Abstract
Objectives
This study aimed to investigate the anti-inflammatory properties and underlying mechanisms of ostruthin, a natural coumarin isolated from the roots of Paramignya trimera. Using an in vitro model of co-stimulation with bacterial and viral mimetics in RAW264.7 macrophages, we examined the effects of ostruthin on the expression of key signaling molecules: Myd88 and Nf-κb1 (mediators of Toll-like receptor signaling), and Arid5a (a stabilizer of IL-6 mRNA). We also assessed the production of inflammatory cytokines, including IL-6, TNF-α, and IL-10.
Methods
Ostruthin was isolated from P. trimera roots collected in Khanh Hoa Province, Vietnam. RAW264.7 cells were pre-treated with various concentrations of ostruthin before being stimulated with lipopolysaccharide (LPS) and poly(I:C). The mRNA expression of Myd88, Nf-κb1, and Arid5a was quantified at 3, 6, and 24 h using qRT-PCR. Cytokine levels were quantified at 24 h using ELISA. Additionally, molecular docking and molecular dynamics simulations were performed to predict direct interactions between ostruthin and the inflammation-related target, IL-6.
Results
Ostruthin significantly and dose-dependently reduced IL-6, TNF-α, and IL-10 secretion in co-stimulated RAW264.7 cells, with IC50 values of 3.0 ± 1.2 µM, 4.8 ± 1.5 µM, and 11.3 ± 0.9 µM, respectively. Treatment with ostruthin (3-60 μM) also markedly downregulated Myd88 (by 36%-79%), Nf-κb1 (by 49%-97%), and Arid5a (by 29%-83%) mRNA expression relative to untreated controls. Docking simulations revealed favorable binding affinities of ostruthin with IL-6 (−5.381 kcal/mol), comparable to or exceeding those of known inhibitors like tocilizumab.
Conclusions
Ostruthin exhibits potent anti-inflammatory activity in RAW264.7 macrophages stimulated with bacterial and viral mimetics, primarily by suppressing pro-inflammatory cytokines and modulating both TLR signaling and IL-6 mRNA stability pathways. These findings suggest that ostruthin may serve as a promising lead compound for the development of anti-inflammatory therapies targeting infections of mixed microbial origin.
Introduction
Polyinosinic:polycytidylic acid (poly(I:C)) is a synthetic analog of double-stranded RNA that mimics RNA viral infection, while lipopolysaccharide (LPS) is a component of the outer membrane of gram-negative bacteria. The combined presence of these pathogen-associated molecular patterns (PAMPs) can trigger a robust immune response, potentially contribute to a cytokine storm. 1 Co-stimulation of Toll-like receptor (TLR) pathways, TLR4 by LPS and TLR3 by poly(I:C), leads to complex crosstalk and modulation of downstream signaling effectors.2–4 TLRs are a family of pattern recognition receptors that detect PAMPs and activate innate immune responses. Among them, TLR3 and TLR4 are key sensors of viral and bacterial components, respectively. TLR3, primarily localized in endosomal membranes, recognizes double-stranded RNA (dsRNA), and activates the TRIF-dependent pathway, leading to the production of type I interferons and inflammatory cytokines. 5 In contrast, TLR4 is expressed on the cell surface and serves as the primary receptor for LPS. TLR4 signals through both MyD88-dependent and TRIF-dependent pathways, resulting in the activation of NF-κB and interferon regulatory factors (IRFs), which drive proinflammatory and antibacterial responses. 6 Upon recognition of a PAMP by TLR4, MyD88 is recruited to the receptor via its Toll/IL-1 receptor (TIR) domain. MyD88 then recruits IRAK (IL-1 receptor-associated kinase) family kinases and TRAF6, forming a signaling complex that activates the NF-κB and MAPK pathways. This cascade leads to the transcription of proinflammatory cytokines, chemokines, and co-stimulatory molecules. 7
Paramignya Wight, a genus in the Rutaceae family, comprises 17 accepted species. 8 The roots of Paramignya trimera have been traditionally used in Vietnamese folk medicine to treat liver diseases, particularly cirrhosis. 9 The first phytochemical investigation of the methanol extract of P. trimera roots, conducted in 2013, identified the presence of mono-coumarins, ostruthin as the major compound and a newly discovered coumarin, ninhvanin. 9 Subsequent studies have confirmed that mono-coumarins10–12 and bis-coumarin glycosides13–15 are the predominant chemical constituents of P. trimera roots. Methanol extracts of P. trimera roots, when administered at a dose of 10 g/kg body weight, have demonstrated hepatoprotective effects comparable to those of silymarin at 50 mg/kg. 16 Ostruthin hase also been shown to suppress LPS-induced iNOS and COX-2 protein expression. 10 Additionally, several reviews have highlighted coumarins as a promising source of anti-inflammatory and antiviral agents.17–19 Therefore, ostruthin was selected for evaluation of its antiviral and anti-inflammatory properties in this study.
