Abstract
Background:
Tailored therapy is crucial for reducing the irrational use of antibiotics and enhancing Helicobacter pylori (H. pylori) eradication rates. Detection of antibiotic resistance gene mutations offers a rapid, accurate, and simple approach for tailored therapy, facilitating its widespread clinical implementation. However, the efficacy, safety, adherence, and cost-effectiveness of tailored therapy guided by genotypic resistance from gastric mucosa samples for first-line H. pylori treatment remain unclear.
Objectives:
This study aims to clarify the efficacy, safety, patient compliance, and cost-effectiveness of genotypic resistance-guided tailored therapy for first-line H. pylori treatment.
Design:
A systematic review and meta-analysis following the PRISMA 2020 guidelines.
Data sources and methods:
The literature was searched in PubMed, Web of Science, and Wanfang databases under the keywords “Helicobacter pylori,” “tailored therapy,” and “genotypic resistance” before March 1, 2025. Two investigators independently performed data extraction and literature quality assessment. Single-arm meta-analysis was performed using the Meta package of the R program to summarize the efficacy, safety, and compliance of tailored therapy, and classical meta-analysis was performed using Review Manager to compare tailored therapy with empirical treatment. Dichotomous outcomes were summarized using relative risks with 95% confidence intervals.
Results:
Of 1524 retrieved articles, 29 (including 4895 patients) met inclusion criteria for single-arm meta-analysis, and 17 (including 6208 patients) were included in the classical meta-analysis by further excluding studies that did not use empirical treatment as control. The single-arm meta-analysis demonstrated satisfactory efficacy of tailored therapy in first-line H. pylori treatment (intention-to-treat analysis: 84.4%, per-protocol analysis: 91.7%), with good safety (adverse event rate: 26.6%) and adherence (98.0%). Tailored therapy showed a higher overall eradication rate than empirical treatment. Subgroup analysis indicated that tailored therapy outperformed standard triple therapy and was comparable to bismuth quadruple therapy and concomitant therapy in terms of efficacy, with similar safety and adherence across groups. Health economic evidence was limited, but most studies suggested that tailored therapy incurred higher costs per successful eradication than empirical treatment.
Conclusion:
Tailored therapy guided by genotypic resistance detected from gastric mucosa samples demonstrates satisfactory eradication efficacy in first-line H. pylori treatment, superior to standard triple therapy and comparable to bismuth quadruple therapy, with comparable safety and adherence. Current evidence supports bismuth quadruple therapy as the optimal first-line treatment for H. pylori infection. Considering factors such as cost-effectiveness and technical accessibility, it is more appropriate to implement tailored therapy in countries and regions with adequate resources and capabilities.
Trial registration:
This study was not registered in PROSPERO, and future systematic reviews and meta-analyses will be registered in advance.
Plain language summary
This is a systematic review and meta-analysis of the efficacy of selecting sensitive antibiotics based on antibiotic resistance information and developing an tailored treatment regimen in patients with initial Helicobacter pylori infection. The results show that tailored therapy can achieve satisfactory efficacy and safety, significantly better than the standard triple therapy, and comparable to the efficacy of the bismuth quadruple therapy and concomitant therapy.
Introduction
Helicobacter pylori infection and associated diseases (gastric cancer, peptic ulcer, chronic gastritis, dyspepsia, etc.) represent significant global health issues. 1 Approximately half of the global population is infected with H. pylori, and numerous guidelines and expert consensus recommend the proactive eradication of H. pylori infection.2,3 Consequently, a substantial number of patients require eradication therapy. However, the rapid and pronounced escalation in antibiotic resistance among H. pylori strains, particularly to clarithromycin, metronidazole, and levofloxacin, has substantially diminished the effectiveness of empirical therapeutic regimens—a consequence inherently linked to empirical treatment strategies. 4 Rational selection of therapeutic agents based on the antibiotic sensitivity of H. pylori strains and the development of individualized treatment regimens can improve eradication efficacy and reduce antibiotic misuse, representing the main direction and trend of future diagnosis and treatment.
Traditional methods of personalized diagnosis and treatment based on H. pylori strain culture and antimicrobial susceptibility testing are difficult to apply widely in clinical practice due to their time-consuming nature, high technical requirements, significant costs, and low success rates. 5 With the rapid development of molecular biology, gene detection technology, and a deeper understanding of the mechanisms of antibiotic resistance in H. pylori eradication therapy, recently emerged genotypic resistance (antibiotic resistance gene mutation) detection offers a simple, rapid, accurate, and cost-effective method for testing antibiotic sensitivity. Current research indicates that obtaining DNA information of H. pylori strains from gastric mucosal samples and detecting their genotypic resistance to clarithromycin and levofloxacin show excellent detection performance and high consistency with phenotypic resistance, suggesting a promising potential for widespread clinical application.6,7 In addition, tailored therapy is more suitable for treatment-naïve patients because the resistance rates to clarithromycin and levofloxacin increase significantly in patients with recurrent H. pylori infection, diminishing the value and significance of detection.8,9 Moreover, a small number of studies have explored the use of stool, gastric juice, and other samples for detecting the genotypic resistance of H. pylori strains, but these methods have yet to be fully validated in terms of DNA extraction and detection consistency.10,11
The inconsistency in existing research results, coupled with high heterogeneity among studies and a lack of comprehensive meta-analyses, makes it unclear how genotypic resistance-guided tailored therapy compares to empirical therapy in terms of efficacy, safety, patient compliance, and cost-effectiveness. This study aims to perform a systematic review and meta-analysis to evaluate the role of genotypic resistance-guided tailored therapy based on gastric mucosal samples in the first-line treatment of H. pylori infection, specifically focusing on its eradication efficacy, safety, patient compliance, and health economic characteristics.
