Abstract
Background
Quantifying and dealing with lack of consistency in administrative databases (namely, under-coding) requires tracking patients longitudinally without compromising anonymity, which is often a challenging task.
Objective
This study aimed to (i) assess and compare different hierarchical clustering methods on the identification of individual patients in an administrative database that does not easily allow tracking of episodes from the same patient; (ii) quantify the frequency of potential under-coding; and (iii) identify factors associated with such phenomena.
Method
We analysed the Portuguese National Hospital Morbidity Dataset, an administrative database registering all hospitalisations occurring in Mainland Portugal between 2011–2015. We applied different approaches of hierarchical clustering methods (either isolated or combined with partitional clustering methods), to identify potential individual patients based on demographic variables and comorbidities. Diagnoses codes were grouped into the Charlson an Elixhauser comorbidity defined groups. The algorithm displaying the best performance was used to quantify potential under-coding. A generalised mixed model (GML) of binomial regression was applied to assess factors associated with such potential under-coding.
Results
We observed that the hierarchical cluster analysis (HCA) + k-means clustering method with comorbidities grouped according to the Charlson defined groups was the algorithm displaying the best performance (with a Rand Index of 0.99997). We identified potential under-coding in all Charlson comorbidity groups, ranging from 3.5% (overall diabetes) to 27.7% (asthma). Overall, being male, having medical admission, dying during hospitalisation or being admitted at more specific and complex hospitals were associated with increased odds of potential under-coding.
Discussion
We assessed several approaches to identify individual patients in an administrative database and, subsequently, by applying HCA + k-means algorithm, we tracked coding inconsistency and potentially improved data quality. We reported consistent potential under-coding in all defined groups of comorbidities and potential factors associated with such lack of completeness.
Conclusion
Our proposed methodological framework could both enhance data quality and act as a reference for other studies relying on databases with similar problems.
Keywords
Introduction
Administrative databases are a relevant source of healthcare secondary data, containing routinely collected administrative, demographic and clinical information obtained when patients use healthcare services. These databases may allow for the assessment of large populations (including on a nationwide or state-wide scope) and for long periods of time without incurring the high costs that would result from studies based on primary data collection (Johnston, 2014; Raghupathi and Raghupathi, 2014). Despite these advantages, administrative databases also display important limitations on their completeness, accuracy and risk of bias (Alonso et al., 2020a; Freitas et al., 2016). In fact, lack of consistency and completeness of these databases is one of the major issues that affect their internal validity (Rothman et al., 2015), related, among others, to a lack of integration of digital systems or the inconsistent practice of coding secondary diagnoses (Peng et al., 2017). The latter may result in potentially relevant under-coding, which may hamper the validity of results obtained when administrative databases are used for research purposes (Raghupathi and Raghupathi, 2014; Sousa-Pinto et al., 2018).
Quantifying and dealing with under-coding in part requires tracking patients longitudinally without compromising anonymity (so as to compare secondary diagnosis codes across multiple episodes of each patient) (Mazzali et al., 2016; Mbizvo et al., 2018). When this is not directly possible (e.g. due to lack of a consistent patient identification number), machine learning methods may be of help. In fact, such algorithms have been applied to administrative databases with multiple goals, from analysing patterns about individual patients to predicting and imputing missing values, data linkage and addressing constraints due to data quality issues (Junior et al., 2018; Souza et al., 2020a, 2020b). Considering clustering methods only, a diverse range of methodologies has been applied to separate administrative data into homogenous groups, including hierarchical cluster analysis (HCA), k-means, partition around medoids (PAM) or exploratory factor analysis (EFA) (Haneef et al., 2021; Ng et al., 2018; Vuik et al., 2016; Weissler et al., 2020; Yan et al., 2019). Despite their different approaches, all of these clustering techniques combine groups of observations according to their similarities (Yan et al., 2019). For example, the HCA starts with individual data points that are subsequently combined into relevant different groups. On the other hand, k-means assigns subjects to the nearest centre (according to a pre-specified number of clusters), minimising distance error. Hence, these different methods are likely to be applied to different scenarios as they tend to display different advantages and disadvantages (Yan et al., 2019). In addition, the performance of the different algorithm approaches depends upon the quantity and quality of available variables. As a result, and given the difficulties in achieving compliance of assumptions in health data, selecting the best clustering algorithm for each different purpose remains a challenge (Embrechts et al., 2013; Jain et al., 1999; Raherison et al., 2018; Sanchez-Rico and Alvarado, 2019).
