Abstract
Background:
Positive cytology has been identified as an independent negative prognostic factor in patients with peritoneal metastases (PM) of colorectal origin. Liquid biopsy in plasma may detect increasing levels of circulating tumor DNA (ctDNA) and could help predict systemic relapse in patients with colorectal cancer, but little is known about the role of liquid biopsy in peritoneal fluid. The aim of this study was to evaluate the prognostic value of peritoneal fluid and plasma liquid biopsy in patients undergoing complete cytoreductive surgery and hyperthermic intraperitoneal chemotherapy (CC-HIPEC).
Methods:
A longitudinal prospective study was designed in patients with KRAS-mutated colorectal or appendiceal primary tumor, including PM of colorectal origin, pseudomyxoma peritonei and patients at high risk of developing PM (selected for second-look surgery). Eleven patients were recruited according to inclusion and exclusion criteria. ctDNA from plasma and peritoneal fluid before and after HIPEC was studied by droplet digital PCR looking for KRAS mutation. A close follow-up was scheduled (mean of 28.5 months) to monitor for systemic and peritoneal recurrences.
Results:
All patients with positive plasma postHIPEC had systemic relapse and four patients died as a result, while those with negative plasma postHIPEC did not relapse. Patients with negative peritoneal ctDNA after CC-HIPEC did not present peritoneal relapse. Of six patients with positive peritoneal ctDNA postHIPEC, two presented peritoneal recurrence and four systemic relapses.
Conclusions:
Treatment with CC-HIPEC does not always neutralize ctDNA in peritoneal fluid, and its persistence after treatment may predict adverse outcome. Despite being a proof of concept, an adequate correlation between liquid biopsy in plasma and peritoneal fluid with both systemic and peritoneal relapse has been observed.
Keywords
Introduction
Colorectal cancer is the third most common cancer worldwide, and represents the second leading cause of cancer mortality. 1 The peritoneum is the third leading metastatic site in patients with colorectal cancer. Combined treatment of peritoneal metastases (PM) with complete cytoreduction and hyperthermic intraperitoneal chemotherapy (CC-HIPEC) is performed with curative intent, and achieves a 16% recurrence-free rate at 5 years and medium overall survival (OS) of 29 months, 2 although some groups have reported OS of up to 42.2–64 months.3,4 Treatment with CC-HIPEC increases survival and offers a chance of cure 5 ; however, peritoneal recurrence rates remain high after this treatment.6,7 As CC-HIPEC does not prevent relapse in all cases, recurrence-associated factors are a pending issue.
The prevalence of positive cytology in the literature in patients with PM varies due, among other factors, to a lack of homogeneity in the collection and sample-processing procedures. 8 Positive cytology has been reported as an independent indicator of poor prognosis in patients with colorectal PM.9,10 Trilling et al. showed that the presence of tumor cells in peritoneal fluid (positive cytology) is an independent factor for poor prognosis, with a strikingly lower survival (19 versus 44 months) 11 despite CC-HIPEC with curative intent.
The biological function of cell-free DNA (cfDNA) in peripheral blood is currently unknown, but it is thought that circulating tumor DNA (ctDNA) might transform non-tumor cells at a distance.12–14 Dissemination and metastasis could be more closely related to ctDNA than to circulating tumor cells. 15
A large number of studies propose the use of plasma liquid biopsy in oncological patients, either to help early diagnosis, detect residual disease, predict the risk of relapse, or monitor response to treatment or the clonal evolution of tumor tissue.16,17 However, its use in clinical practice is not yet standardized. 18
Several studies have described the relationship between ctDNA positivity in plasma liquid biopsy after surgery and tumor relapse, associating it with lower disease-free survival. 19
In this study, the effect of CC-HIPEC on the two components of liquid biopsy of peritoneal fluid has been analyzed: the cellular component by cytology and the ctDNA fraction detecting KRAS mutations by droplet digital PCR (ddPCR). Our aim was to identify the presence of tumor cells and ctDNA in peritoneal fluid, as well as ctDNA in plasma, in patients with peritoneal colorectal cancer metastases, and to analyze their relationship with disease-free survival and OS.
