Abstract
Introduction
The WHO had reported a total of 769 806 130 confirmed cases of COVID-19 globally, with 6 955 497 recorded deaths. Despite the administration of 13 499 767 926 vaccine doses worldwide, the virus persists in affecting populations on a significant scale. 1 These concerning statistics act as a clear reminder of the significant challenge confronting scientists globally. The persistent spread of the virus, highlighted by these figures, emphasizes the immediate and collective action needed from the global scientific community to uncover effective treatments and preventive measures.
In the past 20 years, the adoption of computer-aided drug discovery (CADD) methods has seen a significant upswing in pharmaceutical research and experimentation.2,3 This widespread use can be attributed to several key factors. First, the ability to determine the precise 3D structures of numerous protein targets within the human body. 4 Second, the remarkable progress in computer software and hardware capabilities. 5 Lastly, an enhanced comprehension of the principles of structure–activity relationship (SAR) and quantitative structure–activity relationship. 6 As a result, computational chemistry techniques have been applied to explore various pharmacokinetic and pharmacodynamic properties, enabling the identification of relationships between chemical structure and activity. These methods include pharmacophore identification, 7 homology modeling, 8 docking, 9 molecular dynamics (MD) simulations, 10 absorption, distribution, metabolism, excretion, and toxicity (ADMET) predictions, 11 density functional theory (DFT), 12 structure similarity, 13 and fingerprints. 14
Since the dawn of history, nature has played an essential role in fulfilling basic human requirements. It has not only supplied remedies for medical treatments but has also provided sustenance and played a part in creating a range of cosmetic products.15,16
In our efforts to combat COVID-19, we employed a multistage in silico CADD approach to identify natural inhibitors and recommend the most suitable one against specific COVID-19 enzymes. From a group of 310 antiviral natural metabolites, potential inhibitors for nsp10, 16 the main protease,17,18 and the papain-like protease 19 have been pointed. Additionally, among 3009 food drug administration (FDA)-approved drugs, the most promising inhibitors against the RNA-dependent RNA polymerase 20 and the SARS-CoV-2 nsp16-nsp10 2′-o-methyltransferase complex 21 have been indicated. Also, we conducted an exploration of a collection of 5956 compounds sourced from traditional Chinese medicine with the aim of identifying potential natural inhibitors for the SARS-CoV-2 helicase enzyme. 22
We herein report the structure and ligand-based in silico discovery of potential SARS-CoV-2 helicase inhibitors among 4924 natural African metabolites. The screened compounds were obtained from the African Natural Products Database, which was collected from various natural products databases of several African regions covering from 1962 to 2019. The data set was obtained from local and international African journals besides MSc and PhD theses in African universities. 23 The applied methods were structure similarity, structural fingerprints, molecular docking, ADMET, toxicity, MD simulations (Root Mean Square Deviation [RMSD], Root Mean Square Fluctuation [RMSF], Rg, SASA, and H-bonding) for 200 ns in addition to molecular mechanics-generalized Boltzmann surface area (MM-GBSA), protein–ligand interaction profile (PLIP), and principal component analysis of trajectory (PCAT) studies.
Results and Discussion
Structure Fingerprints Studies
The co-crystallized ligand is a compound with a pronounced affinity for a specific protein, resulting in the formation of a crystalline complex. This complex serves as a valuable source of information revealing the precise structure of the protein through x-ray imaging. 24 Identify intricate interactions between the ligand and the protein, exposing essential structural and chemical features defining their potent binding. The co-crystallized ligand's chemical structure serves as a valuable blueprint for crafting new inhibitors effective in binding to the target protein, advancing the search for potential therapeutic agents.. 25 Analyzing the structural attributes and functional groups of the co-crystallized ligand provides a deeper insight into the key elements driving its strong binding affinity. Utilizing structure fingerprints involves a ligand-based computational approach, calculating chemical and physical descriptors, and comparing them against a specific active molecule. 26 The fingerprints study was employed vastly to expect the similarity and/or dissimilarity of two molecules or more.27,28 The software computes the chemical and physical descriptors of the examined ligand as mathematical symbols and converts them to bit strings. The obtained strings calculate the presence (1) as well as the absence (0) of a specific 2D fragment in both examined and reference compounds.29,30
