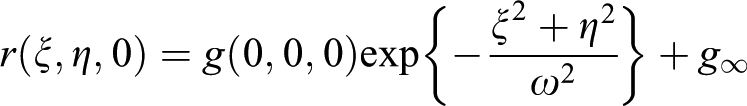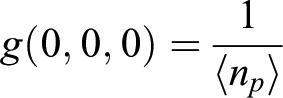Abstract
Liposome–microbubble conjugates are considered as better targeted drug delivery vehicles compared to microbubbles alone. The microbubble in the integrated drug delivery system delivers the drug intracellularly on the target, whereas the liposome component allows loading of high drug dose and extravasation through leaky vasculature. In this work, a new high yielding microbubble production method was used to prepare microbubbles for formulation of the liposome-conjugated drug delivery system. In formulation process, the prepared liposome of 200 nm diameter was attached to the microbubble surface using the avidin–biotin interaction. The analysis of the confocal scanning laser microscope images showed that approximately 8 × 108 microbubbles per millilitre (range: 2–7 μm, mean size 5 ± 0.5 μm) can be efficiently conjugated to the liposomes. The method of conjugation was found to be effective in attaching liposome to microbubbles.
Keywords
Introduction
Currently, commercially available site-specific drug delivery techniques are not efficient in delivering the drug at the target causing adverse reaction to the surrounding healthy tissue. 1 Certain drug delivery carriers like liposomes are known to reduce non-selective exposure of drug to the healthy tissue after systemic administration, and thereby reducing the side effects. However, the slow release of drug from these vehicles limits the drug bioavailability in tumours. 2 One of the strategies of increasing bioavailability of the drug at the target is to attach the tumour-targeting ligands to liposome. Adopting this strategy increases the local concentration of the liposomes in the tumour tissue, but its targeting efficiency is limited by the saturation of targetable receptors. 1 Additionally, the non-specific presence of these tumour receptors in heathy tissue can also result in toxicity due to accumulation of the tumour-targeted particles. Some of the shortcomings of the biochemical method have been overcome by an ultrasound drug delivery system, where the drug is delivered to the target by focussing ultrasound on the pressure-sensitive carriers accumulated at the target site. 1,2 The lipid- or albumin-shelled microbubbles and liposomes are generally used as a pressure-sensitive drug carriers because they are established contrast agents for ultrasound imaging and echogenic at safe exposure levels. 3 When these pressure-sensitive carriers are exposed to acoustic radiation force, they cavitate vigorously to eject the microstream of fluid, which generates transient pore in the membrane of the cells and delivers the drug into the cell. Because of the difference in ultrasound impedance of water and high molecular weight gas core in the microbubble, the acoustic radiation force causes nonlinear compression and decompression, which is responsible for microstreaming and high backscatter useful in ultrasound imaging. These properties of microbubbles allow combination of molecular ultrasound imaging and therapeutics on the same platform. 4 Even though microbubbles are versatile platform for the drug delivery, they have low retention of hydrophobic drug in the large lipid fragments and very short circulation times in vivo. 4
Liposomes are clinically established efficient drug carriers as opposed to microbubbles. They are composed of a lipid bilayer shell that can trap hydrophobic and amphipathic drugs in it and can encapsulate hydrophilic drugs in the aqueous core. Liposomes concentrate drug at the tumour site by 50- to 100-fold as compared to free drug injections. 4 Liposomes can be tailored to an optimized diameter for maximum extravasation into tumour vasculature and fusion with cell membranes. As opposed to microbubbles, the liposomes are weakly echogenic and not readily manipulated with ultrasound radiation force. 1 If any drug delivery carriers are spatially concentrated at the target using ultrasound radiation force, they can achieve better delivery rates in comparison to blood flow–assisted delivery technique alone. 5 Separate targeting of drug-loaded liposomes and microbubbles to the same tissue is unlikely to achieve sufficient proximity and concentration in the tissue for the cavitation shockwave to effectively disrupt the membranes and deliver therapeutic dose. 2 Therefore, the liposome–microbubble conjugates would provide advantages of liposomes such as a superior drug payload capacity, functionalization with targeting moieties, combined delivery of therapeutic/diagnostic molecules and an ability to confine spatially in response to ultrasound radiation force. 5 The microbubble–liposome conjugates having compressible gas core can also serve as ultrasound contrast agents, 5 making it complete theranostic vehicle that combines drug delivery, targeting and molecular imaging on a single platform.