Methods
Patients
Twenty-six patients undergoing CC-HIPEC between April 2016 and May 2017 at Fundación Jiménez Díaz University Hospital (FJD) were included in this study after signing informed consent. The protocol was approved by the Ethics Committee for Clinical Research of this institution (PIC 75/2016_FJD). Eleven patients were recruited according to inclusion and exclusion criteria
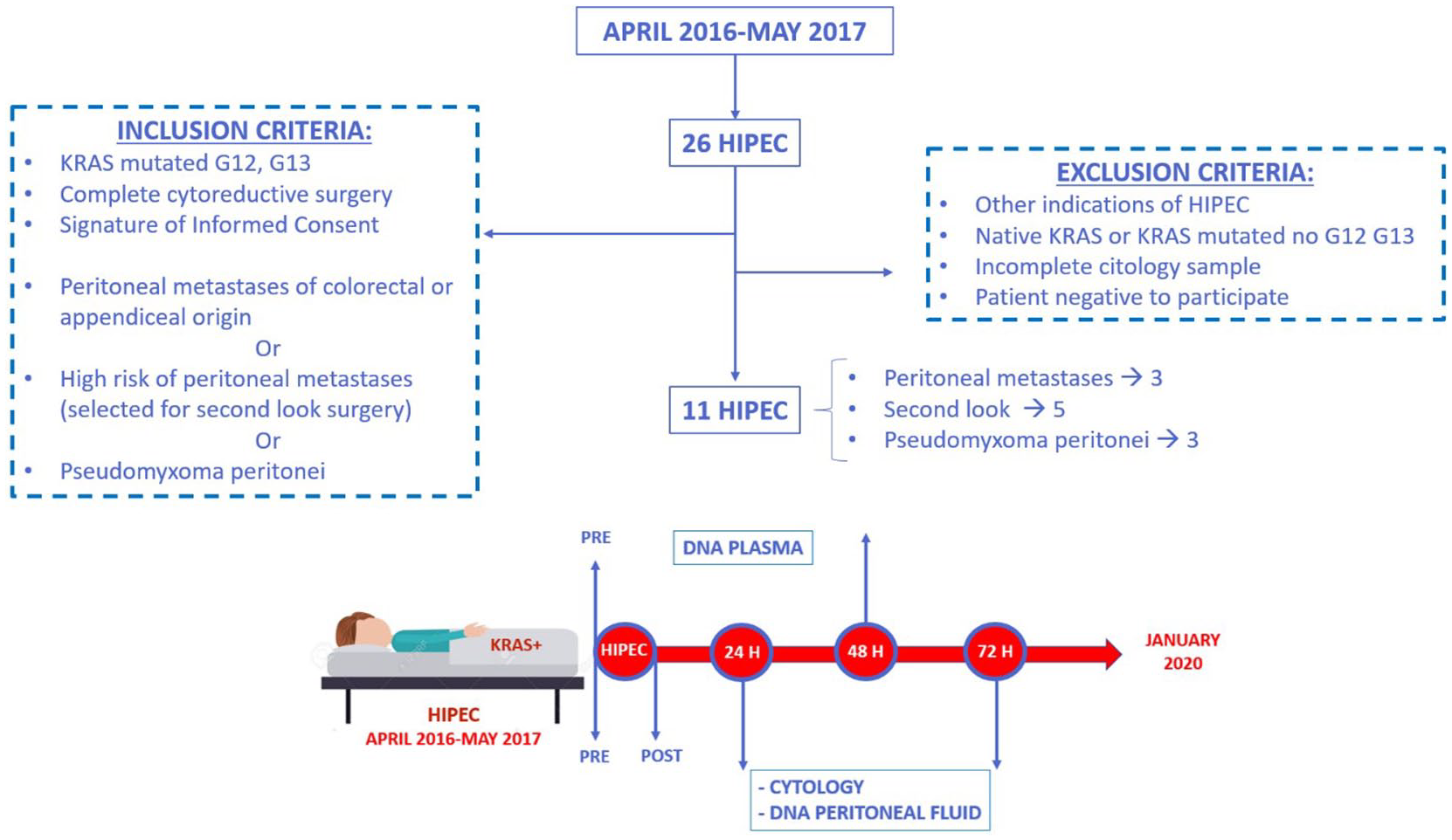
Inclusion and exclusion criteria. Sample collection design. Patients in whom CC-HIPEC was indicated between April 2016 and May 2017 due to suspicion or high risk of colorectal peritoneal metastases or pseudomyxoma peritonei. Sample collection protocol: plasma sample obtained before CC-HIPEC and 48 h after surgery. Peritoneal fluid samples before CC-HIPEC and after cytoreductive surgery and HIPEC (peritoneal lavage), 24 and 72 h after surgery, via abdominal drains.
