Abstract
Residual dipolar coupling (RDC) is a newly emerging technique for obtaining more precise conformational features of molecules in nuclear magnetic resonance (NMR), which can be observed when the molecules are oriented in NMR sample solutions. Unlike the case of solubilization of resveratrol (Res) in aqueous media by surfactants, multiglycosylated resveratrol (GlyRes) was dispersed in aqueous media forming nanoparticles with an extremely small size. In the presence of an NMR alignment medium of bacteriophage, GlyRes molecules were efficiently oriented in the strong magnetic field allowing for observations of RDCs in NMR. Structural refinement using the observed RDC values enabled differentiation of
For determination of three-dimensional structures of biomacromolecules and organic molecules, solution nuclear magnetic resonance (NMR) signals such as nuclear Overhauser effect (NOE), residual dipolar couplings (RDCs), and 3
The prerequisite for the RDC measurement is preparations of alignment media to orient target molecules in NMR samples. As the importance of RDC in protein structural analysis has been revealed, alignment media in aqueous solutions such as a filamentous virus bacteriophage and disk-shaped phospholipid bilayers, the so-called bicelle have been extensively developed. However, as for the RDC analysis of hydrophobic organic natural products, it has been difficult to orient the molecules due to aggregations in the aqueous solutions. Although lyotropic liquid crystals or stretch gels have been developed for alignment media of organic molecules, 5 preparations of these alignment media are much complicated. We should consider the following factors for the solubilization of organic molecules in order to effectively orient the molecules for observation of RDCs. Smaller nanoparticles are better to maintain anisotropy of molecules. Smaller nanoparticles are also advantageous to observe sharp NMR data because nanoparticles having low mobility tend to give broadens NMR data. In addition, it is preferable that molecules for the solubilization are covalently bound to the target molecules not to hamper functions of alignment media coexisting with the target molecules.
So far, we have investigated the glycosylation reaction of hydrophobic molecules using plant culture cells.
6
Through these works, we noticed that glycosylation would be a reasonable way for measurements of RDCs (Figure 1(F)). In this method, small molecules of glucose are covalently conjugated with the target molecules forming water-soluble small nanoparticles. In addition, the glycosylation method by plant culture cell is widely applicable to hydrophobic organic molecules with hydroxyl groups. Unlike the case of solubilization using surfactants which possibly deteriorate alignment media, covalent modification of glucose units is supposed to minimize the unfavorable effects on the alignment media. As a proof-of-concept work, we herein report RDC analysis of a stilbenoid backbone
7
-9
using aqueous dispersions of multiglycosylated resveratrol (GlyRes) with commonly used bacteriophage-based alignment media.
10
Resveratrol (Res) is a suitable model target because we can change the number of modified glucose molecules by biotransformation, and we can compare

Molecular structures of (A) resveratrol (Res), (B) piceid, (C) multiglycosylated Res (GlyRes), (D) Triton-X, and (E) sodium dodecyl sulfate (SDS). (F) Schematic illustration for residual dipolar coupling (RDC) measurement of GlyRes with alignment media of bacteriophage.
First of all, we investigated the dispersibility of GlyRes in water. Figure 2 shows characterizations of aqueous dispersions of Res (Figure 1(A)), Res modified with 1 glucose unit (piceid; Figure 1(B)), GlyRes (Figure 1(C)) and Res solubilized with a representative neutral surfactant of Triton-X (Figure 1(D)) or anionic sodium dodecyl sulfate (SDS; Figure 1(E)) (Triton-X⊃Res and SDS⊃Res, respectively). As can be seen in the result of piceid (Figure 2(B)), modification of 1 glucose unit was not enough for Res (Figure 2(A)) to be water soluble, and the particle size hardly changed as analyzed by dynamic light scattering (DLS) measurements (Figure 2(F)). In contrast, GlyRes was nicely dispersed in water to give a transparent solution (Figure 2(C)), whose particle size was markedly small less than 2 nm (Figure 2(G)) indicative of negligible aggregation. It is noteworthy that the ultrasmall nanoparticles are hard to prepare by surfactant-based solubilizations. Indeed, the samples of Triton-X⊃Res and SDS⊃Res showed cloudy dispersions (Figure 2(D) and (E)). In line with the naked-eye observations, DLS showed larger particle sizes than that of GlyRes indicating Res molecules were somewhat agglutinated (Figure 2(H) and (I)).