Various techniques are used to prepare the liposome–microbubble conjugates. First, the polyethylene glycol (PEG) chain derivatives are incorporated into the shell of microbubbles to pose steric hindrance against coalescence and adsorption of macromolecules to the microbubble surface and improve in vivo circulation persistence through reduction in opsonization. 6 Microbubbles, in the past, have been conjugated to liposomes by post-labelling and incorporating functionalized lipids into the microbubble shell, which allowed conjugation of the liposomes to the microbubble monolayer surface through a bioconjugate. Post-labelling increased loading efficiency by increasing the conjugation of multiple liposomes to a microbubble. The use of the covalent conjugation of liposomes to the microbubble surface can raise the critical issue of liposomes release from the bioconjugate. However, the ultrasound cavitation caused via focussing of the acoustic radiation on the microbubbles opens the cell membranes transiently and releases bound liposomes from the conjugates, permitting them to absorbed within the cells. 7
In this work, the Biotinylated liposomes and microbubbles each were prepared using 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[biotinyl(polyethylene glycol)2000] (DSPE-PEG2kBiotin) as a linker to bind avidin by a previously described method, 5 and Cholesterol is attached to fluorescent dye 7-nitrobenzofurazan (NBD-cholesterol) was used as a fluorophore to investigate the binding efficiency of liposomes to the microbubbles in the hybrid vehicle. Detailed optimization of liposomal formulation and design, for example, addition of negatively charged lipids, targeting ligand, etc., is beyond the scope of this study.
In order to demonstrate the efficiency of microbubble–liposome conjugation and construction of the vehicle, the NBD-cholesterol–labelled lipid was used in the bilayer of the liposomes as a flourophore. 8 The emission intensity from the fluorophore NBD-cholesterol in liposomes was evaluated using the confocal scanning laser microscopy (CSLM), which allowed determining the conjugation efficiency. Furthermore, the formation of the microbubble–liposome construct was studied by labelling lipid on the surface of the microbubbles with the lipid-soluble Sudan IV (red) dye. Using the CSLM, the red coloured microbubbles decorated with numerous green coloured NBD-labelled liposomes on the microbubble surface are evaluated and simultaneously observed.
Materials and methods
Materials
1,2 Distearoyl-sn-glycero-3-phosphatidylcholine (DSPC), 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC), 1,2 distearoyl-sn-glycero-3-phosphoethanolamine-N-[methoxy(polyethylene glycol)-2000] (DSPE-PEG2k), DSPE-PEG2kBiotin and 25-[N-[(7-nitro-2 -1,3-benzoxadiazol-4-yl)methyl]amino]-27-norcholesterol (25-NBD-cholesterol) are purchased from Avanti Polar Lipids Inc. (Alabaster, Alabama, USA). Neutravidin is obtained from Pierce Biotechnology (Rockford, Illinois, USA). All other reagents are of analytical grade and are used as received.
Methods
Microbubble–liposome construct preparation
Biotinylated microbubbles
Many successful methods of preparing microbubbles are now established in research domain. Mainly electrospraying, 9 microfluidics 10 and flow focusing 11 are used frequently for research purpose. The bespoke device used for preparing microbubbles for this work is described elsewhere. 12 The device consisted of a pair of concentric capillaries exiting downwards into a cylindrical Perspex tube chamber that was pressurized using compressed air. The outer steel capillary carrying liquid phase and the inner capillary supplying the core gas were forced out of the orifice of 100 μm. The resulting spray contained microbubbles of required diameter. Resulting size dispersion of the microbubbles is shown in Figure 3.
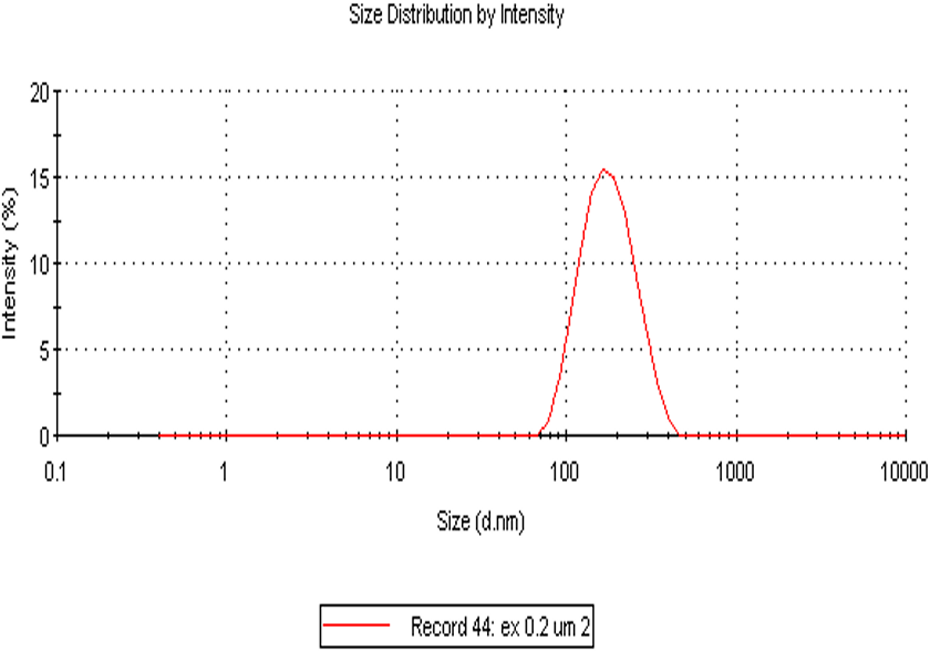
Size distribution of extruded liposomes (average diameter 204 ± 1.559 nm).
