Abstract
Background
Recent technologies enabling the study of spatial biology include multiple high-dimensional spatial imaging methods that have rapidly emerged with different capabilities evaluating tissues at different resolutions for different sample formats. Platforms like Xenium (10x Genomics) and PhenoCycler-Fusion (Akoya Biosciences) enable single-cell resolution analysis of gene and protein expression in archival FFPE tissue slides. However, a key limitation is the absence of systematic methods to ensure tissue quality, marker integrity, and data reproducibility.
Objective
We seek to optimize the technical methods for spatial work by addressing preanalytical challenges with various tissue and tumor types, including a decalcification protocol for processing FFPE bone marrow core specimens to preserve nucleic acids for effective spatial proteomics and transcriptomics. This study characterizes a multicancer tissue microarray (TMA) and a molecular- and protein-friendly decalcification protocol that supports downstream spatial biology investigations.
Methods
We developed a multi-cancer tissue microarray (TMA) and processed bone marrow core samples using a molecular- and protein-friendly decalcification protocol. PhenoCycler high-plex immunohistochemistry (IHC) generated spatial proteomics data, analyzed with QuPath and single-cell analysis. Xenium provided spatial transcriptomics data, analyzed via Xenium Explorer and custom pipelines.
Results
Results showed that PhenoCycler and Xenium platforms applied to TMA sections of tonsil and various tumor types achieved good marker concordance. Bone marrow decalcification with our optimized protocol preserved mRNA and protein markers, allowing Xenium analysis to resolve all major cell types while maintaining tissue morphology.
Conclusions
We have shared our preanalytical verification of tissues and demonstrate that both the PhenoCycler-Fusion high-plex spatial proteomics and Xenium spatial transcriptomics platforms work well on various tumor types, including marrow core biopsies decalcified using a molecular- and protein-friendly decalcificationprotocol. We also demonstrate our laboratory's methods for systematic quality assessment of the spatial proteomic and transcriptomic data from these platforms, such that either platform can provide orthogonal confirmation for the other.
Keywords
Introduction
Multiple high-dimensional spatial imaging methods are rapidly emerging that expand our understanding of spatial biology, defined for this manuscript as the human genome, epigenome, transcriptome, and proteome in their native morphological contexts.1,2 These methods include various multiplex immunohistochemistry (mIHC) platforms based on antibodies labeled with fluorophores (e.g., Akoya PhenoImager and Lunaphore COMET), heavy-metal tags (Imaging Mass Cytometry or Ionpath Multiplexed Ion Beam Imaging [MIBI]), 3 and oligo-tags (Akoya PhenoCycler). Recently, advancements in spatial biology have also transformed our ability to map and quantify increasingly numerous proteins and mRNA transcripts in their native morphological contexts. This has been made possible by the advent of spatial profiling technologies that are permitting the analysis of progressively more numerous protein and mRNA transcripts in parallel with subcellular resolution.1,2 These approaches have provided insight into in situ cell populations,4,5 cellular neighborhoods,4,6 and cell-to-cell interactions that inform treatment response,7–10 disease progression,9,10 and clonal evolution of cancers.11–13
Spatial proteomics is a powerful tool capable of discerning altered cancer pathways and identification of cell types or proteins associated with aggressive colorectal cancer (CRC), which can serve as novel biomarkers. 14 Similarly, the use of spatial transcriptomics has been successful in identifying biomarkers associated with early onset prostate cancers and those with aggressive behaviors enabling novel targeted therapy. 15 Spatial organization of the tumor microenvironment (TME) and the accessibility of this environment to endogenous tumor-specific T cells or transduced cellular therapy products is a critical feature of tumor control and immune escape. 16 For example, CD8 T cells’ proximity to tumors determines whether a patient could benefit from PD-1 checkpoint inhibitor treatment, 17 heralding the need for clinical implementation of spatial based assays for use in the clinic to inform the management of cancer patients. However, aside from a limited number of publications,18,19 a systematic approach to ensure reproducibility and quality assurance/control (QA/QC) has not been addressed in the current literature for spatial proteomic or transcriptomics studies. These observations have resulted in the use of spatial proteomic assays as clinical trial correlative tests to guide patient selection for treatment. 20 Our lab has performed high-plex spatial proteomics for more than five years and has developed workflows for instrument verification, assay validation, and downstream data QA/QC.
Limitations to clinical implementation of spatial proteomic and transcriptomics work have been the lack of preanalytical assessment for quality tissues, the need for a systematic method to select regions of interest while retaining comparable morphology, and difficulties working with decalcified bone or tissue. Routine histological processing of bone marrow core biopsies requires decalcification prior to sectioning for microscopic evaluation. Typical decalcification protocols use powerful acids to remove calcium to soften bone allowing for downstream cutting of the specimen into very thin sections (2–4 µm thick) for morphologic examination.21–23 DNA, RNA, and some proteins degrade rapidly in low pH. Unfortunately, routine decalcification protocols include immersing marrow cores in hydrochloric acid (HCl), severely diminishing nucleic acid yields.24,25 Furthermore, autofluorescence of the bone poses a significant problem for downstream immunofluorescence protocols. 26 After our evaluation of several gentler decalcification methods based on neutral pH chelators, we have developed a protocol that permits the use of decalcified bone marrow samples for spatial proteomics (PhenoCycler) and spatial transcriptomics (Xenium) analysis.
The PhenoCycler-Fusion platform utilizes a self-contained, cyclic workflow to visualize ≥ 50 markers on a single 3–4 µm section of tissue or tissue microarray (TMA) sectioned onto a standard positively charged slide. It employs barcode-conjugated antibodies against protein targets of interest followed by an iterative process of revealing up to three antibodies at a time via fluorescently tagged reporters. Similarly, spatial transcriptomics are based on mRNA detection with spatially barcoded primers or spatially resolved photo-cleavage of mRNA probes (e.g., 10x Genomics Visium, NanoString GeoMx, and Slide-seq). While these spatial transcriptomics methods initially provided capture fields with a spatial resolution of ∼10–300 cells, advancement of these platforms with multiplex mRNA detection based on in situ hybridization (e.g., 10x Genomics Xenium, NanoString CosMx) have enabled subcellular resolution. 2 The Xenium platform currently permits single cell analysis of the expression of 377 genes simultaneously and an additional 100 targeted, custom genes. Three-to-four-micron thick tissues are sectioned onto a specialized slide with an image capture area of 12 mm × 24 mm, and the RNA undergoes probe hybridization and ligation to circularizable DNA probes which are enzymatically amplified via rolling circle amplification creating bright fluorescent signals that can be imaged. DAPI counterstaining of cell nuclei allows for cell segmentation analysis that is integrated with the fluorescent transcript signals. The Xenium workflow also preserves sample morphology, and both hematoxylin & eosin (H&E) and immunofluorescence (IF) staining can be performed on the tissue post run, enabling the direct comparison of Xenium transcriptional data to morphological data.
Our goals for this study were to address preanalytical verification of tissues and demonstrate that both the PhenoCycler-Fusion high-plex IHC and Xenium spatial transcriptomics platforms work well on various tumor types, including marrow core biopsies decalcified using a molecular- and protein-friendly chelation protocol. To do this, we developed a small TMA with different tumor tissues that would allow for assessment of a large variety of biomarkers in the spatial platforms under development. In working with different tumor/tissue types, we also found a preanalytical hurdle and developed a protocol to mitigatethe destruction of nucleic acids during decalcification of boney tissues. Finally, we describe our laboratory's methods for systematic quality assessment of the spatial proteomic and transcriptomic data from these platforms, such that either platform can provide orthogonal confirmation for the other.