We applied Discovery Studio software to explore the degree of similarity of the fingerprints of 4924 natural African metabolites with VXG, the co-crystallized of SARS-CoV-2 helicase (PDB ID: 5RMM). The selection of PDB entry (5RMM) was based on the pivotal role of the helicase in the viral replication, coupled with the simplicity of its co-crystallized ligand (VXG). This choice increases the likelihood of identifying structurally similar compounds through the examined library, aligning with the study's objective of exploring potential inhibitors targeting the SARS-CoV-2 helicase. The fingerprints experiment was adjusted to determine the most similar 200 compounds among the examined compounds (Table S1 in Supplementary data). The fingerprints study described the following descriptors in the fragments as well as atoms of the examined compounds and VXG: H-bond acceptors, 31 H-bond donors, 32 charges, 33 hybridization, 34 positive ionizable atoms, 35 negative ionizable atoms, 36 halogens, 37 aromatic groups, 38 align with the ALogP. 39
Molecular Similarity
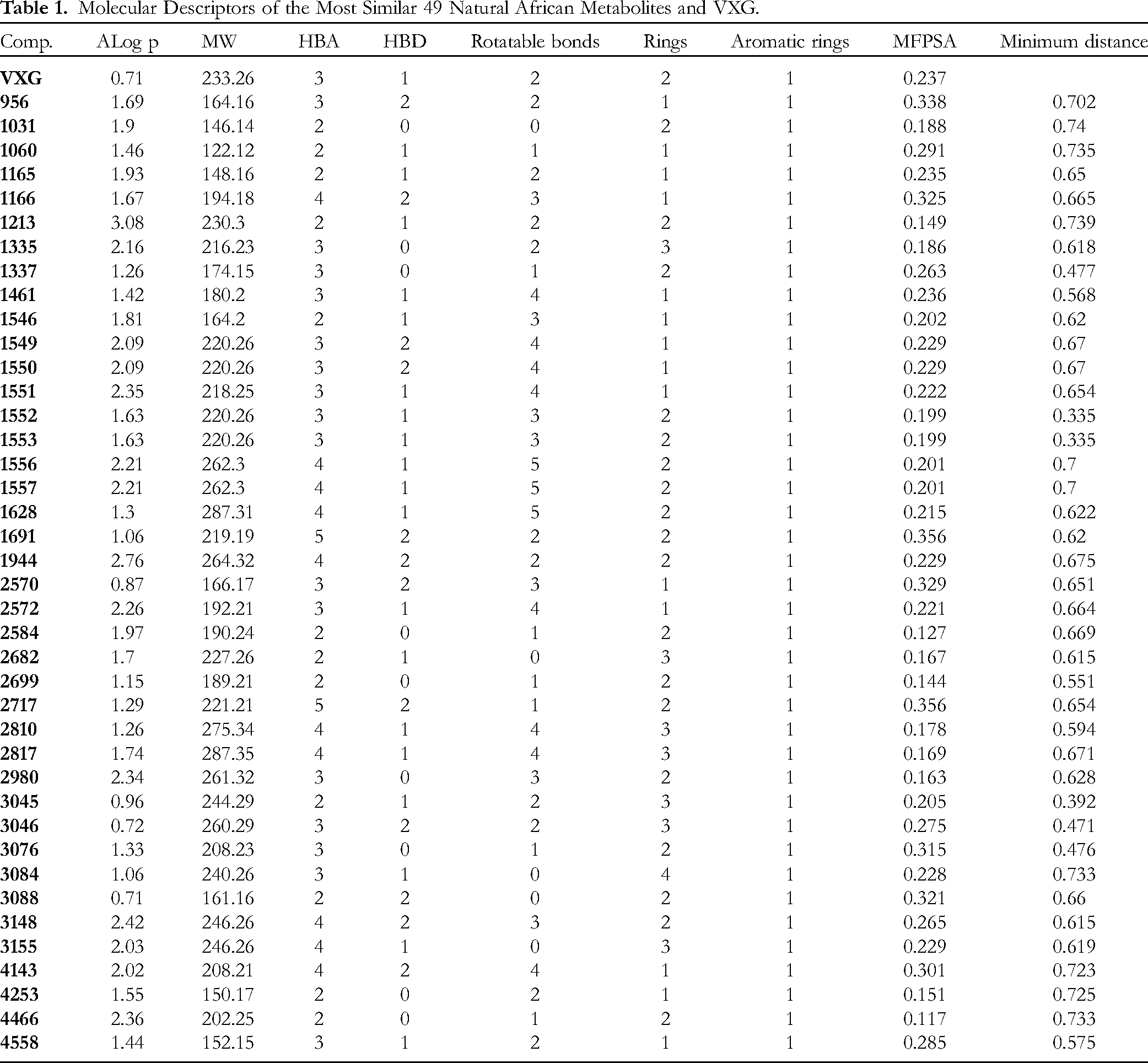
The main difference between fingerprints and molecular similarity is that the first calculates the presence or the absence of certain 2D atomic descriptors in fragments, or substructures. 40 On the other hand, the molecular similarity study computes the molecular descriptors regarding the whole structure of the examined compound. The tested descriptors could be steric, electronic, topological, or physical. 41 We depended on Discovery studio software to examine the molecular similarity of the most similar 200 metabolites against VXG. The tested descriptors in this study (Figure 1 and Table 1) included partition coefficient (ALog p), 42 molecular weight (MW), 43 H-bond donors (HBDs), 44 H-bond acceptors (HBAs), 45 rotatable bonds number, 46 number of rings as well as aromatic rings 47 and minimum distance 48 together with the molecular fractional polar surface area (MFPSA). 49 The similarity study was adjusted to single out the most similar 40 compounds (Table 2 and Figure S1 in the Supplementary data).

The molecular similarity of the most similar 40 natural African metabolites and VXG. The green balls represent the reference ligand (VXG), the red balls represent the most similar compounds with VXG, and the blue balls represent compounds with low similarity with VXG.
Molecular Descriptors of the Most Similar 49 Natural African Metabolites and VXG.
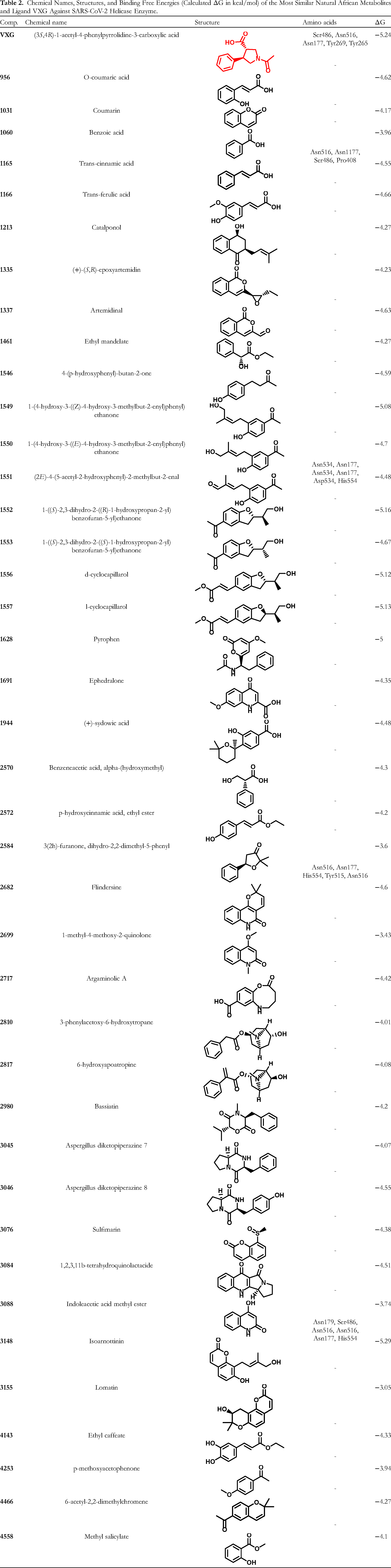
Chemical Names, Structures, and Binding Free Energies (Calculated ΔG in kcal/mol) of the Most Similar Natural African Metabolites and Ligand VXG Against SARS-CoV-2 Helicase Enzyme.
Docking Studies
The most similar 40 natural African metabolites were docked using the MOE software, with the co-crystallized ligand VXG serving as a reference compound. The fundamental purpose of this research was to know more about the orientations and binding modes of the tested compounds at the SARS-CoV-2 helicase enzyme's binding site (PDB ID: 5RMM). The docking method was validated by redocking the co-crystallized ligand in the active site of the SARS-CoV-2 helicase enzyme using the docking algorithm. The validation step established the protocol's applicability for the intended docking study as demonstrated by the small RMSD (1.15 Å) between the redocked pose and the co-crystallized one. Moreover, the ability of the docking algorithm to retrieve the co-crystallized ligand's reported binding mode also indicated the validity of the selected docking algorithm (Figure S2, in the Supplementary data).
The binding free energy (ΔG) between the docked molecules and the active site, as well as the correct binding mode, were considered in the comparison between the tested compounds. Table 2 shows the calculated ΔG (binding free energies) of the tested compounds and the reference molecule (VXG) against the SARS-CoV-2 helicase enzyme. It is noteworthy that the docking results showed that compound 3148 illustrated high-affinity values of −5.29 kcal/mol, which is higher than the redocked ligand VXG (−5.24 kcal/mol). Furthermore, binding energy scores of −5.00 to −5.16 kcal/mol were found in compounds 1628, 1552, 1556, 1557, and 1549, which are comparable to the redocked ligand VXG. The other compounds, on the other hand, had lower affinity values than the ligand VXG.