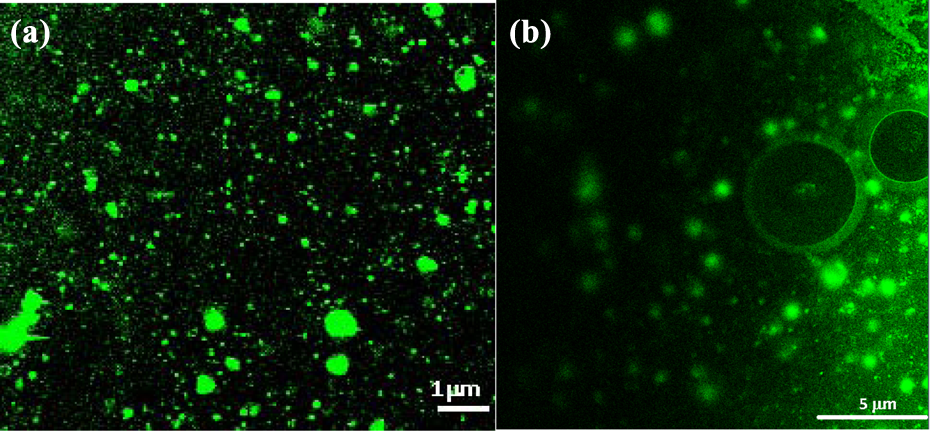

Confocal fluorescence images of (a) liposomes encapsulating 5% NBD with an average diameter of 200 nm (×100 oil; scale bar: 1 μm). (b) NBD-stained free liposomes and microbubbles (average diameter 5 µm) with NBD-labelled liposomes concentrated on their surface (×100 oil; scale bar: 5 μm).
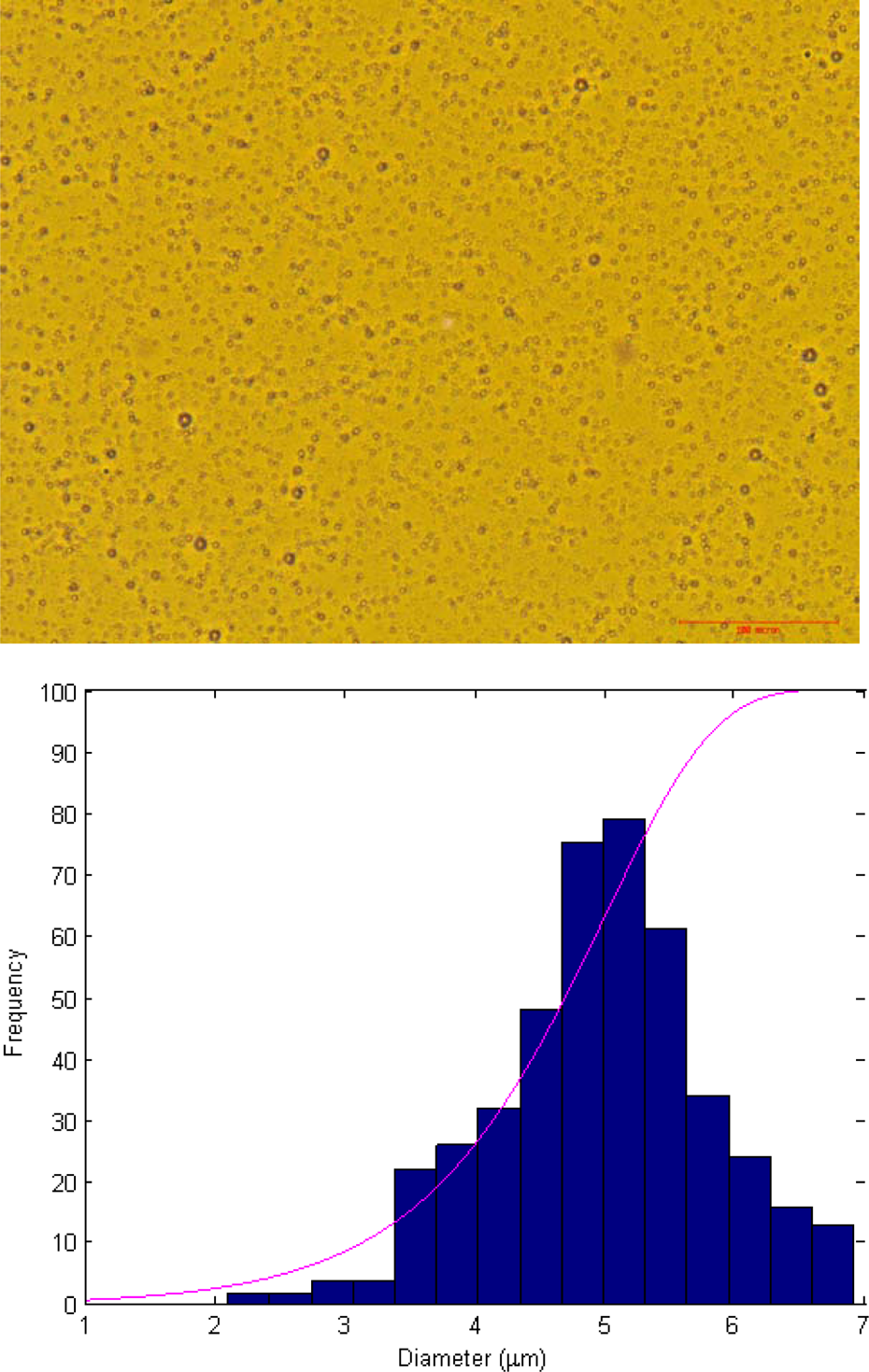
A representative micrograph showing phospholipid shelled microbubbles with an average diameter of 5 µm (scale bar: 100 µm). A size distribution histogram of the representative microbubble population (bottom).
