Abstract
Background
Human Huntington's disease (HD) is a genetic neurodegenerative disorder caused by the mutant HTT gene containing CAG repeat expansions, resulting in motor dysfunction and behavioral deficits. CAG repeats of 40–53 occur in adult HD and 60–120 repeats occur in early onset juvenile HD, differing from the normal range of 5–35 repeats.
Objective
The HTT gene is translated to the huntingtin (HTT) protein that interacts with proteins in the development of HD. There have been few studies of HTT protein interactors in human HD brain. Therefore, this study evaluated the hypothesis that dysregulation of HTT protein interactors occurs in human juvenile HD brains.
Methods
The strategy of this study was to analyze proteomic data of human juvenile HD brain putamen and cortex regions for dysregulation of HTT interacting proteins, using a database that we compiled of HTT interactors identified in HD model systems from yeast to HD mice.
Results
Results showed significant dysregulation of HTT protein interactors of mitochondria, signal transduction, RNA splicing, chromatin organization, translation, membrane trafficking, endocytosis, vesicle, protein modification, granule membrane, and macroautophagy pathways. The majority of downregulated and upregulated HTT interactors occurred in the putamen region compared to cortex. Dysregulation displayed downregulation of mitochondria and signal transduction interactors, combined with upregulation of RNA splicing, chromatin organization, and translational interactors. Network analysis revealed interactions among clusters of HTT interactors.
Conclusions
These findings demonstrate prevalent dysregulation of HTT protein interactors in human juvenile HD brain, especially in the putamen region that controls movement deficits in HD.
Keywords
Introduction
Huntington's disease (HD) is a rare genetic neurodegenerative disorder caused by CAG trinucleotide expansions in the mutant HTT gene, inherited in an autosomal dominant manner. Progression of HD results in chorea movement dysfunction, psychiatric disturbances, cognitive decline, other neurological symptoms, and death.1–5 The severity and age-at-onset of HD negatively correlates with the length of CAG expansions. 5 Juvenile HD is caused by CAG repeat expansions of 60 to 120 repeats and up to 200 repeats which result in early childhood onset of HD.5–7 The most prevalent form of HD is represented by the mutant HTT gene with 40–53 CAG repeats of adult HD. Normal individuals have 5–35 CAG repeats within the HTT gene. Reduced penetrance of the disease occurs in the range of 36–39 CAG repeats. HD dysfunctions involve the brain putamen region of the striatum that participates in motor coordination and cognitive abilities,8,9 and the cortex areas of BA4 and BA6 which regulate complex movements for motor functions.10,11
Expression of the mutant HTT gene results in polyglutamine (polyQ) expansions within the NH2-terminal region of the huntingtin (HTT) protein. Mutant HTT protein with expanded polyQ has been found to be responsible for HD behavioral dysfunctions, HTT inclusions and neuropathology, and neurodegeneration. It has been hypothesized in the field that proteins interacting with mutant HTT mediate neurotoxicity and behavioral deficits of HD.12–15 Proteins that interact with normal HTT and with mutant HTT have been studied in numerous non-human HD models from yeast to transgenic mice, reviewed by Podvin et al. (2022). 16 We recently compiled a database of HTT interactors identified in HD models, consisting of 2971 proteins. 16 Bioinformatics analysis indicated twelve clusters of HTT interacting proteins of biological functions including signal transduction, membrane trafficking, chromatin, mitochondria, translation, and RNA splicing, among others.
The next important question to address is whether these identified HTT interacting proteins, observed in models of HD, undergo dysregulation in human HD brain? Therefore, the goal of this study was to evaluate such HTT interacting proteins in human juvenile HD brain putamen and cortex (BA4 and BA6 regions) by proteomics and bioinformatics analysis. We analyzed these proteomics data from human juvenile HD brains for upregulation and downregulation of HTT protein interactors in human juvenile HD brain regions compared to age-matched controls. Results found that the majority of dysregulated HTT interactors occurred in the putamen region for interactors involved in mitochondria, signal transduction, RNA splicing, translation, chromatin, and membrane trafficking functions. A more modest level of dysregulated HTT interactors occurred in BA4 and BA6 cortical regions compared to putamen. These findings provide insight into the dynamic dysregulation of multiple HTT protein interactors in human juvenile HD brain.
Methods
Brain tissues from human juvenile HD and age-matched control putamen, BA4, and BA6 regions
Proteomics of human juvenile HD and control human brain regions was conducted as we previously reported, 17 and summarized here. Brain tissues consisted of juvenile HD brain regions from five brains each, and age-matched control regions from seven brains each. Brain tissues were obtained from the Harvard Brain Tissue Resource Center (HBTRC), the HDSA Center of Excellence at the University of Washington, the HDSA Center of Excellence at the University of Miami, and the HDSA Center of Excellence at the University of Pittsburgh. The brain banks collected non-identified human brain tissues according to Institutional Review Board protocols. The number of juvenile HD brain tissue samples was limited due to the rare number of cases.
Determination of CAG repeat numbers within the HTT gene in tissue samples
The numbers of CAG repeats within the HTT gene in juvenile HD and control tissue samples were determined by PCR (polymerase chain reaction) amplification of the CAG region in exon 1, followed by DNA sequencing of amplicons, conducted as we previously reported. 17 Brain tissue samples were digested by incubation with proteinase K at 65°C for 24 h, followed by 95°C for 10 min. PCR of the CAG HTT repeat region was conducted with the forward primer 5′-ACCGCCATGGCGACCCTGGA-3′ and reverse primer 5′-AAACTCACGGTCGGTGCAGC-3′, with Taq PCR Core Kit (Qiagen, Valencia, CA) according to the manufacturer's protocol. PCR resulted in DNA amplicons of 265 to >400 bp for CAG repeats in the normal and juvenile HD disease expanded HTT gene domains. PCR-amplified DNA bands were excised from agarose gels and extracted using PureLink™ Quick Gel Extraction Kit (Invitrogen, Waltham, MA) according to the manufacturer's protocol. Determination of the number of CAG repeats was achieved by DNA sequencing conducted by GeneWiz (San Diego, CA) using forward primer 5′-GTTCTGCTTTTACCTGCGGC-3′ and reverser primer 5′-GCTCCCTCACTTGGGTCTTC-3′. Results showed the number of CAG repeats for the two alleles of the HTT gene for brain tissues from juvenile HD and age-matched controls, shown in Table 1.
Human juvenile Huntington's disease (HD) brain samples.
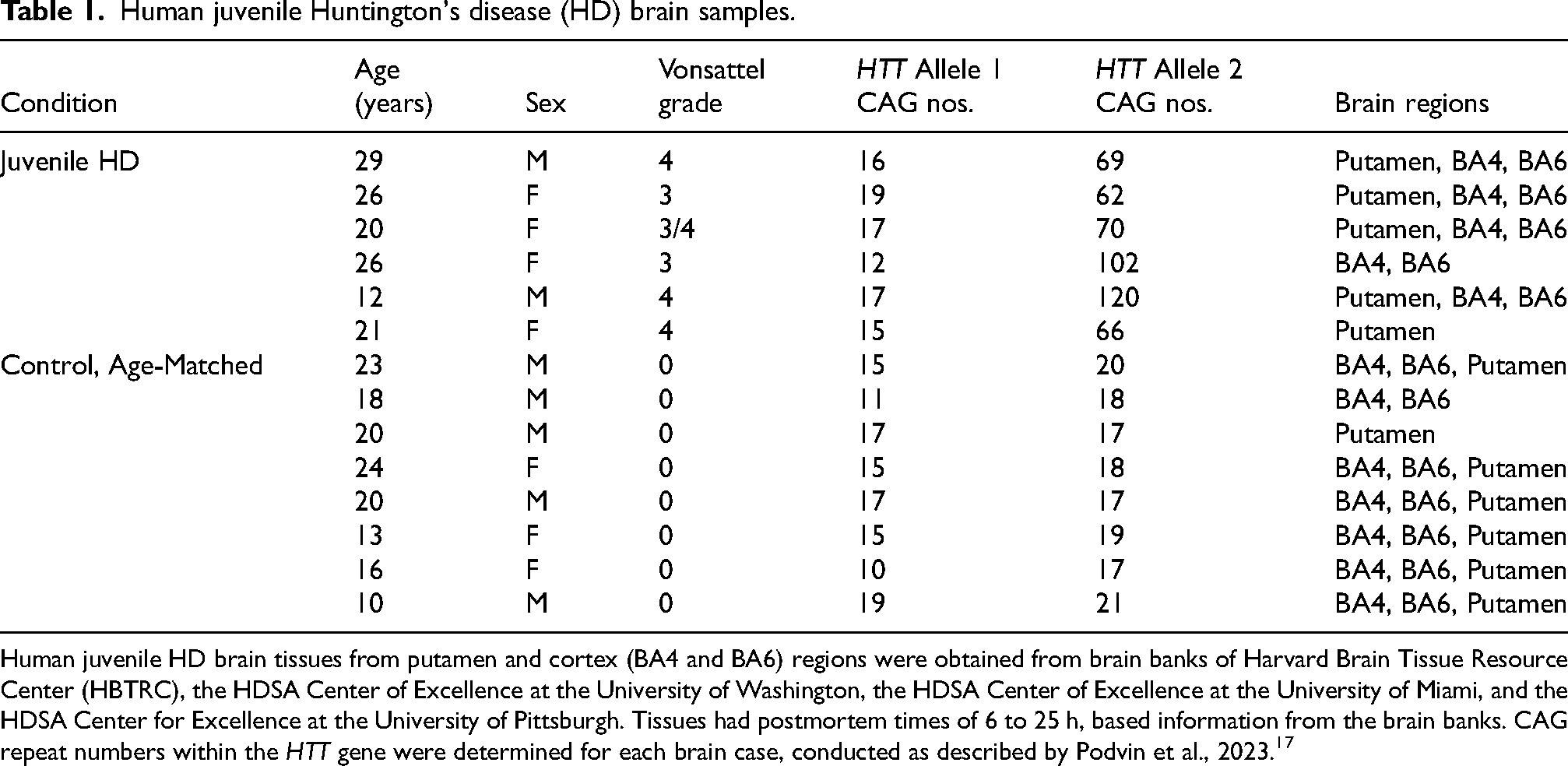
Human juvenile HD brain tissues from putamen and cortex (BA4 and BA6) regions were obtained from brain banks of Harvard Brain Tissue Resource Center (HBTRC), the HDSA Center of Excellence at the University of Washington, the HDSA Center of Excellence at the University of Miami, and the HDSA Center for Excellence at the University of Pittsburgh. Tissues had postmortem times of 6 to 25 h, based information from the brain banks. CAG repeat numbers within the HTT gene were determined for each brain case, conducted as described by Podvin et al., 2023. 17
Proteomics analysis of human brain HD tissues
Proteomics of human juvenile HD and control human brain regions was conducted as we previously reported,17–19 and summarized here.
