Abstract

Introduction
Current diagnosis of oropharyngeal squamous cell carcinoma (OPSCC) involves endoscopy, PET, and biopsy of suspicious lesions. Video nasopharyngolaryngoscopy (NPL) is a common, minimally-invasive data source frequently used to evaluate the location of the primary OPSCC tumor. However, it is difficult for clinicians to visually determine OPSCC from Video NPL alone, especially in the posttreatment setting to evaluate for recurrence.1,2 This leads to many patients needing an invasive, expensive biopsy to confirm OPSCC. An accurate ML model that determines OPSCC from Video NPL could eliminate an invasive procedure for many patients, reduce patient anxiety, and reduce cost of care. Advancements that improve diagnostic capacity with minimal disruption of current clinical workflow would improve the identification of unknown primary tumors and local surveillance after the treatment of OPSCC and other head and neck cancers (HNC).
Over the last decade, advancements in machine learning (ML) combined with collecting large medical datasets have resulted in increased research in the application of ML in medicine, 3 including disease diagnosis, triage and prognostication, clinical decision-making, surgical planning, intraoperative assistance, and patient education. 4 Each year between 2005 and 2019 has had an estimated 61-fold increase in the number of papers applying ML to medicine. The applicability of ML in cancer diagnosis has seen a surge in evidential support with image analysis methods.5-10
Similarly, research on ML applications in HNC has also increased. 3 Studies have used endoscopic data to detect cancer, but this has generally been multispectral narrow band imaging (mNBI). 2 No previous research has used video endoscopy. In this paper, we propose a novel machine learning application in HNC, using deep learning methods to classify whether there is evidence of OPSCC in video NPL. This pilot work lays the groundwork for a model that could be applied to video laryngoscopy as a diagnostic-assist tool, particularly in cases where the identification of a malignancy may not be readily obvious to the clinician.
Methods
Video NPL was provided from The Ohio State University Wexner Medical Center from 85 patients undergoing treatment or follow-up care for OPSCC from January 2019 to April 2022 (Table 1). We included patients with diagnosis of oropharyngeal cancer who had recordings of their NPL either prior to treatment or after the completion of definitive radiation ± chemotherapy. It is our institutional standard to attempt to record all initial and follow-up NPL examinations as part of the medical record. Of the video data classified by clinicians, 17 patients showed no evidence of disease in the video taken posttreatment, 65 patients showed active signs of disease in the pretreatment video, and 3 patients showed recurrence in the posttreatment video totaling 68 patients with evidence of disease. All posttreatment cases followed up within 6 months after ending treatment. All patients were treated only with chemoradiation.
Patient Characteristics.

A 2D Convolutional Neural Network (CNN) model was trained usin

A frame from a patient’s Video NPL that has a visible tumor in the bottom left corner (green arrow added by authors for clarity). NPL, nasopharyngolaryngoscopy.
As each frame is processed independently by the CNN using 2D convolutions, to extract feature-wise dependencies between frames within the video we add a frame-wise dot-product multi-head self-attention layer. To extract frame-wise dependencies important to the classification task, self-attention is an alternative to using a recurrent neural network or 3D convolutional approach. One benefit of this approach is that each frame can be attended to by itself as well as by every other frame making self-attention capable of capturing longer range dependencies. Figure 2 details our proposed CNN architecture, and for clarity, we separate the architecture into 3 separate phases. Phase A shows how we use 2D convolutions on each frame independently followed by ReLU nonlinearity to extract relevant features from each frame. Then, every two frames are averaged together using average pooling before passing it through another 2D convolution + ReLU. We average frames to smooth and reduce the length of the video before sending it into Phase B. As the video is continuous, subsequent frames will be highly correlated and averaging will result in minimal information loss, but large improvements to the inference time of the overall network. Phase B shows how we flatten each frame to pass through a multi-head self-attention mechanism to model frame-wise dependencies across the entire Video NPL. Then, in phase C, after reshaping each frame back to its original shape before the multi-head self-attention layer, every 2 frames are again averaged and passed through another 2D convolution + ReLU. The entire feature space is then flattened and goes through a fully-connected layer for the final prediction of “active cancer” or “no evidence of disease” for the entire Video NPL.