To prepare microbubbles, the lipids DSPC, DSPE-PEG2 k, DSPE-PEG2kBiotin and Sudan IV (red) dye (89.95:5:5:0.05, molar ratio) were dissolved in a round bottom flask using chloroform. After complete dissolution of the lipid, the chloroform was removed under constant flow of nitrogen, which was then followed by evaporation under vacuum in a rotary evaporator (Buchi, Flawil, Switzerland) for 2 h to yield a thin lipid film at the bottom of the round flask. Distilled water:glycerol:propyleneglycol in the volume ratio of 80:10:10 was added to the dried thin lipid film to prepare a suspension with a lipid concentration of 1 mM (1 mg/mL). The resulting lipid suspension was mixed thoroughly using a bath sonicator maintained just above the lipid phase transition temperature of 60°C. Resulting clear lipid suspension was used to prepare microbubbles.
First, the lipid suspension was fed to the device using a syringe pump (Harvard PHD-4400, Hilston, MA, USA), while the air (BOC Chemicals, UK) and core gas – perfluorobutane (F2 Chemicals, Preston, UK) connected to the device via flexible High performance liquid chromatography (HPLC) tubing were also supplied simultaneously. The lipid suspension flow was maintained at 2.4 × 10−5 m3/s, while the air pressure of 200 kPa and the core gas pressure of 150 kPa were maintained for microbubble preparation. Flow of the core gas and formation of the liquid-gas cone was confirmed and monitored by a real-time high-speed camera (pco. 1600; PCO AG, Kelheim, Germany) imaging. 12 The high-speed camera was focused on the Perspex wall of the chamber. The multiphase gas in liquid cone forms in the gap formed between the end of capillaries and orifice in the plate.
Bubbles were collected as they emerged from the aperture for immediate examination by microscope or in a vial stored in ice for future use. The mean size of the microbubbles was determined from sizing 500 microbubbles in the optical microscope (inverted microscope (IX71, Olympus, Tokyo)) images using Infinity Analyze Program (Hatfield, PA, USA [Version 6.0.0]). Quantification of microbubble concentration was accomplished using haemocytometer and the average of triplicate measurements was taken into account for further calculations.
Biotinylated liposomes
Liposomes have been prepared using the thin film hydration procedure and NBD dye–labelled cholesterol. A multilamellar lipid solution consisting of DPPC:DSPE-PEG2kBiotin:25-NBD-cholesterol (90:5:5) was prepared, as described in section ‘Biotinylated microbubbles’. However, the final concentration of the lipid in distilled water was maintained at 12 mM (10 mg/mL). Obtained solution was then extruded through two stacked polycarbonate membranes in a mini extruder (Avanti Polar Lipids Inc., Vancouver, BC) having a pore diameter of 200 nm. During the entire extrusion process, the temperature of the lipid mixture was maintained above its phase transition temperature. The physical stability of the liposomes in solution was checked at 4°C, room temperature and at 50°C by determining the changes in vesicle size over a period of 2 months by dynamic light scattering (DLS) and average of triplicate measurements is taken.
Conjugation of biotinylated liposomes to biotinylated microbubbles via avidin
Approximately 8 × 108 microbubbles per millilitre were washed with 3 mL of air-saturated distilled water and stored in a 5-mL inverted syringe. The syringe was then transferred to a bucket centrifuge to allow centrifugation of the suspension at 300 × g for 3 min to remove the free lipids and dye from the suspension. The centrifugation process forced the bubbles to float at the top of the syringe and allowed the removal of the unincorporated lipids from the bottom of the syringe. Afterwards, the avidin solution with a concentration of 0.022 mg/mL was added to the washed microbubbles before mixing well and incubating for 15 min at a room temperature. During the incubation, the mixture was shaken gently. The avidin-bound biotinylated microbubbles were then separated from the free avidin by washing them four times using air-saturated distilled water. To conjugate the liposome to the microbubble surface, 1 mM of the extruded liposomes (200 nm size) was added to this suspension and incubated for 10 min at room temperature with gentle shaking. Microbubbles carrying liposomes were washed three to four additional times to remove any free liposomes.