Sample collection
Blood samples were collected in EDTA tubes before surgery and 48 h after surgery and processed in less than 2 h. Blood underwent double centrifugation at room temperature (1800 × g and 3000 × g for 10 min, respectively) and plasma was stored at −80°C until use.
Immediately after laparotomy, abdominal lavage with saline solution was performed and this fluid was collected (preHIPEC). Peritoneal fluid contained in drainage bags was also recovered at 24 and 72 h post-surgery. Each peritoneal fluid sample was simultaneously processed for cytologic analysis in accordance with routine practice in our hospital and for ddPCR analyses at our research laboratory. Peritoneal fluid underwent double centrifugation as described above for plasma samples.
KRAS mutation detection
DNA was extracted from plasma and peritoneal fluid with the QIAamp Circulating Nucleic Acid Kit and the DNeasy Blood & Tissue kit was used for cell lines (Qiagen). The starting volume was 3 ml for plasma and 2 ml for peritoneal fluid. ddPCR analyses were performed using the QX200 Droplet Digital PCR System (Bio-Rad). KRAS G12D, G12V and G13D mutations were detected with PrimePCR™ ddPCR™ Mutation Detection Assays (Bio-Rad). DNA from LS-174T, SW480 and HCT-116 human adenocarcinoma cell lines was used as a positive control of these mutations, respectively. SW480 was purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). LS-174T and HCT-116 were kindly provided by the Translational Oncology Division, OncoHealth Institute. KRAS wild-type control DNA was obtained from healthy donor peripheral blood mononuclear cells. No-template controls (NTCs) adding water instead of DNA to the reaction mixture were also included. Results were analyzed using Quantasoft v.1.7 software (Bio-Rad). Two to four replicates of each sample were analyzed.
Statistics
Merged values of concentration (mutated copies per microliter of reaction) were compared between samples and wild-type controls using a Z test, with the assumption that concentrations followed a normal distribution. A cut-off value of p < 0.05 was used for positivity. Those patients whose ctDNA concentration is over this limit are called positive or detectable ctDNA.
Results
Eleven patients met the inclusion and exclusion criteria with a mean follow-up of 28.5 months (range 8–41). Of the 11 patients, three presented PM, three acellullar pseudomyxoma peritonei, and five underwent second-look surgery for high PM risk (one patient with proven peritoneal disease in pathology report).
Mean age at diagnosis was 56.9 years (29–75). Overall, 64% of the cases were women versus 36.3% men. The primary tumor was located in the right colon in 46% of patients, 36% in the appendix, and 18% in the left colon. The origin of pseudomyxomas was low-grade appendiceal mucinous neoplasm (LAMN).
Regarding surgical details, the median peritoneal cancer index (PCI) value was 4 (0–26) with a mean of 7.2. Some 18% presented positive cytology before HIPEC. Overall, 91% of surgeries achieved CC0 cytoreduction, and 9% CC1 due to small bowel involvement. A mean of 3.45 visceral resections were performed per procedure. Three patients out of the 11 presented liver disease that was resected simultaneously. Duration of surgery was 9.5 h on average (7–12 h) with a mean 17.4 days of hospital stay (11–36 days). Complications happened in 45% of the patients; 40% of these complications corresponded to Clavien Dindo severity grade III.