For extraction of proteins from tissues, each brain tissue sample of ∼100 mg wet weight was excised from frozen tissue on a glass plate over dry ice. Each tissue sample was combined with lysis buffer (100 mM Tris-HCl, pH 7.4, 50 mM NaCl, with protease inhibitors 10 µM pepstatin A, 10 µM leupeptin, 10 µM chymostatin, 10 µM E64c, and 100 µM AEBSF) to result in 20% w:v for tissue weight (w) and buffer volume (v). Each brain tissue sample was sonicated 2 times for 10 s on ice, with 30 s on ice between sonication steps. Following protein quantification (Bio-Rad DC protein assay kit, Bio-Rad, Hercules, CA), 200 µg protein was precipitated in 90% methanol on ice for 15 min, then centrifuged at 14,000×g for 15 min at 4°C. The protein pellet was collected and dried in a speed-vac.
Protein pellets were then each reduced in 10 mM tris(2-carboxyethyl)phosphine (TCEP) and alkylated with 2-chloroacetamide (CAA), followed by digestion with trypsin/LysC (Promega, Madison, WI) at 37°C overnight. Tryptic peptides were desalted and isolated by solid-phase extraction (SPE) using Empore C18 membranes (3 M Maplewood, MN). Peptide concentrations were measured using the Pierce total peptide assay kit (ThermoFisher, Waltham, MA).
Peptide samples (2 µg) in 2% ACN, 0.1% TFA were injected into the nano-LC-MS/MS system consisting of a Dionex UltiMate 3000 nano-LC, and tandem mass spectrometry on an Orbitrap Q-Exactive (both from ThermoFisher). Samples were injected in a randomized order with a flow rate of 300 nl/minute using a gradient of 5% to 25% acetonitrile/0.1% TFA for 80 min, and a gradient of 85% to 90% ACN/0.1% TFA for 20 min. MS1 data were acquired in positive ion mode at a resolution of 70,000, AGC target of 3 × 106, maximum inject time of 100 ms and scan range of 310 to 1250 m/z. MS1 microscans were acquired for scan range of 310 to 1200 m/z at resolution of 70,000 at 200 m/z, injection time of 100 ms. Data-dependent MS2 was acquired in centroid mode, at a 1.5 m/z isolation window, scan range of 200 to 2000 m/z, first fixed mass 150 m/z, resolution of 17,500, maximum inject time of 50 ms, automatic gain 1 × 105, intensity threshold of 4 × 103, and HCD cell normalized collision energy of 27 V.
Mass spectrometry MS1 and MS2 spectra were analyzed by the PEAKS (v. 8.5) software for protein identification with database-de Novo bioinformatics, post-translational modifications, and label-free quantitation (LFQ), conducted as we have reported.17–19 Spectra were queried against the UniprotKB/SwissProt human protein database. Monoisotopic precursor mass error tolerance was 20 ppm with fragment mass error tolerance of 0.01 Da. Identification parameters used a false discovery rate (FDR) of 0.9%, peptide identification of -log10(P) ≥ 22, protein identification of -log10(P) ≥ 20, and ≥ 2 unique peptides per protein. A protein was considered identified when present in at least two out of three technical replicates for each sample. A protein was assigned as being present in the juvenile HD group if it was present in at least three out of five biological samples in that group. A protein was assigned to the control group if it was present in at least four out of seven biological control samples.
Proteins were quantified by label-free quantitation (LFQ), determined by PEAKS v 8.5, conducted as we reported.17–19 MS2 peaks were converted to area under the curve (AUC). For each quantifiable protein group, all MS2 peak AUCs for the corresponding peptides were summed. Replicate normalization of LFQ intensity distributions used the LOESS-G method determined by Normalyzer (web application). To compare quantifiable proteins between juvenile HD and control groups, multiple testing was conducted using the Benjamini-Hochberg post-hoc method that assesses the FDR (false-discovery rate) of false-positives at q values of < 0.05 for significance.20,21 Protein quantities different between JHD and control groups were significant at q < 0.05.
Bioinformatics analysis of human juvenile HD brain proteomics data for HTT-interacting proteins identified in non-human HD models
Within the human juvenile HD proteomics data, we analyzed HTT-interacting proteins that have been previously identified in non-human HD models, assembled by us as a database of HTT interactors. 16 Interactors were evaluated for upregulation and downregulation in juvenile HD brain regions of putamen, BA4 cortex, and BA6 cortex compared to age-matched control tissues of the same regions. Protein-protein interaction networks were predicted by STRING-db which uses a database of known protein interactions. 22 Protein interaction networks for upregulated and downregulated HTT-interacting proteins in HD compared to controls were evaluated in putamen, BA4 cortex, and BA6 cortex brain regions.
Results
Strategy for proteomics evaluation of mutant HTT-interacting proteins in human juvenile HD brain using a database of HTT-interacting proteins identified in non-human HD model systems
The goal of this study was to assess the hypothesis that HTT-interacting proteins may be dysregulated in human HD brain. The strategy used to achieve this goal consisted of conducting proteomics analysis of human juvenile HD brain tissues for assessment of HTT-interacting proteins previously identified in animal models of HD (Figure 1). Quantitative proteomics data from human juvenile HD brain was obtained for putamen, BA4 cortex, and BA6 cortex regions and age-matched controls from the same brain regions. These data were analyzed for HTT-interacting proteins, numbering 2,971, identified in non-human HD models from yeast to transgenic HD mice. 16 We previously compiled this HTT-interacting protein database to provide a resource of known HTT interactors for their analysis in human HD and control brains by this study. The HTT interactors were organized into twelve clusters representing functions of protein modification, chromatin organization, RNA splicing, membrane trafficking, signal transduction, mitochondria, granule membrane, macroautophagy, cytoplasmic vesicle lumen, clathrin-mediated endocytosis, ion channel transport, and translation (Figure 2). 16 Importantly, quantitative proteomics data revealed upregulation and downregulation of HTT-interacting proteins among the 12 clusters of HTT interactors in human juvenile HD brains compared to controls. Evaluation of protein interaction networks of these dysregulated HTT interactors was conducted for putamen and cortex (BA4 and BA6) regions of human juvenile HD brains and age-matched control brains.
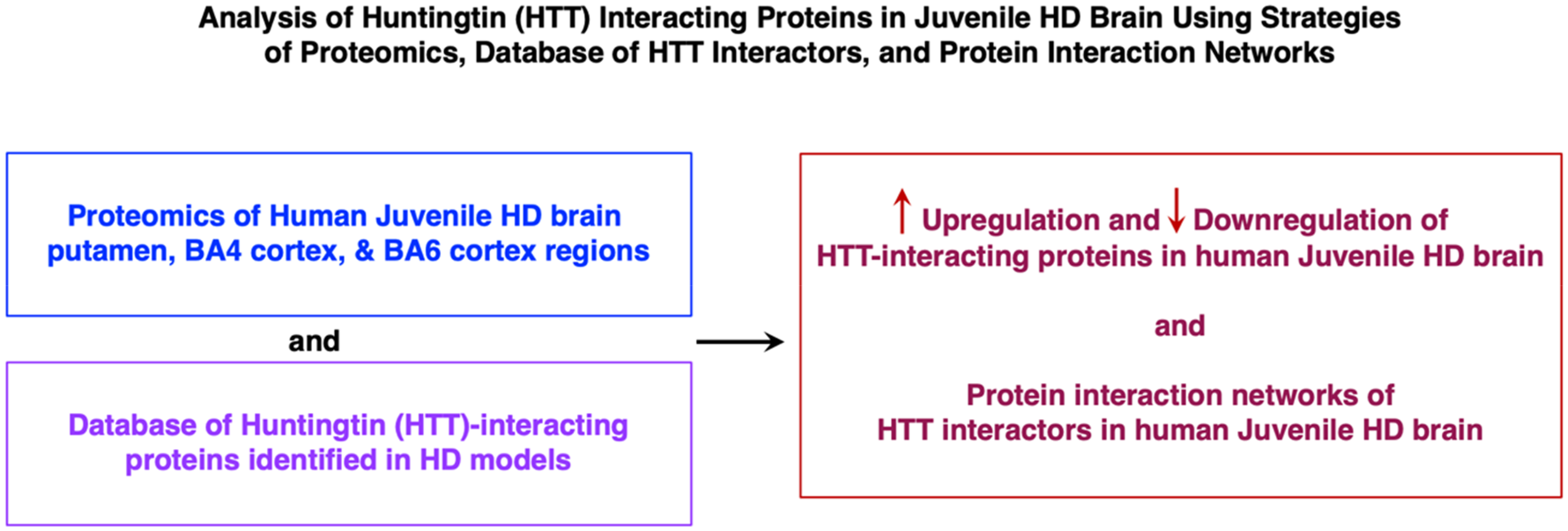
Analysis of huntingtin (HTT) interacting proteins in juvenile HD brain using strategies of proteomics, database of HTT interactors, and protein interaction networks. Juvenile HD brain putamen and cortex (BA4 and BA6 regions), with age-matched controls, were subjected to quantitative proteomics analysis that assessed HTT interactors compiled as a database from those identified in HD model systems.16,17 Significant upregulation and downregulation in HD compared to controls were assessed, combined with bioinformatics analysis of protein interaction networks of dysregulated HTT interactors in human HD brain.