Methods
This systematic review was performed in accordance with the 2020 updated Preferred Reporting Items for Systematic Review and Meta-Analysis (PRISMA) protocol guidelines. 12 See Supplemental Material for the PRISMA item checklist.
Search strategy and inclusion/exclusion criteria
Literature was searched in PubMed, Web of Science, and Wanfang databases using the keywords (“helicobacter pylori” OR “H pylori”) AND (“tailored” OR “personalized” OR “individualized” OR “resistance-guided” OR “susceptibility-guided” OR “PCR-guided”) up to March 1, 2025 by two researchers respectively. See Supplemental Material for a detailed search strategy. Inclusion criteria were: (a) randomized controlled trial (RCTs) or cohort studies comparing genotype resistance-guided tailored therapy with other treatment regimens (for traditional meta-analysis), or single-arm/multi-arm clinical studies of genotype resistance-guided tailored therapy (for single-arm meta-analysis); (b) studies based on gastric mucosal specimens; (c) first-line treatment of H. pylori infection. Exclusion criteria were: (a) tailored therapy based on phenotypic resistance; (b) studies based on specimens other than gastric mucosa (e.g., gastric juice and stool); (c) studies of retreatment or refractory H. pylori infection; (d) reviews, case reports, and other types of articles; (e) articles with insufficient extractable information.
Data extraction
Information was extracted from the included studies independently by two researchers, focusing on the following details: (a) Study source: author, year of publication, country or region; (b) Study content: patient information, H. pylori strain resistance data, methods for detecting resistance genotypes, study type, sample size, eradication regimen, dosage, and duration of treatment; (c) Results: H. pylori eradication rates derived from intention-to-treat (ITT) and per-protocol (PP) analyses, incidence of adverse events, adherence, and health economic information.
Risk of bias assessment
Two researchers (Cailing Li and Kai Zhou) independently assessed the risk of bias in the included studies. In case of discrepancies, a third researcher (Baojun Suo) provided an evaluation. RCTs were assessed using Review Manager software (version 5.4.1), cohort studies were evaluated using the Newcastle-Ottawa Scale, and single-arm studies were evaluated using a methodological index for non-randomized studies (MINORS). 13 No high risk of bias was identified in the included studies. Detailed information can be found in Figure S1 and Tables S1 and S2.
Data analysis
The primary outcome variable of this study was the H. pylori eradication rate, while secondary outcome variables included the incidence of adverse events, patient compliance, and health economic information. Combined analysis of outcome measures was performed using RevMan software (version 5.4.1, The Cochrane Collaboration, 2020, Copenhagen). Relative risks were used to summarize dichotomous outcomes, with corresponding 95% confidence intervals (CIs) provided. Heterogeneity was assessed using I2, with I2 > 50% indicating significant heterogeneity; in such cases, a random effects model was used to pool effect sizes, otherwise a fixed effects model was applied. Single-group eradication rates were combined using the meta package of the R program (version 4.4.3, Vienna). Given the substantial heterogeneity in single-group eradication rates, random effects models were employed to summarize eradication success rates and their respective 95% CIs for both ITT and PP analyses. Meta-analysis results were presented using forest plots. Publication bias was assessed using funnel plots, Egger’s test, and Begg’s test, with asymmetry in the funnel plot or a p-value < 0.05 indicating potential publication bias. Sensitivity analysis was conducted by sequentially excluding each study to evaluate the stability of the synthesized results. The GRADE system was used to assess the evidence quality and the recommendation level.
Results
Literature screening
A total of 1524 studies were retrieved from PubMed, Web of Science, and Wanfang databases up to March 1, 2025. After removing duplicates and studies that did not meet inclusion criteria, 29 studies were selected for a single-arm meta-analysis to summarize the efficacy of tailored therapy. Further exclusion of studies that did not have empirical therapy as a control group resulted in 17 studies being selected for a traditional meta-analysis to compare the efficacy of tailored therapy with empirical therapy. The flowchart is shown in Figure 1. Detailed information on the included studies is provided in Table 1.
