Abstract
The alpha-synuclein seed amplification assay in cerebrospinal fluid is the first validated molecular measurement of alpha-synuclein biology in a living person. The SAA test is transforming our understanding of aging and neurodegenerative diseases by detecting abnormal synuclein biology, and data suggests SAA positivity can occur across Parkinson's disease, Alzheimer's disease, and Dementia with Lewy Bodies. To accelerate development of this important research tool, the Michael J. Fox Foundation proactively funded a community of researchers to work both independently and collaboratively, leading to rapid and iterative progress and validation. The collective validation of the assay across industry and academic groups culminated in a Food and Drug Administration Letter of Support for the test in clinical trials for PD. This article describes the principles that accelerated the development of the assay including patient engagement, collaboration, a commitment to open science through data, sample, and knowledge sharing, and showcases how an international community of experts rallied together towards a common goal.
Plain language summary
Parkinson's disease (PD) research recently reached a pivotal milestone through the validation of a new biomarker test called the alpha-synuclein seed amplification assay (aSyn SAA). Originally derived from methods to detect prion diseases, this assay was adapted through the efforts of a mission-driven research community brought together by the Michael J. Fox Foundation (MJFF). Critical to this success was a commitment to open science—sharing data openly, discussing findings regularly, and collaboratively solving technical challenges. MJFF facilitated access to high-quality, standardized biospecimens, greatly improving test consistency and accessibility. This open approach helped the test earn support from the U.S. Food and Drug Administration, who now recommends its use in selecting participants for clinical trials aimed at slowing or preventing PD. This groundbreaking diagnostic test also enabled, for the first time, a mechanism to define PD in biological terms. Characterizing the disease based on biology (instead of heterogenous clinical symptoms) is paving the way for personalized treatment approaches and earlier, targeted interventions for those at risk.
Keywords
The alpha-Synuclein seed amplification assay (aSyn SAA) builds upon a technology originally developed in the early 2000s for detection of minute concentrations of misfolded prion proteins. Originally coined protein misfolding cyclic amplification (PMCA), with subsequent modifications classified as Real-Time Quaking-Induced Conversion (RT-QuIC) these technologies leverage the inherent propensity of aggregation-prone proteins such as prions, amyloid-beta, and alpha-synuclein (aSyn) to seed the nucleation of fibril formation. 