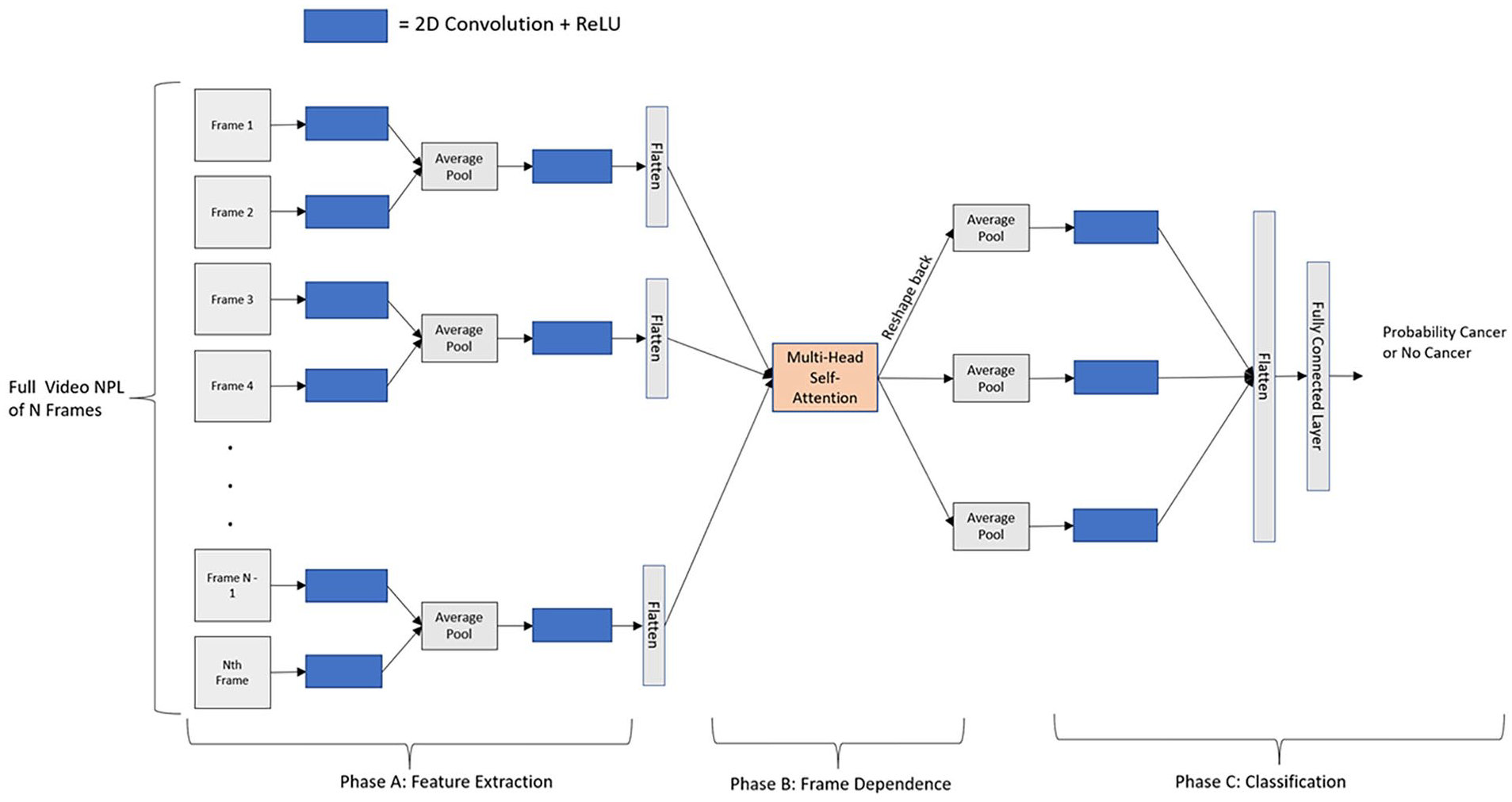
Our proposed CNN Architecture to predict Cancer or No Cancer in Video NPL. We explain the architecture in 3 phases. Phase A extracts features from each frame independently using a 2D Convolution followed by ReLU. Then, every 2 frames are averaged together before applying another 2D Convolution + ReLU. Phase B applies our multi-head self-attention layer to capture the frame-wise dependencies. Phase C then averages every 2 frames together, followed by one more 2D convolution + ReLU. Then, the entire feature space is flattened and passed through fully-connected layer to predict probability of cancer or no cancer. CNN, convolutional neural network; NPL, nasopharyngolaryngoscopy.
The CNN was built in PyTorch and trained using early stopping with a max number of epochs of 100. With the overall objective to minimize the cross-entropy loss, if the validation loss did not improve for 20 epochs, we stopped the training process. For training, the Adam optimizer was used with an initial learning rate of 0.001 and a batch size of 4. No data augmentation methods were used for this study. The model was trained using a single NVIDIA Volta V100 with 32 GB GPU memory, provided by the Ohio Supercomputer Center. After training, when a new, never-before seen Video NPL is input to the model, the CNN outputs a classification of “active cancer” or “no evidence of disease.”
To validate generalization performance, we used fivefold stratified cross-validation and aggregated results across folds. This means for each fold, roughly 68 patients’ Video NPL would be used for training and 17 patients’ Video NPL would be used to assess validation performance. We used a cross-validation strategy to ensure that every patient within our dataset will eventually be used in the validation set during the cross-validation. As our pilot dataset is small (85 patients), if we used a train-test-validation split to evaluate performance, we would not have enough representation in a single validation set to adequately assess the model’s ability to generalize. Additionally, the use of
Results
Due to the class imbalance, we report sensitivity, specificity, AUC, precision, and the F1 score in addition to accuracy in Table 2 to better understand performance.
Cross-Validated Results.