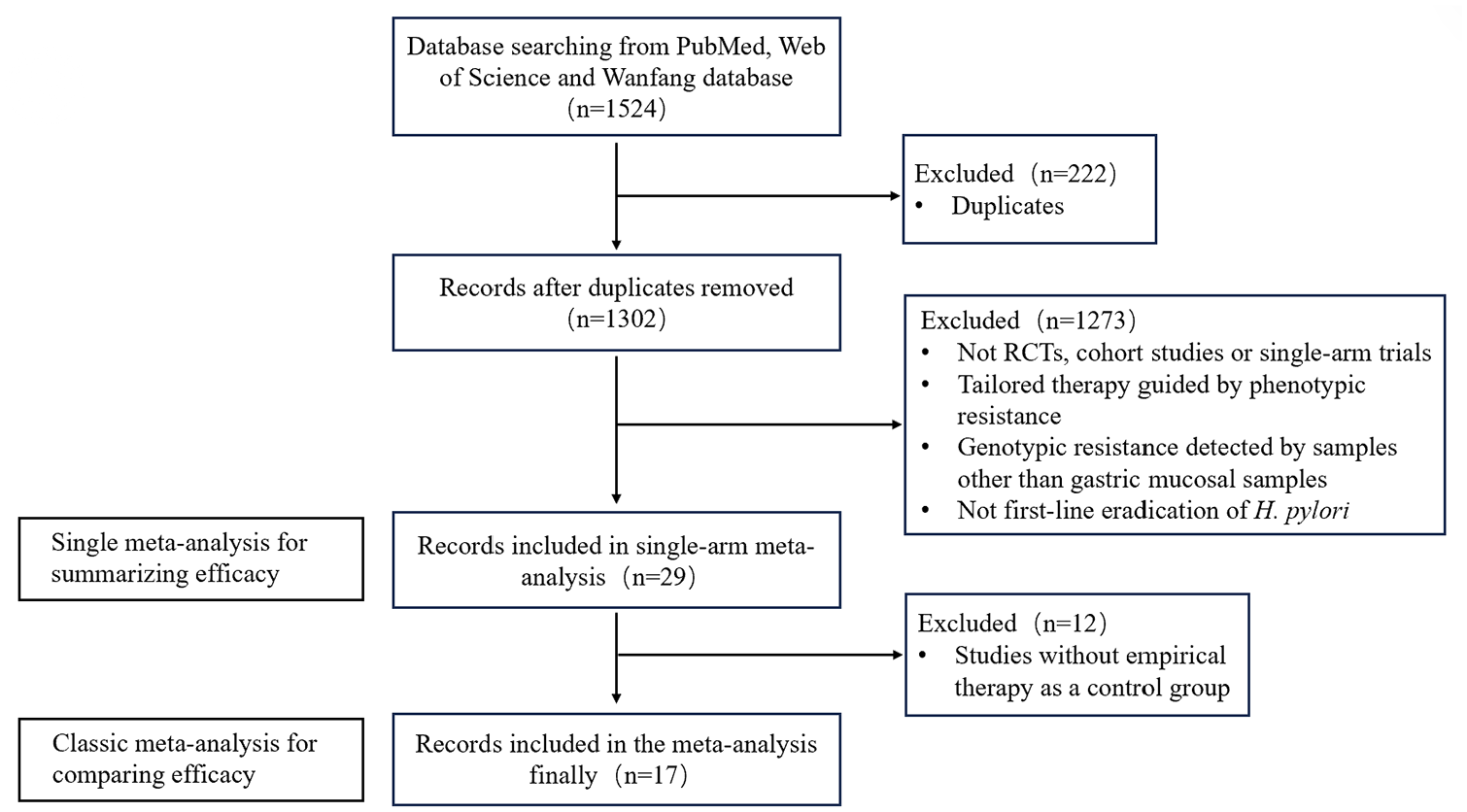
Flow chart of the study.
Characteristics of studies included in the meta-analysis.

AEs, adverse effects; AMO, amoxicillin; BQT, bismuth quadruple therapy; CLA, clarithromycin; CT, concomitant therapy; EPZ, esomeprazole; ET, empirical therapy; FZD, furazolidone; IPZ, ilaprazole; ITT, intention-to-treat; LEV, levofloxacin; LPZ, lansoprazole; MTZ, metronidazole; PP, per-protocol; PPI, proton pump inhibitor; PTZ, pantoprazole; R, resistant.; RCT, randomized controlled trial; RPZ, rabeprazole; S, sensitive; ST, sequential therapy; STT, standard triple therapy; TET, tetracycline; TT, tailored therapy.
Eradication rate analysis
Summary of eradication rates with tailored therapy
A total of 29 studies were included in the single-arm meta-analysis.14 –42 In the ITT analysis, 4895 treatment-naïve H. pylori-infected patients received genotype resistance-guided tailored therapy based on gastric mucosal samples, with 4076 patients achieving successful eradication, resulting in a pooled eradication rate of 84.4% (95% CI 81.9–86.6). In the PP analysis, 4461 treatment-naïve H. pylori-infected patients completed the personalized therapy protocol, with 4044 patients achieving successful eradication, resulting in a pooled eradication rate of 91.7% (95% CI 89.9–93.2), as shown in Figure 2.

The eradication rate of tailored therapy by single-arm meta-analysis (a) ITT analysis and (b) PP analysis.
Among the patient group with genotype-sensitive strains, the eradication regimen incorporating sensitive antibiotics achieved favorable eradication efficacy (91.0%, 95% CI 88.8–92.7). For the patient group with genotype-resistant strains, the eradication regimen was empirically selected, resulting in an eradication rate of 89.0% (95% CI 86.1–91.4). The forest plot is shown in Figure S2.