This study aimed to assess and compare different hierarchical clustering methods on the identification of individual patients within the context of administrative databases, providing a methodological approach that could be applied to future studies dealing with databases of this type. In particular, and as a case study, we assessed the Portuguese National Hospital Morbidity Dataset, which does not easily allow tracking of episodes relating to (or belonging to) the same patient. Additionally, and after identifying hospital admissions belonging to the same individual over a 5-year period, the study aimed to quantify potential under-coding of secondary diagnoses and to identify factors associated with such phenomena.
Method
Database and study design
We analysed hospitalisations data extracted from the Portuguese National Hospital Morbidity Database, which is provided by the Central Administration of the Health System (ACSS – Administração Central do Sistema de Saúde) and contains data from all hospitalisations occurring in public hospitals from Mainland Portugal. This database has, for each hospitalisation episode, administrative information (e.g. admission and discharge dates, and discharge outcome), information on demographic variables of the patient (e.g. sex, birth date and parish of residence) and information on the main and secondary diagnoses, as well as on intervention procedures. Diagnoses and procedures are coded according to The International Classification of Diseases, Ninth Revision, Clinical Modification (ICD-9-CM) standards. In this database, hospital episodes are anonymised, being assigned a unique patient identification number to identify different episodes occurring within the same year from the same patient. This does not link to the clinical registration of the patient (rendering it impossible to keep track of patients’ medical history) and changes annually even for episodes from the same individual. That is, two episodes from the same patient occurring in the same year will be ascribed the same patient identification number, but two episodes from the same patient occurring in different years will be ascribed to different patient identification numbers.
Data analysis
Data analysis encompassed four different steps (depicted in Figure 1). (i) We applied different approaches of hierarchical clustering methods (either isolated or combined with partitional clustering methods) to identify the best approach to group the episodes belonging to the same individuals. To do that, we selected a random sample of 2015 hospitalisation episodes (namely, all 2015 hospitalisations relating to – or belonging to – patients born in five randomly selected birth weeks) from the National Hospital Morbidity Dataset. Results of the clustering methods were then compared with the yearly patient identifier number to calculate the Rand Index as evaluating metric of clustering quality (Krieger and Green, 1999). (ii) Subsequently, in a scaling-up approach, we applied the best-performing classification