Nuclear factor-kappa B (NF-κB) is a key transcription factor involved in the regulation of proinflammatory cytokines, including IL-6.20,21 Several plant-derived-inhibitors of NF-kB, such as resveratrol and quercetin, have demonstrated anti-inflammatory activity in macrophages.22,23 Arid5a, a protein upregulated in response to LPS stimulation, is known to stabilize IL-6 mRNA, thereby enhancing its production during inflammation.24–26 However, the exact mechanism by which ostruthin exerts its anti-inflammatory effects, particularly under co-stimulation with LPS and poly(I:C), remains unclear. In this study, we investigated the effect of ostruthin on the production of inflammatory cytokines (IL-6, TNF-α, and IL-10) in RAW264.7 macrophage cells under co-stimulation with LPS and poly(I:C). We further investigated whether ostruthin influences the expression of Arid5a and explored its potential binding interactions with IL-6 via molecular docking simulations.
Materials and Methods
Plant Materials
The roots of Paramignya trimera were collected in the Ninh Van commune, Khanh Hoa Province, South Vietnam. The plants were identified by Dr Nguyen Quoc Binh, Vietnam National Museum of Nature, VAST, Hanoi, Vietnam. A voucher specimen (C-765) was deposited in the Herbarium of the Institute of Chemistry, VAST, Hanoi, Vietnam.
Apparatus
1H-NMR (600 MHz) and 13C-NMR (150 MHz) data were measured on a Bruker Avance 600 MHz spectrometer. Column chromatography was carried out on silica gel (Si 60 F254, 230-400 mesh, Merck). All solvents were redistilled before use. Precoated TLC plates (Si 60 F254) were used for analytical purposes. Compounds were visualized under UV light (254, 365 nm) and by spraying plates with 10% H2SO4, followed by heating with a heat gun.
Extract and Isolation of Ostruthin
Dried roots of Paramignya trimera (1.0 kg) were extracted with methanol (2 × 3 L) at room temperature and concentrated under reduced pressure to yield a crude methanol extract. The extract was successively partitioned with n-hexane, yielding an n-hexane-soluble residue. The precipitate from the n-hexane fraction (12 g) was washed with n-hexane–acetone (4:1, v/v) and filtered through Whatman filter paper to yield ostruthin as a white solid (200 mg) (Figure 1). The purity of ostruthin is over 98%, determined by HPLC (Figure S2, SI).

Chemical Structure of Ostruthin Isolated from Paramignya trimera Roots.
Cell Culture and Treatments
The RAW264.7 cells (ATCC TIB-71) was kindly provided by Dr T. Kishimoto (Osaka University, Japan). Cells were cultured in RPMI 1640 medium supplemented with 10% FBS, 100 μg/mL penicillin and 100 μg/mL streptomycin. Cells were seeded in 24-well plates at a density of 5 × 104 cells/well. For stimulation, cells were co-treated with 1 µg/mL LPS and 20 µg/mL poly(I:C). 27 In other experiments, the cells were pretreated with ostruthin (3, 10, 30, or 60 μM) for 30 min prior to co-stimulation with 1 µg/mL LPS and 20 µg/mL poly(I:C). Cells were harvested for qPCR analysis at 3, 6, and 24 h, and supernatants were collected for ELISA at 24 h, as previously described. 28
MTT Assay
The RAW264.7 cells (5 × 104 cells/well) were seeded in 96-well plates (Corning) and treated with ostruthin at 3, 10, 30, or 60 μM for 24 h. Cell viability was assessed using the MTT assay. Absorbance was measured at 540 nm using a microplate reader. Each concentration was tested in triplicate. Cell viability was calculated as described previously.