Among the studies included in the single-arm meta-analysis, 22 studies utilized clarithromycin resistance gene mutation detection alone to guide tailored therapy, encompassing 3811 treatment-naïve H. pylori-infected patients. The ITT analysis showed an eradication success rate of 83.4% (95% CI 80.4–86.1), while the PP analysis showed an eradication success rate of 90.9% (95% CI 89.0–92.5). No studies were found that used levofloxacin resistance gene mutation detection alone to guide tailored therapy. Six studies detected resistance gene mutations for both clarithromycin and levofloxacin, involving 973 treatment-naïve H. pylori-infected patients. The ITT and PP analyses demonstrated eradication success rates of 86.8% (95% CI 84.2–89.0) and 93.1% (95% CI 88.9–95.8), respectively. Forest plots are shown in Figures S3 and S4. In addition, one study detected resistance gene mutations for clarithromycin, levofloxacin, amoxicillin, and tetracycline to guide personalized therapy, achieving eradication rates of 92.8% (ITT analysis) and 98.1% (PP analysis) in first-line H. pylori treatment. 27
Comparison of eradication rates between tailored and empirical therapy
After excluding studies without an empirical therapy control group, 17 studies were included in the traditional meta-analysis, encompassing 6208 treatment-naïve H. pylori-infected patients. Among these, 2842 patients received genotype resistance-guided tailored therapy, and 3366 patients received empirical therapy. Overall, tailored therapy demonstrated superior efficacy compared to empirical therapy in first-line H. pylori treatment (ITT analysis: 82.6% vs 74.1%, RR 1.09, 95% CI 1.03–1.16, p = 0.006; PP analysis: 89.7% vs 81.3%, RR 1.09, 95% CI 1.03–1.15, p = 0.003). This study included both RCTs and cohort studies, with 13 RCTs and 4 cohort studies. Further subgroup analysis based on study type revealed that in the RCT subgroup, tailored therapy demonstrated significantly superior efficacy compared to empirical therapy (ITT analysis: 81.4% vs 73.6%, RR 1.10, 95% CI 1.02–1.18, p = 0.01; PP analysis: 89.0% vs 79.7%, RR 1.10, 95% CI 1.03–1.17, p = 0.002), whereas in the cohort study subgroup, the difference in efficacy between the two approaches was not statistically significant(ITT analysis: 86.4% vs 75.9%, RR 1.08, 95% CI 0.95–1.23, p = 0.26; PP analysis: 91.8% vs 86.4%, RR 1.06, 95% CI 0.93–1.21, p = 0.40). The forest plot is shown in Figure 3.

The eradication rate of tailored therapy versus empirical therapy (a) ITT analysis and (b) PP analysis.
Subgroup analysis based on empirical therapeutic regimens
In the included studies, the empirical treatment group consisted of various regimens, primarily including standard triple therapy, bismuth quadruple therapy, and concomitant therapy. Subgroup analysis was conducted to compare each of these regimens with tailored therapy. The results showed that the efficacy of tailored therapy was significantly superior to that of standard triple therapy, while no significant difference was observed when compared with bismuth quadruple therapy and concomitant therapy. Details are as follows, and the forest plot is shown in Figure 4.

Subgroup analysis of the eradication rate of tailored therapy versus empirical therapy (a) ITT analysis and (b) PP analysis.
Tailored therapy versus triple therapy
Six studies compared the efficacy of genotype resistance-guided tailored therapy with the standard triple therapy (PPI + amoxicillin + clarithromycin), encompassing 2381 treatment-naïve H. pylori-infected patients. Of these, 1046 patients received tailored therapy, and 1335 patients received standard triple therapy. In first-line treatment of H. pylori infection, tailored therapy guided by resistance genotype detection from gastric mucosal specimens was superior to standard triple therapy (ITT analysis: 84.0% vs 66.3%, RR: 1.25, 95% CI 1.18–1.32, p < 0.001; PP analysis: 90.1% vs 74.2%, RR: 1.22, 95% CI 1.17–1.27, p < 0.001).
Tailored therapy versus bismuth quadruple therapy
Six studies compared the efficacy of genotype resistance-guided tailored therapy with bismuth quadruple therapy (PPI + bismuth + two antibiotics), encompassing 2010 treatment-naïve H. pylori-infected patients. Among these, 1011 patients received tailored therapy, and 999 patients received empirical bismuth quadruple therapy. In first-line treatment of H. pylori infection, the efficacy of tailored therapy was comparable to bismuth quadruple therapy, with no statistically significant difference (ITT analysis: 80.1% vs 79.2%, RR: 1.01, 95% CI 0.92–1.10, p = 0.88; PP analysis: 88.4% vs 85.7%, RR: 1.02, 95% CI 0.93–1.13, p = 0.65).
Tailored therapy versus concomitant therapy
Four studies compared the efficacy of genotype resistance-guided tailored therapy with concomitant therapy, including 1145 patients. Of these, 576 patients received tailored therapy, and 569 patients received concomitant therapy. In first-line treatment of H. pylori infection, the efficacy of tailored therapy was comparable to concomitant therapy, with no statistically significant difference (ITT analysis: 81.9% vs 83.0%, RR: 0.99, 95% CI 0.94–1.04, p = 0.64; PP analysis: 91.5% vs 91.0%, RR: 1.01, 95% CI 0.97–1.06, p = 0.55).
Subgroup analysis based on tailored therapeutic regimens
In the included studies, the tailored treatment group also comprised various regimens, particularly for patients with genotypic resistance to antibiotics, where the choice of eradication therapy often relied on the physician’s experience. These regimens can be categorized into bismuth quadruple therapy, triple therapy, and dual therapy. Subgroup analysis was conducted based on the different regimens selected for genotypic-resistant patients in the tailored treatment group, aiming to provide references for regimen selection and explore sources of heterogeneity. The results showed that when quadruple therapy or PPI-amoxicillin dual therapy was used for patients with genotypic resistance, the efficacy of tailored therapy was superior to that of empirical treatment. However, when triple therapy was used for genotypic-resistant patients, the efficacy of tailored therapy was comparable to that of empirical treatment. For detailed analysis, see below, and the forest plot is shown in Figure S5.
Tailored quadruple therapy versus empirical therapy
A total of 12 articles (including 13 studies) utilized quadruple therapy for patients with genotypic resistance in the tailored treatment group. Subgroup analysis revealed that when quadruple therapy was used for genotypic-resistant patients, the efficacy of the tailored therapy was superior to that of the empirical treatment (ITT analysis: 81.9% vs 74.3%, RR 1.08, 95% CI 1.01–1.16, p = 0.03; PP analysis: 89.2% vs 82.3%, RR 1.08, 95% CI 1.01–1.16, p = 0.03).