Materials and methods
Patients and tissues
Marrow particle preparations and TMA cores were processed into formalin-fixed paraffin-embedded (FFPE) blocks following CLIA/CAP laboratory standard operating procedures. The collection and use of the archival samples for this study were conducted under protocols approved by the Institutional Review Boards (IRBs) under Fred Hutchinson Cancer Center and the University of Washington. Normal control tissues consist of tonsils (n = 2). Solid tumor tissues (n = 6) were selected from common and antigenically diverse cancers including breast adenocarcinoma (BRC), non-small cell lung carcinoma (NSCLC), colorectal cancer (CRC), melanoma (MEL), hepatocellular carcinoma (HCC), and Merkel cell carcinoma (MCC). Marrows with hematologic malignancies (n = 3) were also included: a chronic myeloid leukemia (CML) marrow core biopsy, particle preparations of a marrow with acute myeloid leukemia (AML), and a marrow with multiple myeloma (MM). The MM sample was collected with the patient's understanding and written consent under the Fred Hutch IRB-approved protocol for which it was collected. Three autopsy marrow core biopsies were obtained to optimize decalcification and nucleic acid extraction protocols. These samples did not undergo additional spatial studies. The remaining samples were collected under a waiver of consent for discarded specimens. All research was conducted in compliance with The Code of Ethics of the World Medical Association (Declaration of Helsinki), printed in the British Medical Journal (18 July 1964).
Optimization of decalcified marrow processing, tissue preservation, and preparation
Bone marrow core biopsies 2 mm in diameter were fixed in 10% neutral-buffered formalin for a ≥ 24 hours to ensure complete fixation before chelation with one of three tested chelation reagents: Immunocal® (American MasterTech Scientific; no. 1414), Versenate Chelating Agent (American MasterTech Scientific; no. DCVER), or EDTA/sucrose decalcifying solution (Newcomer Supply®; no. 1048). After fixation, each specimen was immersed in one of the three tested chelation reagents in a 20:1 ratio per core sample (ideally, the specimen is suspended so it does not contact container surfaces). The specimens incubated in the chelation reagent at room temperature for 24 hours or until sufficiently softened as gauged by physical probing of the bone. The solution was changed at least daily to verify proper sample coverage and to ensure the chelating agent was not depleted by its reaction with calcium. Specimens were placed at 4°C during overnight periods. When sufficiently softened, the specimen was rinsed in DI H2O for 10 minutes before proceeding with routine processing and paraffin embedding.
Nucleic acid extraction
DNA and RNA were extracted from the FFPE chelated marrow core specimens to verify post-chelation nucleic acid quality. Separate manual and automated DNA and RNA extraction procedures (Supplement Table 1) were performed in concordance with the respective kit's product insert. Each extraction's input material consisted of three 15 µm curl sections cut from each FFPE block of decalcified marrow core tissue. Nucleic acid concentration and total yield were quantified using the Qubit dsDNA High Sensitivity assay (ThermoFisher Scientific; no. Q32851) and RNA High Sensitivity assay (ThermoFisher Scientific; no. Q32852). Nucleic acid yield quality was assessed by 260/280 and 260/230 ratios, measured with the NanoDropTM One spectrophotometer (ThermoFisher Scientific; no. ND-ONE-W), and by DNA integrity numbers (DIN) and RNA integrity numbers (RIN), which were attained using genomic DNA ScreenTape® (Agilent; no. 5067-5365) and High Sensitivity RNA ScreenTape® (Agilent; no. 5067-5579) analyses, respectively.
Spatial proteomics with PhenoCycler
Samples were sectioned at 4 µm onto positively charged slides. The slides were baked for 1 hour at 60°C and deparaffinized using standard IHC protocols followed by epitope retrieval with HIER H2 Buffer (Agilent DAKO Target Retrieval Solution pH 9.0) at 95°C for 20 minutes. Subsequent blocking and staining steps were performed using previously reported methods 27 with the following exceptions: B1, B2, B3, and BC4 reagents were substituted with N, J, S, and G blockers (Akoya Biosciences), respectively. The stain fixation and image acquisition steps were performed using Akoya's PhenoCycler-Fusion protocols. The images were acquired with the DAPI, Cy3, Cy5, and Cy7 filters and processed into stitched QPTIFF images.
Spatial transcriptomics with 10x Xenium
The Demonstrated Protocol Xenium In Situ Tissue Preparation Guide (CG000578 Rev A) was followed to place samples onto Xenium slides (10.45 mm × 22.45 mm). Deparaffinization and decrosslinking steps followed (CG000580 Rev C). The Xenium In Situ Gene Expression User Guide (GC000582 Rev C) was used for the remaining workflow beginning with probe hybridization. The slides were loaded in the Xenium Analyzer as directed by the Xenium Analyzer User Guide (CG000584 Rev B). The Xenium workflow takes 2.5 days afterthe tissue is sectioned onto the Xenium slide and approximately 48 hours to run. The two duplicate TMAs were made from archival tissues and consisted of the tumors and normal tissue cores as detailed in Patients and tissues section. The whole array was selected as a single region to decrease the run time, and each core was annotated separately post-run by manually selecting the core's coordinates.
Xenium analysis
Cells were re-segmented using Proseg, a probabilistic segmentation method that uses transcript information directly to infer cell boundaries. 28 Cells with fewer than ten transcripts were removed. Expression was normalized as counts-per-cell, log-transformed, before Uniform Manifold Approximation and Projection (UMAP) projections and unsupervised clustering using Leiden were computed. Cell-type labels were assigned to clusters manually by inspecting the pattern of differentially expressed genes between clusters and, particularly in the case of tumor cells, considering which cores they occur in.
PhenoCycler-Fusion analysis
PhenoCycler images were assessed for staining issues (e.g., focus) and tissue artifacts (e.g., tissue folds). TMA cores and regions of interest that passed the preliminary QC step were selected for cellular analysis. QuPath v0.5.1 29 was used to detect (segment) the individual cells with an Akoya-modified StarDist script which generated a segmentation image for each tissue and the cell object data for each cell containing x, y coordinates and mean cell fluorescence pixel intensity (MFI) values for each marker. Empty barcodes (barcode-conjugated fluors with no corresponding barcode-conjugated antibodies) were included in the staining panel to identify non-specific staining of “sticky” cells and areas, such as necrotic regions, where broken cells have released proteins and other cellular components. The resulting empty barcode MFIs could then be used to exclude false-positive cells. An anatomic pathologist reviewed the segmentation images to visually verify the quality of the cell detections outside of necrotic areas (Figure 1(c) and insert) and cell object data including empty barcodes were exported for further analysis. Dr. Yeung also reviewed each marker stain to assess quality (positive signal above background) and compare it to expected expression. Uniform Manifold Approximation and Projection (UMAP) projections and unsupervised clustering using Leiden were computed. Cell-type clusters were manually labeled by inspecting the pattern of differentially expressed genes across clusters. Mean cell intensities were batch-corrected by fitting independent two-component Log-Normal mixture models to each maker across tonsil cores and scaling marker intensities by the ratio between the respective means values of the lower mixture-model components. These adjusted marker intensities were then log-transformed and visualized by projecting into two-dimensions using UMAP.
