Abstract
Background:
Ukraine remains a high World Health Organization priority country for drug-resistant tuberculosis (TB). Rifampicin-resistant TB (RR-TB) has a more protracted, more complicated, and more expensive treatment. In 2021, Ukraine reported 4025 RR-TB cases – 5.4 times more (751) than all 30 European Union/ European Economic Area countries together.
Objectives:
The objective of the study was to determine the diagnostic accuracy of line probe assay (LPA), AID Autoimmun Diagnostika GmbH, for detecting resistance to anti-TB drugs and its clinical application for selecting treatment regimens.
Design:
A prospective observational cohort study.
Methods:
From May 2019 to June 2020, we consecutively enrolled patients with active TB hospitalized at the Regional Phthisiopulmonology Center (Vinnytsia, Ukraine), aged between 18 and 82 years. The LPA was performed in the Genetic Research Laboratory at National Pirogov Memorial Medical University, Vinnytsia, Ukraine.
Results:
A total of 84 clinical specimens and 97 culture isolates from 126 TB patients were tested during the study. Accuracy (95% confidence interval) of LPA for clinical samples in comparison with phenotypic drug susceptibility test (DST) was 80.1 (68.5–89.0) for isoniazid (H), 74.7 (62.4–84.6) for rifampicin (R), 74.4 (62.5–84.1) for ethambutol, 71.4 (41.9–91.6) for streptomycin, 84.6 (62.4–96.5) for prothionamide/ethionamide, and 84.6 (73.6–92.3) for levofloxacin (Lfx), respectively. We found a significantly higher sensitivity of LPA for H, R, and Lfx for the culture isolates compared to clinical specimens (p < 0.05). LPA detected different mutations in 6 out of 17 (35.5%) patients susceptible to R by Xpert. A shorter treatment regimen with an injectable agent demonstrated a low suitability rate of 5% (8/156) in a cohort of RR-TB patients from Ukraine.
Conclusion:
Initial LPA testing accurately identifies resistance to anti-TB drugs and facilitates the selection of an appropriate treatment regimen, minimizing exposure to empirical therapy.
Plain language summary
Why was the study done? Ukraine has over 4000 patients with tuberculosis (TB) resistant to at least one drug (rifampicin) - five times that of all 30 European Union/European Economic Area countries combined. Unfortunately, only about 60% of such patients have been successfully treated in 2019. At that time, the majority of people suffering from tuberculosis in Ukraine, after checking resistance to rifampicin, initially received standard combinations of the first-line or second-line anti-TB medicines before the result of traditionally used tests (usually few weeks later) became available to individualize the treatment. Alternatively, the sputum could be transported to some overloaded reference laboratories located hundreds of km away from the treatment places.
What did the researchers do? The INNOVA4TB team implemented rapid diagnostics of drug resistance in routine practice, guiding key antibiotics use in TB patients. A total of 181 samples from 126 individuals were tested during 2019-2020.
What did the researchers find? This new diagnostic technology accurately detected resistance to 9 anti-TB drugs in sputum samples. It could be helpful to select appropriate TB treatment regimens, reducing time for decision from 1 month up to 2 days. Recommended at the study time 9-month shorter standardized treatment regimen with injectable agent was suitable only for 5% of patients for whom it was indicated in Vinnytsia region of Ukraine.
What do the findings mean? The study has demonstrated successful implementation of the new molecular diagnostic technology from scratch in a country with restricted resources and limited TB laboratory capacity. This test can facilitate optimal distribution of available wards among patients with different profiles of resistance and correct choice between treatment options.
Introduction
Tuberculosis (TB) remains the deadliest infectious disease in many developing countries, 1 despite being preventable and curable. Recent global progress in the fight against TB has been due to the accelerating implementation of rapid diagnostic methods, 2 new anti-TB drugs, and promising shorter treatment regimens. 3 However, first the COVID-19 pandemic and then the war in Ukraine have reversed these advances. Nowadays, Europe faces one of the biggest waves of migration in history, hosting almost 8 million Ukrainians while there are an estimated 6.5 million internally displaced people within Ukraine. 4 Undoubtedly, the destruction of healthcare facilities throughout Ukraine, combined with war-initiated humanitarian and economic disasters, will have a cumulative effect on TB prevalence beyond Ukraine itself. 5 To prevent new TB outbreaks in these circumstances, it is essential to develop laboratory capacity for accurate TB diagnosis and timely detection of drug resistance.
Multidrug-resistant tuberculosis (MDR-TB) is defined as having resistance documented by a drug susceptibility test (DST) to at least isoniazid (H) and rifampicin (R). In contrast, extensively drug-resistant TB (XDR-TB) is defined as additional resistance of Mycobacterium tuberculosis to fluoroquinolones (FQ) and at least either linezolid (Lzd) or bedaquiline (Bdq). 6 Although monoresistance to R is a rare phenomenon, we also operate with the term rifampicin-resistant TB (RR-TB) in situations when DST to other drugs was not performed or when results are still pending. The previously widely used second-line injectables (SLIs) capreomycin (Cm) and kanamycin (Km) are no longer recommended for the treatment of M/XDR-TB due to an increased risk of unfavorable treatment outcomes. 7 Currently, only two aminoglycosides, amikacin (Am) and streptomycin (S) can be administered to a limited number of patients with M/XDR-TB who cannot be treated with entirely oral regimens.