29
ELISA
The cell culture media were collected, and the concentrations of IL-6 and TNF-α were quantified using ELISA kits (Mabtech) following the manufacturer's protocol. Each treatment was performed in triplicate. IC50 values was determined using ImageJ 1.50i computer software.
Real-Time Quantitative PCR
Total RNA was extracted from RAW264.7 cells using the easy-spin™ Total RNA Extraction Kit (iNtRON) according to the manufacturer's instructions. Briefly, 1 μg of RNA was reverse transcribed into cDNA using the RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher). Relative gene expression was measured using PowerUp SYBR Green Master Mix (Thermo Fisher). Primer sequences are listed in Table 1 and were previously described.30–32
The Sequences of Primers of Studied Genes.

Quantitative PCR was performed using the QuantStudio™ 6 Pro Real-Time PCR System with Design & Analysis Software v2.6.0. Gene expression was quantified using the 2−ΔΔCt method. 33
Molecular Docking Simulations
Proteins predicted as potential targets of ostruthin were subjected to docking simulations to evaluate binding affinity. The x-ray crystal structure of IL-6 was retrieved from the RCSB Protein Data Bank (PDB ID: 1ALU). 34 Protein preparation was performed using the Dock Prep tool in Chimera 1.16, as previously described, which included solvent removal, atom type assignment using the Amber force field, and the addition of polar hydrogens and partial charges.35–38 The 3D structure of ostruthin was generated using MarvinSketch 24.3 and then optimized using the Universal Force Field (UFF) in Avogadro 1.2.39,40 For the docking simulation, the grid box coordinates for IL-6 were defined at x = 0.519, y = −19.83, and z = 9.74. In both cases, the grid box dimensions were set to 24 × 24 × 24 with a spacing of 1 Å. The docking procedure was validated by redocking co-crystallized ligands into their respective binding sites, ensuring that the resulting conformations showed a root-mean-square deviation (RMSD) of less than 2 Å from their native poses. Docking simulations were performed using AutoDock Vina 1.2.3 with the “vina” scoring function, 41 and binding energy (ΔG, kcal/mol) along with key interactions was analyzed.
Molecular Dynamics Simulations
Following docking simulations, the IL-6-ostruthin complex were selected for molecular dynamics (MD) simulations using the GROMACS v2023.3 program to evaluate the binding interactions and the overall conformational stability of the complex in an aqueous environment. 42 Protein structure was prepared using the AMBER99SB-ILDN force field. 43 The ligand was optimized through single-point energy calculations at the B3LYP/6-31 g(d,p) level using Gaussian 09 software, followed by parameterization with the ACPYPE program to generate topology files compatible with the GROMACS program.44,45 The complex was placed in a triclinic simulation box with a minimum distance of 1 nm from the box edges, solvated using TIP3P water, and neutralized by adding counter ions (one Cl− ion). Energy minimization was performed using the steepest descent algorithm, followed by two equilibration phases: NVT (constant volume and temperature) at 300 K and NPT (constant pressure) at 1 atm, each lasting 100 ps with positional restraints applied to the protein backbone and ligand to stabilize the system. A 100 ns production MD simulation was then carried out without restraints using a time step of 2 fs, with trajectory data recorded every 10 ps for analysis. RMSD analysis was employed as a key metric to evaluate the structural stability and dynamic interaction profiles of the protein-ligand complexes.
Data Analysis
Results are expressed as the mean ± standard deviation (SD). Statistical analysis was calculated using Student's t test with p values of below 0.05 considered significant.
Results
Ostruthin Inhibits the LPS- and Poly(I:C)-Costimulated Production of IL-6, TNF-α, and IL-10 in RAW264.7 Cells
The viability of RAW264.7 cells was not affected by ostruthin at concentrations of 0, 3, 10, 30, and 60 μM (100%, 94 ± 5%, 90 ± 3%, 88 ± 5%, and 90 ± 4%, respectively) (Figure 2). To investigate the anti-inflammatory effect of ostruthin, we assessed its ability to inhibit the production of IL-6, TNF-α, and IL-10 in RAW264.7 cells costimulated with LPS and poly(I:C). It was found that ostruthin attenuated the LPS- and poly(I:C)-costimulated cytokine production in a concentration-dependent manner. RAW264.7 cells were pre-treated with various concentrations of ostruthin and then stimulated with 1 µg/mL LPS and 20 µg/mL poly(I:C). Ostruthin significantly inhibited the production of IL-6, TNF-α, and IL-10 with IC50 values of 3.0 ± 1.2 µM, 4.8 ± 1.5 µM and 11.3 ± 0.9 µM, respectively (Figure 3a-c).