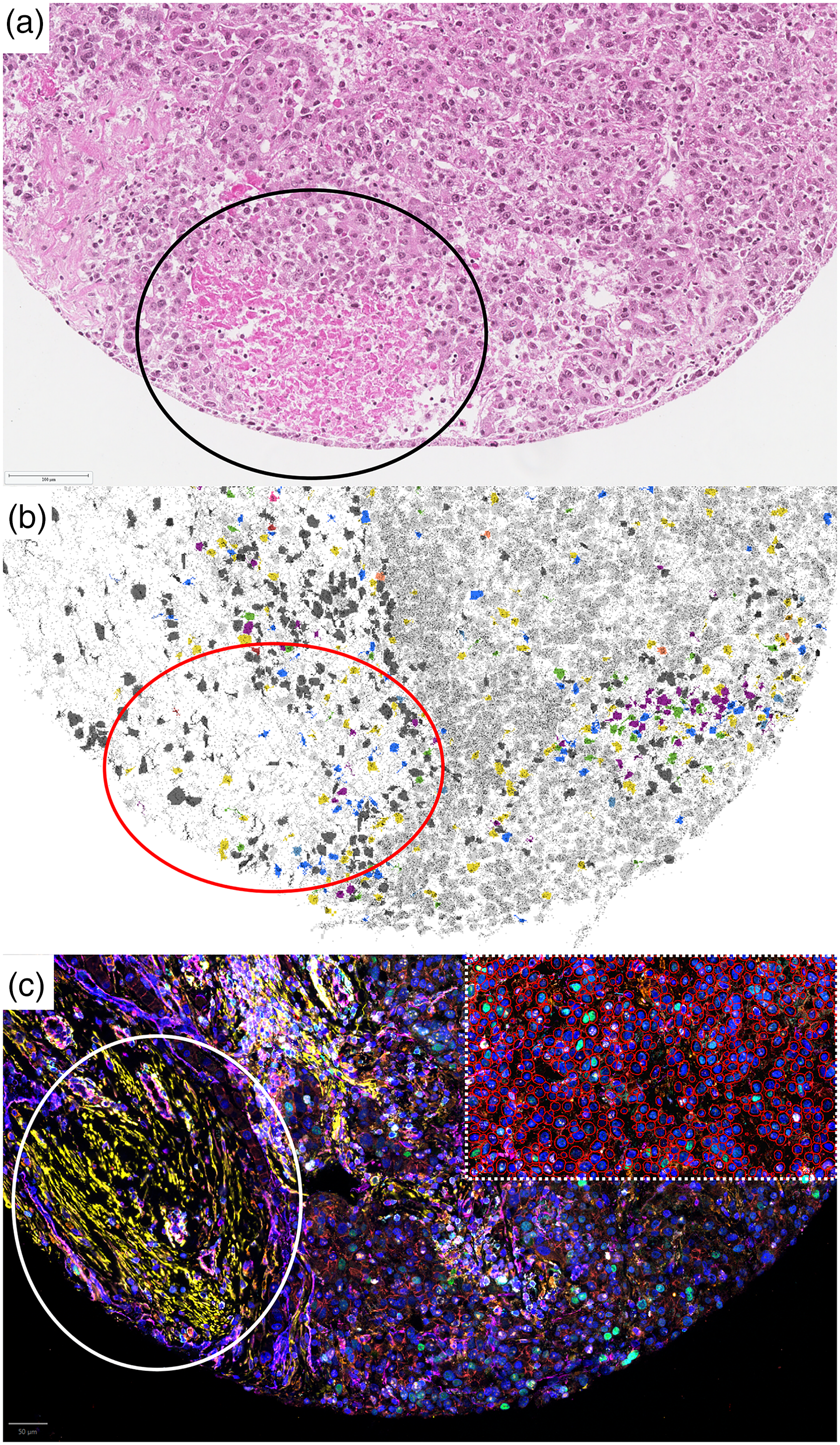
Comparative effect of necrosis on PhenoCycler and Xenium. Matched regions of the HCC show clear patches of necrosis (circled) in the H&E on the top panel (a), Xenium (circled) in the middle panel (b), and PhenoCycler (circled) in the bottom panel (c). (b) The necrotic region shows sparse cellular density as identified using Proseg segmentation in the Xenium image. (c) A combination of residual protein expression and non-specific staining is observed after nuclear degradation. The upper-right insert shows the QuPath-generated nuclear segmentation as cell outlines.
Results
Patient demographics and sample information
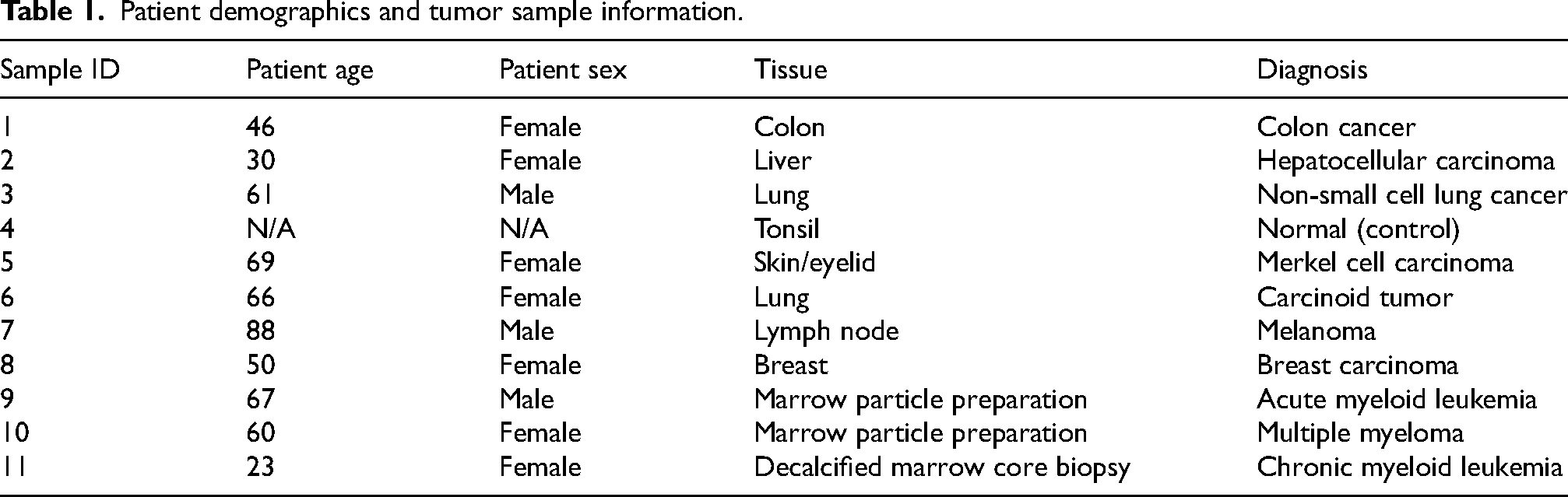
The details of the assayed tissue samples and the associated patient demographics are provided in Table 1.
Patient demographics and tumor sample information.
Quality metrics of samples, decalcified marrow data
Of the three decalcification protocols, the marrow specimens treated with EDTA/sucrose solution consistently produced greater yields of RNA (0.5-fold) and DNA (2-fold). Supplemental table 2 shows the comparative decalcification methods for marrow processing. Although more DNA was purified using the Purigen® instrument than via manual extraction, the RNA protocol was ineffective on the automated instrument requiring manual extractions. The most optimal method was ETDA/sucrose which was used on the CML marrow for Xenium spatial transcriptomics. Quality of each tissue and tumor sample used in the TMA was assessed on fresh-cut H&Es (Supplemental Figure 1) prior to creation of the reference TMA, and regions of interest (ROIs) annotated by an anatomic pathologist (Dr. Lin) were checked by a second anatomic pathologist (Dr. Yeung) ensuring 100% of cores in the TMA show anticipated ROIs. The PhenoCycler-Fusion platform showed increased non-specific background staining with certain cancer types and had more artifacts; the lung carcinoma and hepatocellular tissue both had necrosis (Figure 1(a) – shows an H&E of HCC with necrosis) and demonstrated sparse cells with Xenium data analysis by Proseg (Figure 1(b)). However, PhenoCycler also identified a combination of residual protein expression and non-specific staining in this region (Figure 1(c)). The breast cancer showed edge artifact reactivity with several markers on the PhenoCycler-Fusion platform, whereas the Xenium platform showed relatively uniform coverage and expression throughout the tissue core.