1 Functionally analogous to the polymerase chain reaction used for DNA amplification, the assay involves incubating a human biological sample, typically cerebrospinal fluid (CSF), with an excess of purified recombinant monomeric aSyn. Through repeated cycles of incubation and mechanical disruption, misfolded aSyn seeds induce the conversion of recombinant aSyn substrate into detectable fibrillar aggregates, enabling a highly sensitive measurement. The assay readout is based on florescence upon binding to β-sheet–rich fibrils for real-time monitoring of aggregation (Figure 1).
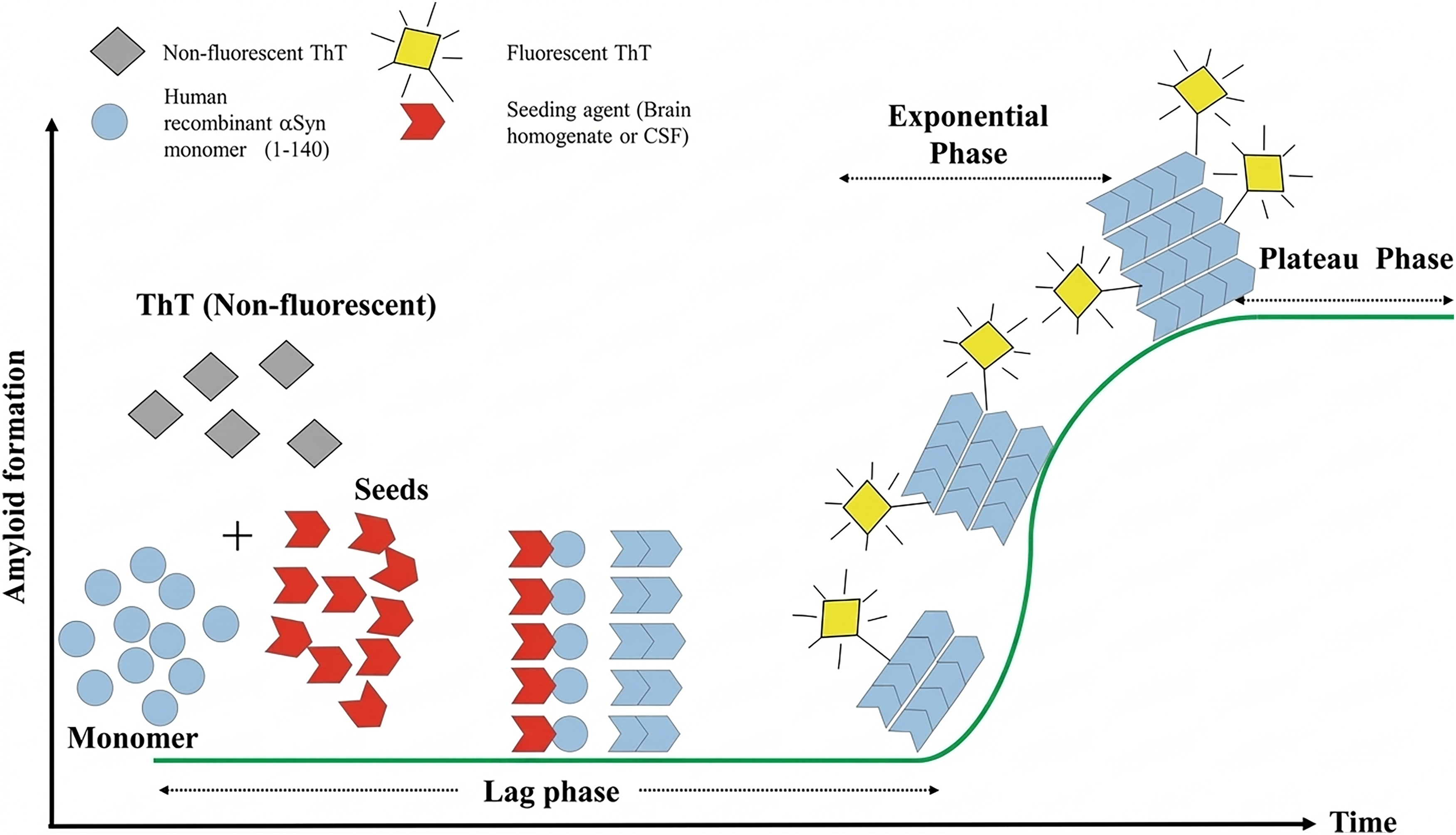
Conceptual schematic of the seed amplification assay (SAA).
The schematic illustrates the core principles underlying α-synuclein SAA. Non-fluorescent thioflavin T (ThT) and recombinant human α-synuclein monomers are incubated with a biological sample containing misfolded α-synuclein seeds (e.g., cerebrospinal fluid or brain homogenate). During the lag phase, pathogenic seeds template the conversion of monomeric α-synuclein into β-sheet–rich fibrillar aggregates. Repeated cycles of incubation and mechanical agitation drive exponential amplification of seeded fibrils, which bind ThT and generate a fluorescence signal. The resulting kinetic curve comprises a lag phase, exponential growth phase, and plateau phase, with fluorescence intensity serving as the assay readout for the presence of seeding-competent α-synuclein.
By 2014 the PMCA technique had been applied to amyloid-beta oligomers. A critical publication from a team in Texas demonstrated both the assay's sensitivity, or high accuracy in correctly identifying individuals with Alzheimer's disease (AD), and its specificity for AD (by also correctly identifying people without AD. 2 Inspired by these findings, scientists at the Michael J. Fox Foundation for Parkinson's Research (MJFF) recognized the potential of this approach for detecting aSyn and quickly contacted the Principal Investigator to provide funding support for its adaptation to Parkinson's disease (PD). Within a few months, the first aSyn seeding assay development grant was initiated, and within a few years, a multicenter study from this group demonstrated that the assay could detect PD in CSF with 88.5% sensitivity and 96.9% specificity. 3 Around the same time, two other teams, one in the UK and another in Montana, pioneered and deployed aSyn-RT-QuICs in local cohorts of not only PD individuals, but also Dementia with Lewy Bodies (DLB) subjects, demonstrating equal or higher sensitivity and specificity values to the Texas group.4,5 These independent studies conducted on different sides of the globe – one using PMCA and two others with RT-QuIC – validated the basic paradigm that misfolded α-synuclein seeds are present in CSF of patients with synucleinopathies and can be amplified for reliable detection. This early engagement reflects a proactive strategy to identify promising technologies across disciplines, a principle that has enabled rapid adaptation of SAA from prion research to aSyn.
