectively (Fig. 3B), while endogenous NOTO was undetectable at this stage. GFP-mRNA reverse transfection showed consistent homogeneous translation of the GFP protein throughout the pellets regardless of the tissue source (Fig. 3C).
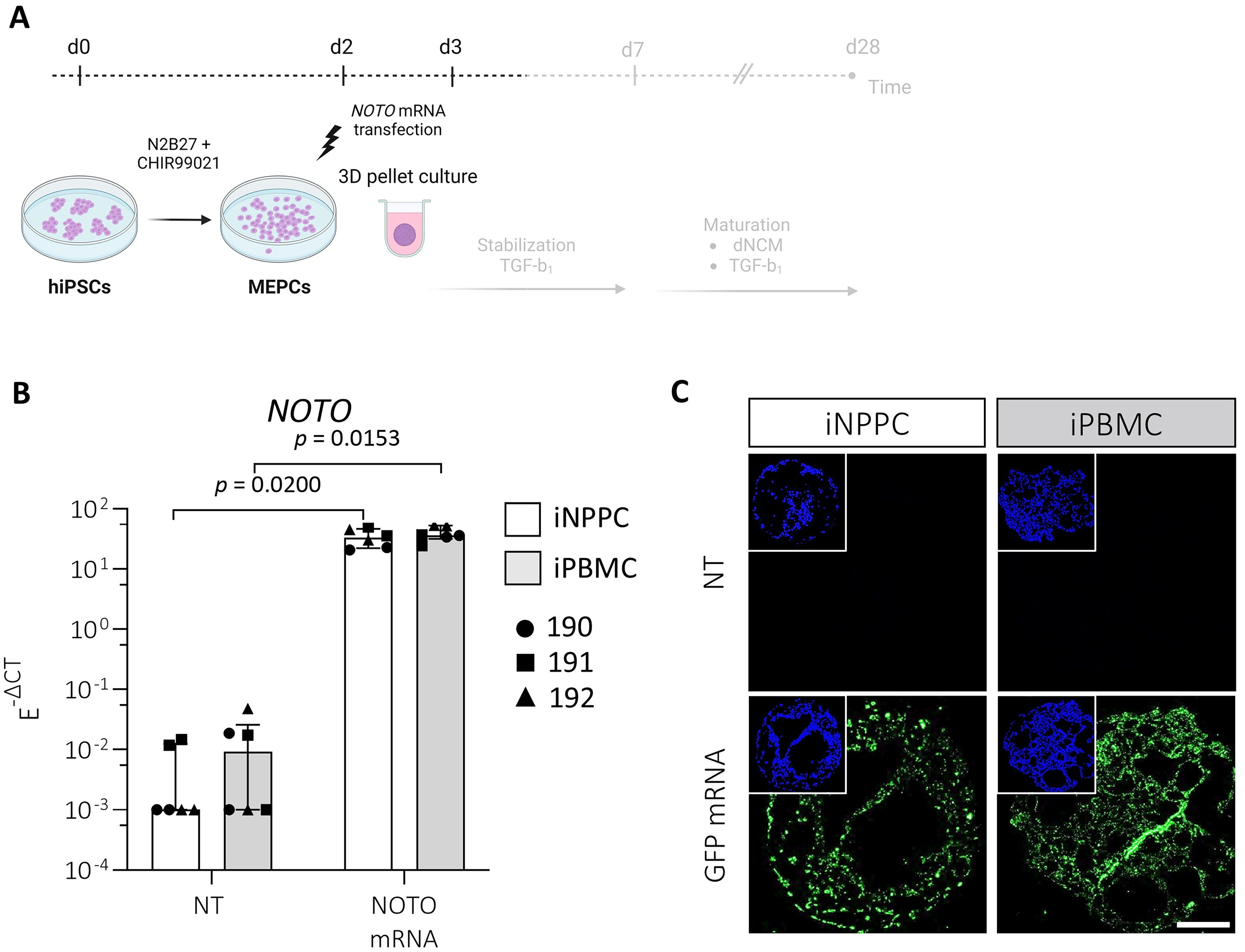
MEPCs obtained from iNPPC and iPBMC lines were successfully transfected with NOTO-mRNA and showed homogeneous GFP protein expression within the 3D cell pellet structures.
iNPPC-pellets instructed by dNCM contain the most GAG-rich ECM
To test the hypothesis that epigenetic memory of the iNPPCs would help them to respond better to the cues found in their tissue of origin, we used dNCM to mimic the healthy NP environment. Since immediate exposure to dNCM hampered MEPC assembly into 3D structures, pellets were stabilized until day 7 in serum-free media comparable with the maturation medium used in previous differentiation studies to promote matrix production.25,26 Thereafter, half of the pellets were switched to serum-free medium with dNCM replacing TGF-β1 (Fig. 4A) while the other half was maintained in TGF-β1. On macroscopic images at day 28, iNPPC-pellets were larger in size compared with iPBMC-pellets (Fig. 4B). This was quantified by estimating the pellets’ size: iNPPC-pellets had a 3-fold (p = 0.029; 11.0) and 2.8-fold (p = 0.059; 10.6) larger volume after TGF-β1 or dNCM stimulation compared with iPBMC-pellets, respectively (Fig. 4C).