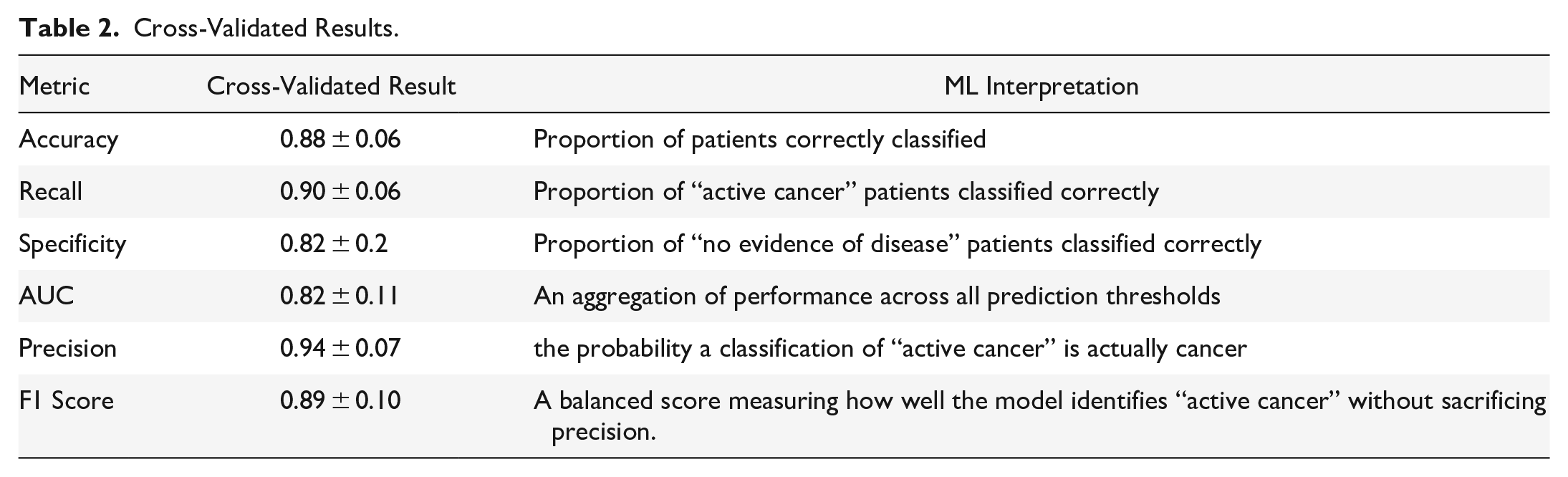
Our proposed model architecture achieved a cross-validated accuracy of 0.88, sensitivity of 0.90, specificity of 0.82, AUC of 0.82, precision of 0.94, and F1 score of 0.89. Such a high precision indicates that we can have confidence when the model predicts evidence of OPSCC. For every prediction of evidence of OPSCC, we have shown it will be a true positive 94% (cross-validated) of the time. Additionally, the high AUC, sensitivity and specificity means that we can accurately predict both classes and that we are not achieving high accuracy by only predicting evidence of OPSCC, but rather can identify patients without evidence of a malignancy as well. In the future, we expect this method to be a useful, real-time clinical tool as the network is lightweight and takes only 0.02 s to classify each Video NPL.
Discussion
Overall, our pilot results show excellent performance in the ability of the ML model to predict active cancer or no evidence of disease in Video NPL, especially considering the small data size. While the results are encouraging, there is some variation in performance across folds, especially in specificity. We expect this and overall performance to improve with a larger dataset.
This is the first study showing potential use of standard recorded Video NPL to detect head and neck cancer through machine learning. The presented model is a pilot proof-of-concept that demonstrates very promising early capabilities but is being validated on a larger dataset. The goal was to develop a diagnostic aid usable by both ENT and non-ENT experts (eg, radiation oncologists or speech and language pathologists) who do not have the ability to biopsy, especially in posttreatment surveillance and identification of the unknown primary tumor for OPSCC. The long-term vision is to develop a suite of algorithms for numerous clinical applications including cancer detection across the larynx and pharynx, swallowing function, vocal cord paralysis, other benign laryngopharyngeal processes, and detection of clinical and subclinical toxicity from surgery and chemoradiation.
Conclusion
Our results validate that ML can be used to predict whether there is active cancer or no evidence of disease in Video NPL. After verification on a larger dataset, this method and model could be applied to video laryngoscopy as a diagnostic-assist tool to help in cases where a biopsy is not possible.
Limitations
While our pilot results indicate that ML can be used to predict evidence of disease, there is further validation that must be performed to substantiate our collected evidence. For this study, we only had data from The Ohio State University Wexner Medical Center; however, assessing the ability for the model to generalize across institutions will help us to better understand potential limitations of our model. Additionally, larger datasets will be required to capture all the potential variability within the patient population, preferably with non-oncology patients.
Footnotes
Acknowledgements
None.
Authors Contributions
RC contributed to the creation of the model architecture, model training, and model evaluation. AR contributed to the preparation of the dataset. SR, KV, and SK contributed to the conception and design of the experiment. All authors contributed to the preparation of the manuscript.
Availability of data and materials
The datasets generated during and/or analyzed during the current study are not publicly available due to containing protected health information.
Consent for publication
Not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics approval and consent to participate
Not applicable.