A historical timeline delineating this initial assay development of aSyn SAA in 2014 through its deployment as a screening tool for individuals at risk for PD in 2025 is detailed in Figure 2. As additional investigators began exploring similar amplification techniques for aSyn using different methodologies, MJFF continued to fund various groups to increase the chances of success toward a reliable synuclein biomarker for PD. This ability to rapidly deploy funding outside of traditional grant cycles has been pivotal in supporting promising leads as soon as data emerged—a flexible funding approach that has proven essential in today's fast-paced scientific landscapes. Multiple laboratories developed and optimized their assays independently, and while they all showed encouraging analytical performance and accuracy in identifying people with PD in smaller, local cohorts, it was unclear how results compared to each other using the same samples. To address this question, MJFF sponsored two round-robin studies aimed at cross-validating and assessing the performance characteristics of these seeding assays in well-characterized PD clinical cohorts.6,7 The first study was conducted using samples from the BioFIND cohort, followed by a second using samples from the Parkinson's Progression Markers Initiative (PPMI). In both studies, made possible by the generosity and continued engagement of patients and healthy volunteers who contributed biosamples and committed to long-term follow-up, matched CSF samples from PD patients and healthy controls were distributed to multiple laboratories. These labs – comprising both academic and industry teams – were blinded to clinical diagnosis and used their own methodological variations of the assay. Some employed RT-QuIC, while others used PMCA-based techniques. Despite technical differences, the results were analyzed independently and showed remarkable consistency across laboratories (Kang et al., 2019; Russo et al., 2021). These pivotal studies demonstrated the robustness and reliability of aSyn seeding assays. Regardless of the specific assay protocols or the criteria used to define positive replicates, the findings were highly concordant, with sensitivity ranging from 86–96% and specificity from 82–100% for PD, depending on the assay used. Importantly, these values were not intended to define a final or sufficient performance threshold for the assay. Rather, the key milestone at this stage was the high degree of concordance observed across independently developed assays, all applied to the same deeply phenotyped BioFIND samples, despite differences in assay protocols, recombinant substrates, and criteria used to define positivity. This concordance provided sufficient justification to proceed with larger-scale validation efforts, with the expectation that estimates of sensitivity and specificity would continue to evolve as the assays matured. In this context, apparent false-positive and false-negative classifications were defined relative to expert clinical diagnosis in well-characterized cohorts and should not necessarily be interpreted as assay failures. Rather, such discordance may reflect underlying biological heterogeneity or differences in assay thresholds and readouts; notably, the same individuals tended to be discordant across multiple assays, supporting a biological rather than technical explanation.
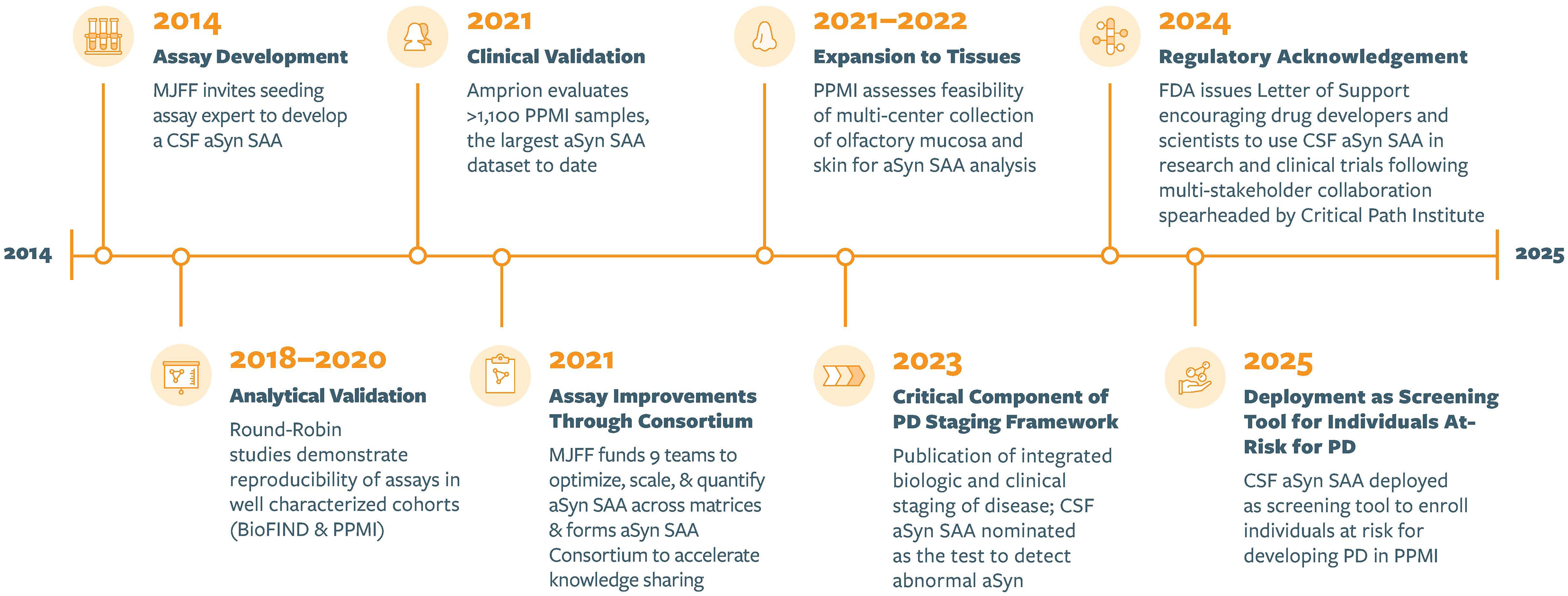