Tailored triple therapy versus empirical therapy
A total of four articles utilized triple therapy for patients with genotypic resistance in the tailored treatment group. Subgroup analysis showed that when triple therapy was used for genotypic-resistant patients, there was no significant difference in efficacy between the tailored treatment group and the empirical treatment group (ITT analysis: 81.8% vs 74.4, RR 1.06, 95% CI 0.95–1.18, p = 0.29; PP analysis: 89.5% vs 80.5%, RR 1.08, 95% CI 0.98–1.19, p = 0.12).
Tailored dual therapy versus empirical therapy
Only one study (Furuta 2007) utilized dual therapy for patients resistant to clarithromycin, and the efficacy of the tailored treatment group was significantly superior to that of empirical triple therapy (ITT analysis: 96.0% vs 70.0%; PP analysis: 96.6% vs 72.9%).
Safety analysis
Among the included studies, 18 publications provided data on the incidence of adverse events associated with tailored therapy in treatment-naïve H. pylori-infected patients, involving a total of 3221 patients. Of these, 1090 patients experienced treatment-related adverse events, resulting in a pooled adverse event incidence rate of 26.6% (95% CI 17.1–38.8). This is illustrated in Figure 5. Fourteen studies compared the safety (adverse event incidence) of tailored therapy versus empirical therapy, with meta-analysis results showing no statistically significant difference between the two (32.2% vs 31.1%, RR: 0.93, 95% CI 0.74–1.17, p = 0.56). The forest plot is shown in Figure 6.

The incidence of adverse reactions of tailored therapy by single-arm meta-analysis.
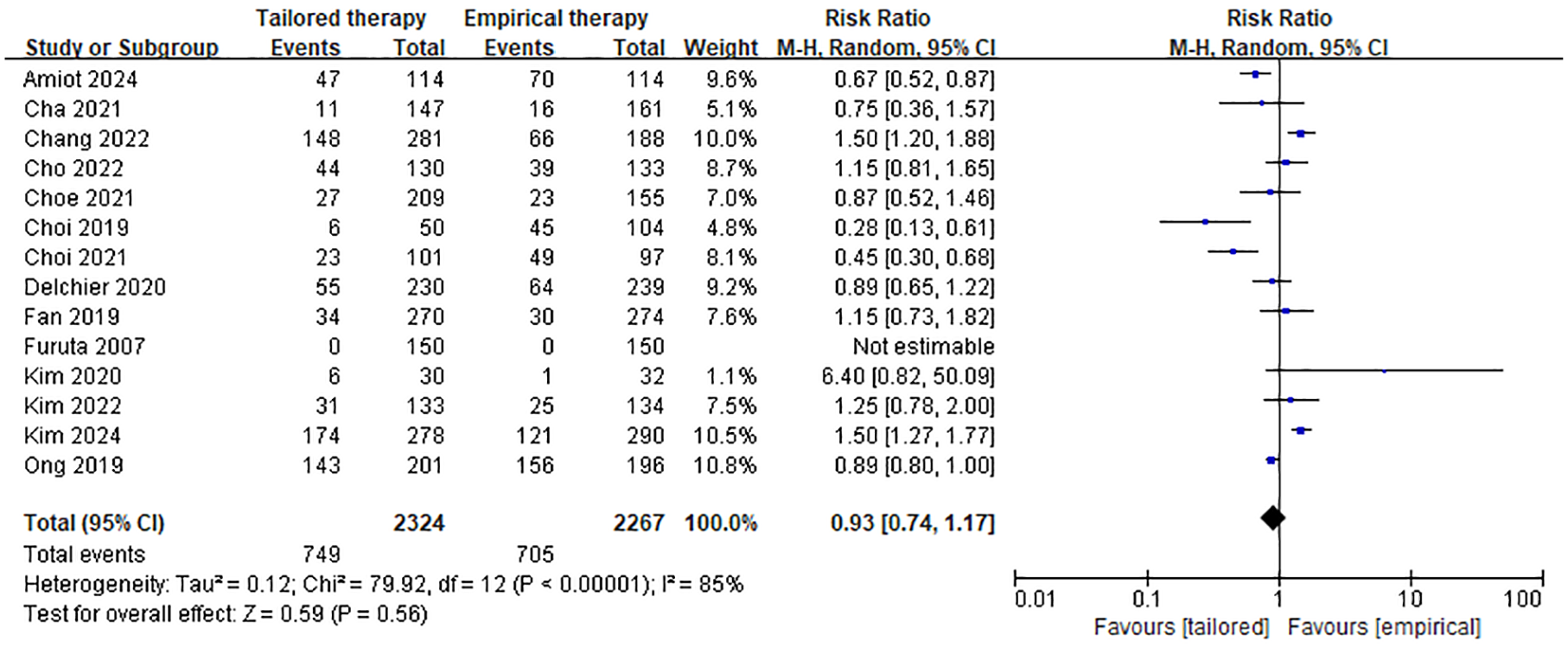
The incidence of adverse reactions to tailored therapy versus empirical therapy.