Effect of Ostruthin Treatment on the Viability of RAW264.7 Cells. Cells Were Treated with Different Concentrations of Ostruthin, and Viability was Assessed Using the MTT Assay. Results are Expressed as the Means ± SD of Three Independent Experiments.

Ostruthin Inhibited the Production of IL-6 (
Ostruthin Inhibited Myd88, Nf-κb1, and Arid5a Expression in LPS-Poly(I:C)-Costimulated RAW264.7 Cells
The production of proinflammatory cytokines in activated macrophages is regulated by various intracellular pathways, including the NF-κB pathway. 25 To elucidate the molecular mechanism by which ostruthin regulates NF-κB-responsive genes such as Myd88 and Nf-κb1, as well as IL-6 regulators such as Arid5a, we examined their expression dynamics under LPS and poly(I:C) costimulation. At the early stimulation phase (0-6 h), Myd88, Nf-κb1, and Arid5a mRNA levels increased, peaking at around 6 h (2.1-fold, 3.5-fold, and 2.5-fold increases, respectively) (Figure 4a-c). This increase was expected, as these genes are key components of the early innate immune signaling cascade—Myd88 and Nf-κb1 mediate NF-κB activation, while Arid5a stabilizes IL-6 mRNA during inflammation. In the subsequent phase, ostruthin reduced the expression of all three genes in a concentration-dependent manner. At concentrations of 3, 10, 30, and 60 μM, ostruthin significantly inhibited Myd88 (by 36%, 51%, 71%, and 79%), Nf-κb1 (49%, 70%, 82%, and 97%), and Arid5a (29%, 51%, 63%, and 83%), respectively (Figure 4a-c). Mechanistically, these results suggest that ostruthin interferes with NF-κB signaling, possibly through inhibition of Myd88 and Nf-κb1, and also downregulates IL-6 mRNA stability via Arid5a suppression. At a later phase (24 h), in untreated cells, the expression of these genes declined from the 6 h peak, likely due to feedback regulation and resolution mechanisms, although residual mRNA remained detectable. In contract, in ostruthin-treated cells, Myd88, Nf-κb1, and Arid5a mRNA became undetectable by 24 h, indicating a more sustained shutdown of these inflammatory pathways. This decrease may reflect both transcriptional inhibition and accelerated mRNA decay. Thus, the increases observed at 3-6 h represent acute inflammatory gene activation in response to LPS and poly (I:C), whereas the disappearance of transcript by 24 h in ostruthin-treated samples reflects potent and durable anti-inflammatory suppression of both NF-κB–dependent transcription and IL-6 mRNA stabilization pathways.