Timeline for the evolution of aSyn SAA. CSF: cerebral spinal fluid; ASyn SAA: Alpha-synuclein Seed Amplification Assay; PPMI: Parkinson’s Progression Markers Initiative; MJFF: The Michael J. Fox Foundation; FDA: Food and Drug Administration; PD: Parkinson's disease.
To accelerate validation of promising synuclein biomarker assays, MJFF established a consortium of grantees funded to develop aSyn biomarkers (including but not limited to SAA) to share knowledge and data regularly as it emerged. For over 7 years, more than 25 field-leading aSyn biomarker experts engaged in spirited discussions via monthly calls, exchanged biosamples, proteins, antibodies, and protocols across continents, convened annually in person for brainstorming workshops, and co-authored data-driven recommendations to align and advance the aSyn biomarker field. 8 Importantly, MJFF actively facilitated collaboration through a guided, milestone-informed funding approach designed to incentivize team collaboration and reduce competitive barriers. This included requirements for timely data sharing prior to publication, participation in regular consortium calls, and engagement in cross-laboratory discussions. The strategic convening of academic, industry, and other relevant stakeholders was instrumental in aligning priorities, reducing silos, and speeding translation from assay discovery to validation. Collaborative efforts such as the round-robin studies highlighted the group's commitment towards a shared goal: to develop robust and reproducible aSyn biomarkers. Over time shared challenges and opportunities for synergy became apparent: the composition and quality of the ‘protein substrate’ was critical to the assay's success, yet each group had a different version of this protein, requiring small-batch synthesis and rigorous biochemical characterization each time. Variables such as shaking frequency and duration, sonication methods, buffer composition, and pH conditions needed to be tailored to the specific protein substrate. Furthermore, commercially available aSyn proteins yielded inconsistent and suboptimal results, and scaling up production posed significant technical and logistical challenges. In response, MJFF launched a large-scale effort to develop affordable, high-quality, commercially available aSyn SAA substrate through a precompetitive collaboration across industry, protein manufacturers, academic SAA experts, and commercial distributors– all for the benefit of the broader research community.
One critical aspect which propelled aSyn SAA forward during this time was a commitment to real-time data sharing. Data deposition from the round-robin efforts described above at PPMI's data repository (the Laboratory of Neuro Imaging - LONI) prior to sample unblinding fostered transparency, minimized duplication, enabled cross-lab learning, and allowed interested researchers from around the world to access the aSyn SAA data along with the wealth of clinical, imaging, and other biomarker information available from these well characterized cohorts. Importantly, aSyn SAA publications are being published under open access permissions to facilitate widespread dissemination, which is a key tenet of MJFF's open science principles. Finally, upon review and discussion of these datasets, and with consensus from the synuclein biomarker consortium, field-leading experts in these assays proposed and aligned on a unified nomenclature for this technology, which was described in the second round-robin publication. 7 The newly coined name aSyn seed amplification assay (aSyn SAA) was quickly adopted and has been substituted for both PMCA and RT-QuIC.