Compliance analysis
Eighteen publications provided data on patient compliance with tailored therapy in treatment-naïve H. pylori-infected patients, involving a total of 2752 patients, of whom 2678 demonstrated good compliance. Using a random effects model analysis, the pooled compliance rate for genotype resistance-guided tailored therapy in first-line H. pylori treatment was 98.0% (95% CI 96.5–98.8). This is illustrated in Figure 7. Nine studies compared patient compliance between tailored therapy and empirical therapy, with no statistically significant difference, found (98.0% vs 97.3%, RR: 1.01, 95% CI 0.99–1.02, p = 0.28). The forest plot is shown in Figure 8.
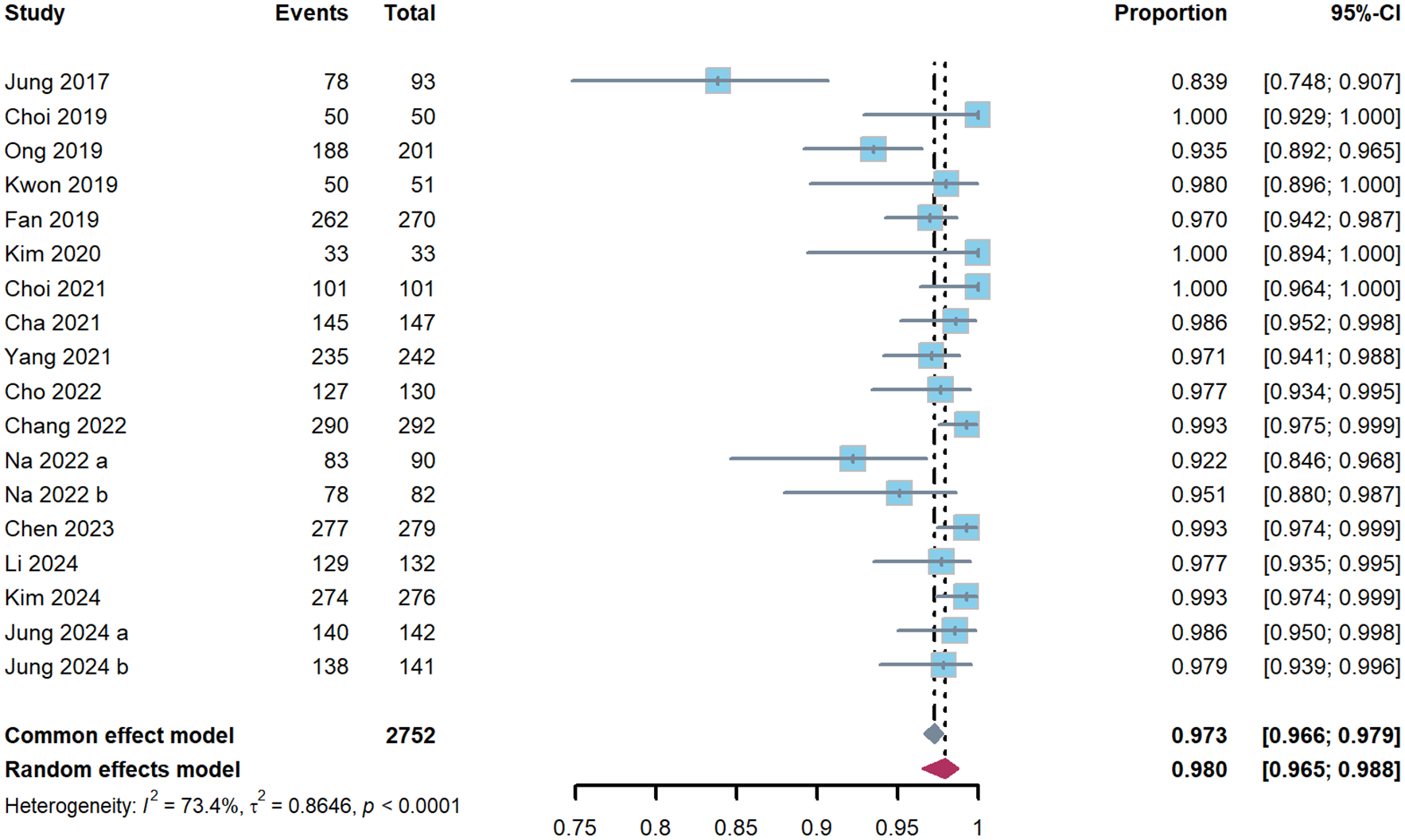
The patient compliance with tailored therapy by single-arm meta-analysis.

The patient compliance with tailored therapy versus empirical therapy.
Cost-effectiveness analysis
Nine studies compared the cost-effectiveness of tailored therapy versus empirical therapy from a health economics perspective, evaluating the cost per successful eradication of an H. pylori infection. Six studies indicated that the cost per successful eradication was higher for tailored therapy compared to empirical therapy, while only three studies demonstrated better cost-effectiveness for tailored therapy. Detailed results are presented in Table 1.
Publication bias analysis and sensitivity analysis
Funnel plots were constructed for the included studies, illustrating asymmetry in the pooled H. pylori eradication rates for both ITT and PP analyses. In addition, both Begg’s and Egger’s tests yielded p-values <0.05, indicating the presence of publication bias. Sensitivity analysis, performed by sequentially removing individual studies, showed no significant changes in the eradication rates and their 95% CIs, suggesting the robustness of the results (see Figures S6 and S7).