The redocked ligand VXG predicted binding mode revealed an affinity value of −5.24 kcal/mol. The formation of three hydrogen bond interactions is most likely responsible for the high binding affinity. One was formed between the C=O group of the carboxylate moiety and Ser486, while the other two were formed with Asn516 and Asn177. Additionally, the nitrogen atom of the pyrrolidine ring formed hydrophobic interactions (pi–cation interactions with Tyr269 and Tyr265. These findings were deemed to be in alignment with the reported data 24 (Figures 2 and S3).

2D binding mode of ligand (VXG) in the active site of SARS-CoV-2 helicase.
With a docking score of −4.55 kcal/mol, compound

2D binding modes of the tested compound in the active site of SARS-CoV-2 helicase. (A) Compound
Compound
The binding mode of compound
The establishment of five hydrogen bonds with the fundamental amino acids in the active site of the SARS-CoV-2 helicase enzyme was shown by the binding mode of compound
ADMET Studies
The approval of any compound to be a drug is based on its pharmacokinetic assessment besides the biological activity. Consequently, the analysis of the ADMET properties of any new compound should be investigated in an early stage of drug discovery to avoid late drug withdrawal. 50
Fifteen compounds (Table 3) showed high binding energy with accepted binding mode against the prospective target (SARS-CoV-2 helicase). These compounds were subjected to ADMET studies. ADMET profile identifies absorption, distribution, metabolism, excretion, besides toxicity. Although the presence of varied in vitro studies that can investigate the ADMET properties, the in silico studies are more advantageous due to the limitations of cost, time, effort in addition to the strict regulations regarding animal lives. 51 The predicted ADMET profiles of the 15 compounds were shown in Table 3 and Figure 4. The applied descriptors were (1) blood–brain barrier penetration (BBB), (2) absorption level from the human intestine (HA), (3) aqueous solubility level (S), (4) CYP2D6 binding and inhibition, and (5) plasma protein binding ability (PPB). Fortunately, the ADMET profiles of all compounds indicated their likeness to be drugs.

The expected ADMET study. ADMET_AlogP98: lipid-water partition coefficient; ADMET_PSA_2D: polar molecular surface area. Two-dimensional polar surface area (PSA_2D) each drug is plotted against their computed atom-type partition coefficient (ALogP98). The area encompassed by the ellipse represents good absorption without any violation of the ADMET properties. Based on Egan et al, 52 absorption model the 95% and 99% confidence limit ellipses corresponding to the blood–brain barrier (BBB) and intestinal absorption models are indicated.
Predicted ADMET Descriptors.
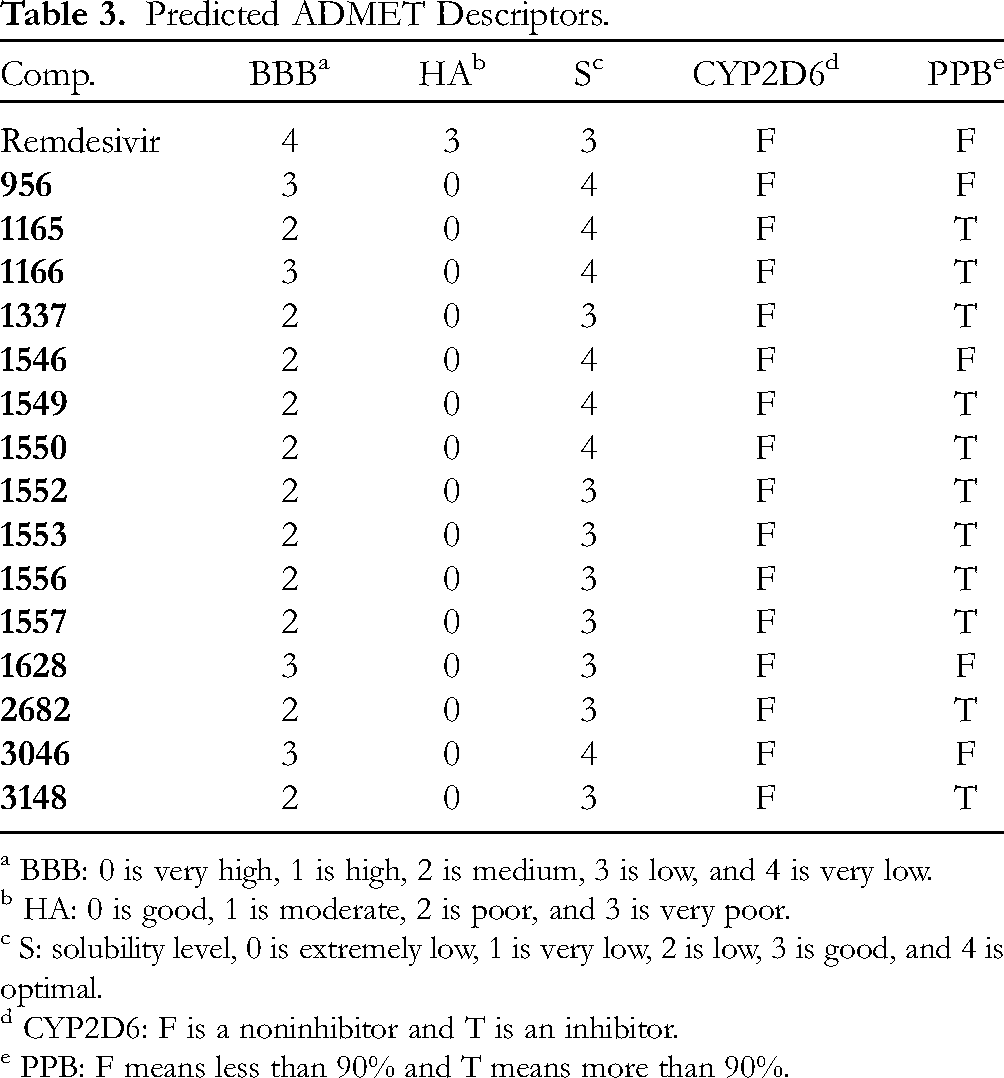
a BBB: 0 is very high, 1 is high, 2 is medium, 3 is low, and 4 is very low.
b HA: 0 is good, 1 is moderate, 2 is poor, and 3 is very poor.
c S: solubility level, 0 is extremely low, 1 is very low, 2 is low, 3 is good, and 4 is optimal.
d CYP2D6: F is a noninhibitor and T is an inhibitor.
e PPB: F means less than 90% and T means more than 90%.