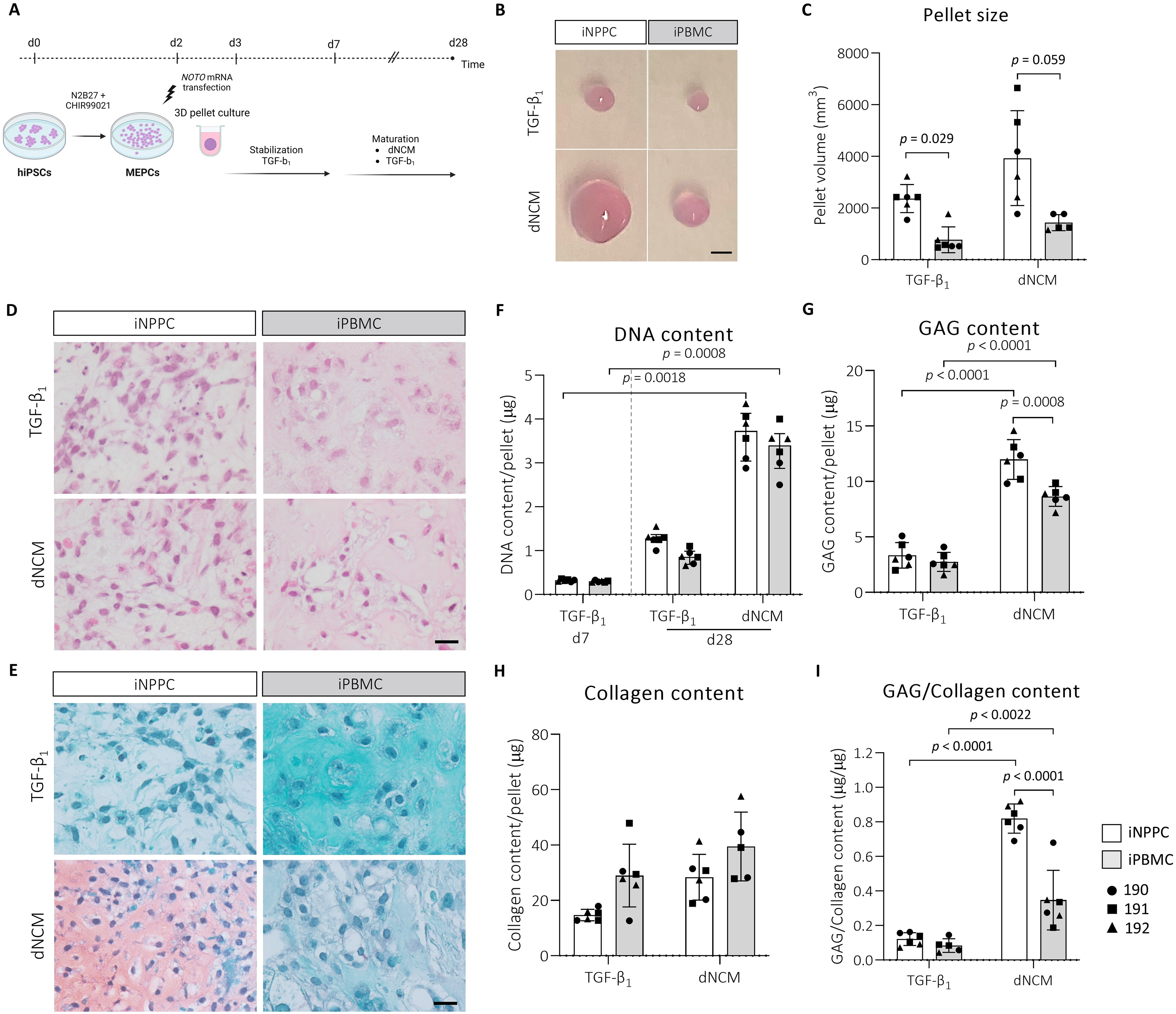
3D pellets of the iNPPCs instructed by dNCM increased in size and contained a higher GAG content compared with the iPBMC-derived pellets.
Mainly dNCM treatment induced heterogeneous iNPPC- and iPBMC-pellets, as shown by H&E-stained sections, containing large cyst-like structures, which were absent in pellets matured with TGF-β1 (Supplementary Fig. S2). iNPPC-pellets appeared to have a relatively higher cell-to-matrix ratio on histology compared with iPBMC-pellets (Fig. 4D, E). Quantifying the pellet DNA content revealed that, starting from a comparable cell density on day 7, only dNCM treatment resulted in a significantly increased DNA content on day 28 by 11.3-fold (p = 0.002; −23.4) and 11-fold (p = 0.001; −24.6) in iNPPC- and iPBMC-pellets, respectively (Fig. 4F).
