Abstract
Apigenin is a natural flavone that possesses excellent biological activities especially against aging and cancer. However, the underlying mode of its action is not yet revealed. The purpose of this study was to examine the pharmacological mechanisms of apigenin using the knowledge of network pharmacology, protein-protein interaction (PPI) databases and biological processes analysis through Cytoscape.
Apigenin targets were retrieved through PASS Prediction and STITCH database and the interactive associations between these targets were studied using STITCH, followed by GO (gene ontology) and pathway enrichment analysis.
As a result of target search, 125 protein targets were retrieved. Moreover, 216 GO terms related to various biological processes, 16 GO terms for various molecular processes, 5 GO terms for the cellular components, and 52 Kyoto Encyclopedia of Genes and Genomes pathway terms were achieved by analyzing gene functional annotation clusters and abundance values of these targets. Most of these terms are strongly associated with inflammation through various pathways, for example, FOXO, mammalian target of rapamycin, tumor necrosis factor, p53, AMP-activated protein kinase, p13K-AKT, and mitogen-activated protein kinase, which play an important role in inflammation, aging and cancer.
Apigenin can be used to treat inflammation, aging, and cancer with an underlying mechanism of inflammation suppression. This study contributed excellent information for a better understanding of the modes of action of apigenin. However, further studies such as docking and MD simulation are required to understand the therapeutic and toxicological roles of these targets of apigenin.
Like other flavones, apigenin (4′,5,7-trihydroxyflavone) 1 is abundantly found in several fruits, such as apples, oranges, and grapefruits, 2 and vegetables, including chamomile, celery, parsley, and onions, 3 which constitute the routine diet of human. Moreover, apigenin also exists in tea, celery, yarrow, tarragon, cilantro, foxglove, coneflower, licorice, flax, passion flower, horehound, spearmint, basil, oregano, red wine, and beer. 4 Apigenin (Figure 1) has been isolated from the plants Matricaria recutita and Gingko biloba as free apigenin 5 and in the form of various acylated derivatives, such as apigenin-7-O-glucoside. 6,7 Mediterranean food is also rich in apigenin, which is considered responsible for lower incidence of cardiovascular disorders, obesity, diabetes, and cancer. 8 -10 Apigenin is synthesized through chemical modification of the flavanone naringenin by inserting a double bond to its ring C. However, apigenin and naringenin have significantly different bioactivities regardless of their structural similarity. In contrast to naringenin, 11 apigenin mediates apoptosis of various cell lines. 12 -14 Apigenin possesses excellent activity against inflammation 15 via activation of the nuclear factor B (NF-B)-mediated pathway. 16 Moreover, there is reduced risk of ovarian risk owing to the consumption of apigenin, as evident from large population-based studies. 8 Besides, several dietary supplements also have apigenin in excess. 17 Therefore, it acts as an excellent natural compound from which to recognize the entire set of human target proteins as an initial step to recognize how dietary phytochemicals provide health advantages.