Binding quantification
The relative level of liposomes bound to microbubbles can be determined by using image correlation spectroscopy (ICS). This method has been applied to cell quantification and drug release quantification via high-speed or Raman imaging. 13 –15 For ICS, the images of liposomes and liposome-bound microbubbles were obtained using confocal microscope (LSM-5; Zeiss, Thornwood, New York, USA) interfaced with an argon–krypton laser (with lines of 405, 488, 555 and 633 nm). For confocal microscopy, an aliquot of 200 μL of liposome-bearing microbubbles at a concentration of 8 × 108 per millilitre was placed on the slide and covered with a coverslip. The coverslip was fixed at the four corners using nail paint to retain the solution of the microbubble-conjugated liposome. Subsequently, the laser beam emitting from a 60× oil immersion objective (NA = 1.3) was focussed on to the microbubble–liposome conjugate via a double dichroic mirror. This focussed laser beam passing though the 488-nm bandpass filter excited the NBD and Sudan Red III simultaneously, which allowed capturing six Z-stacks of images through dual channel. Each Z-stack contained six images with a step size of 4 μm. The relative fluorescence intensity of Sudan Red III was detected at an excitation wavelength of 490 nm and emission wavelength of 512 nm. However, the fluorescent intensity of the liposome carrying NBD-cholesterol was measured at 460 nm excitation and 534 nm emission wavelength, respectively.
Image preprocessing
A correlation function for the obtained series of images was calculated using the Fourier method. As seen in Figure 4(b, c′), to remove streaks from image as seen Figure 4(b, a′), the image was de-convolved to remove the out of focus features. It was important to keep only those features in each image of Z stack, which does not belong to other slice of the volume image. This ensured that each image was related to correct quantity of the liposome. Before de-convolve operation, the PSF was generated using POINT spread function (PSF) generator and image acquisition data. The deconvolution was carried out by imagej plug-in. The non-homogenous features in the background were removed by subtracting mean of all images. Furthermore, all images were cropped using region of interest (ROI) manage in Imagej (Figure 5(b)) to keep important quantifiable microbubble features and watershed segmentation performed to separate overlapping circular features.
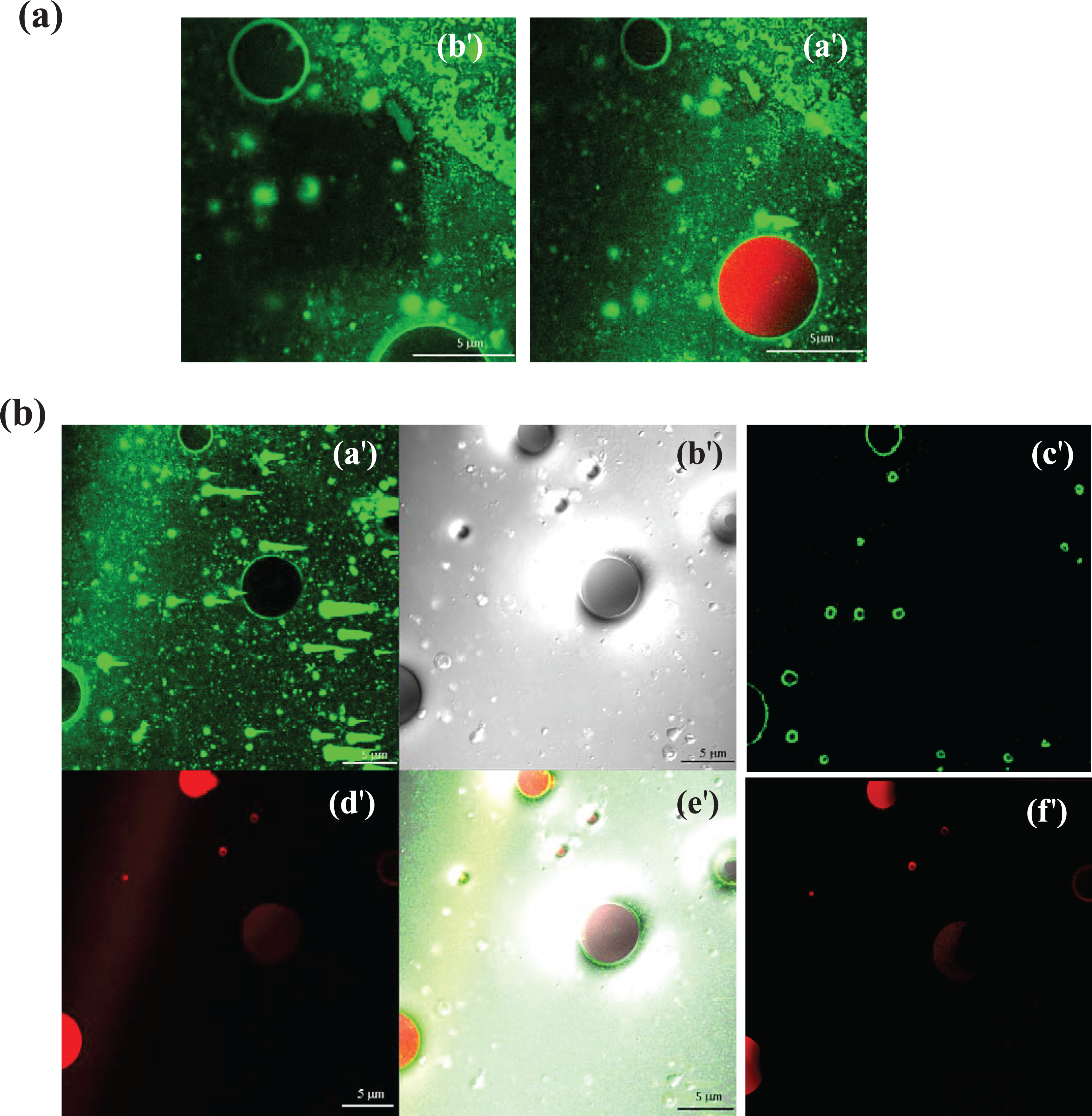