method to a larger random sample of 2015 hospitalisations (namely, all 2015 hospitalisations from patients born in 20 randomly selected birth weeks) from the same database. Results of these clustering methods were again compared with the unique patient identifier number to calculate the Rand Index. (iii) We then selected a random sample of 2011–2015 hospitalisation episodes (all 2011–2015 hospitalisations from patients born in 20 randomly selected birth weeks), in which we applied the best-performing clustering method (Figure 1). (iv) Lastly, based on these results, we identified and quantified potential under-coding of several chronic diseases and assessed factors which were associated with such potential under-coding. To this end, a generalised mixed model (GML) of binomial regression was applied. Representation of the methodological approaches followed in our study.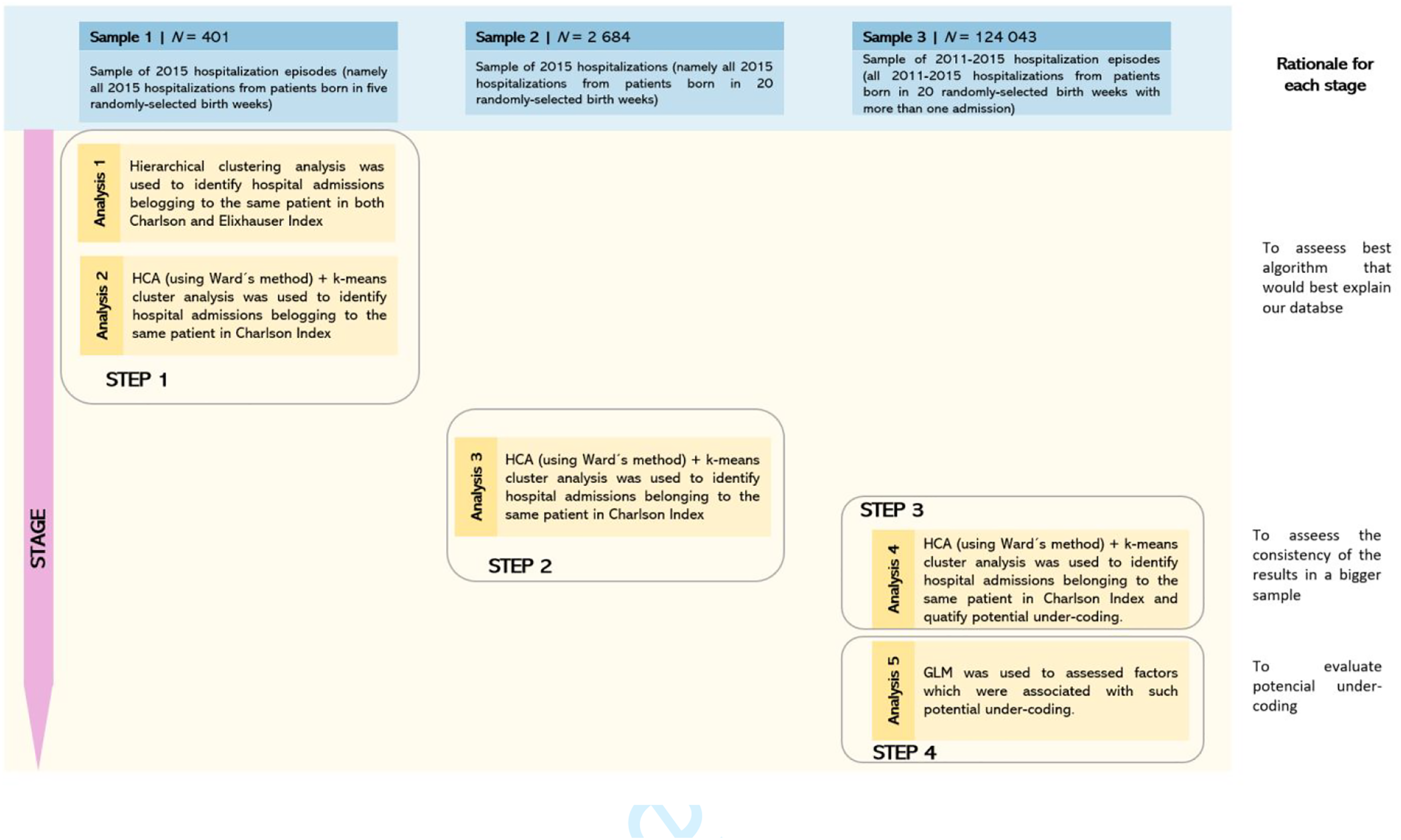
Each sample was randomly obtained based on a gamma distribution which was devised to fit the years of birth density distribution. Data were analysed using SPSS 27.0 for Windows (IBM, Chicago, IL, USA) and R software.
Clustering analysis
We performed clustering analysis based on patients’ sociodemographic variables (gender, date of birth, and residence location defined by the district, municipality and parish) and main and secondary diagnoses. Regarding the latter, the main and secondary diagnoses codes were grouped into the Charlson and Elixhauser comorbidity groups (Charlson et al., 1987; Elixhauser et al., 1998; Freitas et al., 2016), which have been validated in studies that use administrative health data (Dominick et al., 2005; Freitas et al., 2016). Both classification systems are the ones most frequently used for epidemiological studies and health services research (Freitas et al., 2016). The Charlson index originally comprised 19 groups of comorbidities associated with an increased risk of hospital mortality but has subsequently been modified into 17 groups (Charlson et al., 1987; Deyo et al., 1992). The last update on Elixhauser index classifies comorbidities into 31 groups according to their association with increased length of stay, hospital expenses and mortality (Elixhauser et al., 1998; Garland, 2012).
In the first step of our analysis, to calculate the distance between two cases described by variables with different scales, hierarchical-based clustering methods were tested isolated or in combination (hierarchical + partitional clustering) and with different combinations of input variables, including comorbidities grouped either according to the Charlson or the Elixhauser groups of comorbidities. For the subsequent steps of the analysis, we applied the algorithm displaying the best performance (i.e. presenting the highest value of the Rand index (Krieger and Green, 1999), as a proxy of agreement of the clustering algorithm compared to the yearly unique patient identification number), considering both the clustering method and the method for grouping comorbidities.