Chemical structure of apigenin.
In recent times, there is an increasing trend of using network pharmacology to explore the action of herbals. 18,19 Network pharmacology is a systems biology-based approach. Its application in drug discovery has accelerated the process of scientific validation of drug action. 20,21 For instance, Gan et al used network pharmacology involving STITCH and Cytoscape plugin bingo to predict the molecular targets of curcumin that were involved in suppressing inflammation. 22 In addition, network pharmacology is also utilized to highlight the ameliorated drug efficacy by constructing the interactive channels between different signaling pathways. 23 -25
The living body performs various functions through a large number of signaling proteins, which work together as networks. These networks represent protein-protein interactions (PPIs), which work in association with a number of cellular signaling pathways 26,27 to accomplish the biological activities. 28 -31 The gene ontology (GO) project 32 has played an important role in creating the ontologies for gene annotations. GO enrichment analysis is a statistical procedure that can be utilized to predict the association between annotations to GO and proteins. GO enrichment and network analysis together can be supportive to envisage the molecular mode of apigenin activities.
Apigenin has been proved to have anticancer activity, mainly through inhibition of oxidation and inhibition, triggering of cell-cycle arrest and apoptosis, attenuation of proliferation, and provoking of detoxification enzymes. Currently, anticancer activity data of apigenin describes only 1 or 2 pathways and are available in split form, showing the lack of an overall multi-target regulation of apigenin.
Owing to this breach in knowledge, this study was designed to examine the pharmacological mechanisms of apigenin using the knowledge of network pharmacology, PPI databases, and biological processes analysis through Cytoscape. This study will not only act as a reference for clinical trials of apigenin, but also assist to produce analog molecules.
Methodology
Target retrieval through PPI database and network development and analysis were collectively used to investigate molecular targets and biological effects of apigenin. In the first stage of this study, molecular targets of apigenin were searched for using 2 well-renowned databases, i.e. PASS Prediction and STITCH. Targets obtained from both databases were merged, excluding any duplication. Then, an apigenin-target interaction network was constructed and analyzed by using the STITCH database. Finally, GO enrichment analysis and biological processes analysis were carried out using Cytoscape-plugin ClueGO to examine the molecular basis of apigenin activities.
Search and Prediction of Apigenin Targets
PASS Prediction (http://www.pharmaexpert.ru/PASSOnline/) was utilized to retrieve the apigenin targets. 33 PASS Prediction is a database developed to find various types of biological actions of small organic molecules on the basis of their chemical structures, mainly before their chemical synthesis and biological analysis. This database requires “SMILES” or structural formula of a compound in MOLfile format or “SMILES” to identify molecular targets with a probability of Pa >0.7 (probability to be active). In addition, STITCH 5.0 database (http://stitch.embl.de/) 34 was utilized to retrieve known protein targets of apigenin. To explore protein targets of an active organic compound and their interactions, this online database searches several sources of information including experimental confirmations, genomic context, high-throughput experiments (conserved) coexpression, and text mining for a large number of organisms. The database presently is composed of 9 643 763 proteins from 2031 organisms. In addition, the interactions taken from STITCH are of 2 types including direct (physical) and indirect (functional) type. Probabilistic confidence score plays an important role in finding the interactions using the STITCH database, thus a probabilistic confidence score of greater than 0.7 was used to predict protein targets.
Network Construction, GO, and Pathway Enrichment Analysis
After retrieving targets through PASS Prediction and STITCH related to homo sapiens, a network comprising apigenin targets was constructed and analyzed. STITCH database exhibits an excellent feature of network construction and then its analysis. On the basis of biological terms, target genes were divided into a structured hierarchy by using GO biological processes for the identification and analysis of the characteristic features of potential targets. To investigate the modes of apigenin for aging and cancer, pathway enrichment analysis was introduced. GO terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway terms were acquired from STITCH. Moreover, KEGG (http://www.kegg.jp/) database was utilized for more information about these pathways. Afterwards, the apigenin-target network was also analyzed by using ClueGO, a plug-in of Cytoscape to investigate the mode of apigenin and its pharmacodynamics significance. Cytoscape 3.4.0 has an excellent feature of network visualization and analysis and is supplied with numerous plug-ins. 35 Enrichment analysis by ClueGO was conducted by having set the level of significance at 0.05. Besides, a medium network type and 2-sided hypergeometric test with a Bonferroni correction was selected. In addition, the functional network was visualized via an organic layout algorithm.
Results
Retrieval and Identification of Protein Targets
Through PASS Prediction (accessed in April 2017), a total of 105 potential targets of apigenin in human with Pa >Pi (Table 1) were found. On the other hand, 20 potential human protein targets (Table 2) were retrieved via STITCH (accessed in April 2017) setting a confidence score at 0.7. Targets retrieved through both searches were merged, excluding any repeated target. Overall, 125 protein targets with numerous physiological and/or pathological activities were retrieved. These proteins are involved in pharmacokinetic and inflammatory processes.
List of Targets Retrieved Through PASS Prediction Analysis of Apigenin (Pa > 0.7).

Predicted Targets of Apigenin Obtained Through STITCH Database (Confidence View).

Protein Interaction Network
The STITCH 5.0 database was utilized to construct the interaction network of target proteins (encoded by the respective genes) (Figure 2), which were indicated as nodes. These nodes were connected with each other through lines, known as edges leading to the formation of protein pairs. The obtained protein interaction network consisted of 20 proteins and 49 edges. PPI enrichment P-value for the obtained protein-protein network was 0.00642. This small value indicates that the observed number of edges is significant and that the nodes are not random. In addition, the number of edges, if the nodes are randomly selected, is denoted by a term known as the expected number of edges: the value in the presently discussed network is 33. Besides, the network stats indicated that the average node degree and clustering coefficient were 4.9 and 0.688, respectively. At the threshold score, the average of number of interactions of a protein in a network is known as the average node degree, while the extent of connectivity of nodes in a network is denoted as the clustering coefficient. A high value of the clustering coefficient indicates that the network is highly connected A node degree is a quantitative property of a node and reflects the number of interactions of a node. The nodes with a number of interactions higher than the average node degree are called hubs. The degree of each node is given in Table 2. In the obtained network, there are 12 nodes with the average node degree higher than 4.9; thus, this network contains 12 hubs. For instance, the highest degree is noted for TP53 (15), followed by AKT1, vascular endothelial growth factor A, and cyclin D1 having 12, 9, and 9 degrees, respectively, and so on. Most of the protein nodes, especially hubs, in this network are involved in the process of inflammation, aging, and cancer development. Table 3 shows that apigenin activates ESR1, ESR2, CASP3, PARP1, UGT1A1, TP53, ELAVL1, and TNFRSF10B, while ESR1, CDK1, PTGS2, CYP1B1, AKT1, MAOA, SPAM1, VEGFA, MMP9, CCND1, CYP19A1, and HIF1A are inhibited by apigenin. Apigenin undergoes binding with ESR1, PTGS2, CYP1B1, MAOA, ESR2, and CYP19A1. In addition, apigenin plays a role in the expression of ESR1, UGT1A1, AKT1, TP53, VEGFA, CYP19A1, HIF1A, and TNFRSF10B. Their respective scores are mentioned in Table 3.