Quality metrics of PhenoCycler-Fusion and Xenium
Primary PhenoCycler QA/QC involves confirming there were no equipment failures during the run and performing a precursory visual check of the QPTIFF image to verify that tissues were in focus with minimal staining artifacts and exposures issues. A pathologist manually performed qualitative assessment of segmentation accuracy. Figure 2(a) shows ideal image quality was obtained on the CML bone marrow core biopsy processed with our optimized EDTA/sucrose decal protocol, with details that can be discerned from individual markers clearly at single-cell resolution for a PhenoCycler-Fusion assay. Quality metrics for PhenoCycler images include comparison of the cell counts and densities for the duplicate tissues between the two TMAs. Table 2 shows that the densities (cells/µm2) between tissue-type duplicates were consistent regardless of differences between tissue sizes. Al Quality metrics from Xenium, as demonstrated in Figure 2(b), include overall cell count, median transcripts per cell, decoded transcripts per 100 µm2 obtained, and image quality to verify single cellular resolution. TMA A1 had 133,049 cells, with a median transcript of 36 per cell and 47 decoded transcripts per 100 µm2. TMA A2 had 126,770 cells, with a median transcript of 37 per cell and 50.6 decoded transcripts per 100 µm2. The CML marrow had 36,602 cells, with a median transcript of 77 per cell and 51.3 decoded transcripts per 100 µm2. The acute myeloid leukemia (AML) marrow had 195,028 cells, with a median transcript of 28 per cell and 25.8 decoded transcripts per 100 µm2. The multiple myeloma (MM) marrow had 139,791 cells, with a median transcript of 10 per cell and 9.1 decoded transcripts per 100 µm2, which raised a flag for low median transcript count per cell.
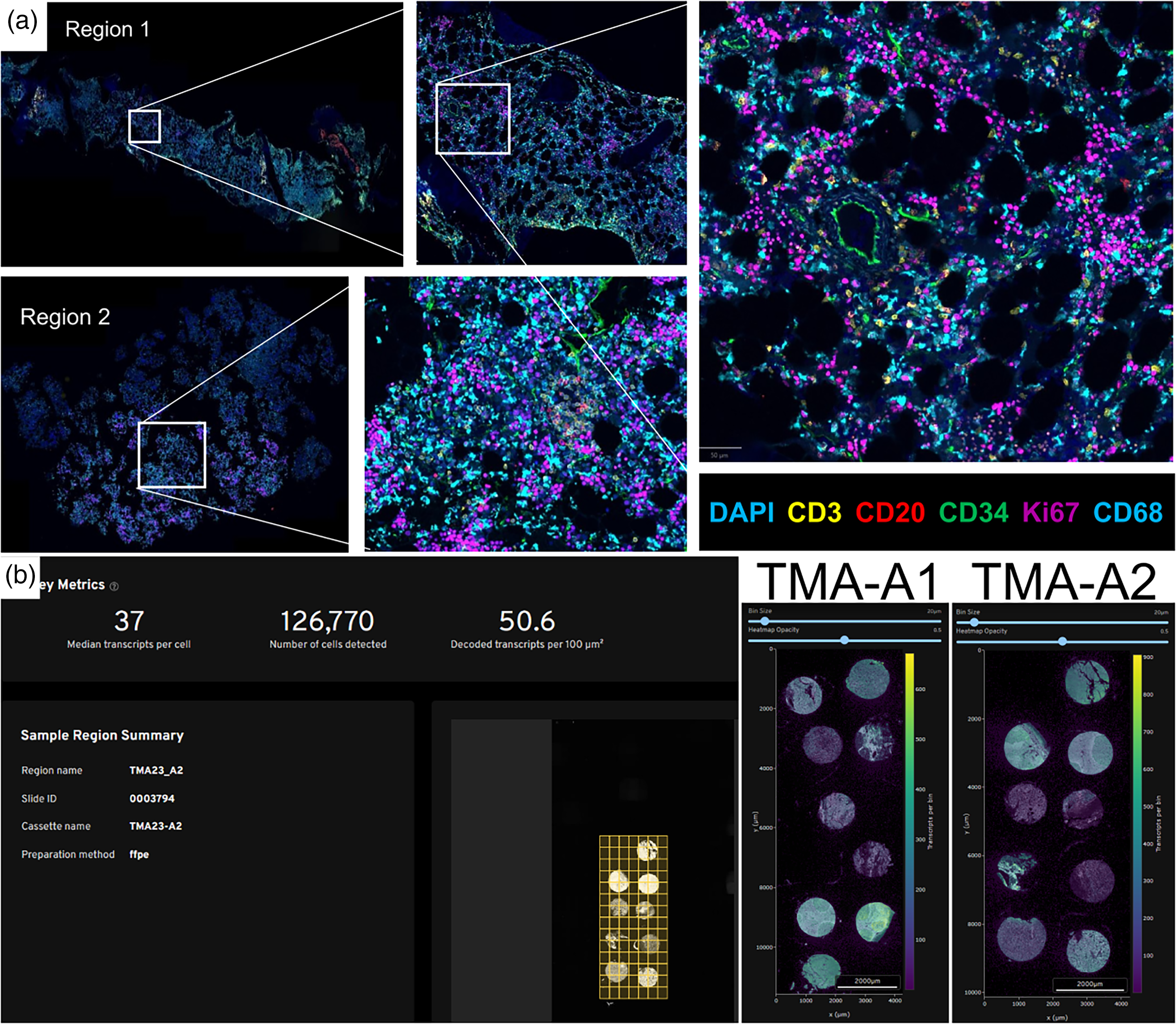
Quality assessment and metrics of PhenoCycler and Xenium. (a) Quality assessment of PhenoCycler-Fusion image ensures focused image capture, single cell resolution, and inspection for artifacts. (b) Quality data provided on the “analysis summary” page of Xenium software providing overall cell count, median transcripts per cell, decoded transcripts per 100 µm2 obtained, and image quality.
Phenocycler cell counts and densities.
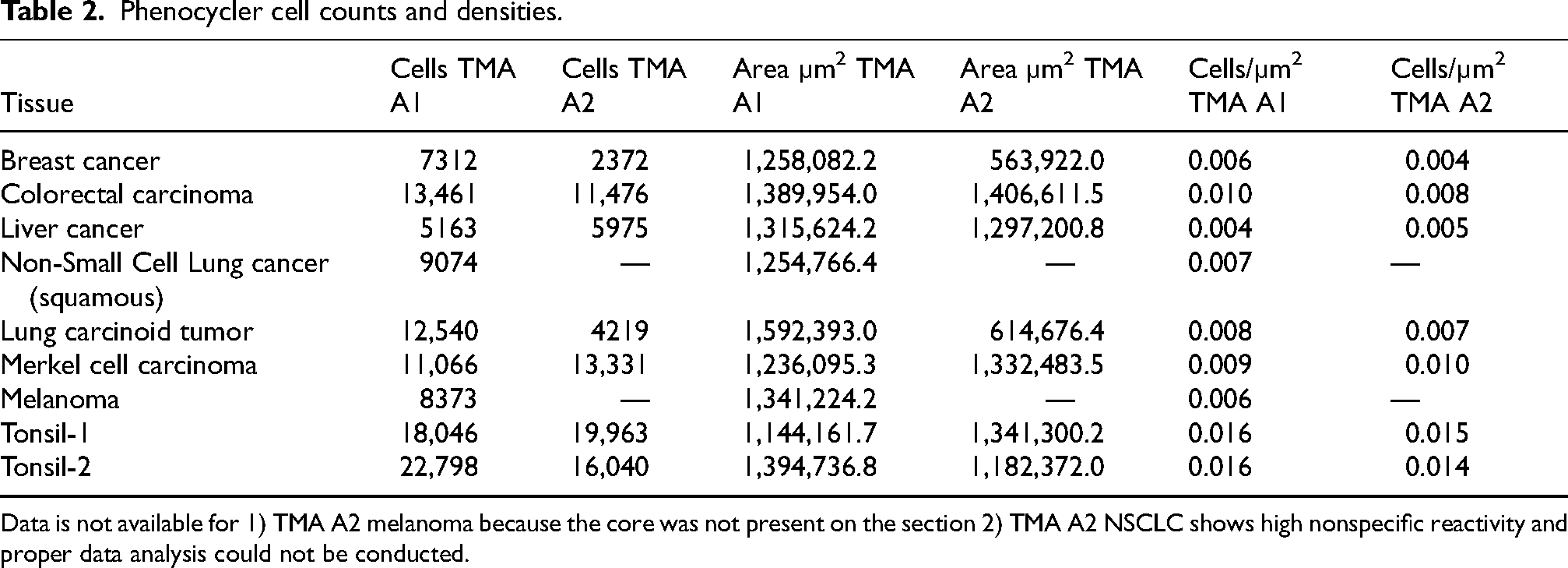
Data is not available for 1) TMA A2 melanoma because the core was not present on the section 2) TMA A2 NSCLC shows high nonspecific reactivity and proper data analysis could not be conducted.