The best clustering method corresponded to a combination of HCA with an agglomerative algorithm (with Ward’s (1963) method using the Euclidean distance) applied to find a pre-specified number of clusters that would better suit the database, followed by a k-means cluster analysis used to partition cases into k number of clusters, maximising between-cluster differences and minimising within-cluster variance on specified variables (Hand and Krzanowski, 2005). In addition, the lowest frequency of misclassification and highest value of the Rand index were found with comorbidities grouped according to the Charlson groups. For subsequent analyses, we considered not only comorbidities of the Charlson comorbidity defined groups, but also six highly prevalent diseases selected mainly through a method of judgement sampling that may not be directly appraised by Charlson defined variables, which we aimed to additionally assess, namely, diabetes (combining with and without chronic complications division), uncomplicated arterial hypertension, asthma, chronic obstructive pulmonary disease, rheumatoid arthritis and chronic renal insufficiency (codes listed in Table S1, online supplement).
Estimation of potential under-coding and regression analysis
For each group of diseases, potential under-coding was calculated based on the absence of ICD-9-CM codes defining the disease group within episodes belonging to the same cluster of hospital admissions (i.e. putatively to the same patient). Therefore, estimation of potential under-coding was only feasible in potential patients identified as having more than one hospital admission over the years. Since the first time a disease is coded may correspond to the diagnosis time and, as such, may influence the definition of potential under-coding, quantification of potential under-coding was performed both (a) irrespective of the first time, a disease group was registered and (b) considering only episodes after the first time, a disease group was registered for that ‘patient’/cluster of hospital admissions.
In order to identify factors associated with potential under-coding for each group of diseases, we applied regression models at the episode level, with the occurrence of potential under-coding as the dependent variable. Independent variables included patients’ sex and age at admission, type of admission (medical or surgical), length of stay, whether the episode was classified as ‘urgent’, type of discharge (normal discharge, discharge against doctor advice or in-hospital death) and hospital complexity (classified into four types based on population-based criteria and healthcare capacity provision) (Saude, 2014). Given the multilevel structure of data, with each patient potentially having multiple hospital admissions, mixed-effects logistic regression models (GML) were applied, clustering hospitalisations data by the presumable patient (identified by the clustering algorithm). Univariable GML models were firstly performed, followed by multivariable GLM regression models simultaneously including all covariates. Analyses were performed with potential under-coding defined both when considering and not considering the first time a diagnosis is coded. Exponentials of the regression coefficients were interpreted as odds ratios (ORs) and their corresponding 95% confidence intervals (CI) were calculated.
Ethics
This is a secondary data study that uses an anonymised database of administrative data collected in all hospitalisations in Portuguese public hospitals. There was no need for an ethical committee approval. The study was conducted in accordance with privacy and data protection principles and regulations.
Results
Identification of episodes from the same patients using clustering methods
Results of the different clustering methods for identification of episodes from the same patient.
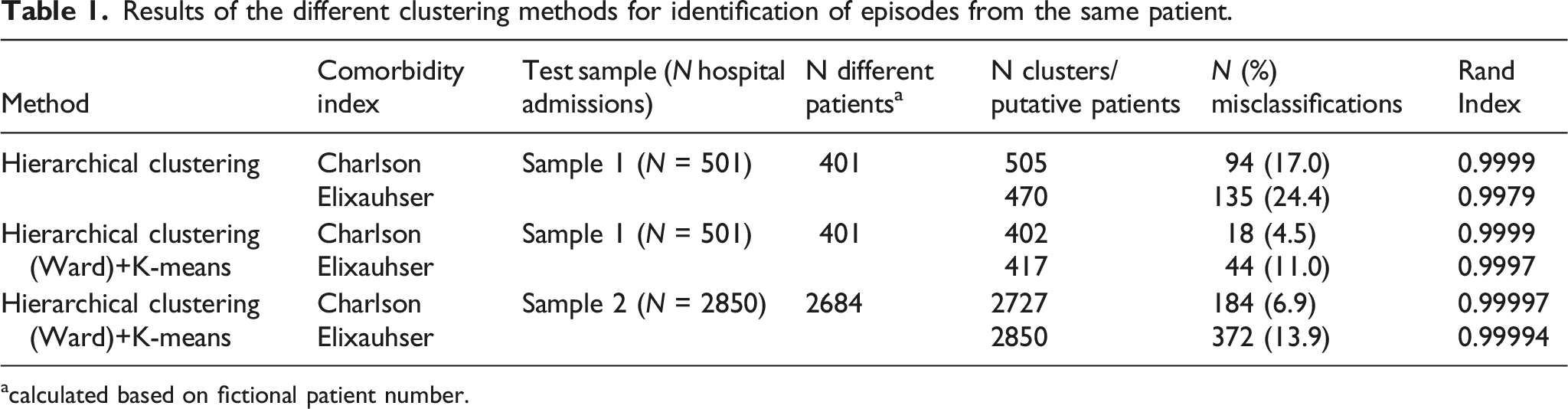
acalculated based on fictional patient number.