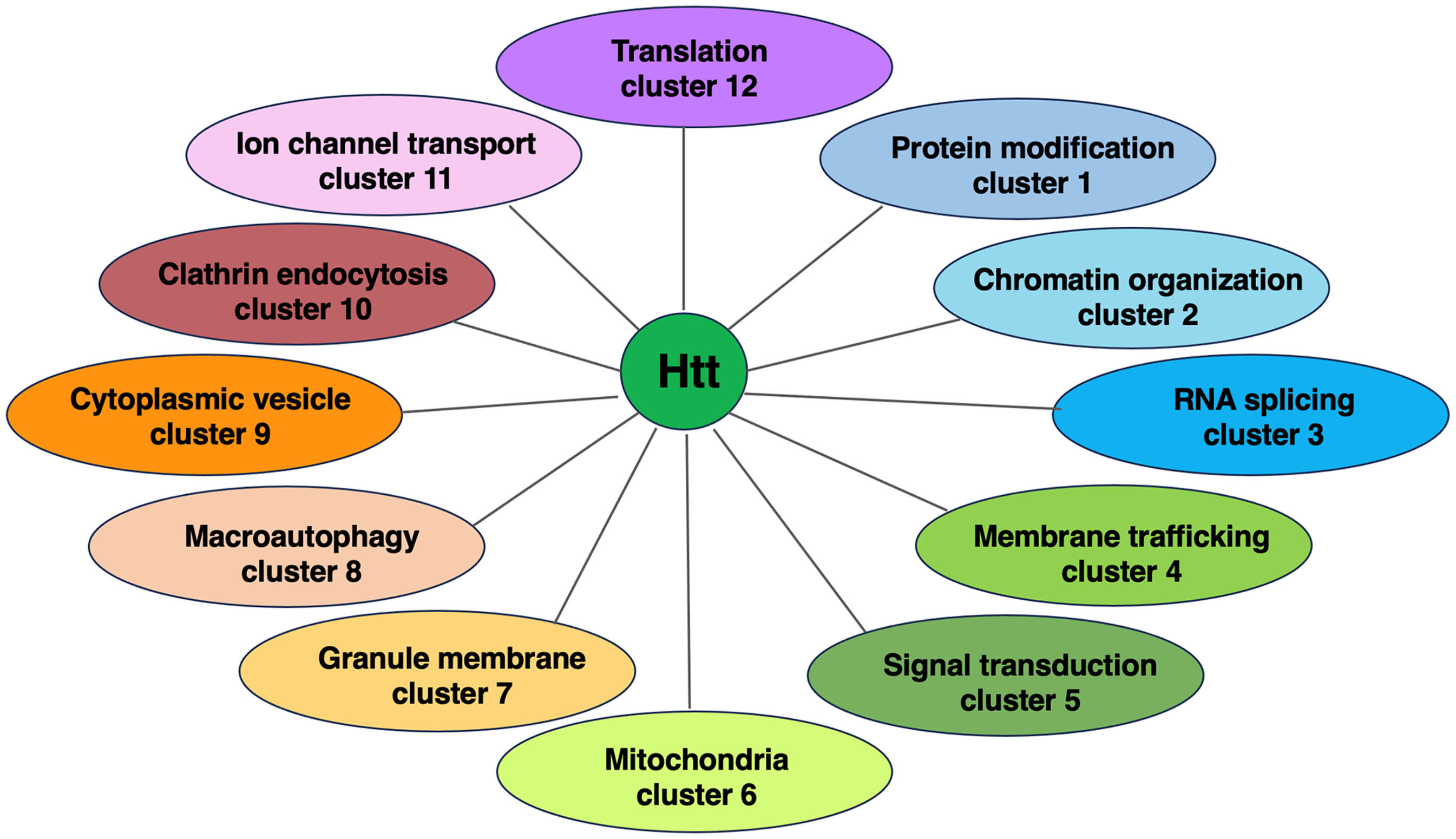
HTT protein interactors clustered by functional biological processes. HTT interactors were compiled from studies in HD model systems, from yeast to transgenic HD mice and clustered by functional categories for biological processes by gene ontology (GO) (16). HTT interactors are indicated as those that were lost with mutant HTT (mHTT), weaker with mHTT, stronger with mHTT, gained interaction with mHTT, or represented a neighbor. This information is from our previous publication of Podvin et al., 2022. 16
Downregulation of the mitochondria cluster of HTT protein interactors in putamen as well as cortex regions of juvenile HD brains
Numerous components of the mitochondria cluster of HTT interactors were significantly downregulated in juvenile HD brains compared to age-matched controls (Figure 3, Table 2). These dysregulated components comprised functional proteins involved in mitochondrial processes of NADH:ubiquinone oxidoreductase (NDUFA3, NDUFA4, NDUFA8, NDUFA9, NDUFA10, NDUFA13, NDUFAF4, NDUFB1, NDUFB6, NDUFB7, NDUFB8, NDUFB9, NDUFB10, NDUFB11, NDUFS2, NDUFS3, NDUFS5, NDUFS7, NDUFV1, NDUFV2), ATP metabolism (ATP5A1, ATP5B, ATP5C1, ATP5F1, ATP5H, ATP5I, ATP5L, ATP5O, ATP6AP1, ATP6V0A1, ATP6V0D1, ATP6V1C1, ATP6V1D), transporters (SLC25A3, SLC25A4, SLC25A6, SLC25A11, SLC25A12, SLC25A13, SLC3A2), cytochrome c oxidase (COX4I1, COX5A, COX5B, COX6B1, COX6C, COX7A2, CYCS), mitochondrially encoded ATP synthase (MT-ATP6, MT-CO1, MT-CO2, MT-ND4, MT-ND5), and ubiquinol-cytochrome C reductase (UQCRC1, UQCRC2, UQCR10, UQCRFS1) (Supplemental Table 1).
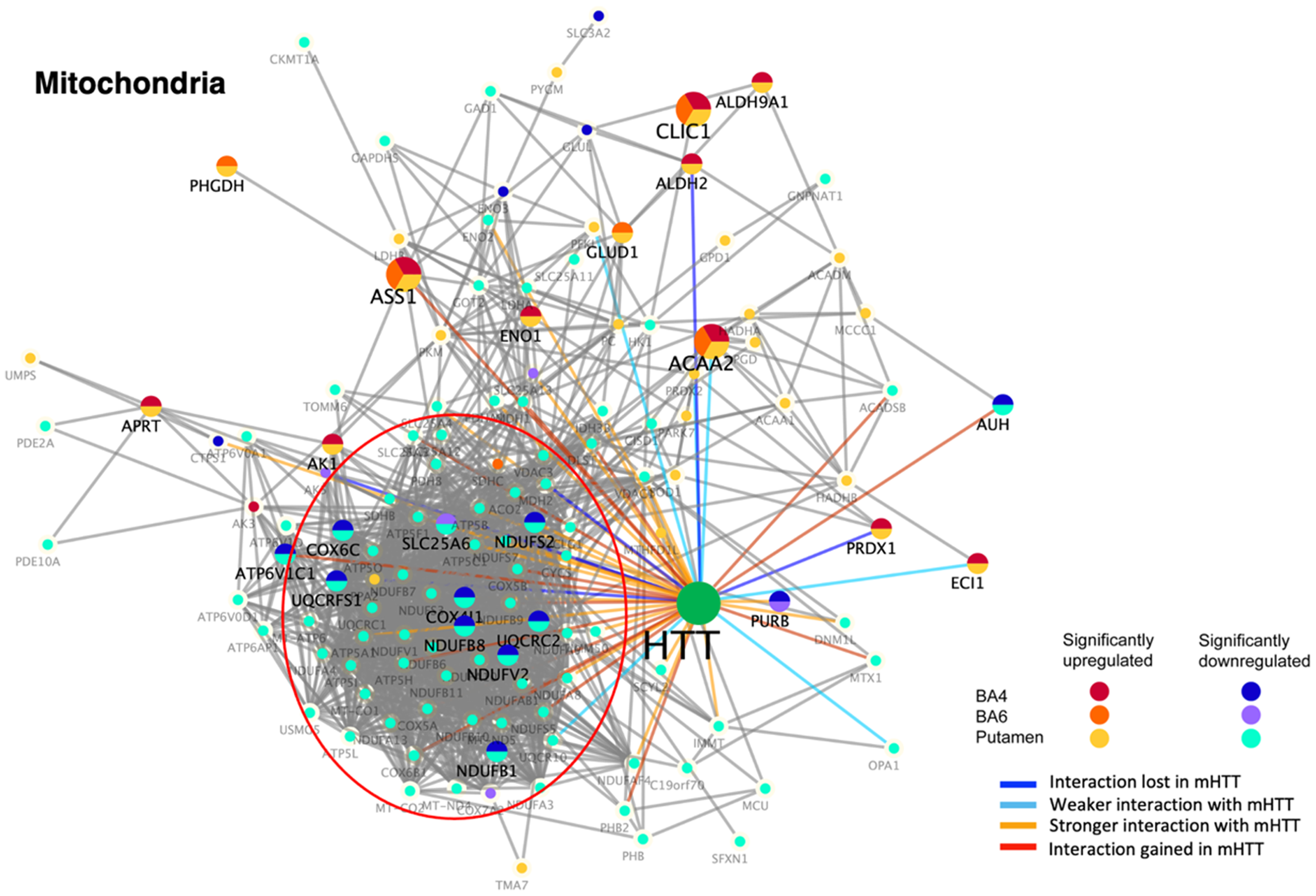
Dysregulation of mitochondrial HTT interactors in putamen and cortex (BA4 and BA6) of juvenile HD brains compared to age-matched controls. HTT interactors of the mitochondrial cluster that were significantly upregulated or downregulated in HD compared to control brains are illustrated for putamen, BA4 cortex, and BA6 cortex regions by the indicated color-coded protein components (shown by gene name). Interactions with mutant HTT are indicated as lost, weaker, stronger, or interaction gained compared to normal HTT, using information from our database of HTT interactors identified in HD model systems. 16
Dysregulated protein interactors of HTT protein in juvenile HD brain regions.