Verification of Xenium and PhenoCycler-Fusion markers
During assay validation, tumor and tissue types were analyzed as individual ROIs for bulk expression of each marker individually and checked to understand if there were underperforming markers. Individual ROIs used in the assay validation from the reference TMA and marrows include breast cancer (BRC), colorectal cancer(CRC), melanoma, hepatocellular carcinoma (HCC), non-small cell lung cancer (NSCLC), Merkel cell carcinoma (MCC), carcinoid tumor, tonsil, AML, CML, and MM for Xenium (n = 11) and BRC, CRC, melanoma, HCC, NSCLC, MCC, carcinoid tumor, and tonsil for PhenoCycler (n = 8). The MFI was calculated for each marker at the individual ROIs based on single-cell mean intensity PhenoCycler and single-cell mean transcript Xenium data provided by QuPath and Xenium Explorer software, respectively. Each marker's expression profile in the tumor and tissue types were compared to expected expression and classified as one of three results: Yes = matches the expected annotation from manufacture, published literature, or public database on NCBI/protein atlas, or per pathologist assessment of the morphology and the included cell type composition; No = does not match expected annotation; Maybe = when one or more tissue/tumor types have insufficient data to support the expression profile. Figure 3(a) shows our quality review where we inspected each of 377 genes in the Xenium panel and matched the observed expression profile to the expected annotation for each gene. 79.6% of the Xenium panel markers performed as per the expected annotations, whereas 6.4% of the markers did not perform as expected. 14.1% of markers were classified as Maybe because these markers showed expression of some tumor/tissue typesas expected, but some tumors did not have enough published literature to support expression information for that marker. Figure 3(b) shows similar data for PhenoCycler-Fusion, which demonstrated a higher percentage of markers did not perform as expected (21.4%). In contrast to Xenium data, a PhenoCycler-Fusion marker may be classified as Maybe due to lack of published literature for the tumor/tissue type, lack of expected positive staining cells, or if there were artifacts in the tissue obscuringproper staining (e.g., necrosis or foreign object). 62.5% of the markers passed quality assessment (resulted as Yes), thoughsome tissues contained artifacts such as tissue tearing, antibody/reagent trapping, or non-specific reactivity with necrosis. When there was artifact that impacted the MFI of a particular marker but did not appear to negate staining upon visual review, the markers were passed as acceptable. Detailed data shown in Supplemental Tables 3 and 4 and Supplemental Figure 2.
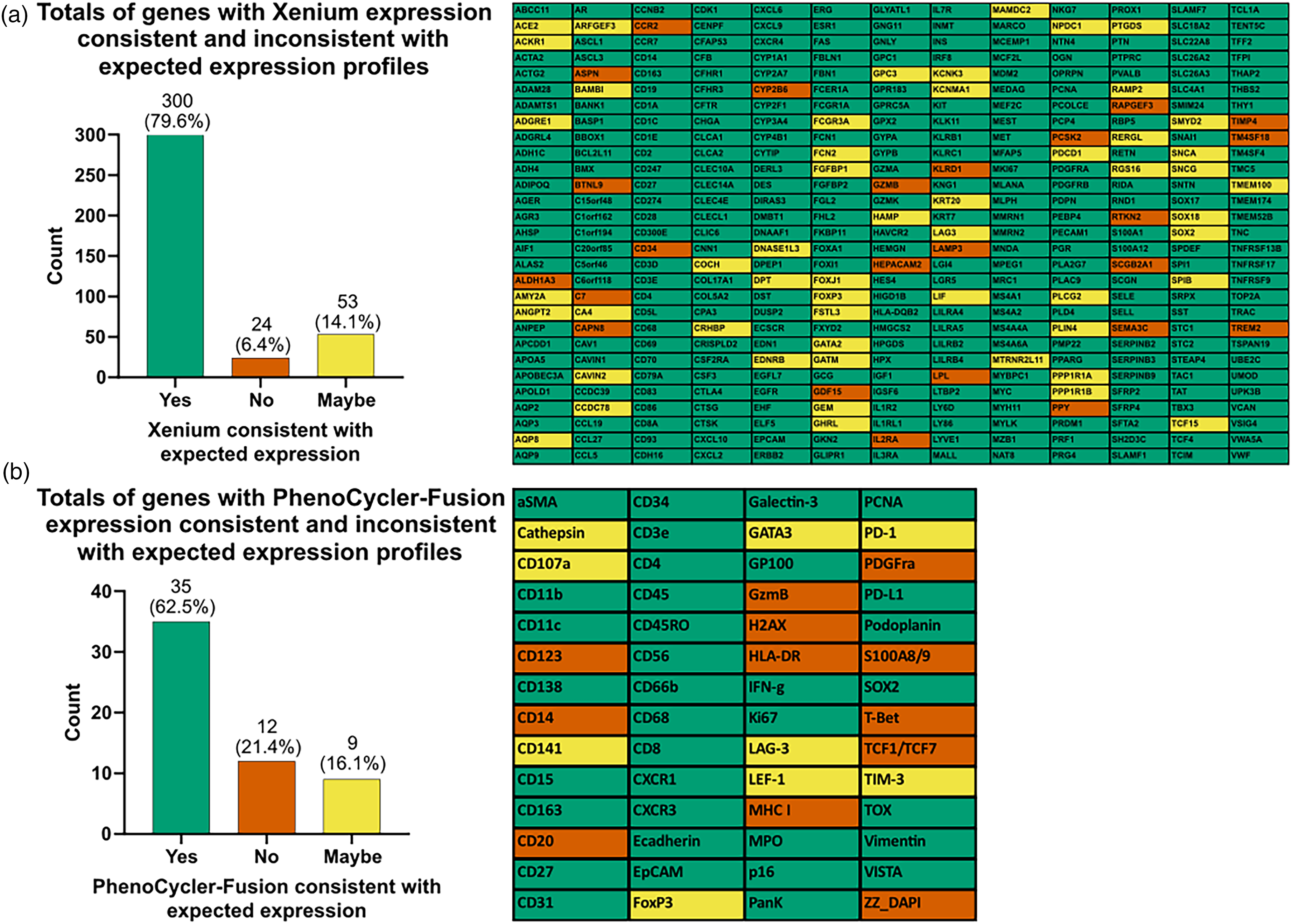
Xenium and PhenoCycler marker expression consistency with expected expression profiles. (a) Quality review data for each of 377 genes in the Xenium panel where we matched with the observed expression profile to the expected profiles. 79.6% of the Xenium panel markers performed as per the expected annotations. (b) Quality review data for each of 56 markers in the PhenoCycler panel where we matched the observed expression profile to the expected profiles. 62.5% of the PhenoCycler panel markers performed as per the expected annotations.
Orthogonal confirmation of spatial profiles between PhenoCycler-Fusion and Xenium platforms
By performing both spatial proteomics and spatial transcriptomics on the same reference TMA, we had comparable results that could be used to cross verify the platforms. Two UMAP analyses were performed. One on the replicate reference TMA slides for PhenoCycler and one for Xenium with a combination of the marrows slides and well as both replicate reference TMAs. UMAPs of the Xenium platform and the PhenoCycler platform were generated by performing cell segmentation and unsupervised clustering which generated unsupervised cell clusters based on cluster gene expression profiles. Cell-type annotations were applied and UMAPs were compared for the ability to identify immune cells versus specific cancer cell types (Figure 4). Because Xenium has more markers, a more refined ability to identify specific immune subsets was possible, however both platforms were capable of clearly separating the different tumor types from immune cells in primary quality assessment. Figure 5(a) shows the orthogonal comparison by linear regression between 25 markers which are shared between the PhenoCycler and Xenium panels, where a subset of markers shows good correlation (e.g., Spearman's Rank correlation coefficient for Ki67 = 0.95, CD4 = 0.98, EpCAM = 0.9, Figure 5). ROIs, which demonstrated the aforementioned tissue quality issues, showed a significant impact on the performance of their markers. This is demonstrated through an orthogonal comparison of the specific tissues individually, shown in Figure 5(b) (e.g., Spearman's Rank correlation coefficient for CRC = 0.89, Lung CA = 0.38).