Confocal fluorescence image (a, a′ AQ: Per style, part labels “I” and “II” have been changed to “a” and “b” and the subparts “A–F” as “a′–f′.” Please check and approve the edits.) is a representative image of microbubbles (red) containing Sudan III dye and carrying liposomes loaded with 5% NBD-cholesterol on their surface. The image is taken using filters for both dyes (average diameter of liposome is 200 nm; scale bar: 5 μm). The image (a, b′) obtained using filter for NBD shows the microbubbles decorated with liposomes with 5% NBD-cholesterol (scale bar: 5 μm). (b) All confocal images display four horizontal planes with either simultaneous or individual monitoring of NBD and Sudan Red III (scale bar: 5 μm). The free liposomes attached to microbubbles and the microbubbles stained with Sudan III dye are seen in (a′) and (d′), respectively. The figure (b′) shows an optical micrograph of microbubbles and liposomes. The figure (e′) shows an optical micrograph of microbubbles with gas core (dark coloured) and tiny-sized liposomes with fluid filled core (light coloured). The image is the simultaneous visualization of both channels for NBD (liposomes) and Sudan red III labels (microbubbles). The (c′) image represents the de-convolved (a′) image, whereas image (f′) shows the de-convolved and filtered (d′) image. These images demonstrate the effect of image preprocessing on artefact removal from the original image.
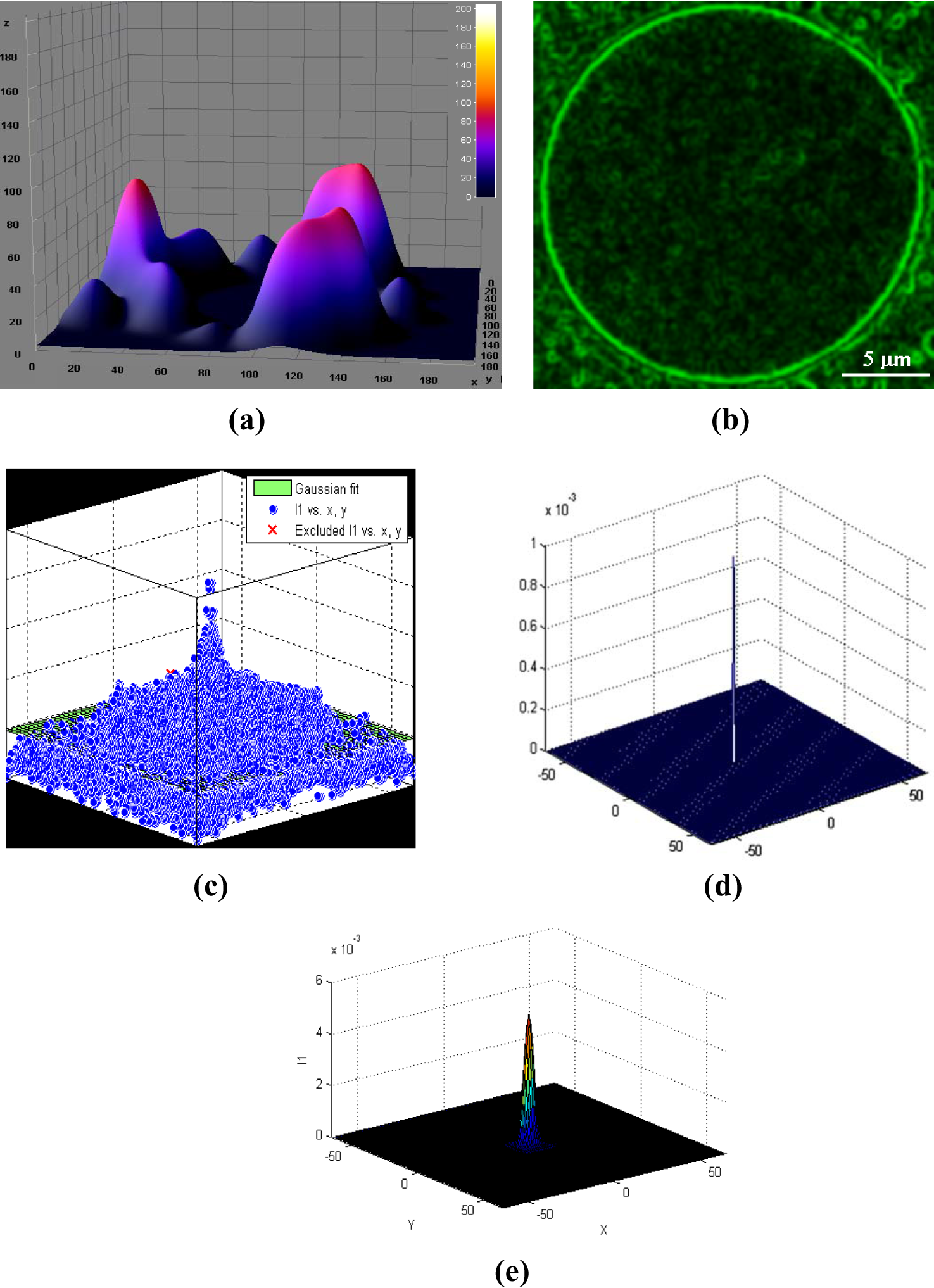
(a) All peaks surrounding the microbubble shows the fluorescent intensity peaks relating to liposome concentration. (b) An example image shows a single microbubble for further analysis. The out of focus lipid fluorescence was removed via preprocessing. (c) The fluorescent peaks related to each liposome-conjugated microbubble are identified and used to obtain the correlation peak. A correlation peak fitted with the Gaussian function is shown. All blue dots represent the raw correlation values corresponding to its spatial positions, whereas green surface represents the fitting function (d). The solitary peak shows the presence of background noise. (e) After removing noise, the Gaussian peak is fitted to obtain correlation peak to count liposomes.
In microbubble images, the liposomes were surrounding the bubbles. Thus, the fluorescent intensity peaks are seen in the circular fashion (Figure 5(a)). These peaks will provide uniform background and intensity, and therefore, each image area corresponding to individual microbubble was cropped to make 64 × 64 or 128 × 128 pixels image for further correlation peaks. Such images provided high resolution to resolve image for features useful for quantification.