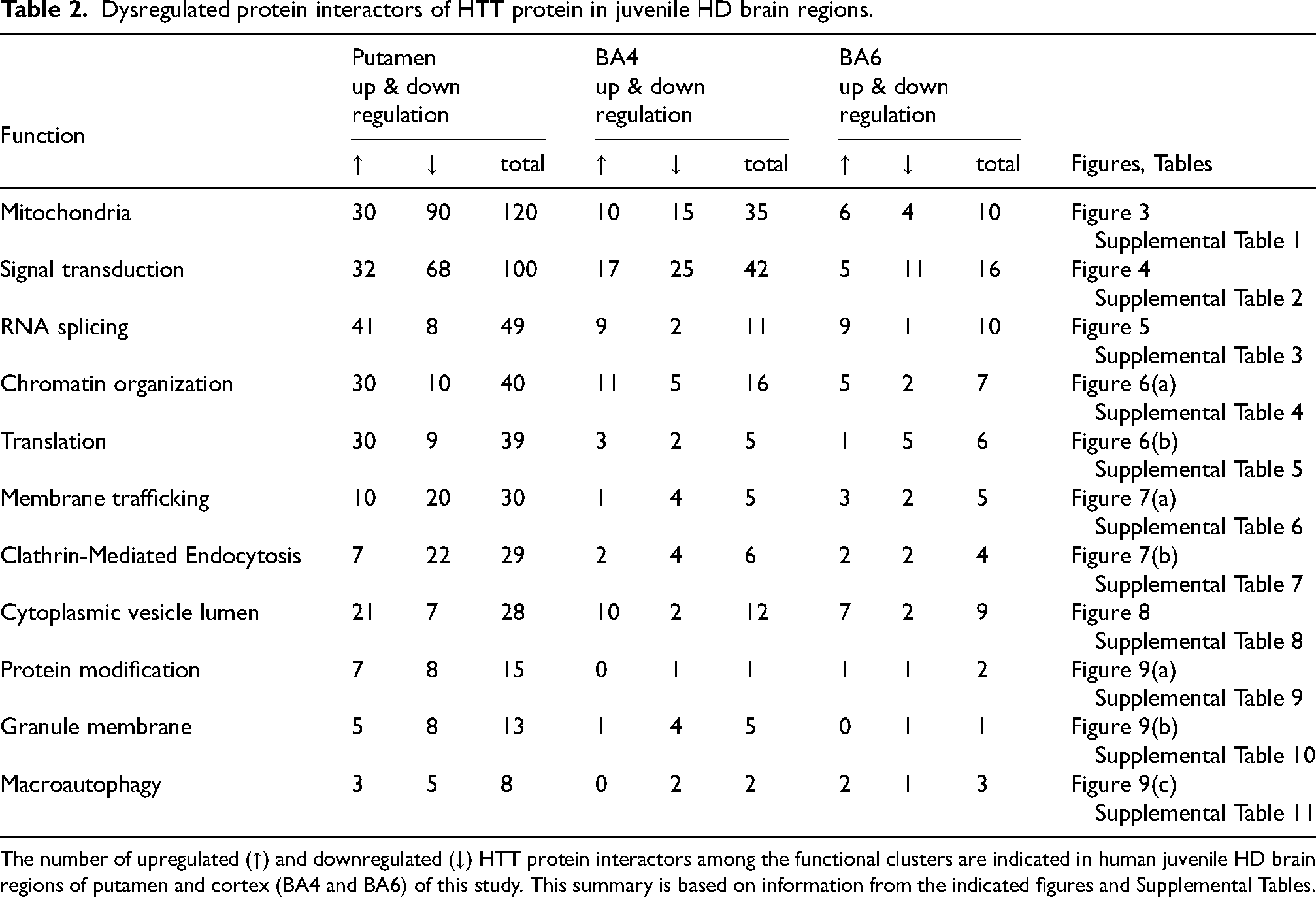
The number of upregulated (↑) and downregulated (↓) HTT protein interactors among the functional clusters are indicated in human juvenile HD brain regions of putamen and cortex (BA4 and BA6) of this study. This summary is based on information from the indicated figures and Supplemental Tables.
In addition, several upregulated mitochondrial HTT protein interactors occurred in putamen and cortex (BA4 and BA6 regions) (Table 2), which included ACAA2, ALDH9A1, ALDH2, ASS1, CLIC1, ENO1, GLUD1, PHGDH that are involved in biochemical metabolism (Supplemental Table 1).
The majority of HTT interactors of the mitochondrial cluster have been shown to display stronger interactions with mutant HTT (compared to normal HTT) (Figure 3), based on our HTT interactor database. 16
These results demonstrate prominent downregulation of mitochondrial components known to interact with HTT, with the majority present in putamen and a lower number of dysregulated HTT interactors in BA4 and BA6 cortex regions. Dysregulation of mitochondria type of HTT interactors may represent involvement of mutant HTT in mechanisms of mitochondrial deficits of HD.23,24
Downregulation of signal transduction interactors of HTT in putamen and cortex of juvenile HD brain
Signal transduction interactors were significantly downregulated, as well as upregulated, in juvenile HD compared to controls (Figure 4, Table 2). The putamen region showed 68 downregulated protein components, BA4 cortex showed 25 downregulated proteins, and BA6 cortex showed 11 downregulated proteins. These downregulated components in putamen and BA4 cortex consisted of protein phosphatases and kinases (CDK5, PIP4K2C (BA4 only), PIP5K1C, PPFIA3, PPP1R9A, PPP1R12A, PPP3CA, PPP3CB, PRKCB, PRKCE, PRKCG), combined with solute carrier proteins (SLC8A1, SLC17A7, and SLC32A1) (Supplemental Table 2). Also, upregulated and downregulated components in juvenile HD were represented by G protein subunits (GNAO1, GNA1, GNA11, GNA13, GNB5, GNG7), dihydropyrimidinase enzymes (DPYSL1, DPYSL3, SPYSL4, DPYSL5), and glutamate receptor NMDA subunits (GRIN1 and GRIN2B) (Supplemental Table 2).
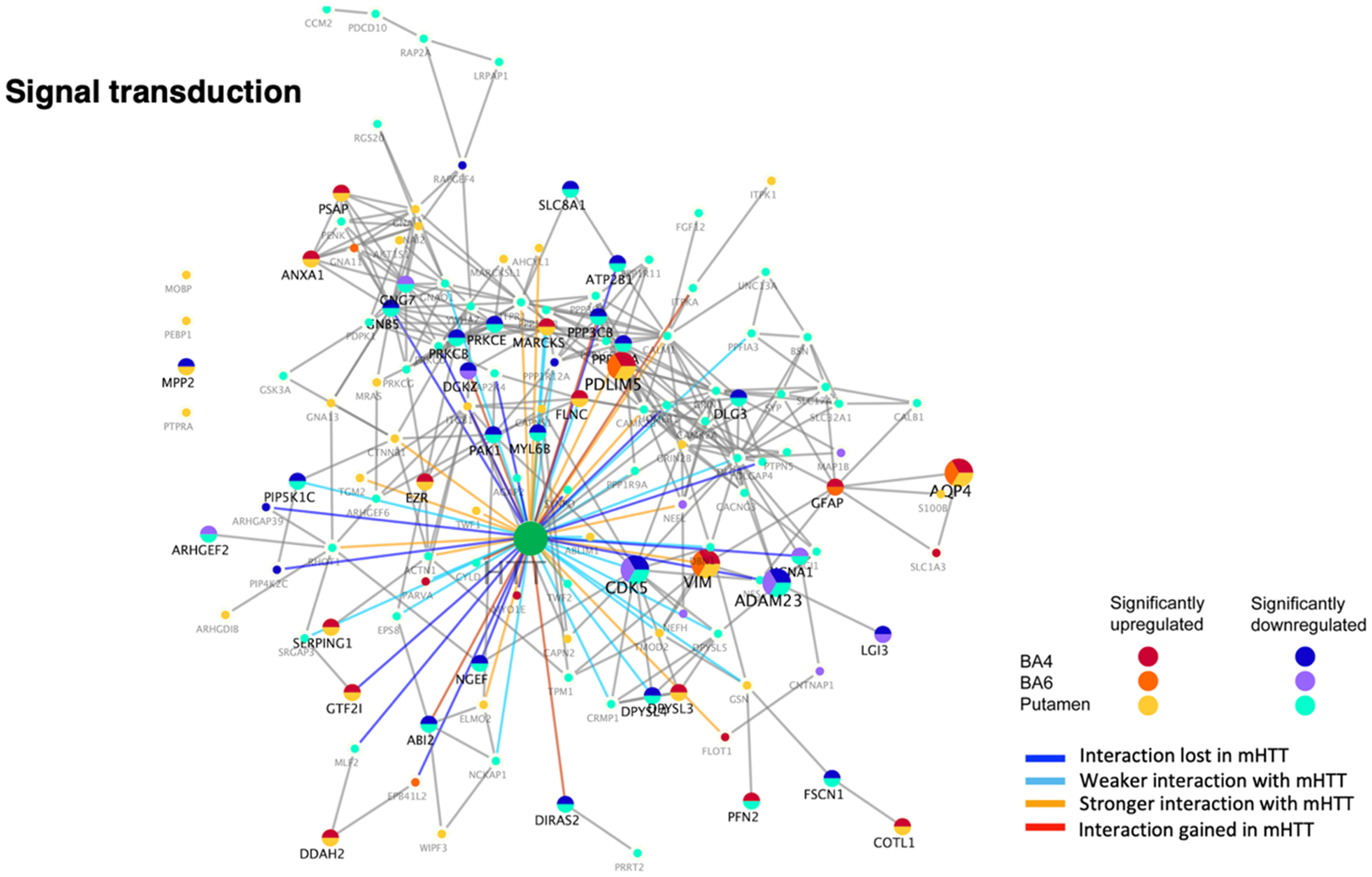
Dysregulation of signal transduction HTT interactors in putamen and cortex (BA4 and BA6) of juvenile HD brains compared to age-matched controls. HTT interactors of the signal transduction cluster that were significantly upregulated or downregulated in HD compared to control brains are illustrated for putamen, BA4 cortex, and BA6 cortex regions by the indicated color-coded protein components (shown by gene name). Interactions with mutant HTT are indicated as lost, weaker, stronger, or interaction gained compared to normal HTT, using information from our database of HTT interactors identified in HD model systems. 16
These dysregulated signal transduction interactors have been found in previous studies to display weaker or lost interactions with mutant HTT compared to normal HTT, as well as stronger interactions with mutant HTT.
These findings demonstrate that dysregulation of HTT interactors with signaling transduction functions participate in human juvenile HD brain.
Upregulation of the RNA splicing cluster of HTT-interacting proteins in putamen and cortex of human juvenile HD brains
Numerous components of the RNA splicing cluster of HTT-interacting proteins were significantly dysregulated via upregulation in juvenile HD brain compared to age-matched controls (Figure 5, Table 2). These data show that the majority of the upregulated proteins were observed in the putamen region of juvenile HD brains, which consisted of ribonucleoproteins (HNRNPA0, HNRNPA2B1, HNRNPH1, HNRNPH2, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, SNRPD2, SNRPE, and SNRPF), RNA binding proteins (RBM39, RBMX, ELAVL1, and SSB), heat shock proteins (HSP90AB1, HSPA1B, HSPA2, HSPB1, HSPB8), and RNA splicing factors (SF3A3, SRSF2, SRSF3, and ERH) (Supplemental Table 3).