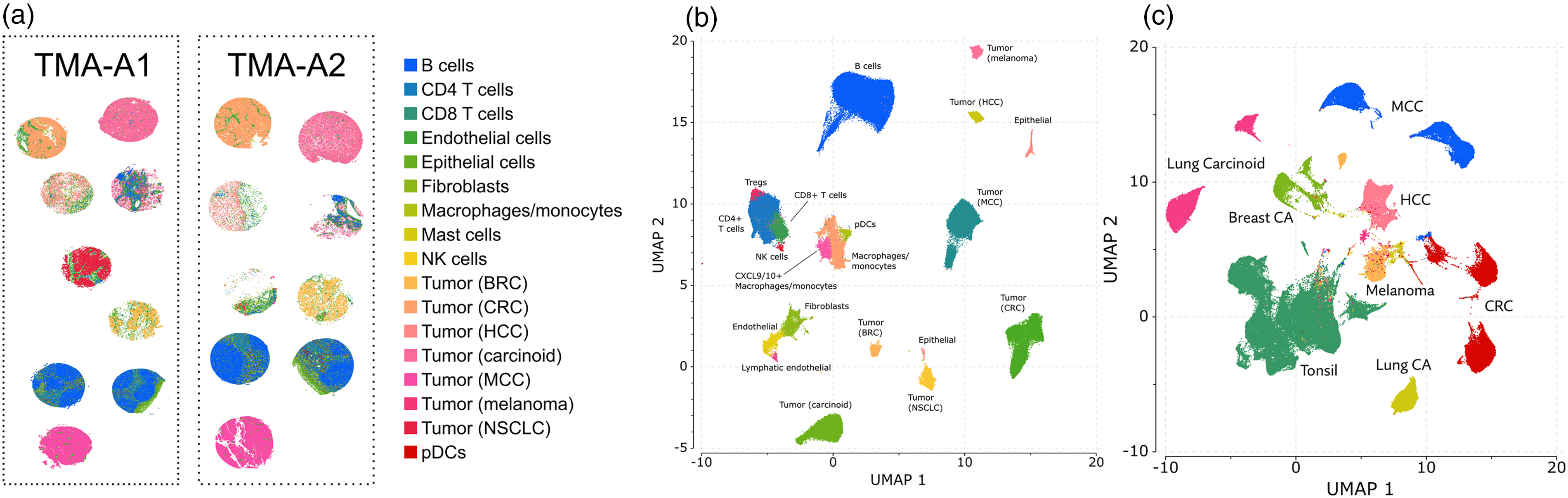
Xenium Proseg cell segmentation and clustering and uniform manifold approximation and projection (UMAP) of Xenium vs PhenoCycler data. (a) Xenium cell segmentation and clustering of cell types with Proseg. Each method affirms these cores are mostly pure tumor, and that each tumor is transcriptionally distinct. (b) UMAP of the Xenium platform analyzed by Proseg vs (c) UMAP of PhenoCycler data analyzed by QuPath of all tumor types show how different tumors separate vs immune cells.
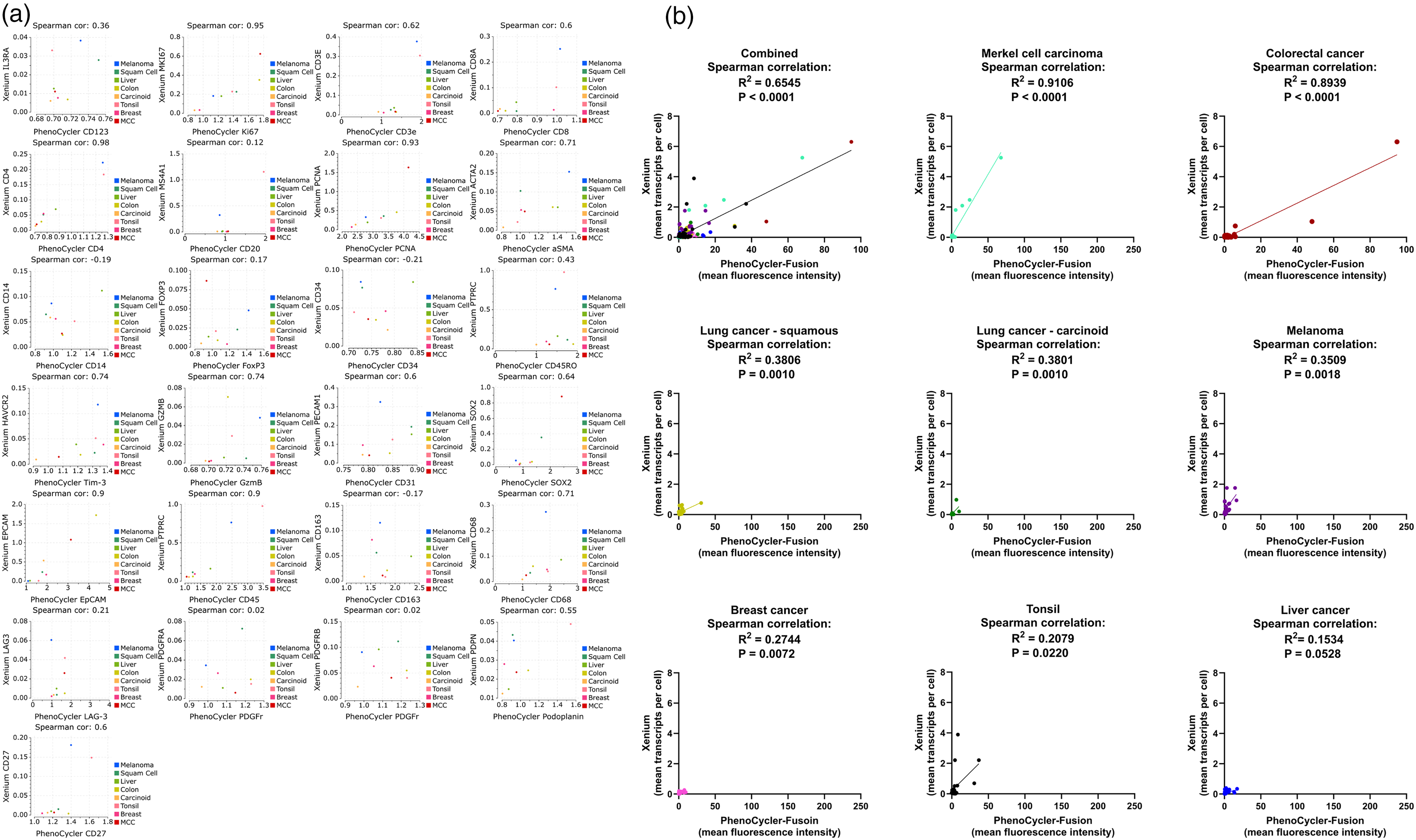
Correlation of screening results between PhenoCycler-Fusion and Xenium platforms. (a) Results faceted by protein/transcript target. (b) Results faceted by tissue. Overall correlation was observed, but with inconsistent significance among tissues.