Clinical details are included in Supplemental Material.
Overall, at the last review of follow-up 46% of patients remained disease-free. Three patients died due to systemic progression, one due to systemic and peritoneal progression, and one due to surgical complications. Relapse occurred within 5.5 months on average after HIPEC, with the earliest relapses for patients with liver metastases resected at surgery. Mean survival after HIPEC was 28.5 months (8–41) with a disease-free period of 20.6 months (2–41).
Main outcomes are summarized in Table 1.
Clinical and molecular features of the patients included in the study and their subsequent clinical outcome.

ctDNA, circulating tumor DNA; LAMN, low-grade appendiceal mucinous neoplasm
Discussion
In all cases, cfDNA was detectable in the fluid collected from peritoneal cavity after lavage with saline solution, performed just after laparotomy (preHIPEC determination). Subsequently, a remarkable increase in total cfDNA in peritoneal fluid from drainage bags was observed 24 h after CC-HIPEC, probably due to the important cellular lysis following CC-HIPEC treatment. For the analysis of KRAS mutations by ddPCR, it was decided to compare preHIPEC with 72 h postHIPEC peritoneal liquid biopsy, discarding 24 h postHIPEC samples because high levels of non-tumoral DNA could mask the small amounts of tumoral DNA.
Liquid biopsy in plasma
Is a positive preHIPEC plasma liquid biopsy associated with systemic disease?
We found preoperative detectable ctDNA in plasma in 45.5% of patients. In line with our results, Baumgartner et al. reported detection of preoperative mutated KRAS ctDNA in blood in 38.8% of patients with peritoneal carcinomatosis of colorectal origin. 20
Five out of the 11 patients were positive in the preHIPEC plasma analysis, but only two had systemic disease (Figure 2). In the three cases without preoperative evidence of systemic disease, ctDNA was detectable in the bloodstream. This could be related to micrometastasis, as it has been previously described for pancreatic cancer. 21

Liquid biopsy in plasma. Five patients out of 11 had detectable mutant KRAS cfDNA preHIPEC. Patients who maintained a positive plasma ctDNA or who became positive after treatment had a worse disease course, compared with those who maintained a negative value or became negative, who remained disease-free.
PreHIPEC liquid biopsy analysis identified two out of the three patients with established preoperative systemic disease among the 11 patients. The third patient, with systemic disease evidenced on radiological imaging but without ctDNA detected preHIPEC, had a very high wild-type KRAS concentration, suggesting contamination with DNA from blood cells (because the sample was processed 3 h after extraction), possibly masking tumor DNA. Of the six patients without mutant ctDNA preHIPEC, the remaining five had no evidence of systemic disease (Table 1). Thus, the preoperative detection of ctDNA appears to be related to systemic metastases and probably to micrometastases.
Can positive postHIPEC plasma liquid biopsy predict systemic progression?
Five patients presented positive postHIPEC ctDNA, four patients had undetectable levels and in the remaining two, postHIPEC blood samples were unavailable.
Of the five patients with detectable ctDNA in plasma preHIPEC, three maintained positivity after treatment, presenting accordingly poor short-term prognosis: one of them with systemic relapse 5 months after treatment and subsequent death at 10 months; the second one with systemic and peritoneal relapse at 22 months of treatment and death at 34 months; and finally the third one presented systemic relapse at 2 months of treatment and death at 10 months (HIPEC 17, HIPEC 18, and HIPEC 21, respectively). In one of the remaining two patients with detectable ctDNA in plasma preHIPEC, 48 h plasma sample was not available, but his outcome was also poor, with systemic relapse 2 months after treatment and peritoneal relapse at 24 months (HIPEC 19) (Figure 2). Regarding the last patient, HIPEC 22, ctDNA was neutralized after treatment and she remained disease-free (systemic and peritoneal) until the end of the follow-up. She died after a long hospital stay, due to surgical complications following surgery for rectal cancer developed de novo 22 months after HIPEC.