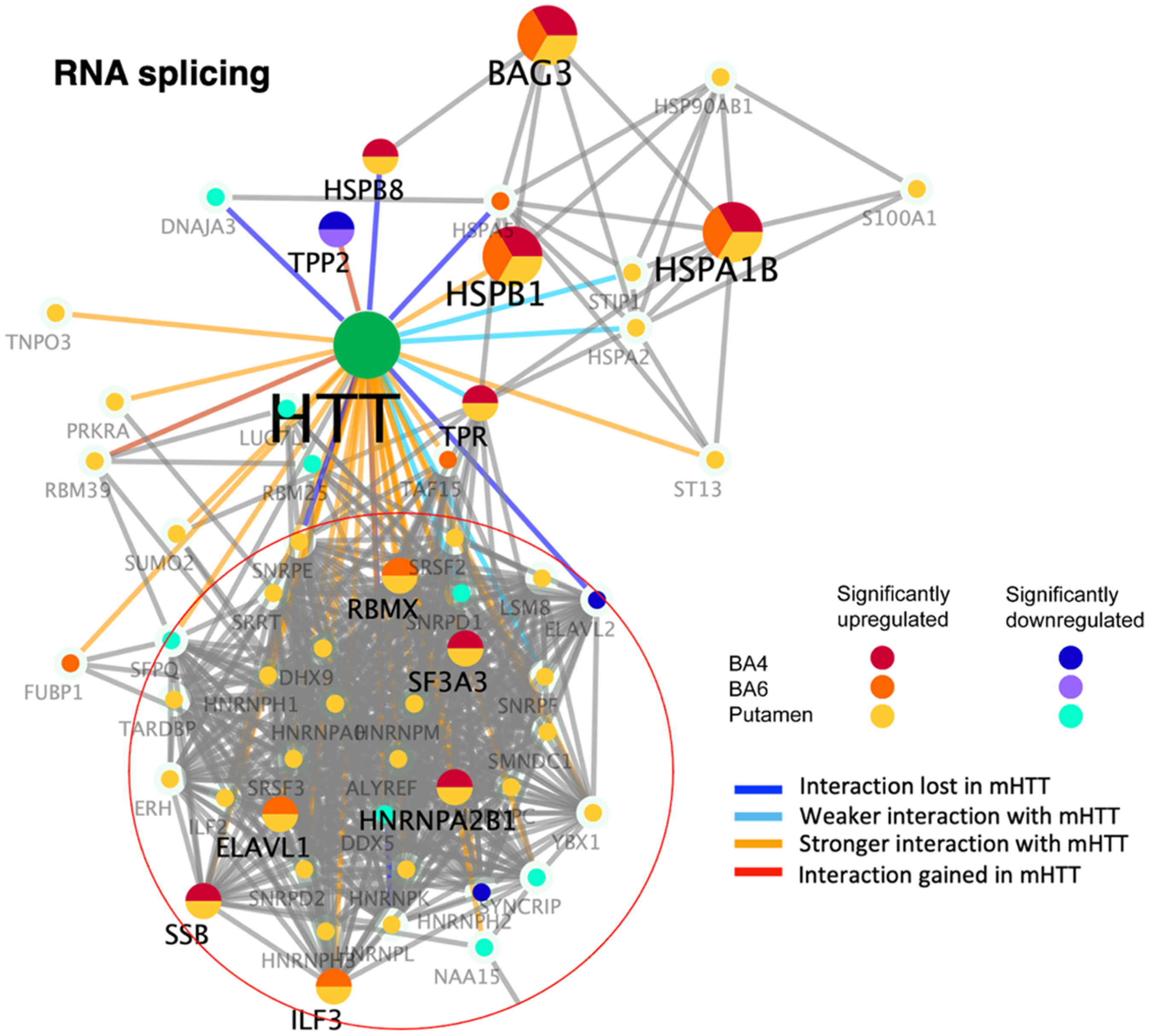
Dysregulation of RNA splicing HTT interactors in putamen and cortex (BA4 and BA6) of juvenile HD brains compared to age-matched controls. HTT protein interactors of the RNA splicing cluster that were significantly upregulated or downregulated in HD compared to control brains are illustrated for putamen, BA4 cortex, and BA6 cortex regions by the indicated color-coded protein components (shown by gene name). Interactions with mutant HTT are indicated as lost, weaker, stronger, or interaction gained compared to normal HTT, using information from our database of HTT interactors identified in HD model systems. 16
The putamen and BA4 cortex both showed upregulation of HNRNPA2B1, HSPB8, SF3A3, SSB, and TPR proteins. The putamen and BA6 cortex showed upregulation of ELAVL1, ILF3, and RBMX proteins. All three regions of putamen, BA4, and BA6 showed upregulation of BAG3, HSPB1, and HSPA1B.
Several downregulated RNA splicing HTT interactors were observed in putamen (DDX5, DNAJA3, LUC7L, NAA15, RBM25, SFPO, SYNCRIP, SNRPD1), BA4 cortex (ELAVL2, HNRNPH2, TPP2), and BA6 cortex (TPP2) (Supplemental Table 3).
Many of the upregulated interactors for RNA splicing have been found to possess stronger interactions with mutant HTT than with normal HTT, and several upregulated interactors have weaker or lost interactions with mutant HTT (Figure 5). 16
These data show that dysregulation of HTT protein interactors of the RNA splicing cluster occurs in juvenile HD brain, consistent with involvement of altered RNA splicing in HD.25–28
Upregulation of the chromatin organization cluster of HTT interactors in putamen and cortex of juvenile HD brain
Numerous components of the chromatin cluster of HTT-interacting proteins were upregulated in putamen of juvenile HD brain (Figure 6(a), Table 2). These upregulated components are represented by histone regulators (ANP32E, H1F0, H2AFX, HDAC1, HIST1H1B, HIST1H1C, RUVBL1) and peptidases (TPP1, ZMPSTE24) (Supplemental Table 4). HTT interactors involved in protein phosphorylation were upregulated (NUCKS1, PRKDC) and downregulated (CAMK4, CSNK2A2, DGKE). Biochemical enzyme types of HTT interactors were also both upregulated (ACSL1, AKR7A2, CBR1, CPT1A, FNTA, PPT1) and downregulated (ACSL4, OGT).
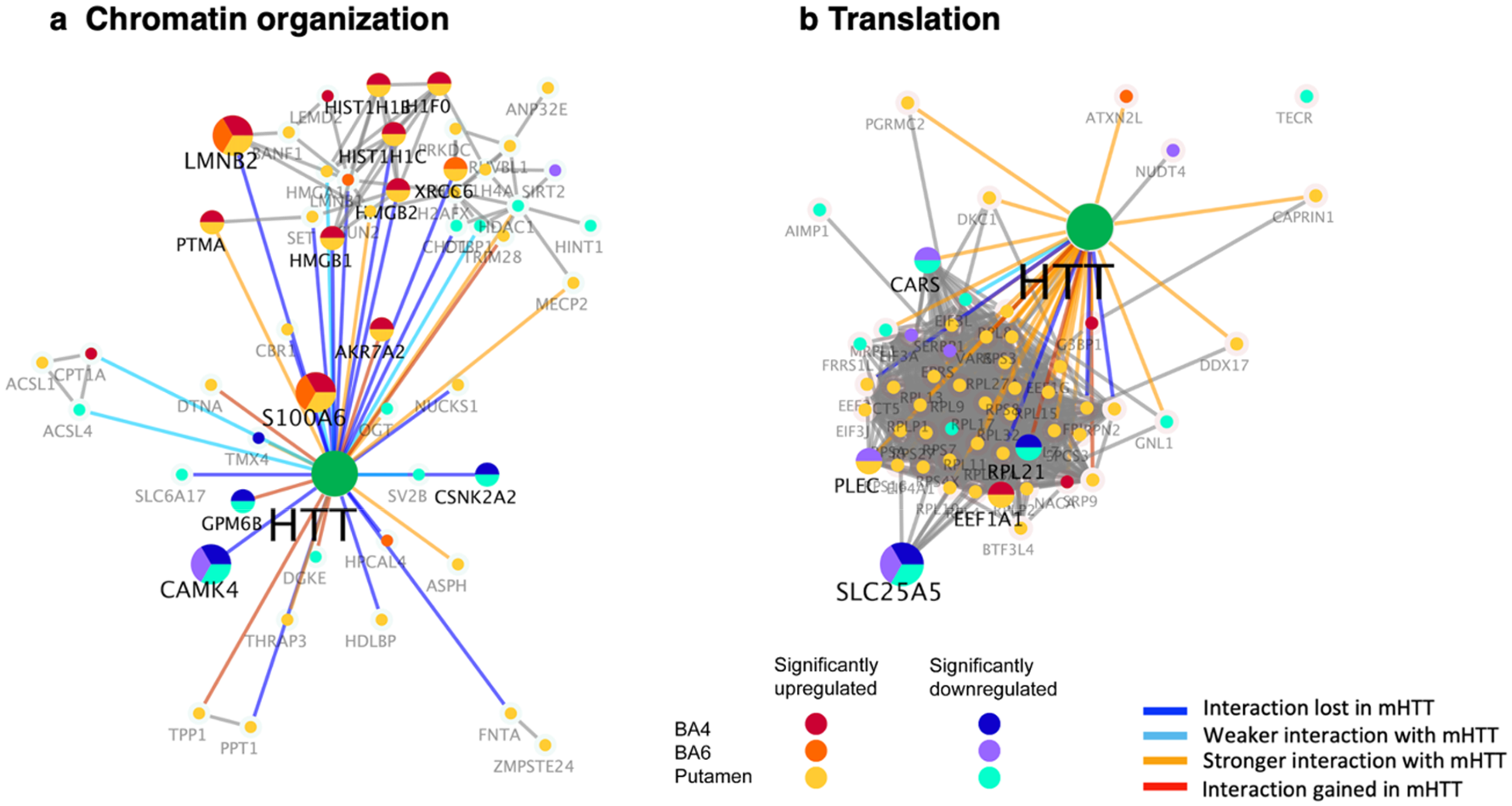
Dysregulation of chromatin organization and translational HTT interactors in putamen and cortex (BA4 and BA6) of juvenile HD brains compared to age-matched controls. HTT interactors of the chromatin organization cluster (panel a) and the translation cluster (panel b) that were significantly upregulated or downregulated in HD compared to control brains are illustrated for putamen, BA4 cortex, and BA6 cortex regions by the indicated color-coded protein components (shown by gene name). Interactions with mutant HTT are indicated as lost, weaker, stronger, or interaction gained compared to normal HTT, using information from our database of HTT interactors identified in HD model systems. 16
These dysregulated chromatin HTT interactors have been shown to display weaker or loss of interactions with mutant HTT compared to normal HTT, as well as stronger interactions with mutant HTT (Figure 6(a)). 16
Dysregulation of HTT interactors with chromatin organization functions suggests involvement of nuclear DNA and associated protein modulators in human juvenile HD brain.
Upregulation of translation cluster components of HTT-interacting proteins in putamen of human juvenile HD brain
HTT protein interactors involved in translational functions were found to be significantly upregulated in putamen of human juvenile HD brain, compared to age-matched controls (Figure 6(b), Table 2). In the putamen region, numerous translational HTT interactors were upregulated, and several of such interactors were upregulated in BA4 and BA6 cortex (Table 2). Upregulated HTT interactors of the translation cluster in HD putamen consisted of ribosomal proteins (RPL4, RPL8, RPL9, RPL11, RPL13, RPL15, RPL17, RPL19, RPL21, RPL27, RPL32, RPLP1, RPLP2, RPN2, RPS3, RPS4X, RPS7, RPS8, RPS27), elongation factors (EEF1A1, EEF1G, EEF2, EIF3A, EIF3J), and signal peptidases (SPCS3, SRP9) (Supplemental Table 5). In addition, upregulated translational HTT interactors in HD BA4 and BA6 cortex included ATXN2L, EEF1A1, NACA, and G3BP1 that possess functions related to ataxin, elongation factors, and transcription.