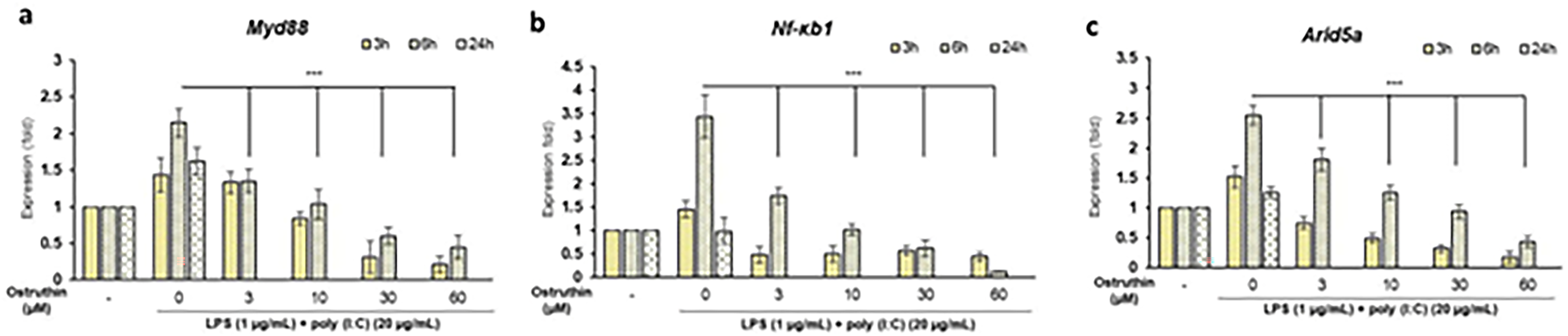
Ostruthin Suppressed the Gene Expression of Myd88 (
Ostruthin can Interact with IL-6 in Silico
Ostruthin significantly reduced Arid5a mRNA expression in RAW264.7 cells stimulated with LPS and poly(I:C). Given the well-established role of Arid5a in stabilizing IL-6 mRNA and enhancing its expression during inflammation, we hypothesized that ostruthin may impair IL-6 production by targeting Arid5a. However, because no crystal structure of Arid5a is currently available, molecular simulations could not be extended to this protein. We therefore focused on IL-6 as a direct molecular target of ostruthin. Tocilizumab, a clinically approved monoclonal antibody that directly targets IL-6 signaling, was selected as a positive control to benchmark the performance of ostruthin. 46 Docking analyses of IL-6 are presented in Figure 5 and Table 2. Ostruthin binds within the same surface pocket occupied by the control ligand (shown by pose superposition), yielding a binding energy of −5.38 kcal.mol−1, compared with −5.94 kcal.mol−1 for tocilizumab (Table 2). To assess the reliability of the docking protocol, a re-docking validation was performed by re-docking co-crystallized ligand into IL-6 protein's active sites. The re-docking results showed RMSD values below 2 Å, confirming that the docking protocol reliably reproduced the native binding poses (Figure S1). 47 This validation supports the predicted interactions and the credibility of ostruthin's binding modes with IL-6. Docked poses and interaction maps (Figure 5) indicated that ostruthin engages IL-6 through hydrophobic and polar interactions: alkyl/π-alkyl contacts with Leu178 (and the adjacent hydrophobic patch), and hydrogen bonds with Arg179 and Arg182. Tocilizumab establishes hydrogen bonds with Gln175 and Arg182 and formed additional π-type contacts involving Leu33 and Leu178.

Docking Interactions of Ostruthin and Tocilizumab in the Active Sites of IL-6.
Binding Affinities and Interactions of Ostruthin and Tocilizumab with IL-6 Protein.

To further evaluate the stability of IL-6 complexes, 100 ns MD simulations were performed for IL-6 in the apo state and in complex with tocilizumab or ostruthin. The RMSD profiles (Figure 6A) revealed that both ligands remained stably bound throughout the trajectory, with deviations below 0.3 nm. Ostruthin displayed slightly higher fluctuations during the initial 30 ns but quickly converged to a stable conformation comparable to tocilizumab. Analysis of residue-level flexibility (RMSF, Figure 6B) indicated that both ligands reduced fluctuations in several loop regions relative to apo IL-6, suggesting a stabilizing effect on the local dynamics of the protein. The overall compactness of the complexes, assessed by the radius of gyration (Rg, Figure 6C), remained consistent at ∼1.6 nm, confirming that neither ligand induced large-scale structural rearrangements. Backbone RMSD comparisons (Figure 6D) demonstrated that apo IL-6 exhibited greater conformational drift, whereas the presence of either ligand significantly stabilized the protein, with ostruthin showing slightly lower RMSD values at later stages of the simulation. Similarly, SASA profiles (Figure 6E) remained steady across all systems, indicating that ligand binding did not markedly alter solvent exposure of IL-6. The interaction profiles confirm that ostruthin forms a stable complex with IL-6, conferring a degree of structural stabilization comparable to tocilizumab during the simulation. Subsequently, MM/GBSA free energy calculations were performed based on 100 ns MD trajectories. For ostruthin, the predicted binding free energy (ΔGbinding) was −24.9 ± 5.7 kcal/mol, indicating a thermodynamically favorable interaction. Decomposition of the energy terms revealed that van der Waals interactions (−27.4 kcal/mol) and electrostatic contributions (−20.0 kcal/mol) were the major stabilizing forces, whereas polar solvation energy (EGB, + 26.3 kcal/mol) strongly opposed binding. Nonpolar solvation (ESURF, −3.8 kcal/mol) provided slight stabilization. These results highlight that ostruthin stabilizes within a partially hydrophobic pocket of IL-6, where residues such as Leu178 and Leu33 contribute hydrophobic packing, while Arg179 and Arg182 mediate electrostatic contacts. In comparison, the clinically approved anti-IL-6 antibody tocilizumab exhibited a weaker predicted binding free energy of −14.4 ± 2.7 kcal/mol under the same protocol. Although tocilizumab benefits from high specificity and multivalent interactions inherent to antibody-antigen recognition, its lower calculated binding affinity reflects the limitations of MM/GBSA in fully capturing protein-protein interfaces. By contrast, the stronger van der Waals stabilization observed for ostruthin suggests that the small molecule fits more tightly into the IL-6 binding pocket. Taken together with docking and MD data, these results support ostruthin as a potential small-molecule modulator of IL-6, while underscoring the different binding mechanisms between chemical ligands and therapeutic antibodies.