From the six patients with no detectable ctDNA preHIPEC, three of them maintained negative levels after treatment, presenting good prognosis, without relapse or deaths during follow-up (Figure 2). The 48 h plasma sample was not available from a fourth patient. On the other hand, in two patients ctDNA postHIPEC turned positive and both presented adverse outcome: HIPEC 11 presented systemic relapse 13 months after treatment and HIPEC 24 presented systemic relapse 6 months and death 8 months after treatment, respectively.
Preoperative detection of plasma ctDNA has been associated with shorter OS and reduced disease-free survival in patients with resectable colorectal cancer. 22 Elevation of cfDNA after surgery has been associated with surgical manipulation, 23 although its postoperative detection has been linked to a worse prognosis 14 and with tumor relapse in patients who initially had non-advanced disease. 24 Reinert et al. described that quantification of ctDNA in plasma had high sensitivity and specificity to predict relapse after colorectal surgery with curative intent.25,26 Postoperative variations in plasma ctDNA, monitored weekly and monthly, have previously been correlated with the appearance of tumor progression or new systemic metastases. 27
In this study, the evolution of plasma liquid biopsies after surgical treatment shows that variations in ctDNA are associated with clinical outcome (Figure 3). Patients who maintained positive or developed positive ctDNA postHIPEC presented systemic relapse. Conversely, patients who maintained negative or neutralized ctDNA postHIPEC remained disease-free during the follow-up period. Plasma liquid biopsy emerges, then, as a prognostic factor for systemic relapse after CC-HIPEC.

Droplet digital PCR results. Graphical results of ddPCR analyses of KRAS mutations in plasma liquid biopsy and post-HIPEC course. Gray events represent double negative droplets, in which there was no amplification. The green cluster represents droplets that only contain non-mutated (non-tumor) cfDNA. The blue cluster is formed by droplets containing only mutated KRAS cfDNA (ctDNA), while the orange cluster shows droplets containing both tumor and non-tumor DNA molecules. In the first case (HIPEC 17), the density of the blue cluster decreases but does not disappear after treatment, remaining positive postHIPEC; the patient had systemic relapse during follow-up. In contrast, in the case below (HIPEC 22), only a few residual events are observed: the difference in the estimated concentration of ctDNA in this sample was not statistically significant compared with the wild-type only control, thus it was classified as negative, and the patient remained disease-free.
Liquid biopsy in peritoneal fluid
Liquid biopsy in peritoneal fluid has been poorly explored. In colorectal cancer, it has been observed that isolation of mRNA in peritoneal fluid could differentiate healthy subjects from colorectal cancer subjects. 28 In gastric cancer, the presence and changes in the mRNA of carcinoembryonic antigen (CEA) isolated in peritoneal lavages have been described as a disease-free survival and recurrence indicator,29,30 while in ovarian cancer, exosomes released from the primary tumor at the peritoneal level have been linked to metastatic seeding. 31 The relationship of cfDNA with HIPEC treatment and the clinical evolution in patients with peritoneal metastasis of colorectal origin has not been previously described.
What is the usefulness of preHIPEC peritoneal liquid biopsy? Does it complement cytology?
Six patients had detectable peritoneal mutant KRAS cfDNA preHIPEC. Three had PM of colorectal origin, two were pseudomyxomas, and one patient was from the second-look group (HIPEC 17) (Table 1). In patient HIPEC 17, ctDNA preHIPEC was detected in peritoneal fluid despite the absence of tumor implants at the second-look procedure. This patient had detectable levels of KRAS mutation both in plasma and peritoneal fluid, probably related to the presence of systemic disease. Thus, it could be hypothesized that ctDNA might have been detectable in all body fluids irrespective of the lack of evidence of local disease. The feasibility of detecting tumor cell-free genetic material in different fluids of the organism in metastatic patients, including ascites, has been previously described. 32
PM was confirmed histologically in seven of the 11 patients. Two of them presented positive cytology, while KRAS mutations in cfDNA obtained from peritoneal lavage fluid were detected in five (Table 2). This detection rate is significantly higher than positive cytology rates previously described in the literature, 8 suggesting that it might have a complementary value to the cytology. In line with this, a previous study has shown that the analysis of methylation markers in cfDNA from ascites adds diagnostic and prognostic information to cytology and the combination of both methods increases the detection rate of malignant ascites. 33 This could be relevant when performing staging laparoscopies in patients in whom a therapeutic decision depends on a positive or negative diagnosis of PM.