Naked-eye observation of dispersions of Res (0.4 wt%) (A), piceid (0.4 wt%) (B), GlyRes (2.0 wt%) (C), Triton-X⊃Res (2.4 wt%) (D), and SDS⊃Res (2.4 wt%) (E). DLS profiles of piceid (F), GlyRes (G), Triton-X⊃Res (H), and SDS⊃Res (I) as compared with Res. DLS, dynamic light scattering; SDS, sodium dodecyl sulfate.
Next, we performed CLIP-HSQC measurement of GlyRes, Triton-X⊃Res, and SDS⊃Res to evaluate the effects of the particle size on NMR profiles. As expected, the sample of GlyRes forming small-sized nanoparticles gave sharp NMR signals (Figure 3, Supplementary Material), while broad NMR peaks were observed form the Triton-X⊃Res sample (Figure 3(B)). In the case of SDS⊃Res, relatively sharp NMR peaks were observed (Figure 3(C)) as well as the GlyRes in spite of the larger particle size in DLS. We envisioned that a portion of SDS⊃Res was dispersed as small nanoparticles because of electrostatic repulsion of charged SDS molecules. We also checked effects on the alignment media of bacteriophage. For evaluating the performance of aligning agents quantitatively, 2H quadrupole splittings for the D2O dispersions in the presence of bacteriophage (0.4 wt%) were measured. As shown in Figure 3(D), quadruple splittings of around 4 Hz were observed from the GlyRes and Triton-X⊃Res samples, which was comparable to the previously reported value of a bacteriophage solution without additive. 10 Nevertheless, the SDS⊃Res dispersion did not show quadrupole splitting, probably because SDS molecules caused denaturation of the bacteriophage to lose the orientation capability. That is to say, GlyRes was ideal for observing sharp NMR peaks in CLIP-HSQC with keeping the orientation capability of bacteriophage.

1H-13C CLIP-HSQC spectra of GlyRes (2.0 wt%) (A), Triton-X⊃Res (2.4 wt%) (B), and SDS⊃Res (2.4 wt%) (C). (D) 2H quadrupole splitting profiles of GlyRes (2.0 wt%), Triton-X⊃Res (2.4 wt%), and SDS⊃Res. (2.4 wt%) in the presence of bacteriophage (0.4 wt%). CLIP-HSQC, clean inphase-heteronuclear single quantum coherence; SDS, sodium dodecyl sulfate.
We then carried out 1H-13C CLIP/CLAP-HSQC for measuring RDCs and evaluated correlations to the