On day 28, dNCM treatment augmented the GAG content of iNPPC-pellets (3.6-fold; p < 0.001; 8.6 [−10.7; −6.6]) and iPBMC-pellets (3.1-fold; p < 0.001; −5.9[−7.9; −3.8]) compared with TGF-β1 (Fig. 4G). Additionally, dNCM resulted in a significant 1.4-fold increase (p = 0.001; 3.3 [1.3; 5.4]) of the GAG content in iNPPC-pellets compared with iPBMC-pellets (Fig. 4G). This was confirmed by the observation that only sections of iNPPC-pellets instructed by dNCM showed distinct GAG-rich areas after Safranin O-Fast Green staining (Fig. 4E). The collagen content did not statistically differ between the tissue source and treatments, but the GAG:collagen ratio was significantly higher after dNCM treatment in iNPPC- (6.8-fold; p < 0.0001; −0.7 [−0.9; −0.5]) and iPBMC-pellets (4.1-fold; p = 0.0022; −0.3[−0.4; −0.1]) compared with TGF-β1 (Fig. 4H, I). Notably, dNCM-instructed iNPPC-pellets exhibited a 2.4-fold higher GAG:collagen ratio (p < 0.001; 0.5 [0.3; 0.6]) compared with iPBMC-pellets, an important functional characteristic of NCs/NPCs 36 (Fig. 4I).
RNA sequencing of iNPPC-pellets instructed by dNCM shows the highest expression of healthy phenotypic and matrisome-related markers
To further investigate the effects of the tissue source on healthy NP-related ECM, phenotypic markers, and differentiation into other cell lineages, we performed bulk RNA sequencing. Principal component analysis revealed a clear separation for the two tissue sources and treatments with minimal donor variation (Fig. 5A). DESeq2 analysis resulted in 2361 down- and 2406 upregulated genes (| log2 fold change | ≥1 and p < 0.05) and 2310 down- and 1453 upregulated genes between iNPPC-pellets compared with iPBMC-pellets when instructed by TGF-β1 or dNCM, respectively (Fig. 5B). Reactome pathway analysis, including the 500 most significantly DEGs between iNPPCs and iPBMCs instructed by dNCM or TGF-β1, revealed regulation of mainly ECM-related pathways (Supplementary Tables S5 and S6). Matrisome analysis showed that glycoproteins and collagens were the categories with the highest overlap percentage after TGF-β1 or dNCM treatment (Fig. 5C). Targeted analysis of specific ECM-associated markers showed that dNCM treatment resulted in a 2-fold increase in the expression of the most important component of the healthy ECM, ACAN, and an 8-fold increase in its linking protein-coding gene, HAPLN1, in iNPPC- compared with iPBMC-pellets (Fig. 5D; Supplementary Fig. S3A). This was confirmed on protein level, since only sections of NPPC-derived hiPSC lines instructed by dNCM showed aggrecan immunopositivity (Fig. 5E), which is also in line with the Safranin O Fast Green staining (Fig. 4E). dNCM treatment induced a 4-fold upregulation of COL2A1 in iPBMC- compared with iNPPC-pellets, whereas on protein level, dNCM treatment resulted in abundant collagen type II staining in both iNPPC- and iPBMC-pellets in contrast to TGF-β1 treatment (Fig. 5D; Supplementary Fig. S3A). COL6A1, present in the human NC-matrisome, 8 was significantly upregulated in iNPPC-pellets instructed by TGF-β1 (1.7-fold) and dNCM (4-fold), compared with iPBMC-pellets (Fig. 5D). TGF-β1 treatment induced the expression of fibrotic matrix marker COL1A1 in iPBMC-pellets (Fig. 5D; Supplementary Fig. S3A) confirmed by IHC (Fig. 5E). Furthermore, in iPBMC-pellets, dNCM treatment resulted in the highest MMP3 expression, whereas TGF-β1 treatment resulted in the highest expression of MMP19 and ADAMTS2 (Fig. 5D), all involved in ECM degradation.