Cytology results versus detection of KRAS mutations in peritoneal liquid biopsy.

ctDNA, circulating tumor DNA; HIPEC, hyperthermic intraperitoneal chemotherapy
Can postHIPEC peritoneal liquid biopsy predict peritoneal relapse and adverse prognosis?
Six patients were positive postHIPEC. It is remarkable that all the peritoneal pseudomyxoma cases were negative, and results were positive only in the PM or second-look group. In terms of cytology, all patients presented negative postHIPEC cytology (Table 2). CC-HIPEC with curative intent aims to eliminate tumor cells from the abdominal cavity and peritoneal fluid, or at least make these cells undetectable by the routine clinical methods. PostHIPEC cytology, as in all cases turns negative, does not allow us to distinguish patients at risk of relapse from patients with a good prognosis.
ctDNA has been isolated in peritoneal fluid of patients with acellular pseudomyxoma peritonei. All patients showed negative peritoneal liquid biopsy after treatment. In a previous study, 34 we detected ctDNA in mucin in the same cases.
Two out of the six positive patients preHIPEC turned negative with treatment, remaining disease-free during follow-up (Figure 4). The other four patients, who remained positive despite treatment, had poor prognosis: two deaths due to systemic relapse (HIPEC 17 at 10 months, HIPEC 21 at 10 months) and two peritoneal relapses (HIPEC 18 at 22 months post-treatment with death at 34 months, and HIPEC 19 at 24 months).

Liquid biopsy in peritoneal fluid. Six of our 11 patients presented ctDNA in peritoneal fluid pretreatment. Four patients who remained positive presented poor prognosis, compared with the two patients whose results were neutralized, who are still disease-free. On the other hand, of the five patients in whom ctDNA was not detected before CC-HIPEC, the two patients who became positive presented adverse outcomes.
ctDNA preHIPEC was not detected in five of the 11 patients. Three remained negative after treatment and disease-free at the closure of follow-up time (HIPEC 22 died due to postoperative complications from a second surgery due to another tumor as explained above). However, the two patients who developed positive peritoneal ctDNA after CC-HIPEC (HIPEC 11 and HIPEC 24) presented poor prognosis: both developed systemic relapse and one died consequently.
These results suggest that the course of peritoneal liquid biopsy may be related to patient prognosis, since patients whose peritoneal liquid biopsy remained or became negative remained disease-free, while all patients whose peritoneal liquid biopsy remained or became positive presented relapses and/or death (Figure 4). None of the cases in our series deviated from these trends or presented incoherent results in terms of ctDNA postHIPEC and subsequent disease progression. Thus, detection of peritoneal ctDNA after surgery may help predict those patients at high risk of early relapse.
To summarize, we found that positive cytology preHIPEC is a factor of poor prognosis according to literature results10,11 (peritoneal relapses occurred only in patients with positive cytology), whereas postHIPEC cytology does not predict patient outcome. Peritoneal liquid biopsy to determine peritoneal ctDNA levels preHIPEC is a better indicator of PM than cytology and is not always removed by the treatment with CC-HIPEC.
Therefore, these results suggest that the variation of peritoneal and plasma ctDNA concentration could be interpreted as an independent prognostic factor which could help detect patients at high risk of systemic and/or peritoneal relapse, as previously described for plasma,19,35 or with a worse perioperative prognosis.