The raw correlation data (Figure 5(c)) displays the raw normalized intensity fluctuations correlation function from the image. The correlation of the background image in relation to first background image determined the source of noise. The decaying profile shows the intensity profile from laser beam profile along with image features. The solitary peak in Figure 5(d) proved the presence of noise in the image. After preprocessing of each image, the correlation peak function was then fit to the 3D Gaussian function using the nonlinear least square method. The Gaussian 2D correlation fit function for nth number of image is given as
where g(0,0,0), ω and g ∞ are the amplitude of the intensity at the zero lag, the correlation radius and the offset. 16 The correlation radius, proportional to the laser-beam horizontal radius, was measured by focussing the laser beam on to the sharp edges. A microbubble–liposome conjugate system can be considered as a non-interacting system. In such case, the mean number of independent liposomes (np ) present in the correlation area under the laser focus can be found by its inverse relationship with zero-lag amplitude g(0,0,0). This can be described as
The peak as shown in Figure 5(e) shows the best fit Gaussian function to raw correlation data. The Gaussian function is not visible in Figure 5(a). The correct fit also confirms the removal of noise from the original image.
Results and discussion
Liposome size and fluorescence intensity
The biotinylated liposomes have been prepared and conjugated to microbubbles employing avidin–biotin adhesion as described by Kheirolomoom et al. 5 The average diameter of extruded liposomes was evaluated using zetasizer (Zetasizer Nano ZS, Malvern Instuments Ltd, Malvern, UK) and was found to be 204 ± 1.559 nm (Figure 1) with a polydispersity index of 0.27 ± 0.026. The size of liposomes after 2 months at 1.1 bar pressure and 4°C temperature remained unchanged (average diameter 187 ± 1.861 nm with a polydispersity index of 0.36 ± 0.010 nm). A minor peak at 5000 nm indicated slight aggregation in the liposomal sample. The liposomal fluorescence intensity was then evaluated using confocal fluorescence microscopy and the extent (numbers) of microbubble–liposome binding by ICS (Figure 2). For a constant concentration of NBD-cholesterol, the fluorescence intensity of the microbubble–liposome construct carrying liposomes with an average diameter of 200 nm was calibrated (Figure 2). The liposome and liposome–microbubble conjugate samples are prepared on glass slides as horizontal smears of both samples in solution. Since Figure 4(b) focusses on microbubbles (5–6 μm) decorated with several liposomes, free liposomes approximately 200 nm in size appear obscure. Hence, the view looks non-uniform.
