Abstract
Objectives
Bacterial resistance to current antibiotics is mounting at an alarming rate, making many conventional antibiotics ineffective and posing a serious worldwide health threat; thus, there is a crucial need for exploration of novel antimicrobial leads from unusual sources, particularly the marine environment, which produces unique chemical compounds with promising therapeutic potential. This study aims to purify antimicrobial compounds from the ethyl extract of the fungus Alternaria rosae DY-79, characterize their chemical structures, and evaluate their antimicrobial effects against pathogenic microbes.
Methods
The antimicrobial fraction of the fungal culture extract was partitioned, and purified, yielding two unreported compounds, alternoic acids A (
Results
Alternoic acids A (
Conclusion
Compound
Introduction
Marine microorganisms are counted as a valuable source of drug candidates, potential drugs and agrochemicals.1-4 Fungi obtained from different marine environments have been investigated due to their ability to thrive in diverse marine habitats, regardless of environmental conditions.5,6 These conditions force the fungi to produce secondary metabolites with unique architecture, revealing a wide range of diverse structural motifs and promising biological effects.5,6 Fungi from the marine environment have been isolated and reported from invertebrates, algae, sediments, and many other marine environments. 7 They can survive in harsh conditions, such as low oxygen levels, low or absence of light, and high pressures, which may lead to the development of unique biological secondary metabolites.8-10 Therefore, these fungi are believed to be an interesting resource of diverse scaffolds needed for the process of drug discovery.8-10 Genomic studies have revealed that they harbor numerous “silent” biosynthetic gene clusters, which have the potential to produce far more compounds than can be cultivated using standard laboratory methods.11,12 This presents a major opportunity for the discovery of new drug precursors.11,12
A recent review in 2024 reported on the analysis of 56 newly isolated natural antibacterial products from marine fungi including polyketides (41.7%), peptides (22.2%), and alkaloids (19.4%). Most of these compounds comes from the genera Penicillium (33.3% of samples) and Aspergillus (29.2%), highlighting their importance as a significant source of bioactive compounds. 13 The review specifically notes the potent antibacterial activity of 16-epiascomylactam B. 14 By detailing these recent discoveries, the review underscores the critical role of marine fungi as a “gold mine” for structurally novel antibiotics needed to address the global health crisis of antimicrobial resistance. 13
Although marine-derived fungi have received less attention compared to their terrestrial counterparts, they have increasingly become recognized as a valuable source of secondary metabolites for pharmaceutical discovery. Considerable progress in this area underscores their importance in the field of natural product research.15-17 Marine organisms produce a diverse and often structurally intricate array of natural products, and the rate of discovery of new chemical entities from marine environments has steadily risen over the last ten years. 18 These marine-derived compounds belong to a wide variety of molecular groups, such as polyketides, peptides, terpenes, phenols, acetogenins, alkaloids, and volatile halogenated hydrocarbons. 19 As a result, natural products have consistently proven to be a critical resource for drug discovery, both in recent years and for the foreseeable future. 20 They are regarded as indispensable tools for pharmacologists and biologists, serving as leading therapeutic options for managing hypercholesterolemia, inflammatory conditions, microbial infections, cancer, as well as for tissue transplantation procedures.21,22
Beyond fungi, marine-derived Actinomycetota have also proven to be a rich source of potent anti-MRSA compounds. Chromomycins A2 and Ap from marine sediment Streptomyces microflavus exhibited MICs as low as 0.05 μM, while lactoquinomycin A from Streptomyces bacillaris showed MIC 0.13 μM and acts via DNA intercalation. Weddellamycin from an Antarctic deep-sea Streptomyces sp. achieved MIC 0.23 μM. Additional leads include citreamycin θA and θB (MIC 0.25 μM) from Streptomyces caelestis, the medermycin analogue G-15F from a sponge-associated strain, and strepoexipimycin D (MIC 0.55 μM) from marine sediment Streptomyces sp. XMA39. These examples underscore the value of marine bioprospecting for novel anti-MRSA scaffolds, complementing the contributions from marine fungi. 23