In addition, several downregulated translational HTT interactors were observed putamen and cortex (BA4 and BA6) which included CARS, EIF3L, FRRS1L, GNL1, RPL21, SLC25A5 (Supplemental Table 5). It is noted that a number of downregulated translational HTT interactors have been found to display stronger interactions with mutant HTT compared to normal HTT (Figure 6(b)). 16
These data demonstrate a role for translational proteins as HTT interactors in human juvenile HD brain.
Downregulation of the membrane trafficking cluster of HTT interactors in putamen and cortex in juvenile HD brains
Downregulation of membrane trafficking interactors of HTT was observed in putamen of juvenile HD brains (Figure 7(a), Table 2). These downregulated components consisted of members of the Ras family of GTPases (RAB3A, RAB3C, RAB3D, RAB33B, RAB39B), ADP ribosylation factors (ARF1, ARL2), kinesins (KIF2A, KIF5A), and tubulin (TUBB8) (Supplemental Table 6). Several of these components were also downregulated in BA4 or BA6 cortex in HD (KIF5A, RAB33B). Many of the downregulated components have been shown to interact weakly with mutant HTT, and to lose interactions with mutant HTT (Figure 7(a)). 16 A small number of components of this cluster were upregulated, involved in various trafficking functions. These findings indicate that HTT interactors with membrane trafficking function are dysregulated in juvenile HD.
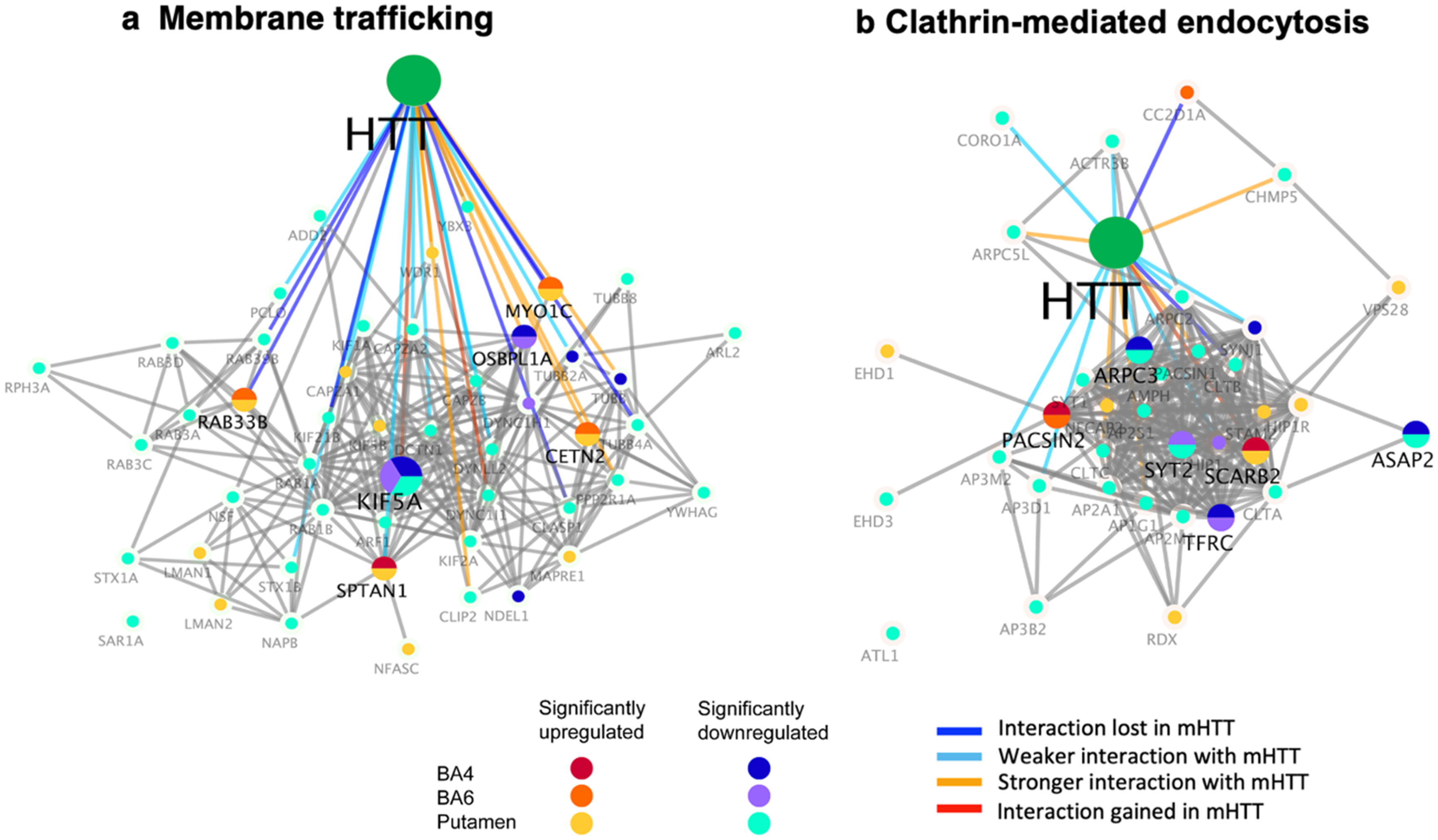
Dysregulation of membrane trafficking cluster and clathrin-mediated endocytosis interactors of HTT in putamen and cortex (BA4 and BA6) of juvenile HD brains compared to age-matched controls. HTT interactors of the membrane trafficking cluster (a) and the clathrin-mediated endocytosis cluster (b) that were significantly upregulated or downregulated in HD compared to control brains are illustrated for putamen, BA4 cortex, and BA6 cortex regions by the indicated color-coded protein components (shown by gene name). Interactions with mutant HTT are indicated as lost, weaker, stronger, or interaction gained compared to normal HTT, using information from our database of HTT interactors identified in HD model systems. 16
Downregulation of the clathrin-mediated endocytosis cluster of HTT protein interactors in putamen and BA4 cortex of juvenile HD brains
The putamen brain region of human juvenile HD brains showed downregulation of HTT-interacting proteins of the clathrin-mediated endocytosis cluster (Figure 7(b), Table 2). The putamen displayed downregulation of 22 components of this endocytic cluster, with a few downregulated components in BA4 and BA6 cortex regions (numbering 4 and 2, respectively) (Table 2). In putamen, downregulated endocytic components consisted of adaptor complex subunits (AP1G1, AP2A1, AP2M1, AP2S1, AP3B2, AP3D1, AP3M2), clathrin proteins (CLTA, CLTB, CLTC), actin-related proteins (AP3M2, ARPC3, ARPC5L), and synaptotagmins (SYT1, SYT2) (Supplemental Table 7). Several components were downregulated in BA4 and BA6 cortex that included ARPC3, ASAP2, and TFRC involved in functions related to actin, vesicle budding, and transferrin receptor, respectively. Many of these downregulated HTT interactors have been previously shown to display weaker interaction with mutant HTT compared to normal HTT (Figure 7(b)). 16
Upregulation of several HTT-interacting proteins with clathrin endocytic functions occurred in putamen (EHD1, HIP1R, PACSIN2, RDX, SCARB2, STAM2, VPS28) as well as in BA4 and BA6 cortex (SCARB2, PACSIN2, and CC2D1A).
Dysregulation of HTT interactors of clathrin-mediated endocytosis implicates deficits in trafficking of cellular proteins from the cell surface to intracellular locations in human juvenile HD brain.