Toxicity Studies
The toxicity estimation in the early stage is of the utmost importance to minimize the failure of the drug in the late development or the clinical stages. 53 The utilization of in silico approaches in toxicity prediction played an indispensable role in drug development to avoid ethical regulations and resources availability besides time wasting in the usual in vitro and in vivo studies. 54 The basic principle of in silico toxicity prediction is the SAR-predictive toxicity. In detail, the computer compares the basic chemical structural properties of the examined molecules against the structures of thousands of compounds of reported safety and or toxicity 55 (see the detailed toxicity report of the examined compounds in the Supporting data). We employed Discovery Studio software to compute the toxicity profiles of the selected molecules against seven different models. The employed models were FDA rat carcinogenicity in mouse-female (FDA),56,57 mouse carcinogenic potency (median toxic dose [TD50]), 58 rat maximum tolerated dose (MTD),59,60 rat oral median lethal dose (LD50), 61 rat chronic lowest observed adverse effect level (LOAEL),62,63 ocular irritancy, and skin irritancy. 64 The obtained results (Table 4) eliminated compounds 1337, 1546, 2682, and 3148 as they computed to have carcinogenic potencies.
Toxicity Properties of Tested Compounds and the Reference.

aN, not a carcinogen; C, carcinogen.
bN, nonirritant; M, mild; D, moderate; S, severe.
Molecular Dynamic Simulation
While molecular docking predicts ligand binding, it has limitations in studying enzyme rigidity. 65 MD simulations, on the other hand, offer detailed insights into atomic-level protein behavior and accurate temporal resolution, 66 capturing conformational changes and energy alterations in the enzyme–ligand complex. 67 MD simulations compute forces on each atom in the ligand, protein, and complex at ultra-short intervals using a force field, reflecting potential energy. 68 Integration follows Newton's motion laws, considering bonded and nonbonded interactions. 69 This atomic-level calculation, repeated billions of times, yields atomic trajectories for a defined time interval. 70
Compound

Shows the (A) helicase RMSD graph, (B1) shows the compound
Additionally, Figure 6A the radius of gyration (RoG) and Figure 6B solvent-accessible surface area (SASA) show that, on average, the two systems exhibit very similar behavior over the whole simulation. From what can be seen in Figure 6C, the average number of H-bonds formed between each ligand and helicase systems shows different values. While VXG shows mainly two H-bonds, compound

(A) Helicase protein’s RoG graph, (B) helicase protein’s SASA graph, (C) change in the hydrogen bonds numbers between each ligand and helicase, (D) helicase protein's RMSF graph, (E) distance from the center of mass of compound
Molecular Mechanics-Generalized Boltzmann Surface Area
The MM-GBSA method was employed to calculate the free binding energy of the SARS-CoV-2 helicase-
The different parts of the MM-GBSA estimated binding free energy are shown in Figure 7. Considering that compound

MM-GBSA energy constituents and their values. Bars represent the standard deviations.

Analysis of SARS-CoV-2 helicase-
Protein–Ligand Interaction Fingerprint Study
Protein–ligand interaction fingerprint (ProLIF) is a crucial technique in the realm of computer-aided drug design, molecular docking, and MD investigations. It plays a central role in thoroughly examining and characterizing the interactions that occur between proteins and ligands. This method involves the creation of interaction fingerprints, which are unique patterns resulting from the interactions between a protein and a ligand. These fingerprints are vital for quantifying both the strength and the nature of these binding interactions.
72
ProLIF enables the quantification of various types of interactions, including significant categories like hydrogen bonds, hydrophobic contacts, and other noncovalent associations. Furthermore, when applied in MD simulations, ProLIF serves as an invaluable tool for tracking the dynamic behaviors of protein–ligand complexes over extended periods. It provides crucial insights into how these interactions between the protein and ligand evolve throughout the simulation, thereby enhancing our comprehension of complex stability and binding affinity.
73
Results from the ProLIF library showed that three amino acids had an incidence of hydrophobic interaction greater than 73%. The three most common amino acids found to interact with compound
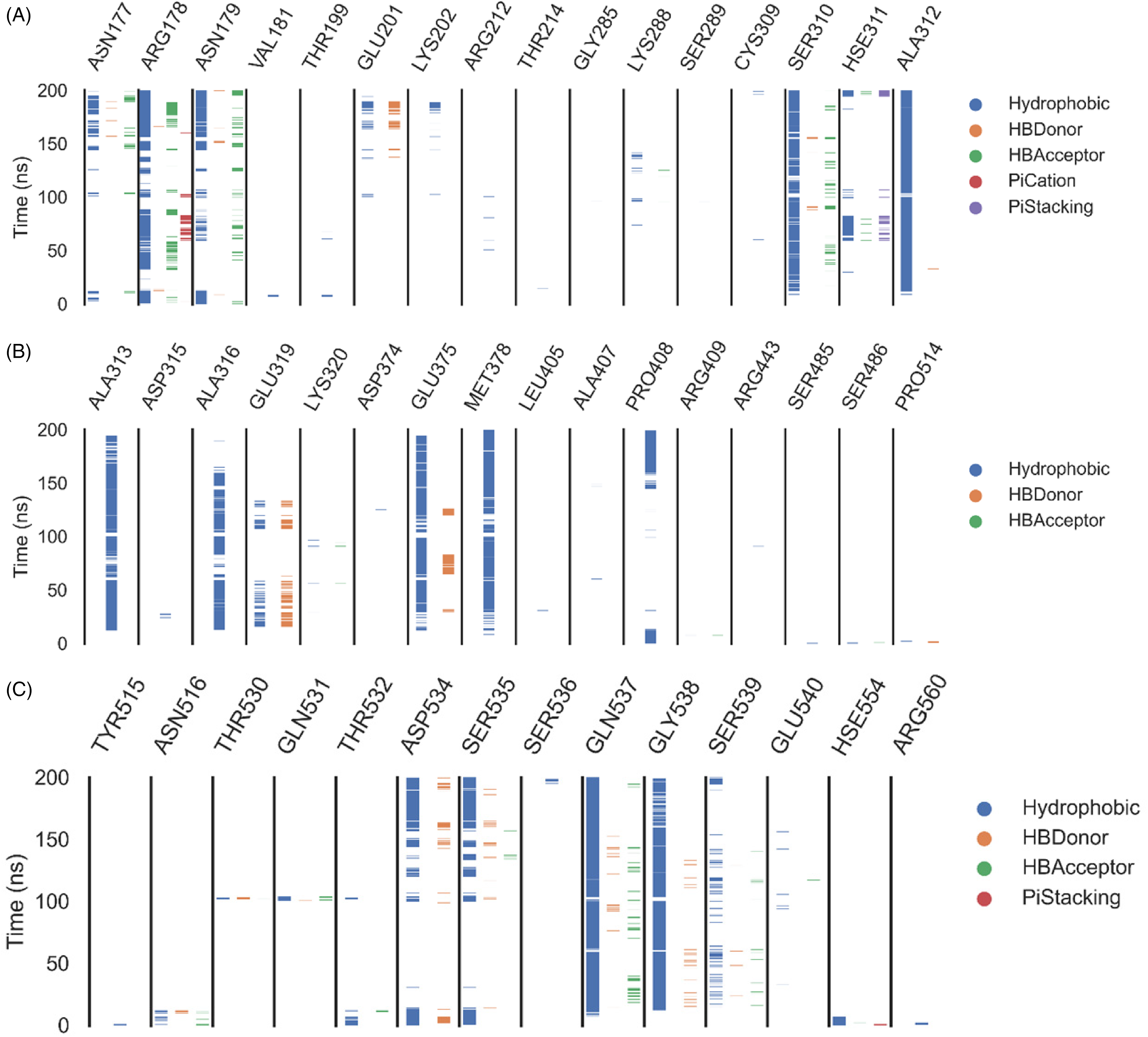
ProLIF python library output: the interacted helicase amino acids with compound
Protein–Ligand Interaction Profiles Study
PLIP, a prominent bioinformatics approach, plays a crucial role in dissecting and illustrating interactions occurring within molecular complexes involving a protein and a ligand. This computational tool offers invaluable insights into the binding interactions and noncovalent bonds established between the ligand and the protein, shedding light on the intricate molecular mechanisms governing ligand–receptor interactions. 74 Given its paramount relevance across diverse domains, including drug discovery, computational biology, and structural bioinformatics, PLIP has gained widespread adoption for the comprehensive examination of protein–ligand complexes and their interactions. When integrated with other computational methods such as molecular docking, MD simulations, and free energy calculations, PLIP enables a profound understanding of ligand–protein interactions. 75 Consequently, PLIP stands as a pivotal method in the realm of rational drug design. In our study, we utilized PLIP to extract 3D binding interactions in .pse file format from the representative frames obtained through clustering. 76 From the clustering-generated representative frames, the 3D binding interactions were extracted as.pse files using PLIP (Figure 10).