Association of liquid biopsy with disease-free and survival
Previously published research describes the relationship between the detection of plasma cell-free ctDNA after colorectal surgery and posterior tumor relapse, as well as a lower disease-free survival,19,36,37 possibly deriving from hidden micrometastases, meaning that the disease was more advanced than suspected on imaging or clinical studies. 38 In addition, the detection of persistent ctDNA after surgery with curative intent could also identify patients with suboptimal surgeries and could be used as a marker of surgical radicality. 39
The levels of ctDNA (regardless of the marker used) have been shown to correlate with survival in patients with metastatic colorectal cancer, implying shorter survival and less disease-free time the higher its blood concentration is. 40 In several prospective cohorts of patients, it has been identified as an independent prognostic factor, not related to tumor stage. 41
The association of the negativization or persistence of mutated KRAS in plasma ctDNA from operated patients with colorectal cancer has been previously related to systemic recurrences. It has been proposed as an independent predictor of recurrence. 35 In our study, all patients who had detectable ctDNA postHIPEC (in plasma or peritoneal fluid) relapsed during follow-up, so persistent ctDNA after surgery was a prognostic marker allowing the identification of patients at higher risk of relapse during the postoperative period, as stated previously.19,26 Patients with detectable ctDNA in plasma or peritoneum after treatment showed lower disease-free survival rates (8.3 months versus 35.4 months in those with undetectable ctDNA, all of which were disease-free). Median survival ranged from 22.8 months for patients with positive liquid biopsy (in plasma or peritoneum) to 35.4 months for patients with no detectable ctDNA in any fluid after treatment. Survival rate at 3 years falls from 80% in patients with negative liquid biopsy (there was one death due to postoperative complications in a second non-related procedure) to 33% in those with ctDNA detection (all deaths occurred due to tumor progression) (Figure 5).

Survival in patients with positive liquid biopsy in plasma or peritoneum fluid postHIPEC compared with those without cell-free genetic material from tumor origin detectable after surgery.
Since this is a proof-of-concept study with a limited number of patients, the usefulness of this variable has yet to be determined.
Unanswered questions and limitations
This study presents several limitations. One major drawback is the number of patients included. Initially, we recruited 26 patients who were subjected to CC-HIPEC in our institution during the period of study. However, only KRAS-mutated patients were included in the final cohort (N = 11), as this biomarker could be reliably tracked using commercially available validated ddPCR assays. Other clinically actionable genes and mutations should be included in this screening to increase its prognostic value.
On the other hand, this is a preliminary study including patients with a remarkable heterogeneity in terms of diseases (second look for high risk of PM of colorectal origin, PM of colorectal origin, and pseudomyxoma peritonei). Our study included patients who underwent second-look surgery for high risk of PM (T4, positive cytology at first surgery, perforated tumors, ovarian metastases at primary tumor surgery), rather than established PM. Moreover, we included second-look patients irrespective of the presence of limited liver disease (resectable in the same procedure), as we subsequently performed an independent analysis of plasma liquid biopsy (for systemic disease) and peritoneal liquid biopsy (for local relapse). All patients with resectable liver and peritoneal disease met the criteria established in the literature 42 presenting PCI less than 12 and fewer than three metastasectomies associated with CC-HIPEC.
When we designed the methodology of this project, we decided to include all these different kinds of CC-HIPEC indications as a first approach to test the potential utility of our strategy in the real-life clinical scenarios that we face every day in the routine clinical practice in our institution. And, although preliminary, our results have shown to be very consistent: a direct relationship between liquid biopsy and the risk of relapse has been found in all cases studied. However, the number of patients with each one of these malignancies is insufficient to draw any definite conclusion. Larger studies are needed in order to validate these results.
It should also be noted that the design of the study has not been previously validated. We believe that the timing of the sampling (24/48/72 h) was appropriate for our purposes. However, it may be interesting to maintain surgical drains for more than 1 week and repeat the peritoneal liquid biopsy once the cfDNA rebound caused by cytolysis in the early postoperative period has passed. Further studies should be performed to optimize these protocols.
We aimed to identify variables influencing the negativization or positivization of post-HIPEC liquid peritoneal and plasma biopsy but did not find any significant factors in this group of patients in terms of histology, PCI, drugs or HIPEC indication. One possible explanation for negativization/positivization is that ctDNA might be present in preHIPEC samples, but in such small concentrations that it could not be detected by our methods.