Structural Dynamics of IL-6 in the apo Form (red) and in complex with Tocilizumab (Blue) or Ostruthin (Yellow) Over a 100 ns Molecular Dynamics Simulation. (A) Backbone RMSD Profiles of IL-6 in the Ligand-Bound States; (B) per-Residue RMSF Fluctuations of IL-6 upon Ligand Binding; (C) Radius of Gyration (Rg) Representing Overall Protein Compactness; (D) Comparison of Backbone RMSD Between the apo and Ligand-Bound Systems; (E) Solvent Accessible Surface Area (SASA) of IL-6 in apo and complex Forms.
Discussion
Previous studies have shown that ostruthin inhibits the production of NO and PGE2 and suppresses the expression of iNOS and COX-2 in LPS-stimulated BV2 cells. 20 In this study, ostruthin exhibited dose-dependent anti-inflammatory effects on the cytokines IL-6, TNF-α, and IL-10 under co-stimulation with LPS and poly(I:C). By reducing the expression of Myd88 and Nf-κb1, ostruthin may play a role in regulating inflammatory genes, including key cytokines such as IL-6, TNF-α, and IL-10. A recent report indicated that the compound murrayfoline A can inhibit the production of IL-6, TNF-α, and IL-10 under similar co-stimulation conditions by targeting the aryl hydrocarbon receptor (Ahr). 32 In contrast, ostruthin did not affect Ahr expression (data not shown), suggesting that it regulates cytokine production in RAW264.7 cells under LPS and poly(I:C) co-stimulation via an Ahr-independent mechanism. Interestingly, ostruthin was found to decrease the expression of Arid5a, a known stabilizer of IL-6 mRNA, under these co-stimulation conditions in RAW264.7 cells. This downregulation may reduce IL-6 mRNA stability, thereby leading to decreased IL-6 levels (Figure 7). Molecular docking and dynamics studies revealed that ostruthin could regulate IL-6 production by binding directly to IL-6. Additionally, while Arid5a stabilizes IL-6 mRNA, it does not affect TNF-α mRNA. 29 Therefore, the precise mechanism by which ostruthin regulates TNF-α and IL-10, potentially through the NF-κB pathway and/or others, requires further investigation. In summary, ostruthin may inhibit LPS and poly(I:C)-induced IL-6, TNF-α, and IL-10 production via downregulation of Myd88, Nf-κb1, and Arid5a mRNA, as well as by directly binding to IL-6 (Figure 5). The RNA-binding protein Arid5a has emerged as a key post-transcriptional regulator of inflammation, particularly by stabilizing IL-6 mRNA and sustaining its production during inflammatory responses. Inhibition of Arid5a expression has therefore been proposed as a therapeutic strategy to suppress IL-6–mediated pathology. In this study, ostruthin showed strong suppression of Arid5a mRNA expression in LPS–poly(I:C)–stimulated RAW264.7 cells. Our findings indicate that ostruthin may inhibit IL-6 stabilization through a comparable mechanism to tocilizumab, but with slightly higher predicted binding affinity, suggesting a potentially stronger or more stable interaction. In addition, ostruthin was evaluated here in a co-stimulation model mimicking both bacterial (LPS) and viral poly(I:C) inflammatory triggers, broadening its potential applicability. Together, these findings position ostruthin as a promising candidate for an IL-6 and Arid5a inhibitor with broader relevance to inflammatory diseases driven by bacterial and viral components.