The largest analysis of aSyn in PD samples to date is being performed in the PPMI study, a longitudinal international natural history biomarker study sponsored by MJFF. PPMI continues to evaluate all research participants for aSyn SAA, with over 5000 CSF SAA assessments performed to date (with data openly available at LONI). After the pilot round-robin in PPMI (described above), a larger analysis in over 1100 PPMI subjects confirmed the previously reported high sensitivity (88%) and specificity (96%) of aSyn SAA to differentiate individuals clinically diagnosed as sporadic PD and healthy controls. This work showcased the ability of the assay to detect SAA in people at risk for PD with high sensitivity: participants without PD but with risk factors of hyposmia and/or REM sleep behavior disorder also demonstrated high positivity for aSyn SAA. Interestingly, these aSyn SAA data highlight the potential molecular heterogeneity of PD, as evidenced by the fact that participants with mutations in the LRRK2 gene (the most common genetic risk factor for PD) consistently demonstrate lower rates of aSyn SAA positivity. 9
These exciting findings, supported by a growing body of literature from cohorts around the world, fostered a longstanding spirit of collaboration which paved the way for a Letter of Support (LOS) issued by the Food and Drug Administration (FDA) in the Summer of 2024 (FDA LOS, n.d.). The request to the FDA for this letter was spearheaded by the Critical Path for Parkinson's and cowritten by an international group of biopharma stakeholders, neurologists, and academics who assessed, distilled, and presented as evidence aSyn SAA data from five established observational cohorts in geographically diverse populations and two recent randomized controlled clinical trials of aSyn targeted therapies. Data underlying this request came from thousands of research participants who consented to share their samples and personal health information, reflecting a deep community commitment to advancing research. Based on this significant body of evidence, including independent confirmation across laboratories and rigorous quality control and analytical validation efforts, the FDA letter of support encourages the use of aSyn SAA as an enrichment biomarker for patient selection in clinical trials investigating aSyn therapies to acquire more data around its utility. This is only the second PD biomarker to receive a LOS from the FDA and the first fluid based, molecular marker to receive this designation. The record speed in which this LOS was issued after its submission (just 3 months without any additional information requested) illustrates the power diverse stakeholders can have when they come together and share data towards a common goal of accelerating the utility of biomarkers in clinical trials.
Despite the progress in aSyn SAA outlined in this article, challenges remain that are being addressed by multi-stakeholder collaborations and data sharing across sectors. Today's aSyn SAA is currently binary, which reads out as either positive or negative. Transitioning toward a quantitative SAA could enable monitoring disease progression, stratifying patients by “pathological load”, or measuring impact of interventions as a pharmacodynamic biomarker. In a coordinated effort supported by philanthropic partners and guided by a consortium of industry experts, MJFF is spearheading the validation and acceleration of quantitative biomarkers for PD, and efforts to achieve a quantitative SAA are already underway. Novel readouts such as digital assays, which provide an absolute titer of ‘seeding units’, are demonstrating early feasibility providing a quantitative readout of fibril levels in spiked samples.10,11 Assay optimization efforts continue to detect and quantify aSyn seeds in native human samples across diverse platforms and methodologies.
A major practical limitation to deploying aSyn SAA at scale is its reliance on CSF collection via lumbar puncture. This invasive procedure restricts the test's feasibility in routine clinical practice, may delay or deter subject recruitment in trials, and is unsuitable for frequent repeated sampling. Encouragingly, the prospects for developing a more accessible and scalable aSyn SAA using less invasive matrices are rapidly expanding. While CSF is likely to remain the gold standard matrix in the near term, aSyn SAA detection in skin, blood, olfactory mucosa, saliva, urine, and tears has been reported.12–18 To accelerate innovation and progress in this area, MJFF and other funders are convening and supporting groups across industry and academia to optimize and scale assay technologies from single-site proof-of-concept tests to multi-site deployment, while facilitating coordinated sample collection through natural history cohorts—laying the groundwork for robust cross-platform validation and integration in clinical studies.