Confidence view of the protein network of apigenin. This view is composed of nodes and edges signifying the proteins and their interactions, respectively. Thick lines specify the stronger associations. Gray and green lines characterize the protein-protein and chemical-protein interactions.
Predicted Targets of Apigenin Obtained Through STITCH Database (Action View).

GO and Pathway Analysis
The STITCH 5.0 database was further used to analyze the biological functions of these potential targets through functional enrichment of the network. As a result of this analysis, 216 GO terms related to various biological processes were retrieved. It is worthy of mention that most of these targets are involved in the multiple biological responses such as positive regulation of reactive oxygen species metabolic process, regulation of cell proliferation, regulation of protein serine/threonine kinase activity, positive regulation of angiogenesis, response to hypoxia, intrinsic apoptotic signaling pathway, and positive regulation of cellular protein metabolic process. Moreover, the retrieval of 16 GO terms associated with various molecular processes was achieved in this analysis. Most of the molecular functions (GO) were linked with enzyme binding, histone acetyltransferase binding, estrogen receptor activity, and nitric-oxide synthase regulator activity. In addition, 5 GO terms related to various cellular components were also achieved. It mainly included the intracellular organelles such as mitochondria. In addition, the retrieval of 52 KEGG pathway terms with a false discovery rate (FDR) <0.05 (Table 4) was achieved in this analysis. Most KEGG pathway terms were related to xenobiotic metabolism, induction of hepatitis, aging, and development of various cancers such as those of the thyroid, prostate, bladder, colorectal, pancreatic, and renal. All these terms had the lowest FDR. These pathways are associated with FOXO, mammalian target of rapamycin, tumor necrosis factor (TNF), p53, AMP-activated protein kinase, p13K-AKT, and mitogen-activated protein kinase (MAPK), which play an important role in inflammation, aging, and cancer.
Functional Enrichments in Network (Kyoto Encyclopedia of Genes and Genomes Pathways).

Additional enrichment analysis of this protein interaction network was conducted by using GO terms (GO terms) for annotation of the biological functions of apigenin-related targets, employing Cytoscape-based plugin ClueGO. Overall, the significant enrichment of 49 GO terms was obtained (Table 5). Figure 3 illustrates the grouping of these GO terms into 20 subclasses, which are largely involved in macrophage differentiation, intracellular estrogen receptor signaling pathway, response to estradiol, estrogen metabolic process, mitotic G1 DNA damage checkpoint and positive regulation of epithelial cell migration. These biological processes help us better comprehend the mechanisms of apigenin action.