Upregulation of the cytoplasmic vesicle cluster of HTT interactors in putamen and cortex of juvenile HD brain
For the cytoplasmic vesicle lumen cluster of HTT interactors, many were upregulated in putamen and BA4 cortex, and several were upregulated in BA6 cortex, of juvenile HD brain (Figure 8, Table 2). Upregulated components included ANXA2 and C1QC in putamen and cortex (BA4 and BA6), combined with FABP5, CYBR3, PYGB, PRDX6, C1QB, CSTB, MT2A, and ORM1 largely in putamen and BA4 cortex (Supplemental Table 8). Such interactors have been found to show stronger and lost interactions with mutant HTT compared to normal HTT. Involvement of HTT interactors with vesicle functions appears consistent with changes in neurotransmitter regulation observed in proteomics studies of juvenile HD brain. 17
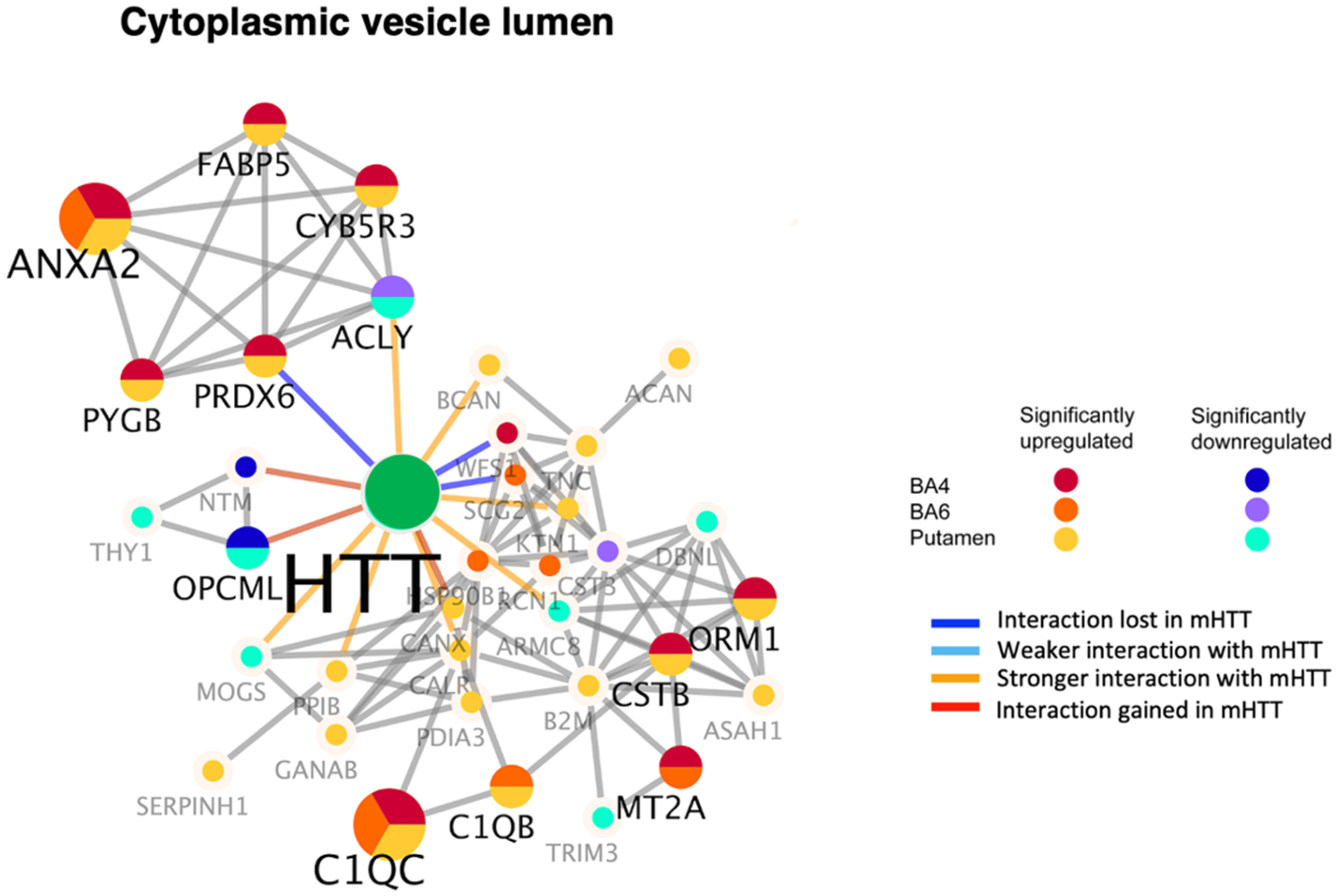
Dysregulation of the cytoplasmic vesicle lumen type of HTT interactors in putamen and cortex (BA4 and BA6) of juvenile HD brains compared to age-matched controls. HTT interactors of the cytoplasmic vesicle lumen cluster of HTT interactors that were significantly upregulated or downregulated in HD compared to control brains are illustrated for putamen, BA4 cortex, and BA6 cortex regions by the indicated color-coded protein components (shown by gene name). Interactions with mutant HTT are indicated as lost, weaker, stronger, or interactions gained compared to normal HTT, using information from our database of HTT interactors identified in HD model systems. 16
Dysregulation of protein modification HTT interactors involving the ubiquitin-proteasome system in putamen and cortex of juvenile HD brain
Dysregulation was observed as downregulation and upregulation of HTT-interacting components involved in protein modification, which occurred in putamen with minor dysregulation in cortex (BA4 and BA6) regions of HD brain (Figure 9(a), Table 2). Downregulation of such HTT interactors in putamen consisted of CUL2, OTUB1, PSMD6, PSMD7, UBE2K, UBL4A, and USP46 (Supplemental Table 7) that regulate ubiquitin-mediated proteasome protein degradation. In addition, PSMD14, a component of the proteasome, was downregulated in BA4 cortex. Further, several components of the ubiquitin-proteasome system were upregulated in putamen consisting of PSMC4, UBA1, UBXN1, and USP10. This HTT interactor cluster of protein modifiers have been shown to display stronger interactions with mutant HTT compared to normal HTT (Figure 9(a)). 16 Dysregulation of HTT interactors of the ubiquitin-mediated proteasome system is consistent with evidence that HTT is ubiquitinated and degraded by proteasome, resulting in clearance of mutant HTT.29,30
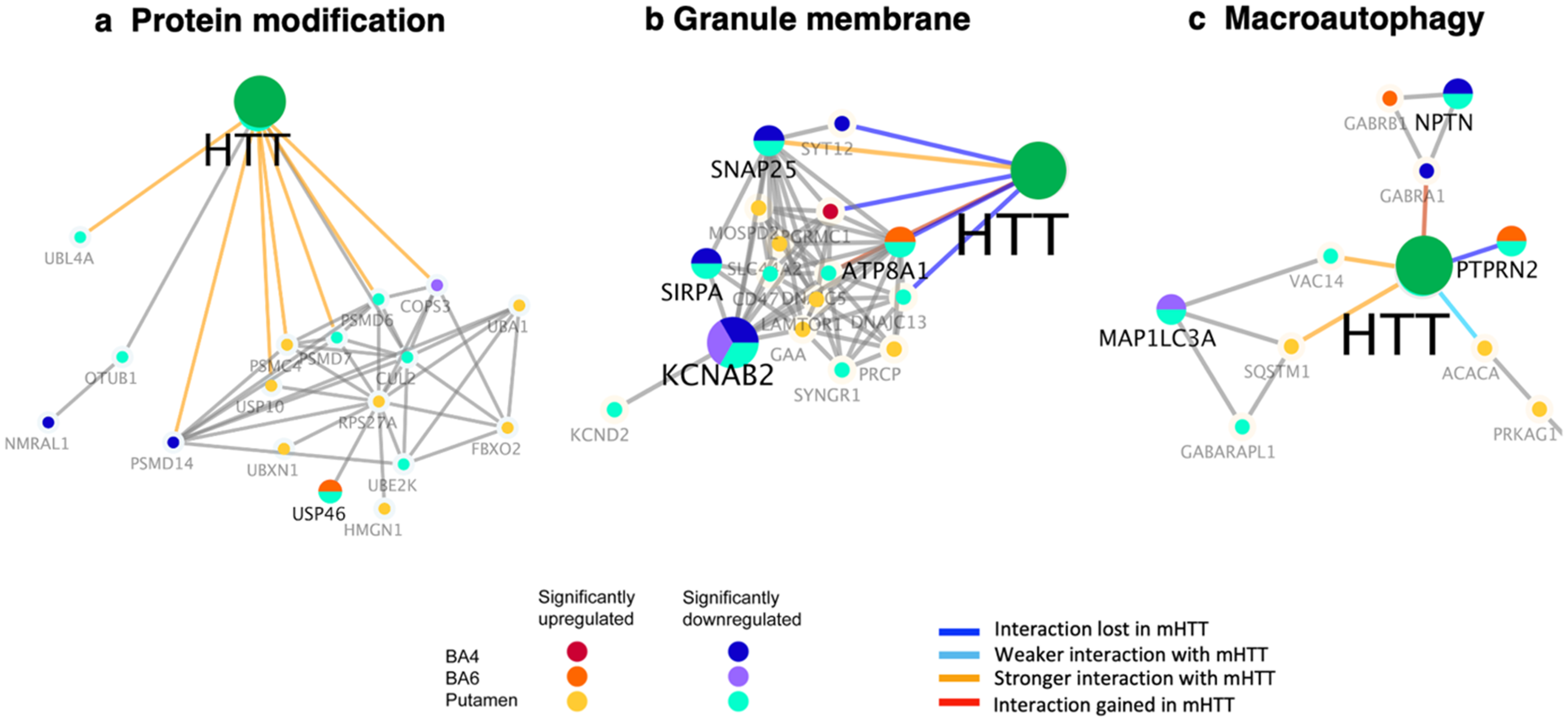
Dysregulation of the protein modification, granule membrane, and macroautophagy clusters of HTT interactors in putamen and cortex (BA4 and BA6) of juvenile HD brains compared to age-matched controls. HTT interactors of the protein modification (a), granule membrane (b), and macroautophagy (c) clusters of HTT interactors that were significantly upregulated or downregulated in HD compared to control brains are illustrated for putamen, BA4 cortex, and BA6 cortex regions by the indicated color-coded protein components (shown by gene name). Interactions with mutant HTT are indicated as lost, weaker, stronger, or interaction gained compared to normal HTT, using information from our database of HTT interactors identified in HD model systems. 16
Dysregulation of granule membrane cluster of HTT interactors in putamen and cortex of juvenile HD brain
The granule membrane cluster of HTT interactors displayed dysregulation in juvenile HD brain regions, shown by upregulated and downregulated components (Figure 9(b), Table 2). Dysregulation was observed for ATP8A1 ATPase function, KCNAB2 and KCND2 involved in potassium channel function, and SIRPA regulatory protein and SNAP25 synaptosome protein (Supplemental Table 10). Such HTT interactors have been found to display weaker or stronger interactions with mutant HTT compared to normal HTT. Altered granule membrane interactors suggests relationships with endocytosis and vesicle functions in HD.
Dysregulation of macroautophagy cluster components of HTT interactors in putamen and cortex of juvenile HD brain
Several macroautophagy HTT interactors showed several dysregulated components (Figure 9(c), Table 2, Supplemental Table 11), consisting of downregulated MAPILC3A, NPTN, GABRA1, GABARAPL1, PTPRN2, VAC14 protein components, and upregulation of ACACA GABR8, PRKAG1, PTPRN2, SQSTM1 in putamen and cortex (BA4 and BA6) regions of HD brain. These components have been shown to display interactions with mutant HTT that are stronger or lost compared to normal HTT. These findings suggest HTT interactor regulation of functions in lysosome-mediated degradation of macroautophagy.31,32
Discussion
Substantial dysregulation of HTT-interacting proteins occurred in human juvenile HD brain, occurring primarily in the putamen region compared to cortex (BA4 and BA6 regions). Juvenile HD putamen displayed prominent downregulation of HTT interactors of mitochondrial and signal transduction functions. Putamen also displayed significant upregulation of HTT interactors involved in RNA splicing, chromatin organization, and translation (Table 2). These findings demonstrate that severe dysregulation of HTT interactors occurs in putamen that undergoes extensive neurodegeneration in HD resulting in compromised motor skills and movement behaviors.1–5 In addition, moderate dysregulation occurred for interactors with vesicular membrane trafficking functions of clathrin-mediated endocytosis, vesicle, and granule membrane functions. The BA4 and BA6 cortex regions of juvenile HD brain showed upregulation and downregulation of HTT interactors like those in putamen, but fewer dysregulated components occurred in these cortex regions compared to putamen. Dysregulation of HTT interactors in putamen combined with cortex BA4 and BA6 regions of juvenile HD brains occurs with compromised motor function regulated by these brain areas.
Investigation of HTT-interacting proteins in human HD brain used the strategy of applying a database of HTT interactors compiled from non-human HD animal models 16 for interrogation of proteomics data acquired from human juvenile HD brain regions of putamen and cortex (BA4 and BA6) regions. 17 The database consisted of 2971 proteins that interact with mutant and normal HTT that were identified in model HD organisms from yeast to transgenic mice. 16 The database includes information on protein interactions with mutant HTT as being weaker, stronger, lost, or gained compared to normal HTT interactions. Use of the HTT interactor database provided a solution to the difficulty of in vivo identification of HTT interactors in human HD brain due to the limited numbers of rare juvenile HD cases and paucity of degenerated putamen, as well as cortex, brain tissues available from human HD brain cases. Analysis of HTT interactors in proteomics data showed that the putamen region from human juvenile HD brain demonstrated prominent dysregulation of HTT interactors indicated by downregulation of mitochondrial and signal transduction with trafficking functions combined with upregulation of chromatin and translational functions.