Quantification and assessment of potential under-coding
Descriptive frequency for potential under-coding in each diagnostic category.
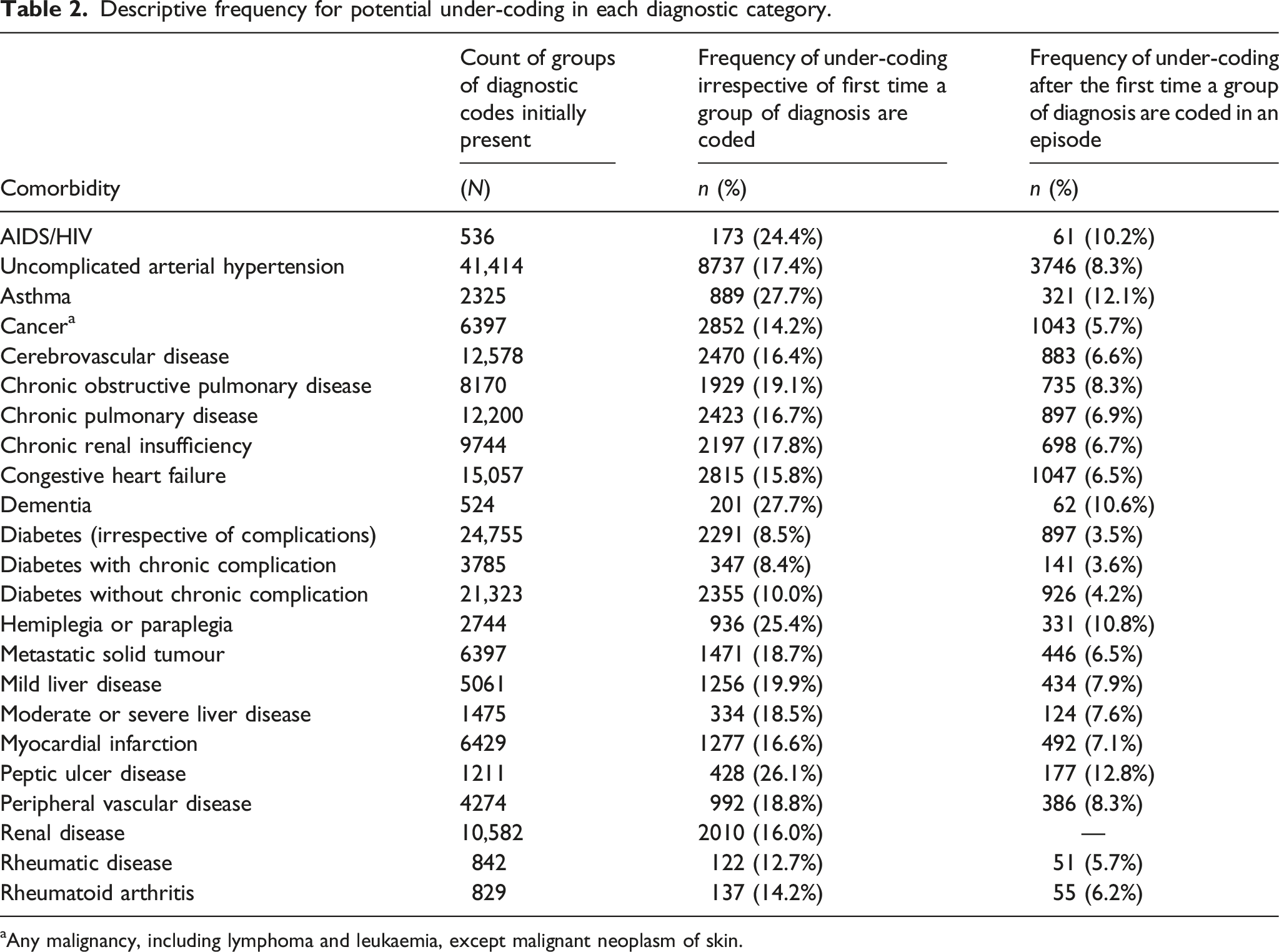
aAny malignancy, including lymphoma and leukaemia, except malignant neoplasm of skin.