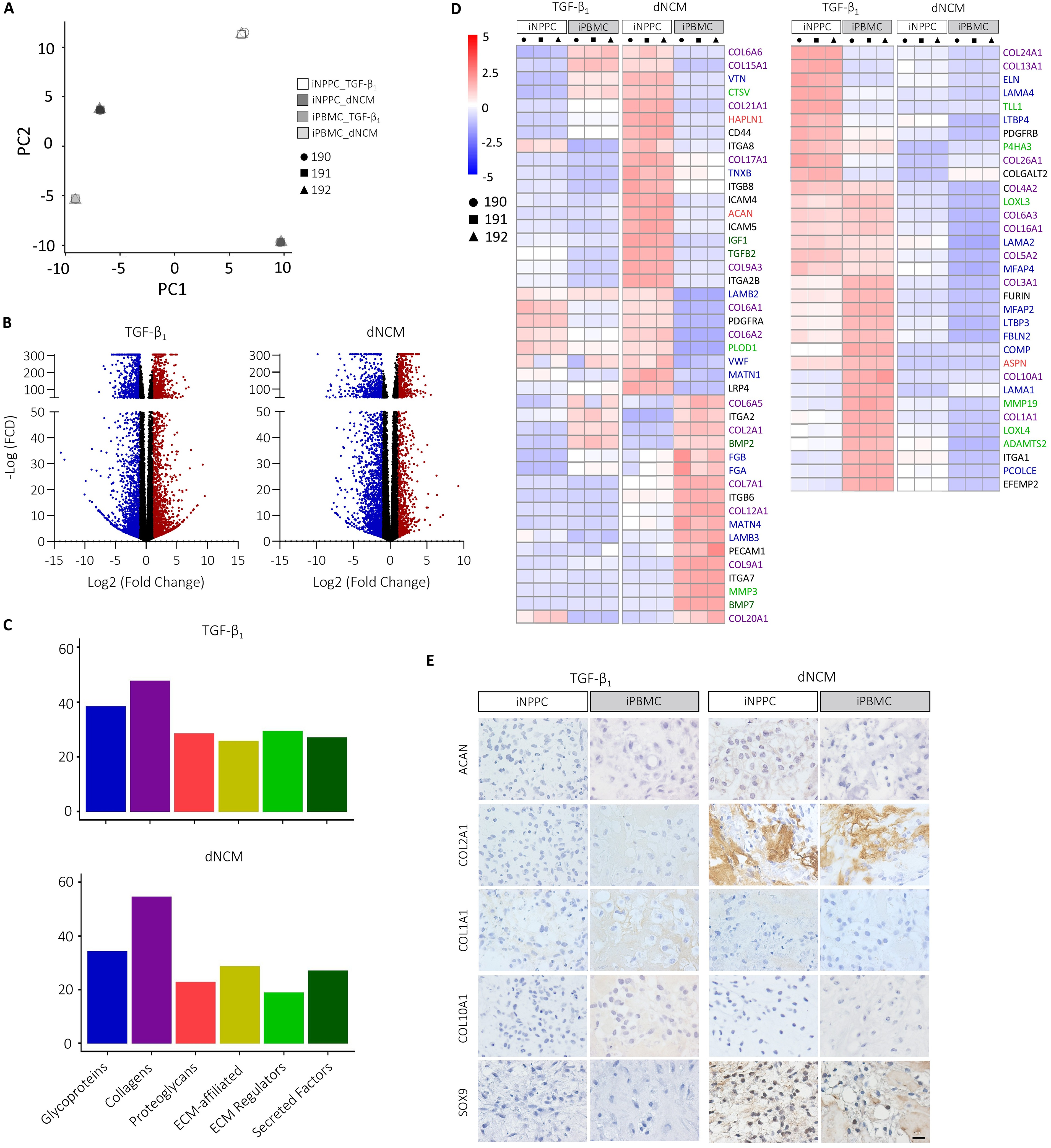
The tissue source and dNCM treatment affected the matrisome at the transcriptional and protein levels.
Notably, dNCM treatment upregulated the expression of TGFB2 (4-fold) and IGF1 (8-fold) in iNPPC-pellets compared with iPBMC-pellets, whereas it downregulated BMP2 (2-fold) and BMP7 (8-fold) expression, indicating less hypertrophic differentiation as confirmed by decreased gene expression of COL10A1 and collagen type X immunopositivity (Fig. 5D, E; Supplementary Figs. S3A and S4A, B). Transcription markers SOX9 and RUNX2 downstream of these pathways were not differentially expressed in iNPPC-pellets compared with iPBMC-pellets treated with dNCM at this timepoint (| log2 fold change | <1; Supplementary Figs. S3A and S4A) and pellets derived from both tissue sources instructed by dNCM showed nuclear SOX9 immunopositivity in contrast to TGF-β1 treatment (Fig. 5E).
There are no specific markers that uniquely and exclusively identify the cells of the human NP in the different stages of development. Therefore, we employed a panel of recommended markers for a healthy matrix-producing phenotype4,8 together with other markers previously found in human NC-rich NP tissue/matrisome4,8,30,38–40 and demonstrated differential expression depending on the tissue source and treatment. Most abundant expression of these markers was detected after dNCM treatment compared with maturation using only TGF-β1, indicating that overall dNCM outperformed TGF-β1 treatment (Supplementary Fig. S5). Gene and protein expression of TBXT were not detected on day 28. dNCM treatment induced the highest expression of GJA1, CAV1, KRT19, AQP6, TEK, and CD44 in iNPPC-pellets, while it resulted in the highest expression of CD24 in iPBMC-pellets (Fig. 6A; Supplementary Fig. S3B). In contrast, CD24 immunopositivity was mainly detected in the nuclei of iNPPC-pellets instructed by dNCM (Fig. 6B), which could be explained by the heterogeneity of the pellets and the limitation of bulk RNA sequencing, providing average expression of all the cells within the pellets. In line with transcriptomic analysis, caveolin-1 (CAV1) and aquaporin-6 (AQP6) immunopositivity were only detected in iNPPC-pellets treated with dNCM. In contrast, TIE2 and cytokeratin 8/18/19 (panKRT) expression was observed in all cell lines and conditions, but with most panKRT expression detected after dNCM treatment (Fig. 6B). Altogether these data indicate that iNPPCs instructed by dNCM expressed the highest healthy phenotypic and ECM markers.