Functionally grouped networks, retrieved via ClueGO analysis to identify the potential targets of apigenin. Each class of targets consists of the most important terms only. The overlapped classes reflect their functional likeness.
GO Terms and Their Associated Genes Obtained Through ClueGO Analysis.
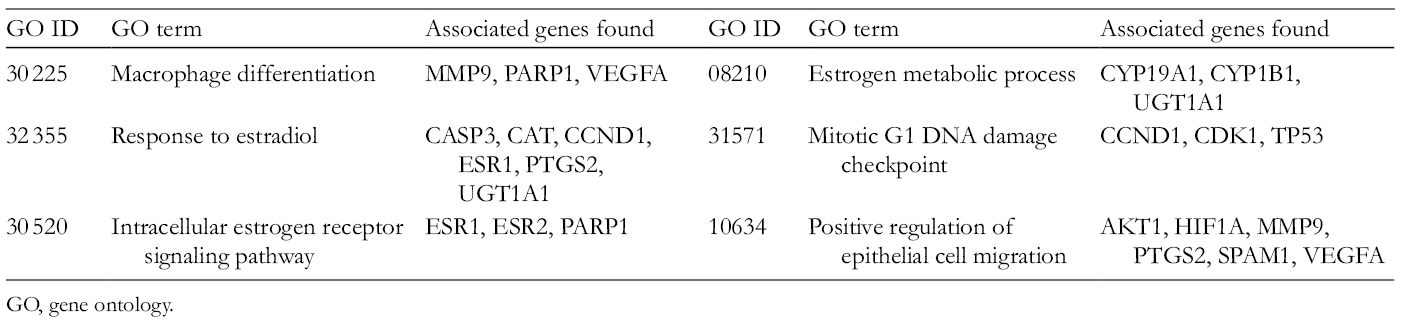
GO, gene ontology.
Discussion
Apigenin possesses excellent antiaging and anticancer activities. 2 However, the precise modes of its action are still uncertain. Therefore, this system pharmacology study was designed and executed via a combination of drug target retrieval, network construction, and pathway analysis.
Our results elaborated a total of 125 potential targets of apigenin. GO analysis of protein targets and target network analysis revealed the excellent effect of apigenin against aging and cancer, largely by inhibiting the inflammatory response. At the same time, the pathway analysis in our work elaborates that apigenin could instantaneously regulate multitargets/pathways coupled with multiple therapeutic modules, for instance, the inhibition of inflammation.
Apigenin has already been documented to mediate various activities against cancer, such as inhibition of oxidation and inhibition, triggering of cell-cycle arrest and apoptosis, attenuation of proliferation, and provoking of detoxification enzymes. Apigenin has been proved to have an inhibitory effect on (i) mutagenicity against nitropyrene-induced genotoxicity in Chinese hamster ovary cells, 5 benzoapyrene, and 2-aminoanthracene-induced bacterial mutagenesis, 5 (ii) carcinoma of skin and colon, 3,7 (iii) ornithine decarboxylase, 36 and (iv) UV-induced cancer. 3 In addition, apigenin-mediated accumulation of intracellular glutathione enhances the endogenous defense against oxidative stress. 15 Apigenin exerts its effect against inflammation through inhibition of cyclooxygenase-2, 15 nitric oxide synthase-2 activity, 37 and TNF-induced nuclear factor (NF)-ĸB. 38 Moreover, literature study reveals multiple molecular signaling effects of apigenin, 39 for instance, inhibition of protein kinase C activity, MAPK, 40,41 protein-tyrosine kinases and peroxisome proliferation regulated kinase (ERK), 42 Na+/Ca2+-exchanger, 43 phosphorylated EGFR tyrosine kinase and nuclear substrate c-myc, 44 casein kinase 2, 45 and p34 (cdc2) kinase activity. 46,47 Antiapoptotic activity of apigenin is mediated through triggering of WAF1/p21, 48 altering the Bax/Bcl-2 ratio, 49 regulating protease production, 50 inhibiting TNF-induced intracellular adhesion molecule-1 upregulation, 51 inhibiting expression of HIF-1, 52 VEGF, 53 aromatase, 54 17β-hydroxysteroid dehydrogenase, 55 and IGF-I. 56 Many other proteins also act as targets of apigenin, for example heat shock proteins, 57 telomerase, 58 fatty acid synthase, 59 matrix metalloproteinases, 60 aryl hydrocarbon receptor activity, 58 and HER2. 43 These proteins play an important role in developing cancer. However, reliable scientific confirmations are really required to associate these genes with the system effects of apigenin.
It is also evident from the literature that apigenin could suppress inflammatory responses and treat the impairment caused by inflammation via inhibiting the expression of IL-6 and TNF-α and ameliorating the expression of IL-2, IL-10, and IFN-γ. 5 In this study, GO enrichment, network, and pathway analyses showed that apigenin considerably enriches target genes involved in suppressing the inflammation response and ameliorates the effects of aging and cancer. Figure 4 states the effect of apigenin on aging and cancer, where the anticancer feature of apigenin has been described mainly through the regulation of cellular response to oxidative stress, DNA damage, inhibition of inflammation, suppression of angiogenesis, reduction in cell proliferation, triggering of autophagy, and apoptosis. Out of all these, p53-mediated apoptosis and promotion of cell-cycle arrest is the well-described mode of apigenin.

Effect of apigenin on aging and cancer. NF-kB, nuclear factor kB; MAPK, mitogen-activated protein kinase.
Conclusions
This study reveals the dynamic targets and pathways of apigenin. These results support the conclusion that the modes of apigenin in aging and cancer largely include suppressing the inflammation response. This study not only contributes to an improved understanding of the modes of apigenin, but also suggests an approach to reveal novel drug candidates at a network pharmacology level. As a limitation of such in silico studies, this study could only retrieve apigenin targets that are available in published studies. Thus, additional experimentation in future studies is required to confirm the rationality of these findings as well as to investigate its clinical use in patients.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