We then performed a GLM regression to analyse which factors may be associated with potential under-coding of each group of comorbidities (Table S2.1 and Table S2.2. online supplement). Consistent results across comorbidity groups were found for sex, age, type of admission and type of discharge. Overall, being male, having medical admission, dying during hospitalisation or being admitted at more specific and complex hospitals were associated with increased odds of potential under-coding (i.e. hospital admissions with such characteristics were found to be potentially less extensively and properly coded). In-hospital death was the only variable displaying different patterns according to whether the time up to the first code registration was or was not considered, associating with higher chances of potential under-coding when such time was considered. No obvious pattern was found regarding hospitalisation planning or the number of admission days, although longer stays tended to be associated with increased odds of potential under-coding.
Discussion
This study assessed different types of hierarchical clustering techniques to identify episodes from the same patient, applying HCA + k-means as the best-performing approach to quantify potential under-coding of secondary diagnoses in administrative databases. The use of administrative data allows researchers, hospital managers and policymakers to analyse data from large population samples for long periods of time. However, studies based on administrative data must be carefully designed and their results cautiously interpreted, as they may contain inaccuracies and be prone to bias due to underreporting or misclassification. Our results suggested that regarding Portuguese hospitalisations, potential under-coding may be quite frequent. Given that under-coding has been reported in other countries, our results may be relevant to other scenarios (Peng et al., 2017; Souza et al., 2019, 2020b).
Applying machine learning algorithms such as clustering methods in administrative databases can be challenging due to the different nature of available variables (Lopez-Arevalo et al., 2020) (with the coexistence of categorical and continuous variables) and to the possibility that some variables are strongly correlated. We used predefined groups to classify comorbidities, since any benefit from considering the individual codes for diagnoses would have been overcome by (a) the difficulty that models would have in adequately dealing with a high number of variables and (b) the potential increased risk of collinearity. We observed better performance of models when comorbidities were grouped according to the Charlson groups (instead of the Elixhauser groups), which may be explained by the fact that it groups diseases into a smaller number of classes. A clustering approach based on an HCA agglomerative scheme with Ward’s method using the Euclidean distance continued with a traditional k-means clustering technique was applied to improve and stabilise the cluster performance (Berzal and Matín, 2002). The Euclidean distance proved to be useful in our study, since we aimed to maximise similarities between homogeneous groups and detect differences with adequate granularity. To the best of our knowledge, our study is the first to use such an approach simultaneously to cluster episodes from different patients in the same administrative database and to tackle potential under-coding. Previous studies using cluster analysis methods in administrative databases have mostly focused on segmentation of a general patient population into homogenous groups (Vuik et al., 2016); linked data from different databases without any unique patient identifier (Junior et al., 2018); while Freitas et al.(2016) applied deterministic approaches to internally link same-patient hospitalisation episodes by using indirect identifiers (Souza et al., 2020b).
On the other hand, there are previous studies that have solely addressed under-coding. For example, Cappetta et al. (2020) used longitudinal data to assess the long-term patterns and consistency of coding of dementia in Australia over time, from the first-coded hospital admission in each patient, relying on a unique patient identification number for data linkage. However, Harron et al. (2017) drew attention to issues concerning data linkage in administrative databases due to insufficient identifying information, with implications for safeguarding personal data and research. By contrast, Kumar et al. (2020) applied a combination of five machine learning techniques (tree-based XGboost [balanced-data-model]; logistic regression; random forest; decision tree and linear SVC) to characterise the incidence of self-harm and to identify factors associated with coding bias, while Peng et al. (2017) addressed under-coding of chronic diseases by using logistic regression models via least absolute shrinkage and selection operator (LASSO) to estimate coding validity. However, both Kumar et al. (2020) and Peng et al. (2017) relied on human coding audit as the gold-standard for model performance. In the absence of historical or current coding audit data, we applied an unsupervised learning methodology, which has already proved capable of identifying subgroups of patients while measuring clinical and meaningful differences in capability (Yan et al., 2019).