Microbubble size and concentration
Quantification of microbubble counts was carried out in triplicate using a haemocytometer. The number of microbubbles per millilitre of solution was found to be approximately 8 × 108. Their initial diameters were determined using image analysis and was found to be in the range of 2–7 μm (mean size of 5 μm ± 0.5 μm standard deviation; Figure 3). Microbubbles have been prepared using PEG2000 and PEGkBiotin at concentrations optimized for stability. 5 The CSLM images (Figure 4) show insignificant change in the size distribution of the microbubble population. However a reduction in microbubble numbers was observed. Bubbles prepared using our device have shown appreciable stability.12 Considering the aforementioned observations, the drop in the microbubble count was the consequence of washings for removal of unincorporated lipids. The size of the microbubbles showed insignificant change in the images (Figure 4), which indicates no cross-linking between microbubbles after the conjugation procedure.
Liposome binding efficiency
Measurement of the mean fluorescence intensity of the liposomes in the resultant microbubble–liposome construct (Figure 4(b), c′) was used to estimate the relative number of liposomes bound to microbubbles and was measured using CSLM.
All six stacks were used to calculate the correlation function value of g(0,0,0). An example of such correlation peak for one of the representative images is shown in Figure 5(e). Some of the representative liposome-loaded microbubble images are shown in Figure 4(b), c′ and f′. In these images, the bubble appears red, while liposomes appear green. For image analysis, the images obtained via green channel were used to calculate the correlation peak. These images filtered out the red colour of the Sudan red and thus allowed calculation of the correlation peak related exclusively to the liposomes concentration. To determine the value of peak at zero lag, the Gaussian function was fit to the correlation peak (Figure 5(e)). Since g(0,0,0) values are inversely proportional to the number of liposomes, the number of liposomes bound to one microbubble was estimated using the inverse of the total fluorescent intensity. The values for the number of particles for the first three highest peaks were found to be 16,666 ± 310, 28,654 ± 621 and 22,966 ± 755. The total number of liposomes around the bubbles was found to be 1.01 × 105 ± 11,425. This mean value was derived by combining the number of liposomes present in each peak of every image in a stack.
For validation of this method, the estimated numbers by image analysis were compared with the numbers obtained experimentally. The experimental method used was based on haemocytometry. In haemocytometry, the liposome–microbubble was diluted in equal volume of buffer solution. The bubbles were excited to emit fluorescence, which was detected in an appropriate channel. Results were expressed as mean fluorescence per microbubble using unloaded microbubbles as a blank. The average diameter for size distribution of conjugated microbubbles was higher than the unloaded microbubbles. Considering liposome as 200 nm spheres, the thickness of liposome coating on microbubbles was used to calculate the number of liposomes.15
The number of liposomes calculated using ICS was found to be +6% to −2% of the estimated number calculated via haemocytometry. The errors between these two methods were of acceptable level, which validated the estimation of liposomes using the ICS method. Using neutravidin, the binding of liposomes with an average diameter of 200 nm approached the number of 1.01 × 105 liposomes/microbubble. The estimated number of liposome per microbubble was consistent with the estimation published earlier by Kheirolomoom et al. 5 Kheirolomoom et al. showed estimated approximately 1 × 105 for 100 nm liposomes. However, their microbubble diameter was in the range of 1.6 μm that was approximately half the average diameter of the microbubble used in our experiment. Our higher surface area might have contributed to the estimation of large number of liposomes. Moreover, our 2D analysis did not take the 3D attachment of the liposome into consideration. However, it can be argued that filtering the red colour from the image may have allowed the appearance of some liposomes attached on the top and bottom surface of the microbubble. Considering this argument, we can conclude that the estimation via ICS may have been 5%–10% higher than that expected.
Conclusion
Microbubbles in the concentration range 8 × 108 microbubbles/mL and size distribution (2–7 μm) were prepared and conjugated to the liposomes for potential theranostic application. The conjugation of liposomes to the microbubbles was evaluated using ICS to show the efficacy of the conjugation method. The image analysis method for non-interacting particles was applied to estimate the number of liposomes conjugated to the microbubble surface. The estimation of liposome numbers using the image analysis method was found to be slightly higher than the experimental counting method. This suggests that the image analysis method can potentially be used for the approximation of lipid counts.
Footnotes
Acknowledgement
Dr. Ketan Pancholi was previously affiliated with Robert Gordon University, School of Engineering, Garthdee, Aberdeen - AB10 7GJ, UK, where this research was conducted.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research received financial support from the grant Nanoporation FP7-PEOPLE-IAPP. Ritu Malik was funded for research from the same grant at IMSAT.