One of the most profound impacts of aSyn SAA to date is the way it has enabled the field to redefine PD in biological terms. The ability to detect abnormal aSyn in living people coupled with neuroimaging approaches that characterize loss of neurotransmitters (e.g., dopaminergic imaging) is shifting the field from a description of disease based on clinical symptom presentation to one that integrates measurement of biologic changes alongside clinical manifestations. Two recently proposed frameworks for defining disease based on the accumulation of pathological aSyn, neurotransmitter deficits (such as dopamine loss), and clinical presentation19,20 reflect this exciting transformation that is occurring across neurodegenerative diseases including in Alzheimer's and Huntington's.21,22 The development of laboratory assays (like aSyn SAA) and neuroimaging tools provides insight into the temporal molecular changes that occur throughout the disease course, including prior to manifestation of clinical symptoms.
Frameworks of disease definition are expected to evolve and improve in precision with increasing understanding of disease biology in parallel with the development of tools detecting and quantifying disease biology. Building strong evidence across multiple global cohorts, including data from natural history studies and interventional trials will be critical to refine the biological definitions. MJFF has established a global consortium to facilitate data and knowledge sharing on molecular biomarkers across neurodegenerative diseases including PD, DLB, MSA, and AD. The opportunity to increase sample sizes by combining data across cohorts and incentivizing co-publication creates a platform for rapid and frequent sharing of data between investigators. In the coming years, these collective efforts are poised to answer pressing research questions such as the temporal relationship between SAA positivity and dopaminergic dysfunction, the clinical trajectory of SAA-negative PD, biological correlates of resilience in asymptomatic SAA-positive individuals, and the significance of distinct molecular subtypes of co-pathologies for clinical interventions. Importantly, these efforts are sustained on principles of transparent data sharing, method harmonization, and community-wide commitment to reproducibility.
Defining diseases based on biological signals will accelerate therapeutic development by enabling informative clinical trial design, more precise patient enrollment and stratification, and better alignment between molecularly targeted therapies and the patients most likely to benefit. Treatments currently in clinical trials are targeting several proposed molecular underpinnings of disease including aSyn misfolding, LRRK2 activity, and GBA dysfunction (summarized in MJFF's Parkinson's Priority Therapeutic Clinical Pipeline Report). The application of objective tools that detect and quantify disease biology in clinical trials will improve the likelihood of success by ensuring appropriate patient matching. Notably, assays such as the aSyn SAA, which can detect abnormal aSyn prior to the onset of motor symptoms, offer the exciting potential to intervene earlier in the disease course—potentially preserving neuronal function before significant degeneration occurs. This mirrors the evolution seen in oncology, where the development of molecular diagnostics has led to more tailored and effective treatment strategies. 23 The PD field is now embracing an unprecedented ecosystem of collaboration, where pharmaceutical companies, academic investigators, and non-profits are aligning to accelerate biomarker-driven innovation. This synergy will continue to be crucial for planning the next generation of interventional trials. The vision of precision medicine in synucleinopathies is becoming increasingly tangible: such biologically tailored decision-making was unimaginable just a decade ago and is now within reach due to sustained investment in open-science collaborations and community-wide commitment to αSyn SAA development. One of the neuropathological hallmarks of PD is accumulation of aSyn in the form of Lewy bodies in the brain. Thus, it is not surprising that aSyn biomarkers are perhaps the most sought after and advanced fluid assays to date, with aSyn SAA being the front runner in detecting abnormal protein misfolding biology in PD. However, it is hypothesized that PD is multi-factorial and many biological pathways including neuroimmune, lysosomal, and mitochondrial function are impaired. Going forward, it will be critical to continue to push forward in developing tools to measure these disrupted biologic pathways, as this will have lasting implications for both clinical trials and future iterations of disease staging (for an overview of the current state of the field of PD biomarkers, see the Parkinson's Biomarkers Pipeline Report). The principles described in this article around early high-risk investments, open data sharing, multi-institutional collaborations, and quick allocation of resources are also being applied to biomarker tool development for these pathways, and provide a transferable roadmap for biomarker development and validation across neurodegenerative diseases.