Under-coding is a well-established major issue for administrative health data (Hua-Gen Li et al., 2019; Peng et al., 2017; Quan et al., 2008). Quan et al. (2002) found that administrative data tended to underestimate the frequency of diseases/conditions compared with patient chart data. Peng. et al. (2017) assessed coding validity of four conditions with high prevalence (hypertension, diabetes, obesity and depression) with findings suggesting that under-coding is closely related to the perception of the clinical importance of a condition and its influence on the length of stay, care received or treatment during hospitalisation. In addition, previous studies reported likely suboptimal treatment on under-coded comorbidities such as obesity (Bilsker et al., 2007) and dementia (Cappetta et al., 2020), as they may fail to draw physician attention. In our study, diseases probably perceived to have a higher impact on the severity or complexity of the admission appeared to have lower frequencies of potential under-coding (such as diabetes). Freitas et al. (2016) found that from 2000 to 2010, the increasing trend on reporting comorbidities was not equal for all comorbidities (Souza et al., 2020b). In contrast to complicated diabetes, they reported a decrease in coding AIDS/HIV and peptic ulcer disease coding (Freitas et al., 2016; Souza et al., 2020b). Such trends may reflect distinct coding patterns (due to internal payment factors or variations on the complexity of clinical coding systems) among different hospitals and medical specialties. In addition, coding standards may also be reflected in differential coding for specific diseases/conditions. In fact, some conditions (such as asthma or peptic ulcer disease) are more typically registered in the personal history of the patient, being coded only when actively associated with acute events (Alonso et al., 2020a; Cappetta et al., 2020; Rollason et al., 2009; Souza et al., 2019). Of note, the coding standards in Portugal follow the American Official Guidelines ICD-9-CM 2011 (where only secondary diagnosis that may impact the admission should be coded) (CDC, 1991). Nevertheless, in this specific study, the impact of such a practice may have been attenuated, as we focused on Charlson comorbidity groups, which mostly tend to impact either the treatment or the admission in some way.
After estimating the frequency of potential under-coding, we further explored which factors could partly explain potential under-coding in each group of diseases. Findings from our study suggest that in contrast to being female, being of older age was associated with a higher frequency of potential under-coding. Regarding the effect of age, it is possible to hypothesise that since older people (a) are more likely to have multiple diseases and (b) tend to have more hospital admissions over time caused by decompensation of major diseases (García-Pérez et al., 2011), this may have led to an increased probability of health professionals failing any related coding (Kripalani et al., 2014). We also observed that increased hospital complexity was associated with higher frequency of potential under-coding. It is known that the availability of medical specialties appears to influence coding patterns (Payne et al., 2012). In addition, we hypothesised that, despite more complex hospitals being subject to higher investment in audits and training for coding awareness, the higher frequency of potential under-coding may be associated to the fact that these hospitals tended to have more complex patients with a larger number of comorbidities and, therefore, increasing the chances that some specific individual conditions may end up not being coded. Having a surgical admission was associated with decreased odds of potential under-coding. Two factors may explain this finding, namely, (a) the fact that a large set of such admissions concern otherwise healthy patients (e.g. fractures or deliveries in healthy young adults) and (b) a straightforward incentive for health professionals in surgical admissions in Portugal, as payment is directly linked to clinical coding, which motivates coding and may even lead to over-coding. Souza et al. (2020a) reported examples of possible incentives for upcoding practices such as switching pneumonia-related conditions from principal to secondary diagnosis, which could increase hospital financial compensation. All of our hypotheses regarding the factors underlying potential under-coding were solely based on clinical knowledge and previous literature. An audit of electronic health records could further contribute to provide potential relevant insight in determining reasons for under-coding, even though electronic health records themselves may suffer from lack of information, unclear documentation and variability in their quality, which can compromise the accuracy of any audit (Alonso et al., 2020b).