Spatial transcriptomics analysis of bone marrow samples
After selecting EDTA/sucrose as a method for bone marrow biopsy decalcification, a single CML bone marrow biopsy sample was run in parallel with two bone marrow aspirate (particle-prep) samples: one from an AML patient, and one from an MM patient. After performing similar data preprocessing as described above, the Proseg segmentation algorithm was used to identify single cells from each of these Xenium analysis samples. UMAP and clustering analysis were used to identify major cell populations (Figure 6(a)), including abundant AML tumor cells present in the AML particle-prep sample. Images were then constructed by coloring each cell by the corresponding cell cluster (Figure 6(b)–(e)). These images show good preservation of bone marrow architecture and cellularity in the CML bone marrow biopsy. Note that this sample was obtained from a post-treatment CML patient with very low frequencies of blasts (0.06% confirmed by clinical flow cytometry) detected in contrast to the AML particle-prep sample which contained a high percentage of AML leukemic blasts (as shown in Figure 6(e)). Limited comparative data could be performed as the CML bone marrow was stained with a small panel by the PhenoCycler-Fusion to check the quality of the tissue as well as preservation of the markers (Figure 2(a), markers listed in the figure). The AML and MM marrows did not undergo PhenoCycler testing.
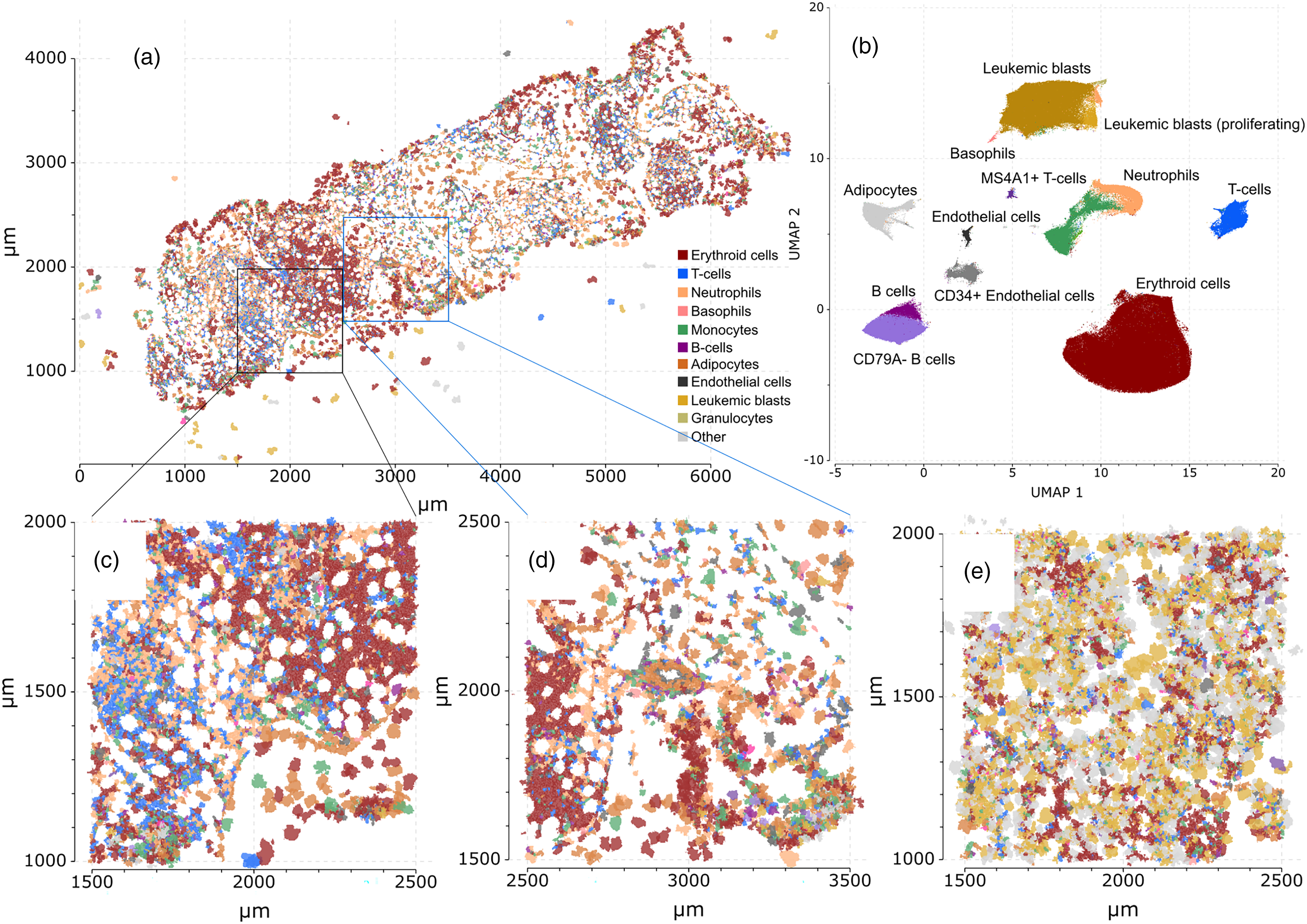
Bone marrow analysis with Xenium (a) UMAP plot of all cells segmented using Proseg from one CML bone marrow biopsy, one AML bone marrow aspirate and one multiple myeloma bone marrow aspirate sample run in parallel using Xenium. (b) Shows an enlarged area of the Xenium image where there is a T-cell rich region admixed with erythroid and maturing neutrophils. (c) is a contrasting region centered around large arteriole and shows a diverse immune population with mixed monocytes, neutrophils, andT cells.(d) shows a combined UMAP with cells represented from the CML, AML and MM marrow samples and thus shows a large blast population (corresponding to the AML sample), and a large CD79a population (corresponding to the MM sample). (e) shows the Xenium Proseg image of the AML sample with high blast percentage.
Spatial proteomics analysis of solid tumor samples
It was noted that the immune cells from the tonsil and the tumor clustered together on UMAP analysis while the separate solid tumor types separated as expected. However, despite normalization on normal tonsil controls, there was an observable batch effect between the duplicate TMA tumor cores on TMA 1 and TMA 2 (Figure 4(c)). The PhenoCycler data clustered differently by both tissue type and TMA image whereas the Xenium transcripts were more consistent. The PhenoCycler TMA slides were processed and stained in parallel and run concurrently, since the platform is limited to running one slide at a time. Although some of the PhenoCycler data differences could be attributed to batch effects during the acquisition runs, proteins are more variable than transcripts by their very nature. Cells with the same transcripts can have different protein expressions due to factors such as post-transcriptional modifications, translation efficiency, and feedback loops. Each TMA tissue core was pulled from a different area of the original samples, and these differing tissue microenvironment neighborhoods can have different ecosystems resulting in different protein expressions. The separated clusters serve as a reminder of the heterogeneity of these tissues.
Discussion
Spatial proteomics and transcriptomics analysis of cancer tissues have demonstrated great utility in several different aspects: demonstrating cancer cell to immune cell interactions which improves our understanding of cancer biology, paired pre/post treatment tissue analysis have demonstrated specific immune cell trafficking patterns which can serve as predictive biomarkers for treatment efficacy,7,30 and spatial analysis of tumor tissues can be prognostic providing tumor heterogeneity scores 6 in addition to immune composition. 31 After repetitive verifications of the mIHC, PhenoCycler-Keyence (aka CODEX), and PhenoCycler-Fusion platforms, we felt a need to develop a systemic pathway for instrument and assay verification. This study's objectives were two-fold: first, to describe a systemic pathway for instrument and assay verification with a multi-cancer reference TMA; second, to develop an approach to decalcify specimens that would allow for preservation of nucleic acid for spatial transcriptomics and preservation of protein for spatial proteomics data generation. Using new computational methods Maxspin 32 and Proseg 28 developed in the Newell Lab, we leverage the spatial transcriptomics data to show efficient classification of tumors and the identification of spatially varying genes in specific cell clusters. Furthermore, these tools were extended to the PhenoCycler data to demonstrate similar analysis could be performed and generate comparative data. With this workflow, a spatial quality assurance program can be established for both platforms which can cross validate.