Diagram of the Anti-Inflammatory Activity of Ostruthin in LPS-Poly(I:C)-Costimulated RAW264.7 Cells.
While this study provides novel insights into the anti-inflammatory mechanisms of ostruthin in LPS- and poly(I:C)-stimulated RAW264.7 cells, several limitations should be acknowledged, which also point to directions for future research. First, the findings are based solely on in vitro experiments using a murine macrophage cell line (RAW264.7). Although this model is widely used to investigate macrophage-mediated inflammatory responses, it does not fully capture the complexity of immune responses in vivo. Validation of ostruthin's anti-inflammatory effects in appropriate animal models or in primary human macrophages would significantly enhance the translational relevance of these results. Second, although ostruthin was shown to reduce IL-6 production, likely through modulation of Arid5a at the post-transcriptional level, the precise molecular mechanism remains unclear. Specifically, how ostruthin downregulates Arid5a expression and whether it directly disrupts the Arid5a–IL-6 mRNA interaction is yet to be determined. Future studies should aim to comprehensively characterize the molecular targets and pathways affected by ostruthin to better understand its role in modulating inflammation.
Conclusion
Ostruthin, a major bioactive coumarin isolated from the roots of Paramignya trimera, demonstrates significant anti-inflammatory activity in murine RAW264.7 macrophages co-stimulated with bacterial LPS and viral mimic poly(I:C). It effectively downregulated the production of key inflammatory cytokines (IL-6, TNF-α, and IL-10) and the expression of pivotal signaling molecules involved in the Toll-like receptor pathway, including Myd88, Nf-κb1, and Arid5a, an RNA-binding protein known to stabilize IL-6 mRNA. Molecular docking simulations indicated that ostruthin can directly interact with the IL-6 cytokine itself, suggesting a dual mechanism of action that disrupts IL-6 regulation. Molecular dynamics simulations were subsequently performed to confirm the interaction of ostruthin with IL-6. We found that it maintains a stable binding interaction with IL-6 throughout the simulation, with IL-6 exhibiting slightly enhanced structural stability, while MM/GBSA free energy calculations supported a thermodynamically favorable binding, which may correlate well with experimental results. These findings highlight ostruthin as a promising lead compound for the development of novel anti-inflammatory agents targeting bacterial- and viral-induced inflammation. Furthermore, this work underscores the pharmacological potential of P. trimera as a rich natural source for drug discovery and therapeutic applications in the management of inflammatory diseases.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X251385033 - Supplemental material for Ostruthin Targets IL-6 and Downregulates Arid5a Expression to Inhibit Cytokine Production Under TLR3-TLR4 Cotriggering: In vitro and in silico Studies
Supplemental material, sj-docx-1-npx-10.1177_1934578X251385033 for Ostruthin Targets IL-6 and Downregulates Arid5a Expression to Inhibit Cytokine Production Under TLR3-TLR4 Cotriggering: In vitro and in silico Studies by Tran Thu Trang, Ta Thi Minh Huyen, Vu Thi Tien, To Minh Nhat, Vu Thanh Loc, Tran Thu Huong, Nguyen Xuan Ha , Hai Pham-The, Nguyen Manh Cuong and Nguyen Trung Nam in Natural Product Communications
Footnotes
Acknowledgments
The auhors are thankful to the Vietnam Academy of Science and Technology, Hanoi, Vietnam.
ORCID iDs
Author Contributions/CRediT
Tran Thu Trang: Writing—original draft, Methodology, Funding acquisition. Nguyen Trung Nam: Conceptualization, Supervision, Writing—review and editing. Vu Thi Tien, To Minh Nhat, Ta Thi Minh Huyen: Technical assistance. Vu Thanh Loc: Writing—original draft, investigation. Tran Thu Huong: Resources. Nguyen Xuan Ha: Software, Formal analysis. Hai Pham-The: Data curation, Conceptualization. Nguyen Manh Cuong: Funding acquisition, writing—review and editing.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Grants from the Vietnam Academy of Science and Technology (code: DL0000.05/22-23 and CT0000.07/22-24).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