Discussion
To reduce the irrational use of antibiotics and improve the eradication rates of H. pylori infection, tailored therapy guided by molecular biology techniques for detecting genotypic resistance is the future direction. The detection of resistance gene mutations to clarithromycin and levofloxacin in H. pylori strains from gastric mucosal samples to guide personalized therapy shows good feasibility and application prospects. A comprehensive literature search indicates that current studies on tailored therapy are still limited, mostly originating from Asian regions such as Korea, Japan, and China, with fewer studies from Europe and North America. Large-scale, multicenter, high-quality RCTs are particularly scarce, necessitating further research to explore and validate the optimal regimen and efficacy of tailored therapy. Most of the existing research focuses on clarithromycin and levofloxacin, with good concordance between genotypic and phenotypic resistance. 43 Some studies have also investigated resistance gene mutation for other antibiotics. However, due to factors such as resistance mechanisms beyond gene mutations, scattered mutation sites, and low resistance rates, the correlation between genotypic and phenotypic resistance for metronidazole, tetracycline, and rifabutin remains poor or has not yet been definitively confirmed. 44
Single-arm meta-analysis indicates that genotypic resistance-guided tailored therapy in first-line H. pylori treatment yields pooled eradication rates of 84.4% and 91.7% based on ITT and PP analyses, respectively. According to the eradication efficacy standards proposed by Professor Graham, the PP analysis results can be classified as “good.” 45 For genotype-sensitive strains, most studies employed a 7-day course, while some used 10-day or 14-day courses, achieving a pooled eradication rate of 91.0%, suggesting good eradication efficacy for tailored therapy. In genotype-resistant patients, the eradication rate was 89.0%, largely dependent on the efficacy of the chosen empirical regimen. Careful comparison of various studies reveals higher eradication rates in genotype-resistant patients when using classical bismuth quadruple regimens,20,22,26,29,34 PPI-amoxicillin dual therapy, 14 or bismuth quadruple regimens containing sensitive antibiotics like furazolidone.16,32 Conversely, lower eradication rates were observed with triple therapy, particularly when using antibiotics with high resistance rates like metronidazole.17,21,25 This underscores the importance of considering local resistance patterns, drug availability, and patient medication history when selecting appropriate empirical treatment regimens for clarithromycin and levofloxacin-resistant cases to achieve satisfactory eradication outcomes. Subgroup analysis based on resistance detection sites shows that tailored therapy targeting only clarithromycin resistance genes achieved a pooled eradication rate of 90.9% while targeting both clarithromycin and levofloxacin resistance genes achieved a pooled eradication rate of 93.1%. In addition, a study that tested for resistance to clarithromycin, levofloxacin, amoxicillin, and tetracycline simultaneously achieved an eradication rate of 98.1%. 27 This indicates that increasing the number of drug resistance detection points can improve eradication rates to some extent. In clinical practice, detecting more drug targets allows more patients to benefit from tailored therapy with sensitive antibiotics. A single-arm study found that 52.3% of patients had clarithromycin-sensitive genotypes, and among clarithromycin-resistant patients, 61.9% had levofloxacin-sensitive genotypes. Concurrent detection of clarithromycin and levofloxacin resistance genotypes enabled 81.8% of patients to receive truly personalized therapy. 39 Future research on the resistance mechanisms of commonly used antibiotics and the development of more drug detection targets are expected to further enhance the efficacy of tailored therapy.
Through a classic meta-analysis that included both RCTs and cohort studies comparing tailored therapy with empirical therapy, it was found that genotypic resistance-guided tailored therapy based on gastric mucosal specimen detection exhibited superior eradication efficacy in first-line H. pylori treatment compared to empirical treatment regimens. Subgroup analysis based on different empirical treatment regimens revealed that tailored therapy was superior to triple therapy, while no statistically significant differences were found in eradication rates when compared with bismuth quadruple therapy and concomitant therapy. The following factors may explain these findings: (a) In some studies, patients with genotype-sensitive strains in the tailored therapy group received shorter treatment durations, whereas genotype-resistant patients and those in the empirical therapy group received longer treatment durations overall20,31,35,36; (b) in the tailored therapy group, genotype-resistant patients often received empirically selected treatment regimens, with some studies using triple therapy containing metronidazole, which is less effective, thus lowering the overall eradication success rate of the tailored therapy group21,25; (c) the eradication efficacy in the empirical therapy group largely depended on the chosen treatment regimen, with some studies opting for highly effective classical quadruple therapy, making it difficult for tailored therapy to demonstrate a clear advantage.20,30,36 The classical bismuth quadruple regimen is difficult to use widely, given the poor accessibility of tetracycline medications, larger antibiotic dosages, and more adverse effects. If tailored treatment regimens fully consider resistance genotype information, regional resistance patterns, and patients’ medication history, more satisfactory eradication rates may be achieved. In terms of safety and patient adherence, there were no significant differences between tailored therapy and empirical therapy. Despite differing treatment modalities, the selected drug types, dosages, and durations were somewhat similar, and both approaches showed satisfactory safety and adherence, making it difficult to detect meaningful statistical differences. Analyzing genotype-sensitive patients separately might yield better safety and adherence outcomes due to fewer drug types and shorter treatment durations, but relevant data could not be extracted from existing studies. In terms of cost-effectiveness, few studies addressed this aspect, with most indicating that the cost per successfully eradicated H. pylori infection was higher in the tailored therapy group compared to the empirical therapy group. This is likely due to the additional costs associated with resistance gene mutation detection. As molecular biology techniques such as PCR become more widespread, detection costs decrease, and as tailored regimens are further optimized to achieve higher eradication rates, tailored therapy may achieve better cost-effectiveness in the future.