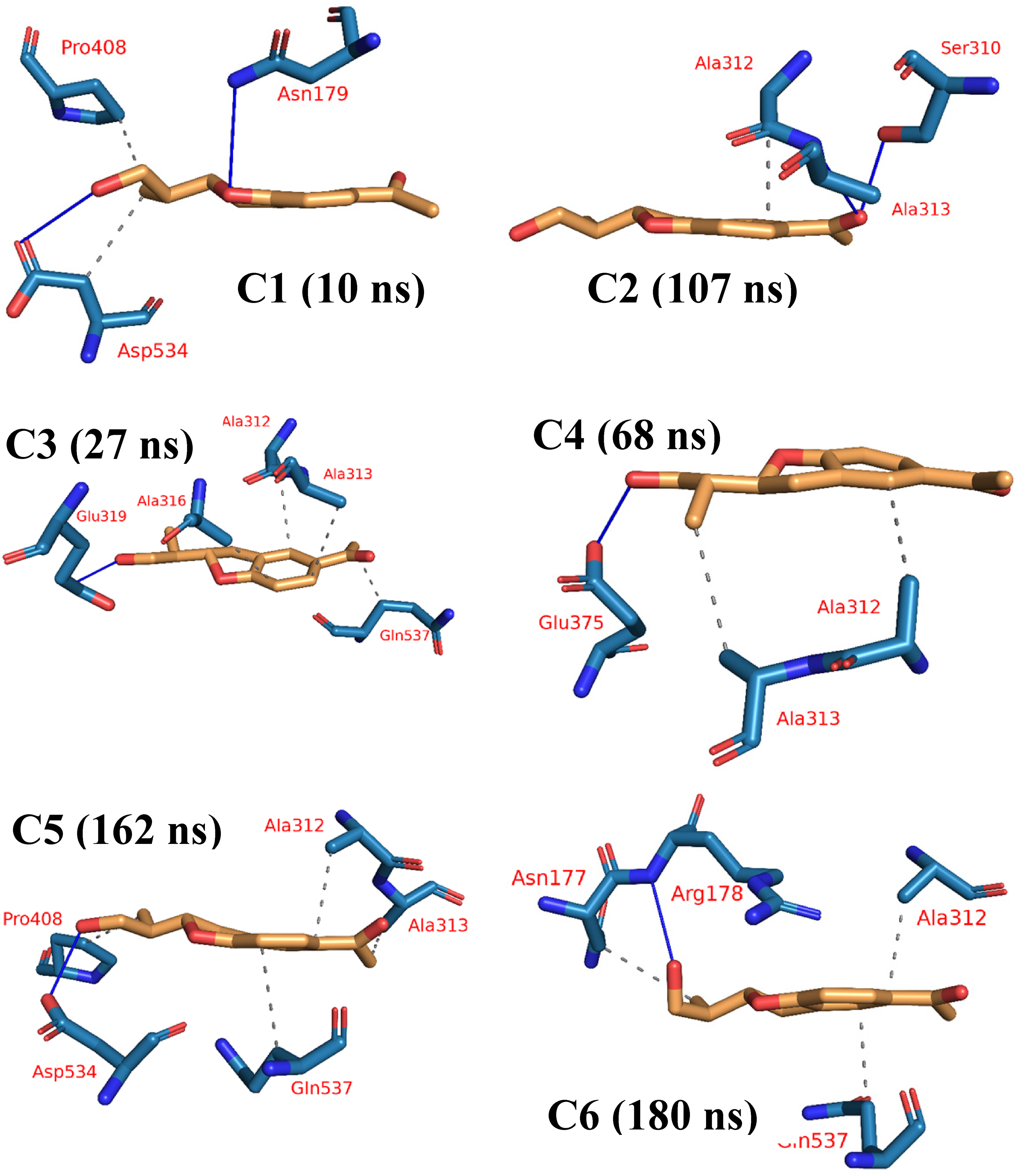
The six representative clusters obtained from TTClust analysis of compound
Principal Component Analysis of Trajectories Studies
PCAT, a well-established computational technique widely employed in the realm of MD simulations, plays a pivotal role in the analysis and extraction of essential collective motions from trajectory data. During an MD simulation, precise records of atomic positions within the complex biomolecular system are continuously logged over time, resulting in a comprehensive trajectory that intricately captures the system's dynamic behavior. 77 At this juncture, PCAT's intervention becomes indispensable, as it simplifies this intricate trajectory. This simplification is achieved by projecting the trajectory onto a lower-dimensional space while preserving the most critical motions. By orchestrating this simplification, PCAT effectively highlights the dominant motions governing the system's behavior and functionality, providing invaluable insights into the fundamental collective motions that underlie the biomolecular system's dynamics. This coordinated motion analysis was performed using principal component analysis (PCA).78,79
PCAT was utilized to identify the coordinated motions of the systems. According to the “Methods” section, we used the scree plot, the eigenvector distribution, and the variance preserved by each additional eigenvector (cumulative total) to determine the appropriate size of the reduced subspace. According to the scree plot, the slope seems to flatten out at the third PC. Additionally, the first eigenvector alone contained around 65.5% of the variation, while the first three eigenvectors together maintained over 79% of the overall variance (Figure 11). The first four principal components were shown to have a non-Gaussian distribution (Figure 12). Therefore, we chose the top three eigenvectors to determine the essential subspace.