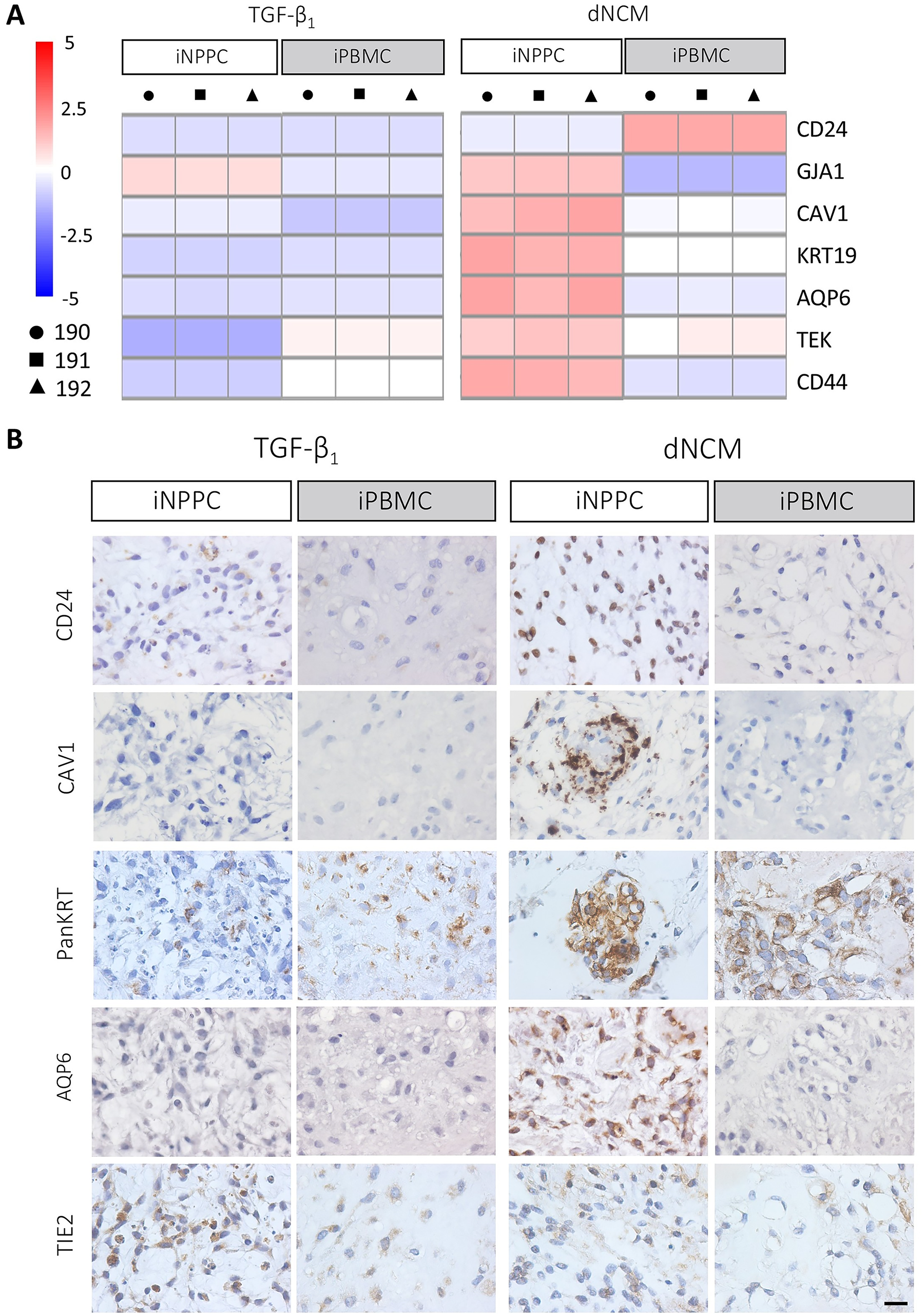
iNPPC lines instructed by dNCM showed the most expression of healthy phenotypic cell markers.