Members of the genus Alternaria are ubiquitous in the environment, with spores commonly present in air and soil worldwide. 24 Beside their ecological existence, exposure to Alternaria spores can trigger asthma in sensitive individuals as well as allergic reactions. Individuals of this genus represent an important source of phytotoxins, which can be harmful to their host organisms but exhibit diverse bioactivities. 25
Fungi belong to Alternaria are widely distributed in nature, functioning as saprophytes, endophytes, and plant pathogens (including those affecting marine algae).26-29 Fungi of the genus Alternaria are known for producing a vast number of structurally diverse compounds, such as dibenzo-α-pyrones, 30 terpenoids, 31 and polyketides.32,33 These compounds exhibited several biological effects, including notable antibacterial effects,30,34 anti-inflammatory,31,35 acetylcholinesterase inhibitory, 31 and cytotoxic effects.33,36
A recent review summarized the chemical and biological diversity of Alternaria-derived metabolites. Over the past two decades, marine-derived Alternaria fungi, primarily isolated from Chinese coastal waters, have proven to be a productive source of over 300 structurally diverse compounds, spanning nitrogenous compounds, pyranones, quinones, terpenoids, steroids, and phenolics, with notable compounds like alternariol and its derivatives exhibiting broad antibacterial, antifungal, antiviral, immunomodulatory, and anticancer effects, approximately 56% of which demonstrate significant bioactivities, highlighting their substantial potential for drug discovery and biotechnology. 37
As part of our constant work to recognize bioactive compounds from fungi obtained from the Red Sea,37-39 the antimicrobial organic extract of the culture of the fungus Alternaria rosae DY-79, which was recovered from Red Sea sediment, was selected for further chemical examination. Partition of the ethyl acetate extract of the fungal culture and final purification of the compounds from the antimicrobial fractions afforded the new compounds, alternoic acids A and B (
Material and Methods
Statistical Analysis
All antimicrobial tests (Disk diffusion and MICs value determinations) were conducted in triplicate, and the results reported as mean ± SD, analyzed using two-way ANOVA followed by Bonferroni post-tests to compare replicate means using GraphPad Prism 5®.
General Experimental Procedures
Optical rotation measurements were carried out at 25 °C on a JASCO DIP-370 digital polarimeter using the sodium D emission line (589 nm). Bruker Avance (DRX 800) MHz spectrometer was used to acquire the NMR spectra. High-resolution electrospray mass spectrometric data [(+)-HRESIMS] were acquired on a Micromass Q-Tof system equipped with a leucine-enkephalin lock-spray source, employing the ion at m/z 556.2771 [M + H]+ as the calibration reference. HPLC analysis was carried out on a C18 column (4.6 × 150 mm, 2.5 μm, Waters Atlantis®, Massachusetts, USA), with a CH3CN/H2O gradient as the mobile phase, monitored at 220 nm and a flow rate of 2.0 mL/min.
Purification of the Fungus Alternaria rosae DY-79
The sediment sample used for isolation of the fungal strain was collected at a depth of 23 meters in a sterile bag using SCUBA diving off Al-Lith (N019o56′31.6″ E040o08′17.3″) at the Saudi Red Sea. For enrichment cultivation, the sediment was inoculated under sterile conditions into three different sterile solid media: malt agar (MA), Czapek-Dox yeast agar (CZY), and Sabouraud dextrose agar (SDA). The media were supplemented with 0.25% chloramphenicol and 2% NaCl. After sealing with parafilm, the dishes were incubated at 30 °C until fungal colonies with distinct morphologies emerged. Fungal strains were isolated by sub-culturing onto fresh solid media, repeating the process until pure isolates were found.
Identification of Fungus Alternaria rosae DY-79
Since the fungal strain was obtained from the Czapek-Dox yeast agar (CZY) plates, the pure isolate (Figure 1) was cultured in CZY medium at 28 °C for 5 days. Collection of mycelia was achieved using filtration under vacuum and gently drying between paper towels. The mycelial mass was powdered after freezing in liquid nitrogen. The Mini Kit (QIAamp DNA) was applied for extraction of the Fungal DNA. The amplification of the fungal ITS-rDNA fragment of the fungal isolate, Alternaria rosae DY-79 was performed as previously reported.
40
The fungal ITS-rDNA sequence of Alternaria rosae DY-79 was compared with associated sequences available in the database of NCBI. Using Clustal X (version 1.83), the achieved ITS-rDNA sequences were edited and aligned with the top n-BLAST hits from GenBank,
41
followed by manual refinement in BioEdit software.
42
Calculation of the composition of the base of fungal sequences was achieved using the MEGA5 program.