Eigenvalues variation with increasing the eigenvectors (blue) with the cumulative variance retained in the eigenvectors represented (red).
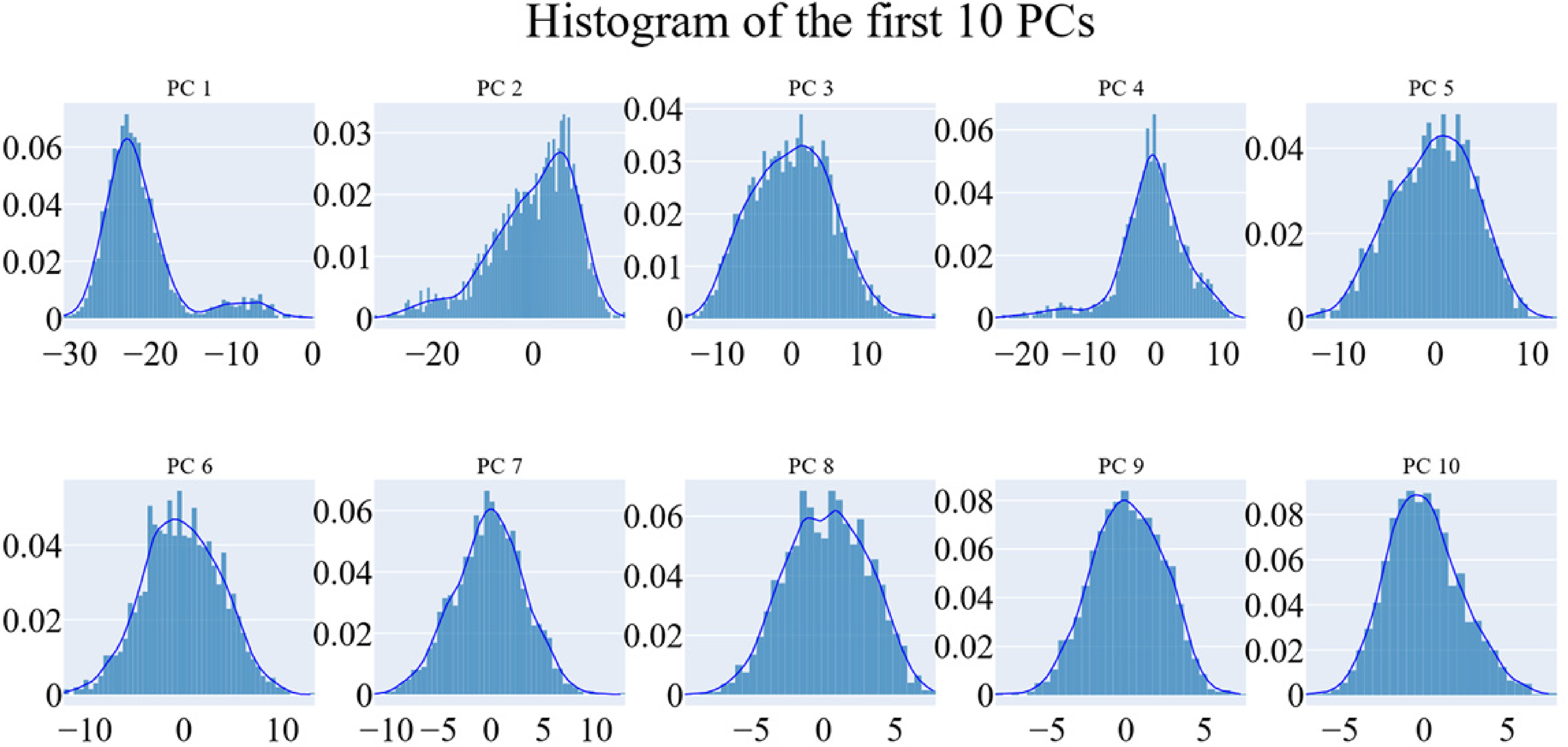
The distribution of the first 10 eigenvectors.
Both the apo helicase and holo helicase simulations had their cosine content calculated, and the randomness of the first 10 eigenvectors was evaluated. In both the apo helicase and holo helicase proteins, the cosine content of the first 10 eigenvectors was below 0.2 (Figure 13). The similarity between the two subspaces (the first three eigenvectors) is 34.6% according to the Root Mean Square Inner Product (RMSIP), showing that the two trajectories are more or less comparable. In addition, the RMSIP indicated that the C matrices are 48.7% identical proving the similarity between the two trajectories.
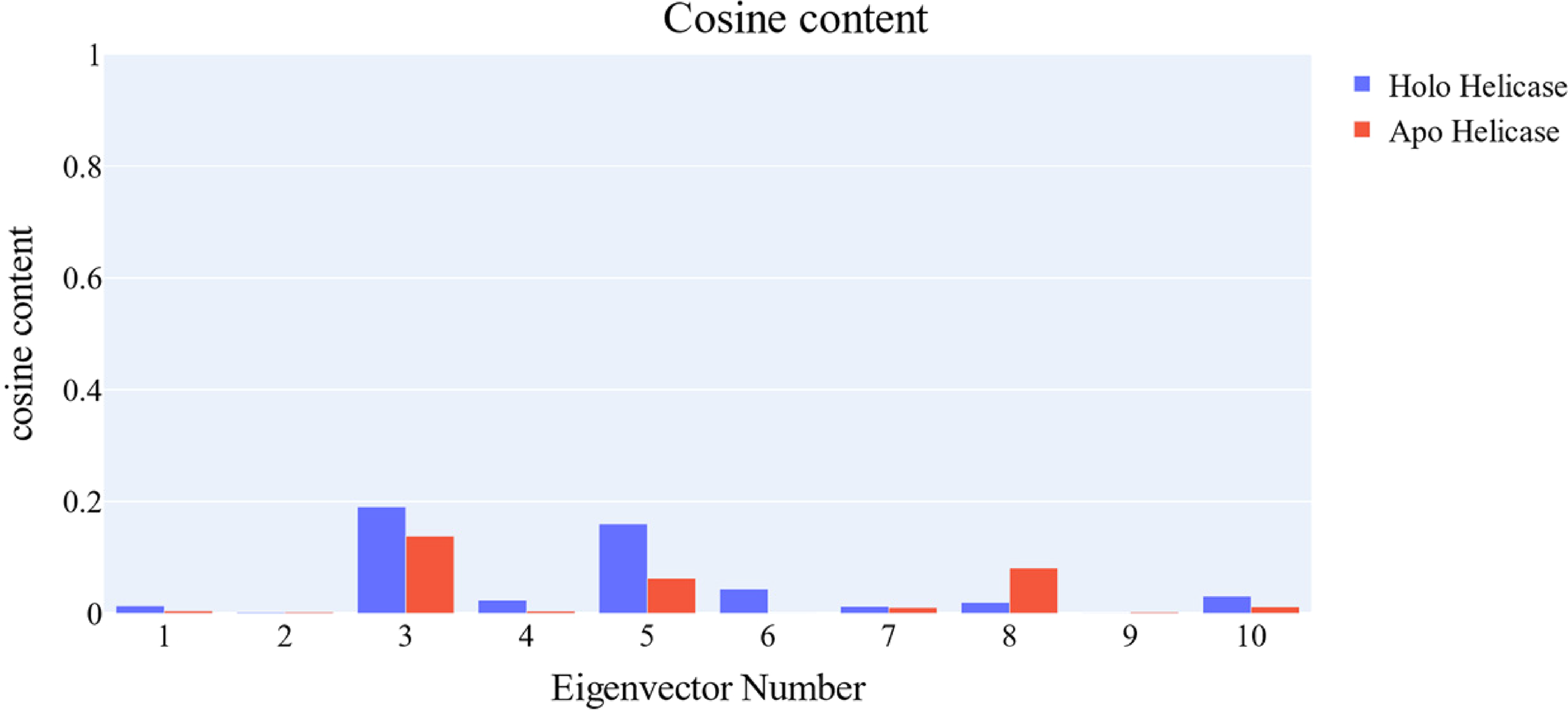
Cosine content of the first 10 eigenvectors for the two (apo helicase and holo helicase proteins) trajectories.