iPBMC-pellets show increased differentiation into other cell lineages
iPBMC-pellets instructed by dNCM showed the highest expression of pluripotency markers ALPL, L1TD1, and OCT4 (Supplementary Fig. S6), indicating that more cells remained in their pluripotency state than in iNPPC-pellets. Pathway analysis showed differences in the regulation of interferon- and cytokine-signaling and neural cell adhesion molecule between iNPPC- and iPBMC-pellets instructed by dNCM (Supplementary Table S5). When mapping panels of DEGs related to the neural and immune/hematopoietic41–45 system, we observed that dNCM treatment induced differentiation toward these lineages to a higher extent than TGF-β1, predominantly in iPBMC-pellets (Fig. 7A, B; Supplementary Fig. S3C). The expression of pan-neuronal marker TUBB3, which is expressed almost exclusively in neurons and is induced early upon the in vitro differentiation of hiPSCs into neurons, 46 was confirmed on the protein level in iPBMC-pellets matured with dNCM (Fig. 7C). Additionally, iPBMC-pellets showed the highest gene expression of CD45, one of the most abundant leukocyte cell surface glycoproteins and exclusively expressed by cells of the hematopoietic system, which was also confirmed on the protein level 47 (Fig. 7B, C). SOX2, which encodes a transcription factor that is expressed by pluripotent stem cells but also neural and hematopoietic stem cells,48,49 was present both on gene and protein level (nuclear stain indicated by red arrowheads) in iPBMC-pellets instructed by dNCM (Fig. 7A–C).
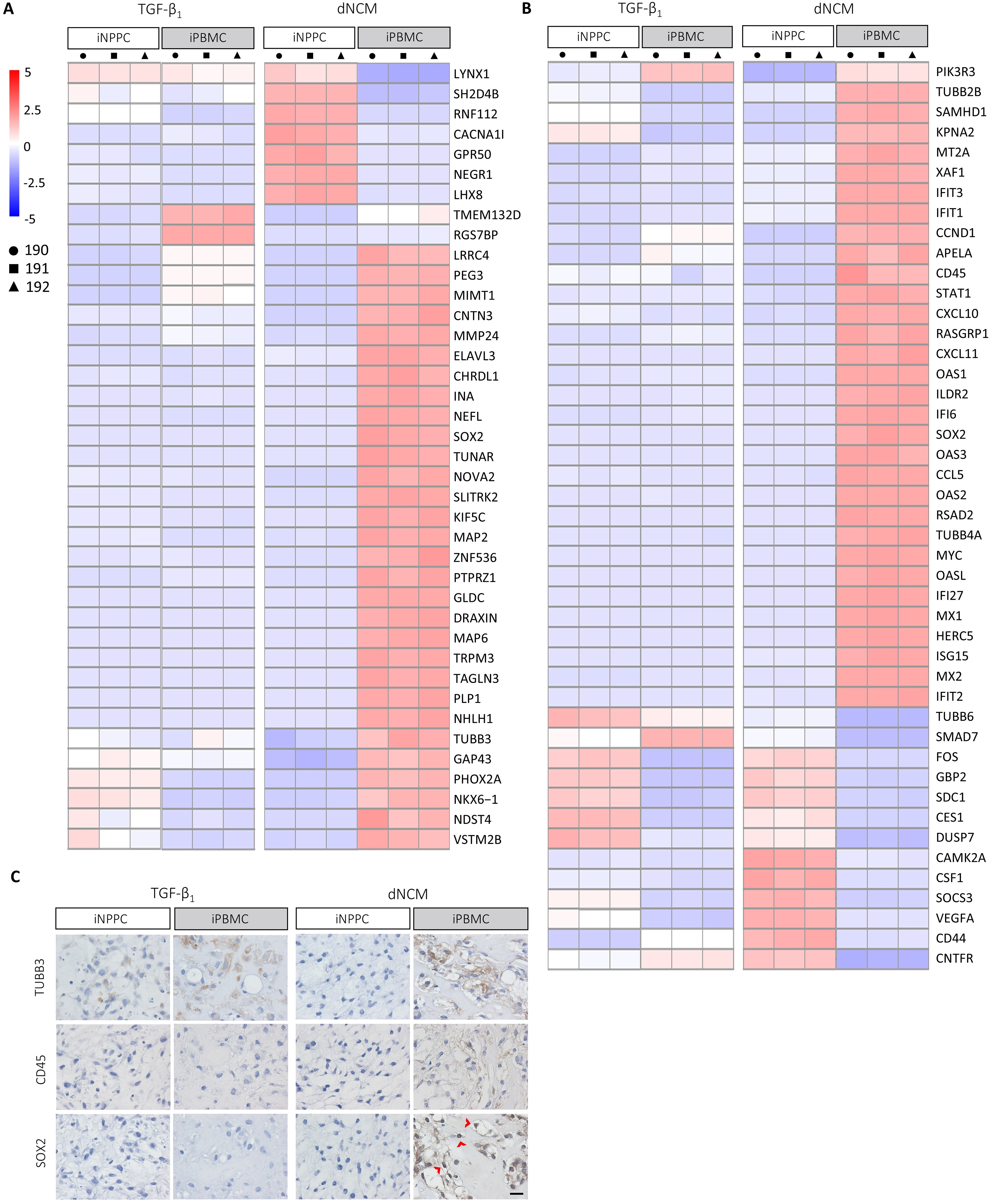
iPBMC-pellets instructed by dNCM showed differentiation toward neural and hematopoietic lineages. Heat map of DEGs related to
Discussion
Key results of this study show that differentiated hiPSCs remained considerably heterogeneous, but the tissue-specific memory enabled the iNPPCs to respond more desirably to the healthy NP environmental cues provided by dNCM than donor-matched iPBMCs, as indicated by enhanced functional and phenotypic properties and tempered differentiation into other cell lineages.