43
The sequence analysis of the fungal isolate DY-79 displayed 100% sequence similarity with Alternaria rosae KSPM 134 (NCBI: PX457691.1) in BLAST. The sequence of the fungal isolate DY-79 was submitted in the GenBank under Accession Number NCBI: PZ193169 on 24 March 2026. Colony of the sediment-derived fungus Alternaria rosae DY-79
Preparation of Culture of the Fungus Alternaria rosae DY-79
The fungus was cultured on solid CZY medium until growth was complete. Spores were then transferred into ten flasks (2.5 L, each) with 1 L of CZY liquid medium. The flasks were incubated and shaken under the same conditions for 14 days. To halt fungal growth, 100 mL of ethyl acetate was poured to each flask. The mixture was then filtered under vacuum to discrete broth from mycelia. The broth was further partitioned against ethyl acetate (10 × 1 L), and the followed by evaporation under vacuum, to yield a dried residue weighing 1.0081 g.
Purification of Compounds 1 and 2
The residue (1.0081 g) was partitioned on ODS VLC using H2O-MeOH gradients starting from H2O (100%) through MeOH (100%) with 20% increase in MeOH each fraction to afford 6 fractions. The fraction eluted with 60% MeOH (437.4 mg) was subjected on a Sephadex (LH-20) column eluted with MeOH-H2O (1:1), yielding 7 fractions. The antimicrobial-active fraction (Fr. F, 176.4 mg) was then separated using a Sep-Pak C18 Cartridge (Waters, 10 g) eluted with H2O-MeOH gradients starting from H2O (100%) through MeOH (100%) with 20% increase in MeOH each fraction to afford 6 fractions. Fraction eluted with 80% MeOH (129.7 mg) was purified on C18 column using a gradient system from 10% CH3CN to 60% CH3CN over 60 minutes to yield
Evaluation of the Purity of the Purified Compounds 1 and 2
The purity of compounds
Spectroscopic Data of Compounds 1 and 2
Alternoic acid A (
Alternoic acid B (
Evaluation of the Antimicrobial Effects of the Compounds
Disk Diffusion Assay
The antimicrobial effect of the compounds was assessed in vitro using a standard disc diffusion assay, following established methodologies reported in the literature.44-50 The test panel consisted of Staphylococcus aureus (ATCC 43300; methicillin-resistant), Escherichia coli (ATCC 35218), and Candida albicans (ATCC 76615). Microbial suspensions were prepared to match a 0.5 McFarland turbidity standard and spread uniformly across Mueller–Hinton agar. Disks containing the test compounds (50 μg) were positioned on the seeded media, alongside solvent controls to verify that the vehicle had no intrinsic effect. Ciprofloxacin served as the antibacterial benchmark, whereas clotrimazole was used as the antifungal comparator. After incubation, antimicrobial performance was evaluated by measuring the diameters of the resulting growth-inhibition zones, reported as mean values from duplicate trials.
Evaluation of the Minimum Inhibitory Concentrations (MICs)
The minimum inhibitory concentrations (MICs) of the tested compounds were assessed using a standardized broth microdilution approach consistent with CLSI guidelines (M02 and M07).44,45 Serial dilutions of each compound were prepared in Mueller–Hinton broth to generate a broad concentration range for evaluation, and these dilutions were dispensed into microplate wells along with appropriately adjusted microbial inocula. Each assay included both growth and sterility controls to ensure data reliability, and ciprofloxacin and clotrimazole were incorporated as reference antimicrobial agents within their established concentration ranges. Following incubation under recommended CLSI conditions, microbial growth was quantified spectrophotometrically at 600 nm, and the MIC was identified as the lowest concentration at which no detectable growth occurred.