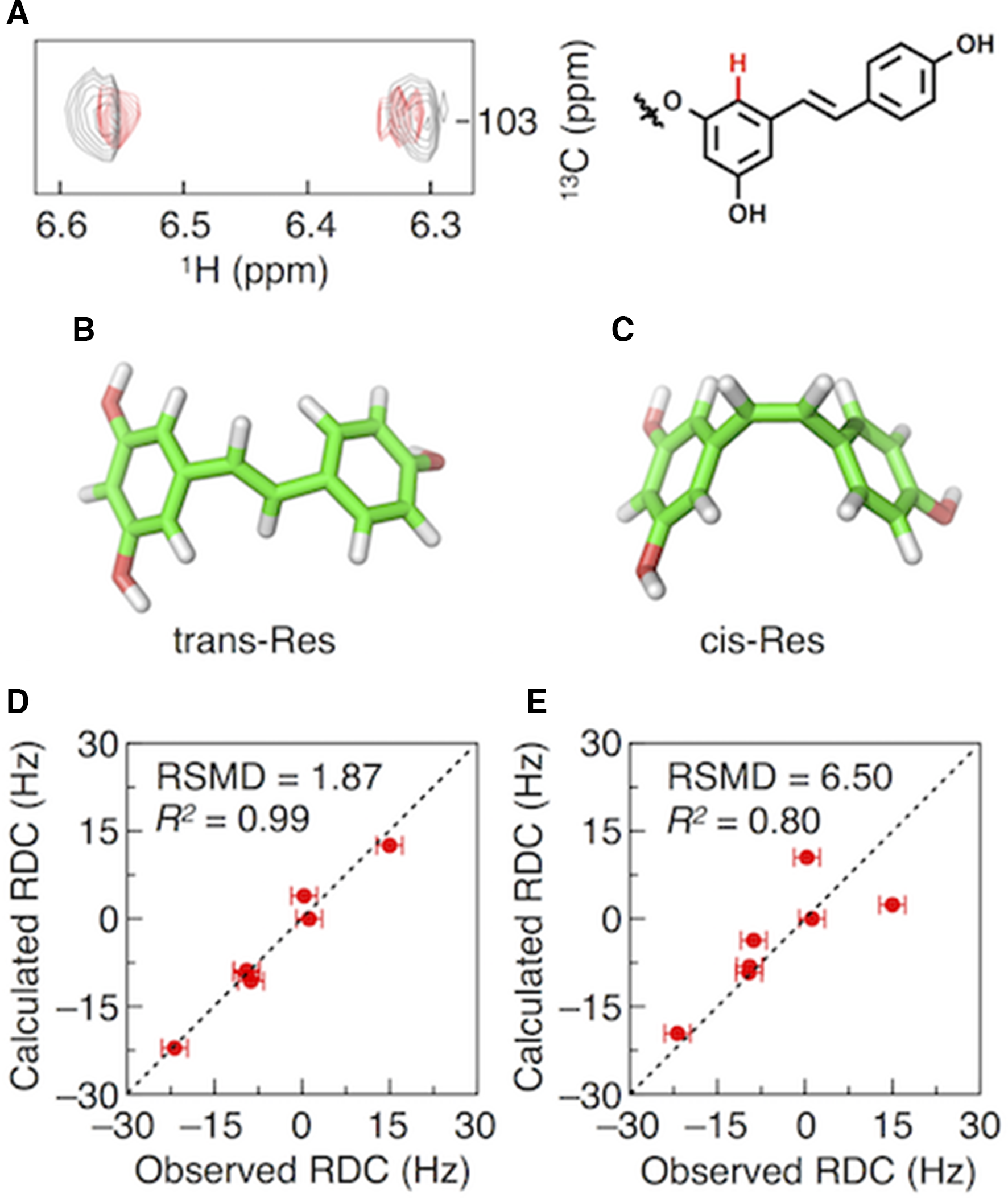
(A) Nondecoupled 1H-13C subspectra of GlyRes without (gray) and with (red) bacteriophage (0.4 wt%) at the chemical-shift of a C-H bond highlighted as red part in the molecular structure of GlyRes. Molecular models of
In summary, we have reported the application of multiglycosylated GlyRes to RDC-based NMR analysis. GlyRes formed ultrasmall nanoparticles suitable for RDC measurements with keeping the orientation capability of alignment media of bacteriophage. GlyRes oriented in the presence of bacteriophage gave sufficient RDC signals in CLIP/CLAP-HSQC spectra allowing for differentiation of
Experimental
General
NMR spectra were recorded on a Bruker model AVANCE-600 (600 MHz). Ultrasonication was performed by using a QSonica model ultrasonic homogenizer. Particle sizes were measured by using a Malvern model Zetasizer Nano ZSP zeta potential analyzer (DLS). Res, Triton-X, and SDS were purchased from TCI. Pf1 bacteriophage was purchased from ASLA Biotech. GlyRes was obtained by biotransformation using plant culture cells. The number of modified glucose as calculated by high-performance liquid chromatography was 2.7 (Supplementary Materials).
Preparation of Triton-X-Res and SDS-Res Nanoparticles
Res (0.4 wt%) was mixed with Triton-X (2.0 wt%) or SDS (2.0 wt%) in water and ultrasonicated at 50 W for 2 minutes to disperse homogeneously.
NMR Analysis
To obtain 1H-13C RDCs of GlyRes (2 wt%), CLIP/CLAP-HSQC
12
(o1p/o2p [ppm] =6.5/90, SW [ppm] =15/180, TD = 8 k/1 k, DS/NS = 16/4) were performed for 3.5 hours in aqueous solutions with and without bacteriophage (0.4 wt%), and peak positions were determined by using Sparky program. Molecular models of
Supplemental Material
Supplementary Material 1 - Supplemental material for Water-Soluble Glycosylated Resveratrol for Measuring Residual Dipolar Coupling in NMR
Supplemental material, Supplementary Material 1, for Water-Soluble Glycosylated Resveratrol for Measuring Residual Dipolar Coupling in NMR by Hiroki Hamada, Masayoshi Yanagi, Kei Shimoda and Noriyuki Uchida in Natural Product Communications
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