As with any platform development, pre-analytical sample variables play a significant role in the success of the data generated. Our efforts have developed a molecular-friendly protocol that works for preserving RNA transcripts for spatial transcriptomics and preserves protein for spatial proteomics. As noted by numerous previous publications,33,34 decalcification protocols typically use powerful acids to remove calcium to soften bone to allow for downstream cutting of the specimen into very thin sections (2–4 µm thick sections for morphologic examination).35,36 We show that our optimized protocol with EDTA/sucrose solution consistently produced greater yields of nuclei acid, which corresponds with other independent groups' who have used EDTA-based decalcification protocols.21,22,25,37–39 Many have used EDTA alone or EDTA with another DNA stabilizing agent; our lab has chosen to use sucrose, which was previously described as protective osmolyte sugar and shown to have stabilizing effects on protein nucleic acid interactions. 40 The combination of EDTA and sucrose in the decalcification protocol has proven to preserve both protein and mRNA as shown in our Xenium and PhenoCycler data.
Our preliminary comparative work on PhenoCycler-Fusion and Xenium shows that both platforms independently perform well on various tumor types, from solid tumors to heme malignancies. Furthermore, between the PhenoCycler and Xenium panels, we were able to use overlapping markers to orthogonally cross validate the platforms, but we could only show correlation between RNA and protein expression profiles of some makers and only in some tissues. We created multi-tissue/cancer TMAs to serve as reference material for developing new assays, verifying platform performance, and performing cross-platform validations and have learned many lessons through this study. Although we systematically reviewed and selected areas with good tumor admixed with immune filtrates, there were still spots of necrosis and collections of ghost cells and/or red blood cells causing artifactual staining by PhenoCycler-Fusion and by Xenium causes some irregular cell clustering. The artifactual staining on the PhenoCycler-Fusion assay was likely due to tissue autofluorescence and staining artifacts generated by the unspecific binding of some barcoded fluorophores which are negative or nominal in most tissues but can be problematic in necrotic or lower quality samples. 41 The melanoma core we selected showed a good mix of tumor cells with immune cells, but the tissue was lost during sectioning onto one of the two PhenoCycler-Fusion TMA slides. While we thought inclusion of uncommon tumors such as carcinoid tumor and Merkel cell carcinoma would help to provide contrast, the lack of expression data for the rare tumors proved to be a challenge. However, once expression has been established across multiple collaborating labs, this could negate the lack of known expression data. We included two tonsil cores as controls, one with epithelium and one with more germinal center, which may be why they did not align as well when we combined the data. Consideration for a multicancer reference TMA should encompass tumors and a tonsil control tissue so that there is a good mixture of tumor cells and immune cells, a known gene expression profile or have collaborating laboratories to develop a consensus “expected” annotation, and a lack of necrosis/ghost cells/pools of red blood cells.
This study was devised as proof-of-concept and development of a quality assurance program for our spatial proteomics and transcriptomics workflows. This type of study includes inherent limitations including a small number of cases which were used to create the reference tissue microarray as the intent of this TMA was to be placed onto the slide alongside primary testing tissue to ensure reproducible performance in every run. Another limitation is the inability to have large numbers of reproducibility runs as these are extremely expensive assays, but we do have limited reproducibility data as two reference TMAs were created with the same patient set and demonstrate comparable data. The intent of this manuscript is to describe a proposal for a potential quality assurance program encompassing samples and biomarkers centered around the use of reference TMAs and additional control decal marrow tissues.
Conclusions
We have demonstrated that both the PhenoCycler-Fusion and Xenium platforms work well on various tumor types including decalcified marrow cores following the molecular- and protein-friendly decalcification protocol (EDTA /sucrose). Our data shows the EDTA/sucrose decalcification protocol is compatible with Xenium spatial transcriptomics and preserves protein for PhenoCycler high-plex spatial proteomics. We demonstrate initial proof-of-concept work on the development of a reference TMA that can be used to validate spatial platforms/assays and share the lessons learned during this process on how we will improve in the future. Finally, we show how our laboratory performs systematic quality assessment of both the spatial proteomic and spatial transcriptomics data and how these platforms can serve as orthogonal confirmation for each other.
Supplemental Material
sj-xlsx-1-cbm-10.1177_18758592241308757 - Supplemental material for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow
Supplemental material, sj-xlsx-1-cbm-10.1177_18758592241308757 for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow by Cecilia CS Yeung, Daniel C Jones, David W. Woolston, Brandon Seaton, Elizabeth Lawless Donato, Minggang Lin, Coral Backman, Vivian Oehler, Kristin L Robinson, Kristen Shimp, Rima Kulikauskas, Annalyssa N Long, David Sowerby, Anna E Elz, Kimberly S Smythe and Evan W Newell in Cancer Biomarkers
Supplemental Material
sj-xlsx-2-cbm-10.1177_18758592241308757 - Supplemental material for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow
Supplemental material, sj-xlsx-2-cbm-10.1177_18758592241308757 for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow by Cecilia CS Yeung, Daniel C Jones, David W. Woolston, Brandon Seaton, Elizabeth Lawless Donato, Minggang Lin, Coral Backman, Vivian Oehler, Kristin L Robinson, Kristen Shimp, Rima Kulikauskas, Annalyssa N Long, David Sowerby, Anna E Elz, Kimberly S Smythe and Evan W Newell in Cancer Biomarkers
Supplemental Material
sj-xlsx-3-cbm-10.1177_18758592241308757 - Supplemental material for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow
Supplemental material, sj-xlsx-3-cbm-10.1177_18758592241308757 for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow by Cecilia CS Yeung, Daniel C Jones, David W. Woolston, Brandon Seaton, Elizabeth Lawless Donato, Minggang Lin, Coral Backman, Vivian Oehler, Kristin L Robinson, Kristen Shimp, Rima Kulikauskas, Annalyssa N Long, David Sowerby, Anna E Elz, Kimberly S Smythe and Evan W Newell in Cancer Biomarkers
Supplemental Material
sj-xlsx-4-cbm-10.1177_18758592241308757 - Supplemental material for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow
Supplemental material, sj-xlsx-4-cbm-10.1177_18758592241308757 for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow by Cecilia CS Yeung, Daniel C Jones, David W. Woolston, Brandon Seaton, Elizabeth Lawless Donato, Minggang Lin, Coral Backman, Vivian Oehler, Kristin L Robinson, Kristen Shimp, Rima Kulikauskas, Annalyssa N Long, David Sowerby, Anna E Elz, Kimberly S Smythe and Evan W Newell in Cancer Biomarkers
Supplemental Material
sj-xlsx-5-cbm-10.1177_18758592241308757 - Supplemental material for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow
Supplemental material, sj-xlsx-5-cbm-10.1177_18758592241308757 for Spatial proteomics and transcriptomics characterization of tissue and multiple cancer types including decalcified marrow by Cecilia CS Yeung, Daniel C Jones, David W. Woolston, Brandon Seaton, Elizabeth Lawless Donato, Minggang Lin, Coral Backman, Vivian Oehler, Kristin L Robinson, Kristen Shimp, Rima Kulikauskas, Annalyssa N Long, David Sowerby, Anna E Elz, Kimberly S Smythe and Evan W Newell in Cancer Biomarkers
Footnotes
Author contributions
For every author, his or her contribution to the manuscript needs to be provided using the following categories:
Conception: CY, EN
Interpretation or analysis of data: All
Preparation of the manuscript: All
Revision for important intellectual content: All
Supervision: CY, EN
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by funding from the National Institutes of Health (NIH), National Cancer Institute (NCI) U2C CA271902, NIH, NCI R01 CA264646, NIH, NCI P01 CA225517, the NIH Human Immunology Project Consortium U19AI128914, and the Immunotherapy Integrated Research Center of the Fred Hutchinson Cancer Center.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
This is an active project with ongoing enrollment of subjects for the study, thus the data will be made publicly available after the study has ended. However, data presented in this study will be made available upon request to the corresponding author, please contact cyeung@fredhutch.org.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