Results
Structural Determination of Alternoic Acid A (
1
)
The molecular formula (C12H18O4) of compound Chemical structures of alternoic acids A ( NMR Spectral Data of Alternoic Acid A (

The COSY spectrum of 1H-1H COSY, 1H-13C HMBC and 1H-1H NOESY correlations of alternoic acid A (
Finally, the E configurations at Δ4,5 and Δ6,7 were supported from the NOESY (Figure S8) cross-peaks between H-5 and H3-9 and between H-5 and H3-8. An additional correlation in the NOESY between H3-10 and H3-12 established this interpretation. Accordingly, compound
Structural Determination of Alternoic Acid B (2 )
The molecular formula (C12H18O4) of compound 1H-1H COSY, 1H-13C HMBC and 1H-1H NOESY correlations of alternoic acid B ( NMR Spectral Data of Alternoic Acid B (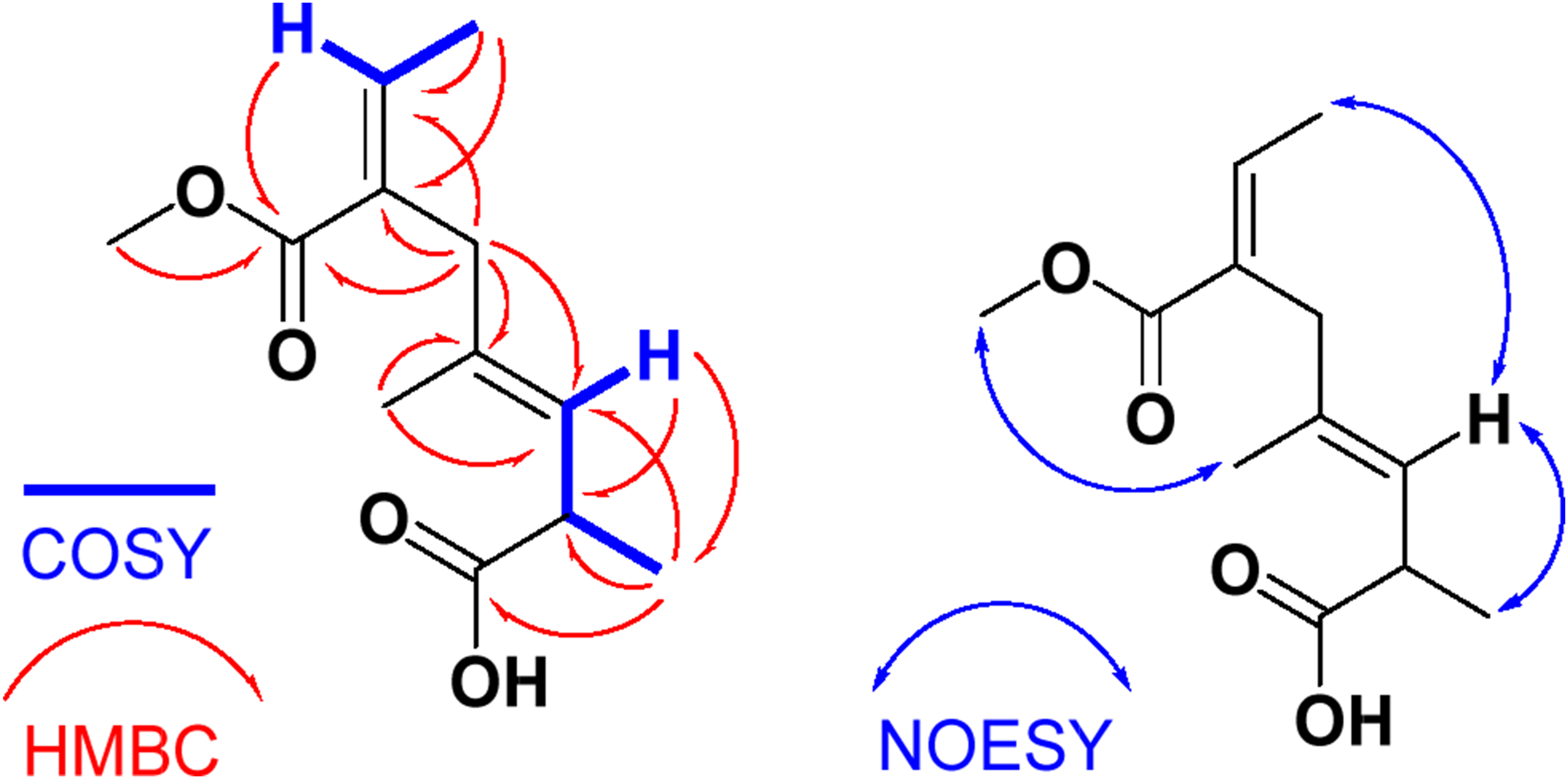

Interpretation of the 13C (Figure S9) and 1H NMR (Figure S8) combined with 2D NMR (Figures S10-S14) spectra supported the assignment of the compound as 6-(methoxycarbonyl)-2,4-dimethylocta-3,6-dienoic acid. Two coupling spin-systems, including the fragments (H-3−H-2−H3-9) and (H-7−H3-8), were traced from the COSY experiment, suggesting the shift of the olefinic moiety from C-4/C-5 in compound
In a similar way, the E configurations at Δ3,4 and Δ6,7 were configured from a NOESY experiment (Figure S14). Significant NOESY between H-3 and H3-9, between H3-8 and H-3, and between H3-10 and H3-12 secured this assignment. Thus, compound
Evaluation of the Antimicrobial Activities of the Compounds
Antimicrobial Effects of Compounds

aPositive antibacterial control (10 µg/disk).
bPositive antifungal control (10 µg/disk); NT = not tested. All tests were conducted in triplicate, and the results reported as mean ± SD, analyzed using two-way ANOVA followed by Bonferroni post-tests to compare replicate means using GraphPad Prism 5®.