Figures 14--16 analysis of the results obtained of projecting each trajectory onto the first three eigenvectors of the new C matrix. Larger dots in these graphs indicate average structures for the indicated trajectories. Figure 14, a projection on the first two eigenvectors, reveals that the two trajectories have separate average structures and that the frames represent unique sampling with just a little amount of overlap before (pale red and white dots) and throughout the simulation (gray and orange dots). As can be seen in Figure 15, the two trajectories show approximate complete overlap and have roughly similar average structures. Figure 16 displays the projection on the second and third eigenvectors, which shows a similar trend to the first projection with just a little amount of overlap between the two trajectories.
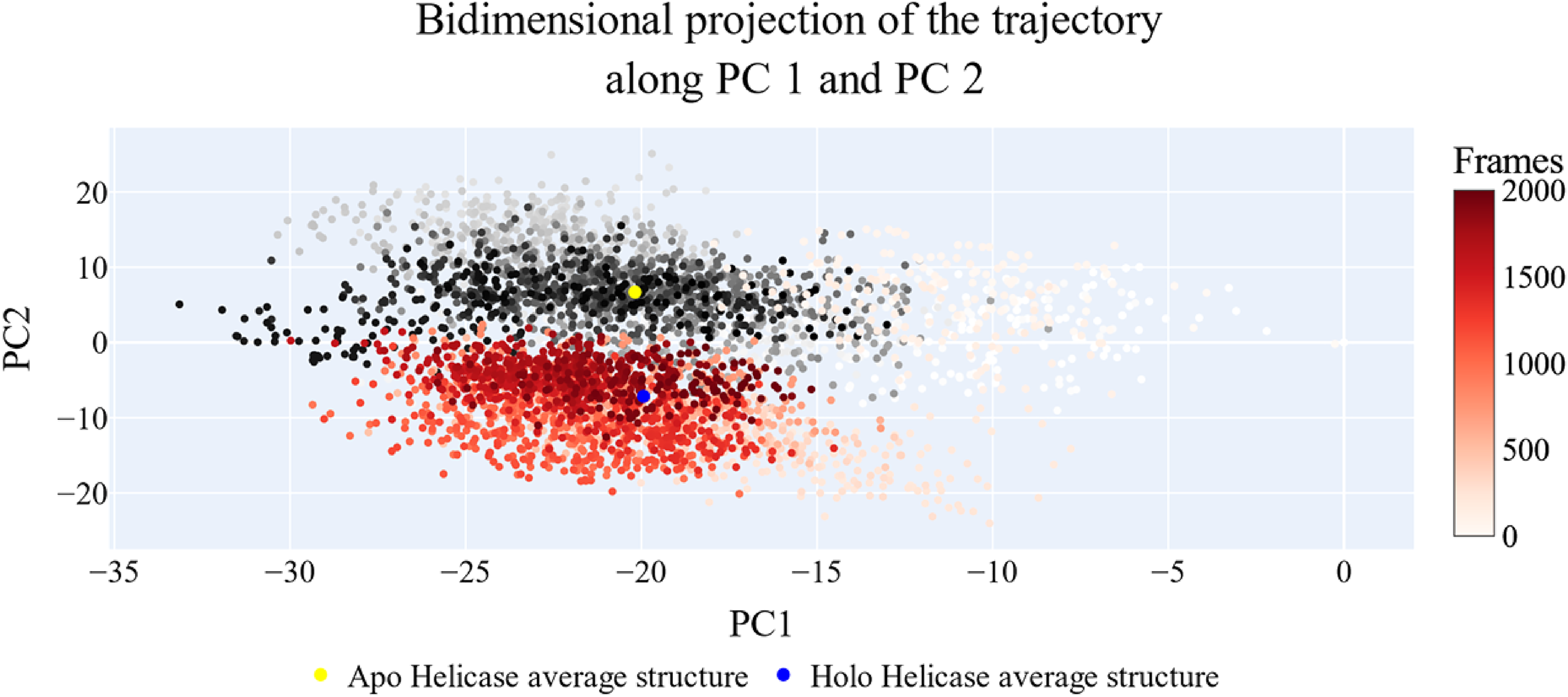
First two eigenvectors projection.

First and third eigenvectors projection.

Second and third eigenvectors projection.
The results obtained from the study are promising and showcase significant potential in the drug discovery of anti-COVID-19 treatments. However, it is imperative to acknowledge that the study primarily relies on in silico methodologies, involving computational simulations and predictions. While these computational techniques provide valuable insights and serve as a crucial initial step, their outcomes necessitate validation through actual experimental confirmation. Recognizing this, it becomes crucial to translate the computational predictions into tangible experimental evidence to ensure the reliability and applicability of the identified compounds especially compound
Method
Molecular Similarity Detection
The molecular similarity of the 4924 natural African metabolites was assessed using Discovery Studio 4.0, with VXG as the reference molecule to find the most similar 200 compounds among them to the VXG. The analysis included quantitative parameters such as the number of rotatable bonds, rings, aromatic rings, hydrogen bond donors, hydrogen bond acceptors, ALog p, molecular weight, and MFPSA. Following compound preparation, the small molecules panel with library analysis was employed. Similar molecules were identified by numeric properties, using the prepared compounds as input ligands and VXG as the reference. The task was set to find the most similar molecules (30 in total) based on Euclidean distance. The output chart of structural similarity was visualized to interpret the results, as elucidated in greater detail within the Supplementary Data section.