Both iNPPC and iPBMC lines showed successful and comparable differentiation into MEPCs upon CHIR stimulation, which is favorable to the emergence of the notochordal lineage. 50 Additionally, the percentage of TBXT+/FOXA2+ MEPCs generated from these patient-derived cell lines was consistent with other hiPSC lines using the same protocol. 30 However, the hiPSC tissue source affected maturation of the MEPCs, resulting in increased expression of key healthy phenotypic and matrisome markers compared with donor-matched iPBMC lines. This aligns with studies in other fields, for example, beta cell-derived hiPSCs showed enhanced ability to differentiate into functional insulin-producing cells 17 and cardiac progenitor cell-derived hiPSCs into beating cardiomyocytes. 12 Interestingly, the latter study used donor-matched fibroblast-derived hiPSCs as a comparator but only observed the differences in early passages and found that the epigenetic memory of the cardiac-derived hiPSCs was eliminated over time (>40 passages), negating any differences in vitro or in vivo. 12 In the present study, we used a relatively early but stable passage (∼p15–30), which is recommended for clinical application to minimize the risk of passage-induced mutations,51,52 increasing the possibility of an effect of the epigenetic memory of the hiPSC tissue source.
Differentiating iPSCs into a pure population of mature and functional cells is still a challenging process requiring a balance between essential biochemical and physical cues. dNCM might provide this ultimate combination because of the bioactive factors it contains as well as ECM components. 4 Mimicking the healthy NP microenvironmental cues using dNCM resulted in improved differentiation into functional matrix-producing cells, compared with instructing the cells with one specific growth factor, especially in the iNPPCs. In line with this, the unique mix of cues provided by non-decellularized NC-rich porcine NP tissue already showed the potency to stimulate the NC phenotype in a commercial human fibroblast-derived iPSC line, although they did not quantify the proteoglycan and collagen content, lacking proper functional characterization of the differentiated cells. 53 Interestingly, similar to our study, the cells lost their TBXT expression after maturation in a 3D culture model, which is considered a key NC marker. 8
The interpretation of the expression of phenotypic cell markers is context-dependent and therefore challenging, especially in in vitro studies using pluripotent stem cells with the potency to differentiate into almost all somatic cell types, stressing a limitation of this model. For example, TIE2, a surface marker of NPPCs,21,22 is also expressed in endothelial, hematopoietic, and neural stem cells. 54 Similarly, SOX9 is a well-known chondrogenic transcription factor, 55 but it also differentiates cells derived from all three germ layers into a large variety of other specialized tissues and organs. 56 The lack of exclusive markers for human NPCs from vacuolated embryonic to mature NCs to nonvacuolated NPCs still capable of producing the appropriate ECM hampers the exact determination of the differentiation efficiency.
From a potency perspective, a head-to-head comparison with native human NCs is lacking due to their scarcity. We focused on the matrisome as a functional readout imperative for regenerative capacity. iNPPCs instructed by dNCM expressed multiple healthy matrisome-related genes. Although the HYP assay does not distinguish between the different types of collagens, IHC showed no collagen type I and X deposition but only collagen type II as well as aggrecan immunopositivity in iNPPC-pellets, the two most important ECM components of the healthy NP. Altogether, at the biochemical level, iNPPCs instructed by dNCM obtained the highest GAG:collagen ratio, which is an essential functional feature of healthy NPCs for attracting water to build sufficient intradiscal pressure and restore the biomechanical function of the IVD. 36 However, as these characteristics were not accompanied by the appearance of the typical vacuolated NC morphology and because of the lack of TBXT expression, the obtained cells should not be identified as NCs but rather healthy matrix-producing NP-like cells.
iPBMC-pellets showed differentiation into the hematopoietic and neural cell lineages to a higher extent when instructed by dNCM, with augmented gene expression of multiple neuronal markers, compared with iNPPC-pellets. This was corroborated by expression of TUBB3 only in iPBMC-pellets, a pan-neuronal marker also expressed during in vitro differentiation of hiPSCs into neurons 46 while maintaining higher expression of pluripotency markers. Interestingly, in the study of Warin et al., 30 single-cell RNA sequencing also revealed variation in cell lines derived from different tissue sources, with iPBMC-derived cell lines showing enhanced expression of markers of the neural lineage, for example, TUBB3, as well as SOX2 expression. Hence, the iPSC tissue source in combination with the microenvironmental stimuli within a specific experimental model affects lineage commitment. Although the parameters we assessed on days 2 and 7 did not reveal significant differences between the tissue sources, RNA sequencing at these time points could further support that the variation in cell population we observed on day 28 was primarily due to the different response of the iNPPCs and iPBMCs to the instructive stimuli within the 3D environment. Follow-up studies, exploring the underlying epigenetic differences between these donor-matched iPSC lines derived from two different tissue sources, could provide valuable insights into the retainment of their epigenetic memory upon reprogramming and associate this with the differential response to the instructions provided by dNCM.