Inhibition zone diameters of compound

MIC values of compounds
Discussion
Stereochemical Considerations and Future Directions
Although the planar structures and olefin geometries of alternoic acids A (
To avoid overinterpretation, the structures are depicted in Figure 2 with a wavy bond at C-2, and the text explicitly notes that the C-2 absolute configuration is unassigned. This approach follows standard practice in natural product reports where definitive methods (e.g., single-crystal X-ray diffraction, Mosher ester analysis, or computational chiroptical spectroscopy) cannot be applied due to limited material.
Should sufficient quantities of
Such computational workflows (e.g., MacroModel conformer search followed by Jaguar DFT calculations for ECD/VCD and NMR chemical shift prediction) are well established and could be implemented in future work. For now, the structures of
Evaluation of the Antimicrobial Activities of the Compounds
Compared to the positive controls, imipenem showed significantly stronger antibacterial activity (MRSA MIC = 0.75 ± 0.02 µg/mL; S. aureus inhibition zone = 29.33 ± 1.15 mm) (P < 0.001). The inhibition zones of 17.17 mm (compound
Conclusions
This study successfully identified and characterized two new compounds, alternoic acids A (
The marked activity difference between alternoic acid B (
These results identify alternoic acid B as interesting starting points for further structural optimization and biological evaluation, rather than validated drug leads. The current study has several limitations, including the absence of cytotoxicity/selectivity profiling against mammalian cell lines, lack of bactericidal/fungicidal endpoint analysis (MBC/MFC), and no mechanistic investigations into the mode of action.
To advance these findings, future investigations should first focus on mechanism of action studies. These include fluorescent membrane integrity assays, macromolecular synthesis profiling using radiolabeled precursors, and chemical proteomics to identify the molecular targets of alternoic acid B in MRSA and both compounds in C. albicans, along with transcriptomic analysis to reveal dysregulated pathways. Second, in vivo efficacy should be evaluated using a neutropenic murine thigh infection model for MRSA and a disseminated candidiasis model for antifungal activity, with bacterial and fungal burden, survival, and histopathology as endpoints. Acute toxicity studies with dose escalation are required prior to efficacy testing to determine the maximum tolerated dose and LD50. Third, pharmacokinetic and safety profiling should determine key parameters such as C max , half-life, AUC, and bioavailability after intravenous and oral administration, as well as plasma protein binding and tissue distribution, alongside mammalian cytotoxicity and hemolytic activity to establish a therapeutic index. Furthermore, future work should examine synergistic interactions with existing antibiotics using checkerboard assays and explore structural analogues via semi-synthesis to optimize potency and confirm the role of the shifted olefinic moiety in MRSA selectivity. Overall, this study enhances the understanding of secondary metabolites from the genus Alternaria and underscores their potential to address the escalating challenge of antimicrobial resistance, while recognizing that substantial preclinical development remains necessary.
Supplemental Material
Supplemental Material - Alternoic Acids A and B: Antimicrobial Esters of Dienoic Fatty Acid From the Marine Sediment-Derived Fungus Alternaria rosae DY-79
Supplemental Material for Alternoic Acids A and B: Antimicrobial Esters of Dienoic Fatty Acid From the Marine Sediment-Derived Fungus Alternaria rosae DY-79 by Diaa T. A. Youssef, Areej S. Alqarni, Mostafa E. Rateb, Sana A. Fadil and Lamiaa A. Shaala in Natural Product Communications.
Footnotes
Acknowledgments
This research work was Deanship of Scientific Research (DSR) under grant no. (KEP-MSc: 50-166-1443). The authors, therefore, gratefully acknowledge technical and financial support from King Abdulaziz University, DSR, Jeddah, Saudi Arabia.
Ethical Considerations
Ethical Approval is not applicable for this article.
Consent for Publication
All authors have read and approved the final manuscript and consent to its publication.
Author Contributions
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. This work was supported by the DSR at King Abdulaziz University, (Grant No. KEP-MSc: 50-166-1443).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data related to this manuscript are available in the manuscript and the supplemental Material.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