Fingerprint Studies
The fingerprints of the tested 200 compounds, with VXG as the reference molecule, were assessed using Discovery Studio 4.0. These fingerprints were based on parameters such as the number of charges, hybridization, hydrogen bond acceptors, hydrogen bond donors, positive and negative ionizable atoms, halogen atoms, aromatic rings, and ALogP (octanol–water partition coefficient). The protocol involved preparing the compounds, activating the small molecules panel, enabling the library analysis option, and selecting similar molecules by fingerprints. The input ligands were the prepared compounds and VXG as the reference. The task was set to find the most similar molecules (15 in total) using the Tanimoto coefficient, and the visualized output provided the results of the protocol as elucidated in greater detail within the Supplementary Data section.
Docking Studies
The SARS-CoV-2 helicase crystal structure (PDB ID: 5RMM) was prepared for docking by removing water molecules and retaining one protein chain with the co-crystallized ligand VXG. Protonation and energy minimization were performed using MOE 2019 and MMFF94x, defining the active site based on residues within 5 Å of VXG. 2D structures of compounds and VXG were converted to 3D structures after protonation and energy minimization using MM2 force field. MOE 2019 conducted protein–ligand docking, validated through VXG redocking. The accuracy of the protocol was confirmed by comparing the redocked VXG RMSD with the co-crystallized reference. For tested compounds, 30 docked solutions were generated, and Discovery Studio 4.0 visualized and analyzed results against the VXG reference 80 as elucidated in greater detail within the Supplementary Data section.
ADMET Studies
ADMET descriptors for compound evaluation were determined using Discovery Studio 4.0. The CHARMM force field was initially applied, followed by the preparation and minimization of the compounds. Models covered human intestinal absorption, aqueous solubility, blood–brain barrier penetration, plasma protein binding, CYP2D6 inhibition, and hepatotoxicity. Molecules were filtered based on SMARTS® rules. After compound preparation, the small molecules panel activated the ADMET descriptors option. Prepared compounds were selected as input ligands, and all relevant ADMET parameters were chosen. The protocol output was visualized, generating an interpretable ADMET chart 81 as elucidated in greater detail within the Supplementary Data section.
Toxicity Studies
Discovery Studio 4.0 was utilized for computing toxicity parameters of selected compounds, followed by a comprehensive evaluation using the Toxicity Prediction (Extensible) Protocol (TOPKAT). TOPKAT employs validated quantitative structure toxicity relationship models to assess the compounds’ toxicity based solely on molecular structure. After compound preparation, the small molecules panel activated the toxicity prediction (extensible) option. Prepared compounds served as input ligands, and various toxicity models were selected. The similarity search task was enabled, and the detailed report task was set to generate a PDF file. The protocol output was visualized, presenting a concise toxicity report 82 as elucidated in greater detail within the Supplementary Data section.
Molecular Dynamics Simulations
In this study, we conducted a 200 nanoseconds (ns) standard unbiased MD simulation using GROMACS 2021. The aim was to assess the stability of helicase alone, the Helicase_VXG complex, and the Helicase_Compound 1552 complex while examining structural changes between the two complexes. Input files for GROMACS were generated via the CHARMM-GUI server's solution builder module. The solvated system, comprising both the docked complex and apoprotein, occupied an 11.8 nm cube using the TIP3P water model, with a 1 nm padding from the farthest atom. Charge neutralization was achieved by adding 0.154 M NaCl ions. The CHARMM36m force field defined amino acid parameters for the helicase protein, TIP3P water model, and ions. For compound 1552 and VXG, the CHARMM general force field (CGenFF) was employed for parameterization 83 as elucidated in greater detail within the Supplementary Data section.
Molecular Mechanics-Generalized Born Surface Area
The ligand's binding energy was calculated using the molecular mechanics-generalized Born surface area (MM-GBSA) method, implemented in the gmx_MMPBSA program. To assess the contribution of amino acids within a 1 nm proximity of the ligand, a decomposition analysis was performed.14,15 The ionic strength was adjusted to 0.154 M, and the solvation method (igb) was set to 5. The internal dielectric constant was set to 1.0, and the external dielectric constant to 78.5 84 as elucidated in greater detail within the Supplementary Data section.
Principal Component Analysis of Trajectories
PCAT was applied to the mass-weighted covariance matrix (C) obtained from atomic positions in MD trajectories. This approach has the potential to reveal patterns of coordinated motion within the system. Specifically, PCA was employed to investigate the patterns of alpha carbons within the helicase. The analysis focused on amino acids spanning from Gln11 to Asp583, excluding terminal amino acids 77 as elucidated in greater detail within the Supplementary Data section.
Conclusion
In conclusion, our comprehensive CADD study, utilizing 4924 African natural metabolites, has yielded promising results as through a rigorous selection process, we identified 11 metabolites, including O-coumaric acid, trans-cinnamic acid, trans-ferulic acid, and others, demonstrating potent inhibitory activity against the critical SARS-CoV-2 helicase enzyme. Our extensive MD simulations and energy calculations confirmed the stable binding of the most promising candidate, 1-((S)-2,3-dihydro-2-((R)-1-hydroxypropan-2-yl)benzofuran-5-yl)ethenone, to the SARS-CoV-2 helicase with low energy and optimal dynamics. These results were provided by MD simulations, MM-GBSA, PLIP, ProLIF, and PCAT studies over 200 ns giving valuable insights into the molecular mechanisms underlying their inhibitory activity. These highly encouraging findings serve as a strong foundation for the next phase of our research, which will involve in vitro and in vivo studies to validate the efficacy and safety of these selected compounds. While more work lies ahead, our study raises hope for the development of potential COVID-19 therapeutics and contributes to the global effort to combat this pandemic.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X241246738 - Supplemental material for Computer-Assisted Drug Discovery of Potential African Anti-SARS-CoV-2 Natural Products Targeting the Helicase Protein
Supplemental material, sj-docx-1-npx-10.1177_1934578X241246738 for Computer-Assisted Drug Discovery of Potential African Anti-SARS-CoV-2 Natural Products Targeting the Helicase Protein by Ahmed M. Metwaly, Mohamed S. Alesawy, Bshra A. Alsfouk, Ibrahim M. Ibrahim, Eslam B. Elkaeed and Ibrahim H. Eissa in Natural Product Communications
Footnotes
Acknowledgments
The authors would like to express sincere gratitude to AlMaarefa University, Riyadh, Saudi Arabia, for supporting this research.
Declaration of Conflicting Interests
All author(s) of this manuscript declare that they have no conflicts of interest to disclose. This includes, but is not limited to, any financial interests, professional relationships, or personal affiliations that might be perceived as influencing the research, analysis, or presentation of the findings contained within this work. The integrity and objectivity of this research remain paramount, and there are no competing interests that could compromise the impartiality of the study or its results.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2024R142), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Ethical Approval
Not applicable for this article as no experiments or studies involving human or animal participants were conducted as part of this research.
Statement of Human and Animal Rights
This article does not include any research involving human or animal participants.
Statement of Informed Consent
Not available as no experiments or studies involving human or animal participants were conducted as part of this research.
Trial Registration Number/Date
Not available.
Grant Number
PNURSP2024R142.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