From a clinical translation perspective, the application of iNPPCs entails more challenges involving additional ethical considerations, cell sorting steps, and quality controls during their production, as opposed to iPBMCs, which can be collected in a minimally invasive manner and in larger quantities. Hence, evaluating the potential of this hiPSC-based treatment approach in models that mimic the in vivo situation as closely as possible is of utmost importance to define if the advantage of the NPPC epigenetic signature outweighs the challenges. This is especially relevant for the IVD as it is avascular with limited nutrient supply and waste removal, 10 creating a challenging environment for the cells to survive in. Since NCM already proved its potential to facilitate tissue repair in vivo after intradiscal injection, 4 employing dNCM as an anabolic and instructive stimulus for resident NPCs, as well as injecting hiPSC-derived NPC-like cells, represents a promising approach to obtain long-term restoration of NP homeostasis and functionality. Another important point required to facilitate clinical application is purifying the obtained population, for example, by removing remnant undifferentiated iPSCs based on the lectin probe rBC2LCN, which binds specifically to their cell surface, 57 and enriching for the desired differentiated cells by unraveling cell surface markers linked to the NC phenotype, for example, CD166 and CD109. 30 This study was performed using isogenic iPSC lines, which is considered to have a higher power than case–control studies. 28 However, from a translational perspective, future studies incorporating more donors and iPSCs derived from various tissue sources are required to corroborate the present findings.
In conclusion, we demonstrate that NPPC-derived hiPSCs instructed by a mix of natural cues provided by dNCM allow for an improved differentiation outcome, stressing the potential influence of the epigenetic memory. Follow-up studies, focusing on understanding how the in vivo microenvironmental cues will direct differentiation, maturation, and possible off-target differentiation, are imperative to refine the differentiation protocol, also taking into account the hiPSC-derived cell functionality and survival within the challenging degenerated IVD environment.
Authors’ Contributions
L.T.L.: Conceptualization, design, acquisition of data—lab work, data analysis, interpretation of data, drafting the work, and revision of the article. D.W.L.P.L.: Conceptualization, design, data analysis, interpretation of data, and revision of the article. R.D.: Acquisition of data—lab work, data analysis, interpretation of data, and revision of the article. F.M.R.: Design, data analysis, interpretation of data, drafting the work, and revision of the article. C.C.W.: Acquisition of data—lab work, data analysis, interpretation of data, and revision of the article. M.S.: Acquisition of data—lab work and data analysis. P.B.: Revision of the article. K.I.: Conceptualization and revision of the article. O.M.: Acquisition of data—lab work, data analysis, interpretation of data, and revision of the article. A.C.: Conceptualization, interpretation of data, and revision of the article. B.G.: Interpretation of data and revision of the article. J.-M.L.: Conceptualization and revision of the article. M.A.T.: Conceptualization, design, data analysis, interpretation of data, drafting the work, and revision of the article.
Footnotes
Acknowledgments
The authors acknowledge the Utrecht Sequencing Facility (USEQ) for providing sequencing service and data. USEQ is subsidized by the University Medical Center Utrecht and the Netherlands X-omics Initiative (NWO project 184.034.019). The authors would like to thank Lianne Snel and Floor Verdonk (Utrecht University) for helping with the execution of experiments and Tara Schmitz (Eindhoven University of Technology) for kindly providing us with the dNCM. They further acknowledge the Orthopedic surgeons that helped to acquire the primary cells of trauma patients, namely, Sonja Häckel, Christoph Albers, Department for Traumatology and Orthopaedics, Insel University Hospital, Bern, Switzerland. Furthermore, the authors thank the Flow Cytometry and Cell Sorting Core Facility of the Department for BioMedical Research for providing cell sorting of TIE2+ live cells. They thank Julien Guerrero for isolating the primary TIE2+ cells. Furthermore, they would like to thank the group of Femke Broere for the advice on the immune-related markers and the group of Niels Eijkelkamp and Paul Coffer for kindly providing the TUBB3 and CD45 antibodies, respectively. Figures 1–![]() were all (partly) created with BioRender.com.
were all (partly) created with BioRender.com.
Funding Information
This project has received funding from the European Union’s Horizon 2020 research and innovation program under grant agreement no. 825925 named “iPSpine” (https://ipspine.eu and ![]() ) and the Dutch Arthritis Society (LLP22).
) and the Dutch Arthritis Society (LLP22).
Disclosure Statement
K.I. reports a relationship with NC Biomatrix that includes Chief Scientific Officer of NC Biomatrix, a company developing dNCM as a commercial product, which holds a license to the IP for its production process (United States Patent Application 20190022278). The other authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this article.
Supplemental Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