Currently, several meta-analyses have been conducted to summarize the efficacy of tailored therapy in H. pylori infection. However, existing meta-analyses still have limitations. Some studies combine tailored therapy based on phenotypic resistance and genotypic resistance for analysis, but phenotypic resistance is difficult to apply widely in clinical practice.46,47 Two recent meta-analyses reviewed the application of genotypic resistance-guided tailored therapy in H. pylori infection. A single-arm meta-analysis indicated that the eradication rates of tailored therapy were 86.9% in ITT analysis and 91.5% in PP analysis, with a PP eradication rate of 92.0% in treatment-naïve patients. This suggests that tailored therapy guided by resistance gene mutation detection can achieve good efficacy in the first-line treatment of H. pylori infection; however, as a single-arm meta-analysis, this study did not compare the effectiveness of tailored therapy with empirical therapy, nor did it address the safety, patient adherence, or cost-effectiveness evaluation of tailored therapy. 48 Another meta-analysis compared the efficacy of tailored therapy with empirical therapy, showing that tailored therapy was more effective than the empirical triple regimen but less effective than the empirical quadruple regimen. 49 However, these two meta-analyses included studies on tailored therapy guided by resistance gene mutation detection from gastric mucosa, gastric juice, and stool samples in both treatment-naïve and previously treated patients. Given that resistance rates to antibiotics such as clarithromycin and levofloxacin are significantly higher in previously treated and refractory patients, the cost-effectiveness of detecting resistance in these patients is lower. 9 Furthermore, the effectiveness of extracting H. pylori DNA and detecting resistance gene mutations from gastric juice and stool samples is not as reliable as from gastric mucosal specimens.10,11 Therefore, this study focuses on the application of tailored therapy guided by resistance gene mutation detection from gastric mucosal samples in the first-line treatment of H. pylori infection. Unlike existing meta-analyses, this study simultaneously incorporates single-arm studies, cohort studies, and RCTs. It conducts both a single-arm meta-analysis to summarize the efficacy, safety, and patient compliance of tailored therapy and a classic meta-analysis to compare the efficacy, safety, and adherence of tailored therapy versus empirical therapy. In addition, the literature search has been updated to March 2025, including more studies to demonstrate the advantages of tailored therapy from multiple dimensions.
This study has several limitations: (a) the current number of studies on genotypic resistance-guided tailored therapy based on gastric mucosal specimens for first-line treatment of H. pylori infection is limited. There is a lack of large-sample, high-quality, multicenter RCTs, and significant heterogeneity exists among the existing studies. Further research is needed to explore the efficacy of tailored therapy; (b) according to the available studies, tailored therapy has not yet demonstrated better cost-effectiveness, necessitating further evaluation of its health economics; (c) after obtaining resistance genotype information, there is no optimal recommendation for tailored treatment regimens (including drug selection, dosage, and duration). Future understanding of the resistance mechanisms of other commonly used antibiotics may enable the detection of more drug targets, allowing more patients to receive personalized therapy with sensitive antibiotics. In genotype-resistant patient populations, it is also essential to consider local resistance patterns, previous medication history, and drug availability to select the appropriate regimen, thereby maximizing the eradication efficacy of tailored therapy. (d) I have evaluated the evidence quality and recommendation level using the GRADE system. Although the included studies were primarily RCTs, and no high risk of bias was identified using the risk of bias assessment tool, significant heterogeneity was observed among the studies. In addition, publication bias was detected through funnel plots, Egger’s test, and Begg’s test. Therefore, further research is likely to alter the effect estimates, resulting in an evidence-quality assessment of “low” and a recommendation level of “weak recommendation.”
Overall, this study summarizes the efficacy, safety, and patient compliance of genotype resistance-guided tailored therapy based on gastric mucosal samples in first-line treatment of H. pylori infection through single-arm meta-analysis, suggesting satisfactory outcomes in these domains. Additionally, the classic meta-analysis comparing tailored therapy with empirical therapy in first-line H. pylori treatment indicates that tailored therapy is more effective than empirical therapy, particularly superior to empirical triple therapy while showing no significant differences in safety and adherence. Genotypic resistance-guided tailored therapy based on gastric mucosal samples holds promise for first-line treatment of H. pylori infection; however, given cost-effectiveness challenges and limited access to molecular diagnostics in resource-limited settings, this approach is more appropriately prioritized in developed regions with established infrastructure.
Conclusion
Tailored therapy guided by genotypic resistance detection from gastric mucosal samples demonstrates satisfactory eradication efficacy in first-line H. pylori treatment and is superior to standard triple therapy and comparable to bismuth quadruple therapy and concomitant therapy. Tailored therapy and empirical therapy have similar safety and patient compliance. There are few studies involving health economic evaluations, and existing research has not demonstrated that tailored therapy is more cost-effective than empirical therapy. Current evidence supports bismuth quadruple therapy as the optimal first-line treatment for H. pylori infection. Considering factors such as cost-effectiveness and technical accessibility, tailored therapy is better suited for implementation in regions with established medical infrastructure and resources.
Supplemental Material
sj-docx-1-tag-10.1177_17562848251348826 – Supplemental material for Tailored therapy guided by genotypic resistance from gastric mucosa samples in the first-line treatment of Helicobacter pylori infection: a systematic review and meta-analysis
Supplemental material, sj-docx-1-tag-10.1177_17562848251348826 for Tailored therapy guided by genotypic resistance from gastric mucosa samples in the first-line treatment of Helicobacter pylori infection: a systematic review and meta-analysis by Cailing Li, Kai Zhou, Baojun Suo, Yanyan Shi, Guangjie Ping, Liya Zhou and Zhiqiang Song in Therapeutic Advances in Gastroenterology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