Protein interactions with mutant HTT is proposed in the field as important mechanisms for motor deficits and neurodegeneration of HD caused by expanded CAG repeats within the mutant HTT gene. Such protein interactors may bind directly to HTT or may be components of protein complexes that bind to or associate with HTT in the HD disease condition. Importantly, results from this study show that the clusters of proteins interacting with mutant HTT are dysregulated in human juvenile brain. The numerous functional clusters that display dysregulation in HD demonstrate a complex spectrum of interactors that involve transcription to translational functions with subsequent signaling mechanisms involving membrane and vesicular trafficking processes. The dysregulated HTT protein interaction complexes occurring in human juvenile HD brain may be proposed to participate in HD behavioral deficits and neurodegeneration. It will be of interest in future studies to assess the roles of prominently dysregulated clusters of HTT-interacting proteins in HD deficits.
Mutant HTT in human juvenile HD brain results in dysregulation of proteomes of mitochondrial and synaptic neurotransmitter systems compared to age-matched controls. 17 Proteomics studies showed that cortex (BA4 and BA6 regions) and putamen from juvenile HD brains displayed deficits in mitochondrial translation and ribosome proteome systems with compromised mitochondrial matrix systems. 17 Furthermore, dysregulation of synaptic neurotransmission proteome components occurred in human juvenile HD brain regions of putamen, as well as cortex. However, it is not yet known which of the mutant HTT-interacting proteins participate as key regulators of HD deficits in mitochondrial, synaptic, and related neurodegeneration systems that result in motor and behavioral deficits. It will be important for future studies to define the primary HTT interactors that control key proteome components in the regulation of motor dysfunction and behaviors in HD.
The changes in the spectrum of HTT protein interactors in cortex and putamen (within the striatum) of human juvenile HD brains represent outcomes of mutant HTT gene expression in the cortical-striatal pathway that regulates motor movement.33–35 Cortical neurons project excitatory glutamatergic inputs to striatal neurons which provide inhibitory GABAergic input to the globus pallidus (Figure 10). Inhibitory GABAergic projections from the globus pallidus to the thalamus regulate the striatum and motor cortex through excitatory glutamatergic innervation. But in HD, these neural connections are lost due to neurodegeneration that result in motor dysfunction. The degenerating cortex and putamen regions in HD display dysregulation of HTT interacting proteins. It will be important in future studies to assess the mechanisms of altered HTT interactome complexes in the degeneration and pathogenesis of HD.
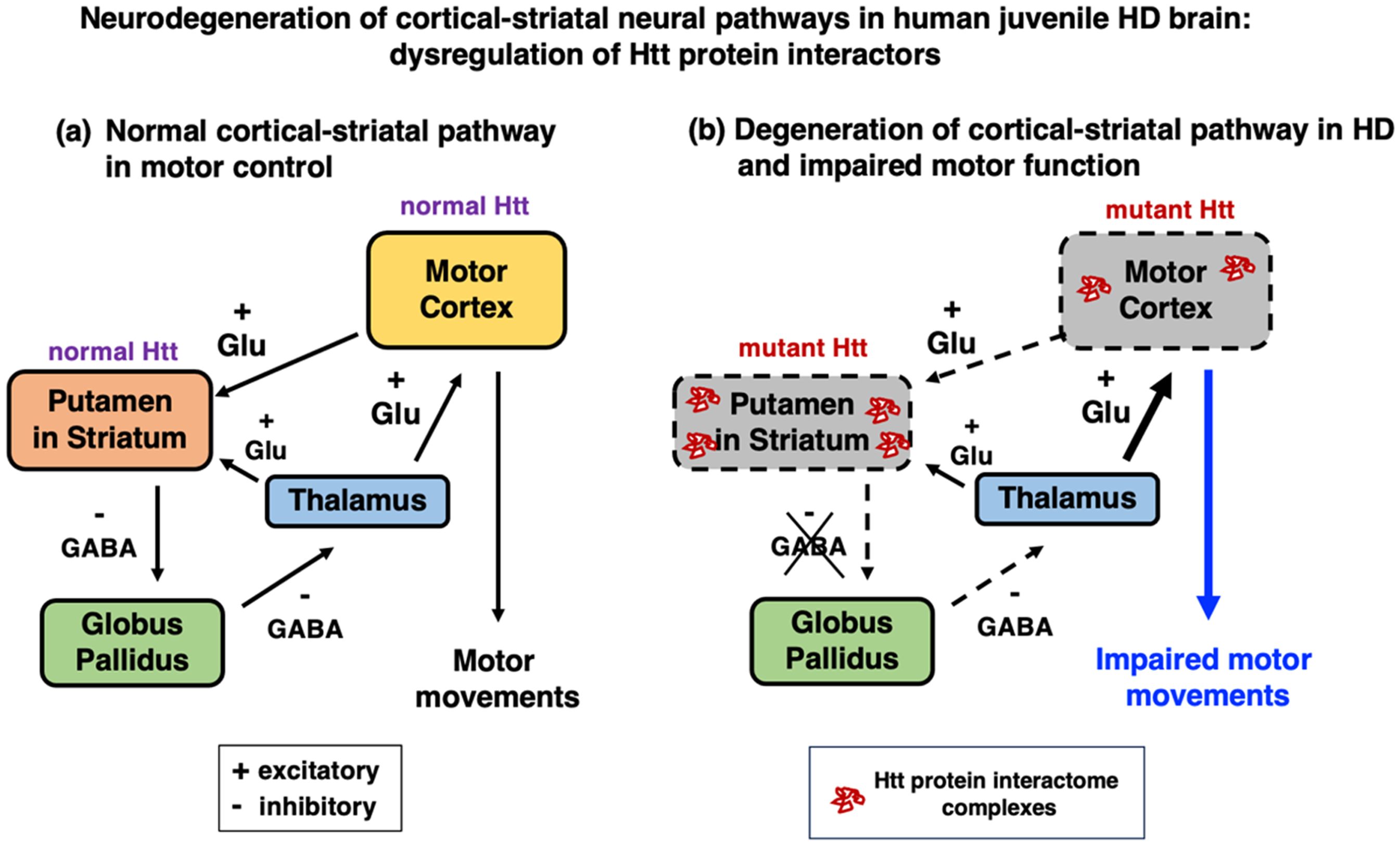
Neurodegeneration of cortical-striatal neural pathways in human juvenile HD brain: dysregulation of HTT protein interactors. (a) Normal cortical-striatal brain pathway for regulation of motor movement. The motor cortex regulates motor movements by excitatory glutamatergic innervation of the striatum containing the putamen region. Striatal neurons provide inhibitory GABAergic innervation to the globus pallidus, which regulates the thalamus's excitatory glutamatergic input to the cortex.33–35 (b) In HD, mutant HTT gene results in degeneration of cortical-striatal pathway regions and changes in the spectrum of HTT protein interactors. The mutant HTT gene of HD results in degeneration of the cortical-striatal pathway, with loss of cortical-striatal connections and altered thalamus excitatory input to the cortex.33–35 These HD degenerating cortical-striatal pathway regions of cortex (BA4 and BA6 regions) and putamen, an area within the striatum, contain changes in the spectrum of HTT interactome protein complexes as shown by this study.
It is important to recognize the value of non-human HD models, from yeast to mammals, for identification of candidate HTT interactors. 16 These data of HTT interactors in animal models advances knowledge of human HD mechanisms, as shown in this study by the dysregulation of such HTT interactors in human HD brain in this study.
This study provides a first step in a series of future investigations to determine the functional roles of dysregulated HTT interactors in human HD pathogenesis. Future utilization of orthogonal multi-disciplinary approaches will benefit the next phases of research to gain understanding of how HTT interactors regulate HD deficits. For example, studies which directly identify dysregulated HTT-interacting proteins in human HD brains may be achieved by separation and isolation of HTT protein complexes, with and without crosslinking of HTT with interacting proteins, followed by proteomics analysis. Further, to determine the cell type involved in HTT-interacting protein functions, co-immunofluorescence imaging of HTT and its interactors can assess whether neuronal or glial cell types participate in dysregulation of HTT interactors. Importantly, functional studies in emerging animal models and model human iPSC HD neurons and/or glia cells36–47 will be important to evaluate the role of HTT interactors in cellular mechanisms of neurodegeneration, pathology, and behavioral deficits. These and other orthogonal approaches will be important in the next phase of studies to bridge the limitations of the current study towards understanding the mechanistic functions of HTT interactors in HD deficits.
Overall, results of this study found that a complex spectrum of HTT protein interactors are dysregulated in human juvenile HD brain regions of putamen and cortex (BA4 and BA6 regions) of the cortical-striatal pathway that regulates motor functions. These findings of downregulated and upregulated HTT interactors were investigated in juvenile HD brains of Vonsattel grade 3–4, representing a late stage of HD. It will be important for future studies to compare HTT interactor regulation during early and later stages of the prevalent adult HD condition; such studies will provide insight into the early disease process that progresses to behavioral deficits and neurodegeneration. Investigation of HTT interactor regulation of protein networks responsible for HD deficits can lead to novel drug targets for future HD therapeutics discovery.
Supplemental Material
sj-docx-1-hun-10.1177_18796397251358348 - Supplemental material for Dysregulation of huntingtin interacting protein networks in human juvenile Huntington's disease brain
Supplemental material, sj-docx-1-hun-10.1177_18796397251358348 for Dysregulation of huntingtin interacting protein networks in human juvenile Huntington's disease brain by Sonia Podvin, Brin Rosenthal, Charles Mosier, Enlin Wei, Kathleen M Fisch and Vivian Hook in Journal of Huntington's Disease
Footnotes
Acknowledgments
We appreciate gifts of human brain tissues for juvenile HD and age-matched controls from the Harvard Brain Tissue Resource Center (HBTRC), the HDSA Center of Excellence at the University of Washington, the HDSA Center of Excellence at the University of Miami, and the HDSA Center for Excellence at the University of Pittsburgh. Review by the Institutional Review Board for human subjects has indicated that the use of unidentified human tissue samples is exempt for this research.
Ethical considerations
The use of non-identified human brain tissues followed protocol of the Institutional Review Board for use of human tissues obtained from brain banks.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by a grant from the Huntington's Disease Society of America (HDSA) to S. Podvin, C. Mosier, and V. Hook.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