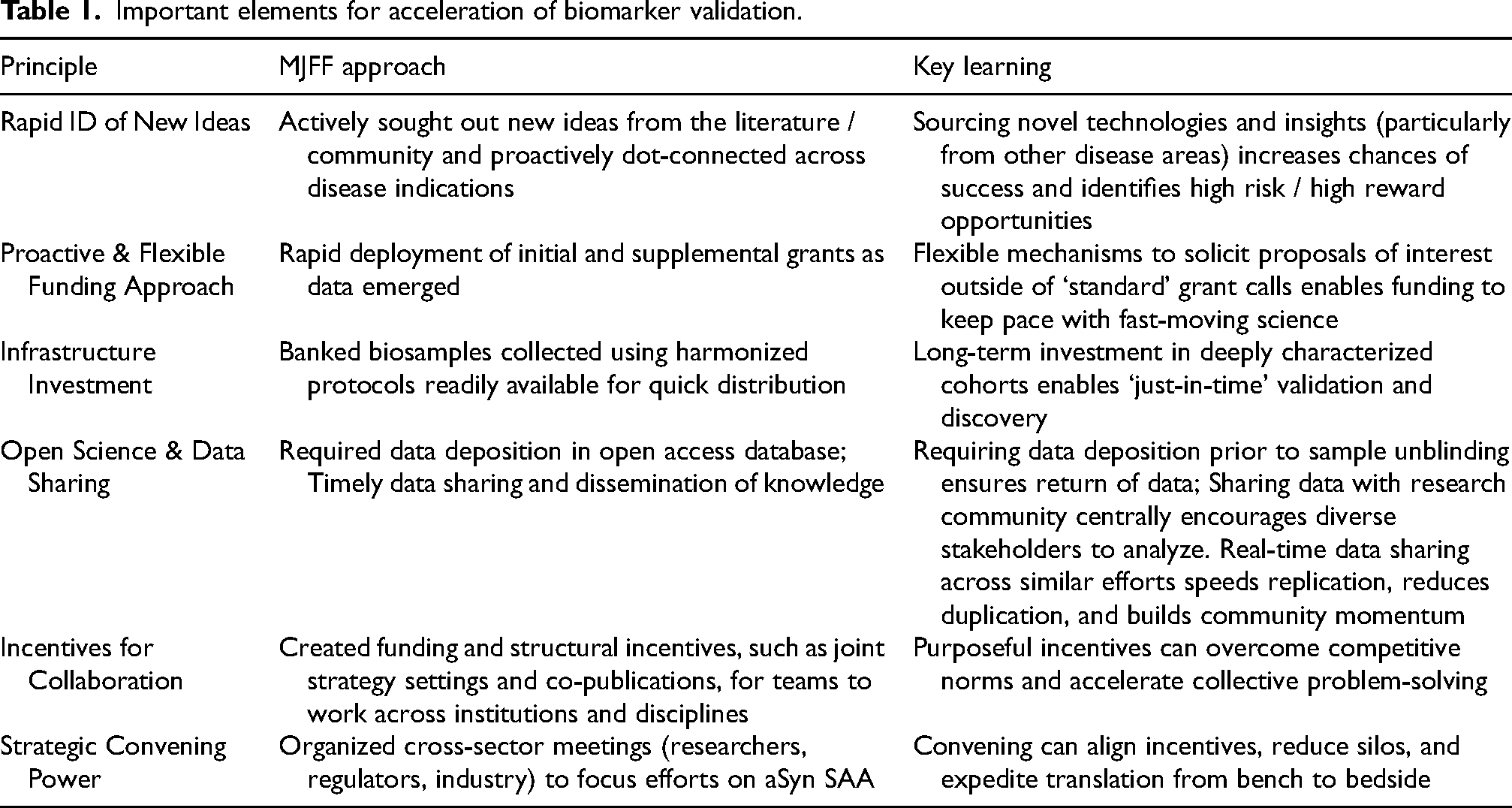
In summary, the journey of aSyn SAA from concept to clinical validation represents not only a major scientific milestone, but also a model of what is possible through a sustained, coordinated, and mission-driven collaborative approach (Table 1). This achievement was made possible by a diverse coalition of stakeholders: scientists, industry partners, foundations, and most importantly, people living with Parkinson's disease. The PPMI study served as a cornerstone of this effort, providing critical validation of aSyn SAA and exemplifying the power of open science and community-driven research. The contributions of PD research volunteers—who generously provided biological samples and shared their lived experiences over more than a decade—were foundational to this success and remain central to the progress ahead. The story of aSyn SAA is still unfolding but given the rapid strides so far and the continued collaborative spirit across multiple stakeholders, there is genuine optimism that aSyn SAA will play a central role in the next generation of breakthroughs in PD research toward a future where precision diagnostics and treatments can meaningfully alter the course of disease.
Important elements for acceleration of biomarker validation.
Footnotes
Acknowledgements
The authors thank Kate Brolley and Brooke Harman for their generous help with designing Figure 2.
Author contributions
All authors contributed to manuscript conception, writing, and review.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