Strengths and limitations
Besides its novelty and the possibility of clustering data from patients over time, one of the most important strengths of this study concerns the nationwide scope of the database used. Moreover, our findings have several practical implications, given they represent an opportunity to (a) improve data quality; (b) properly address some limitations of administrative databases during research and (c) impact policies on clinical coding of secondary diagnosis. However, our study also had some limitations. The database was originally developed for billing purposes and few patients’ demographic variables were available. This could have limited the models’ performance. Such paucity of variables may also be observed in other real-world administrative databases, potentially increasing the applicability of these models to other databases. Additionally, for each episode, we were not able to compare ICD-9-CM codes with electronic health record data, meaning our estimates of potential under-coding should be interpreted cautiously (and the reason for our reference to these estimates of coding inconsistency as ‘potential under-coding’ and not simply as ‘under-coding’). In the absence of a gold-standard, it was difficult to distinguish under-coding from wrong coding (either from isolated unintentional coding errors or from over-coding due to payment incentives). Both over-coding and wrong coding could have impacted on our findings and biased the results in the direction of overestimating under-coding, as coding a wrong diagnosis once can lead to wrong under-coding identification in subsequent patient admissions. Nevertheless, the frequency of potential under-coding was assessed both considering and not considering the date of the first time a certain comorbidity was registered. While such an approach tries to compensate for any overestimation of under-coding, some constraints remain as to whether such diseases had already been diagnosed in primary care service (Cappetta et al., 2020; Peng et al., 2017). Potential under-coding may also have been underestimated, namely, by patients who (a) only had one admission in the entire study period or (b) had several episodes but never had registered a certain comorbidity. Since our approach would not be applicable in such scenarios, under-coding was only estimated taking into account patients with more than one admission in the selected sample. The impact of such limitations can only be quantified through an audit of electronic health records, which can be costly and time-consuming and may even be insufficient as electronic health records themselves be incomplete or heterogeneously filled (Alonso et al., 2020b).
As for limitations in the clustering methods, the possibility of collinearity due to a high number of highly correlated dichotomous variables giving redundant information to the algorithm (Sanchez-Rico and Alvarado, 2019) was partly solved due to the application of grouped diagnoses into comorbidity defined groups. However, this procedure may partially have resulted in loss of granularity (as comorbidity groups encompass several codes and, often, different diseases), potentially leading to underestimation of under-coding (e.g. asthma and chronic obstructive pulmonary disease [COPD] are both grouped in the ‘chronic pulmonary disease’ group. A patient with both conditions and having one admission in which only asthma was reported and another in which only COPD was reported would not have been detected as a case where under-coding had occurred). Additionally, highly under-coded admissions may also have impacted clustering performance, as it limits the algorithm to group admissions (both considering properly coded and highly under-coded admissions) in homogenous groups of the same individual, leading to optimistic under-coding frequencies. Finally, another important limitation concerns the lack of computational power to assess the algorithm performance on the fully available data for the year 2015, and to apply it for the full 2011–2015 database.
Conclusions
Although preliminary, our study provides guidance for improving data quality for observational studies using secondary administrative databases. We assessed several approaches to identify individual patients in an administrative database and, subsequently, by applying HCA + k-means algorithm, we tracked coding inconsistency and potentially improved data quality. We reported consistent potential under-coding in all defined groups of comorbidities and potential factors associated with such lack of completeness. Despite discussed limitations, we recommend adopting this methodological framework, which could also act as a reference for other studies relying on databases with similar problems. However, further longitudinal refinement and validation of the applied approaches and the observed findings and techniques are needed.
Supplemental Material
Supplemental Material - Unsupervised algorithms to identify potential under-coding of secondary diagnoses in hospitalisations databases in Portugal
Supplemental Material for Unsupervised algorithms to identify potential under-coding of secondary diagnoses in hospitalisations databases in Portugal by Diana Portela, Rita Amaral, Pedro P Rodrigues, Alberto Freitas, Elísio Costa, João Almeida Fonseca, and Bernardo S Pinto in Health Information Management Journal
Footnotes
Acknowledgements
We acknowledge Professor Fernando Lopes for his useful indication on the codes used in this paper.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