Last but not least, patients with the highest tumor burden (systemic disease, elevated PCI, etc.) and those with poor prognosis (relapses or death) had positive liquid biopsy in blood and peritoneal fluid. This occurred in patients HIPEC 11, HIPEC 17, HIPEC 18, HIPEC 21, HIPEC 24 and presumably HIPEC 19 (48 h postHIPEC plasma not available) and all of them presented systemic/peritoneal relapses. Could this mean that for patients with a high tumor burden, the peritoneum acts only as an exchange barrier? Might ctDNA be detected in any fluid studied in patients with a high tumor burden? These questions are still unanswered.
Conclusions
The usefulness of liquid biopsy in colorectal cancer has been studied extensively in the literature. To our knowledge, this study is the first report of peritoneal liquid biopsy to detect ctDNA in peritoneal fluid in colorectal cancer patients treated by cytoreduction and HIPEC and the first attempt to correlate perioperative changes in its levels with the clinical prognosis.
Even though this is a proof-of-concept study, CC-HIPEC seems to be effective removing free tumor cells from peritoneal fluid, but not neutralizing ctDNA. Poor outcomes were observed in patients who remained or those who became positive for mutant KRAS cfDNA measured in plasma or peritoneal fluid after treatment. All of these patients presented systemic or peritoneal relapse or died, in contrast to those who maintained negative ctDNA or became negative after treatment, remained disease-free after.
Based on this association, the evolution of liquid biopsy in plasma and/or peritoneal liquid after CC-HIPEC could identify those patients at risk of systemic or peritoneal relapse. According to these results, ctDNA in peritoneal fluid and plasma postHIPEC emerges as a prognostic factor for the development of future peritoneal and distant relapses. However, this is a proof-of-concept study including a limited number of patients with heterogeneous pathological conditions. Further studies are required to confirm these early results in a larger sample.
Supplemental Material
sj-pdf-1-tam-10.1177_1758835920981351 – Supplemental material for Liquid biopsy in peritoneal fluid and plasma as a prognostic factor in advanced colorectal and appendiceal tumors after complete cytoreduction and hyperthermic intraperitoneal chemotherapy
Supplemental material, sj-pdf-1-tam-10.1177_1758835920981351 for Liquid biopsy in peritoneal fluid and plasma as a prognostic factor in advanced colorectal and appendiceal tumors after complete cytoreduction and hyperthermic intraperitoneal chemotherapy by Irene López-Rojo-Rojo, Susana Olmedillas-López, Pedro Villarejo Campos, Víctor Domínguez Prieto, Javier Barambio Buendía, Delia Cortés Guiral, Mariano García-Arranz and Damián García-Olmo in Therapeutic Advances in Medical Oncology
Footnotes
Acknowledgements
The authors would like to acknowledge Dr. Ignacio Mahíllo-Fernández for his help on statistical analysis. A special acknowledgment to Dr. Ismael Guijo Castellano and Dr. Alfredo Badía Yébenes for surgical procedures.
Author Contributions
DGO, MGA and SOL, had a continuous access to all of the data in the study and take responsibility for their integrity and the accuracy of the data analysis. Concept and design: DGO, MGA, DCG. Sample acquisition, analysis, or interpretation of data: DCG, ILR, PVC, SOL, VDP, JBB, MGA, DGO. Drafting of the manuscript: ILR, SOL. Critical revision of the manuscript for important intellectual content: ALL AUTHORS.
Conflict of interest statement
ILR and DCG are currently former surgeons of the Department of Surgery, Fundación Jiménez Díaz University Hospital, Madrid, Spain. The authors declare that there is no other conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by a grant from the “FIS-ISCIII-FEDER” [Fondo de Investigaciones Sanitarias-Instituto de Salud Carlos III-Fondo Europeo de Desarrollo Regional (European Regional Development Fund)], Ministry of Health, Spain (grant number PI17/01233).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