Ukraine is ranked as one of the countries with the highest M/XDR-TB burdens. 1 In 2021, there were reported 4025 RR-TB cases – in 5.4 times more (751) than all 30 European Union/ European Economic Area countries together. 8 In 2022, Ukraine’s RR-TB incidence was firstly increased since many years corresponding to 9.5 per 100,000 population. 9 However, patients with TB have limited access to next-generation sequencing for a DST, and only 4 out of 34 (12%) third-level TB microbiological laboratories have World Health Organization (WHO)-recommended line probe assay (LPA) technology GenoType MTBDRsl (Hain Lifescience GmbH, Nehren, Germany). Shipment of clinical samples to reference laboratories with LPA delays test results, particularly during irregular supply of reagents and consumables. Since 2021, the GeneXpert MTB/XDR cartridges (Cepheid, Sunnyvale, CA, USA) have been used in Ukraine for the prediction of resistance to H, FQ [levofloxacin (Lfx) and moxifloxacin (Mfx)], Am, and prothionamide/ethionamide (Pt/Et). However, programmatic assessment of this assay in routine practice and comparison with LPA has yet to be performed in Ukraine.
Conventional methods for culture isolation of M. tuberculosis from clinical samples and phenotypic DST (pDST) using critical concentrations (CC) to anti-TB drugs have a long turnaround time (TAT). It is worth mentioning that patients with rpoB mutations detected by Xpert MTB/RIF usually shared ward space with other individuals with pending RR-TB results for pDST to other anti-TB drugs. With growing knowledge about genetic mutations, 10 and the development of the WHO Consolidated Guidelines on Tuberculosis 11 describing clinical interpretation of molecular assay results, the genetic DST (gDST) was intended as an initial test for aiding treatment selection. Several studies analyzed the specificity and sensitivity of LPA in clinical samples,12–15 but the clinical utility of this diagnostic tool for recruitment of patients for a 9–11-month short treatment regimen with an injectable agent (4–6Km-Mfx-Pt-Cfz-Z-Hhd-E/5Mfx-Cfz-Z-E) as was recommended by the WHO 16 is still unknown.
For this purpose, we have conducted a prospective observational cohort study (ClinicalTrials.gov Identifier: NCT04002869) with the primary objective of evaluating the diagnostic accuracy of LPA (AID Autoimmun Diagnostika GmbH, Strassberg, Germany) for detecting resistance to anti-TB drugs. We have performed a large-scale within-study comparison of LPA testing on clinical specimens and culture isolates from the same cohort of patients. The second objective of our study was to analyze the clinical utility of initial LPA testing as a decision-making tool to select treatment regimens for patients with RR-TB according to the current WHO recommendations for clinical practice at the time of our study, which are individualized longer 18–20-month regimens consisting of old or new/repurposed medicines 17 and standardized shorter (9–11 month) regimens with an injectable agent. 18 We also studied whether the simultaneous presence of rpoB, katG, and inhA mutations in M. tuberculosis isolates could be associated with FQ resistance in our cohort. 19
Methods
Study design and setting
This prospective observational cohort study evaluated the diagnostic accuracy of the LPA testing of 181 samples (84 clinical specimens and 97 culture isolates) compared to a reference standard (pDST) at the Genetic Research Laboratory at National Pirogov Memorial Medical University (Vinnytsia, Ukraine).
The study results were reported by the ‘Standards for Reporting Diagnostic Accuracy Studies’ 20 (see Supplemental Checklist).
Study population
From May 2019 to June 2020, we enrolled patients with active TB hospitalized at a TB dispensary (Vinnytsia, Ukraine) aged between 18 and 82 years. The main eligibility criteria were a positive sputum Xpert test with M. tuberculosis complex (MTBC) DNA detected and voluntary willingness to participate in the study. The patients were consecutively recruited into the study until the desired sample size was reached. Individuals with a negative culture result and no pDST for comparison with LPA were retrospectively excluded from the analysis.
Study procedures
Morning sputum specimens were collected in an isolated room following infection control precautions and were sent to the laboratory for further processing. Sputum collection was performed within the first week after admission to minimize the effect of treatment on bacterial load and cough productivity. If the person had been on treatment for more than 7 days before agreeing to participate in the study, or we could not organize sputum collection, then only culture isolate, not a direct clinical sample, was collected for LPA testing. Only spontaneously expectorated sputum samples were collected. However, one pleural fluid specimen was also included. A standard procedure using N-acetyl-L-cysteine and NaOH was used for decontamination and homogenization of sputum specimens before they were routinely analyzed by smear microscopy, culture on solid medium Löwenstein–Jensen and liquid medium with BACTEC Mycobacteria Growth Indicator Tube (MGIT) 960 system (Becton Dickinson Diagnostic Systems, Sparks, MD, USA), and by Xpert MTB/RIF platform (Cepheid). LPA analysis was performed on the clinical isolates and on the decontaminated sputum samples. Phenotypical DST based on CC was performed on positive cultures following the WHO guidelines and national protocols. 21 The National TB Reference Laboratory performed regular external quality controls on the regional microbiology laboratory.
LPAs are reverse-hybridization DNA strip-based tests involving the following consecutive steps: DNA extraction from biological sample, the polymerase chain reaction (PCR) amplification followed by reverse hybridization with sequence-specific oligonucleotide probes and visual analysis of the nitrocellulose strip recording detection of signals of wild-type (WT) probes and/or probes corresponding to the most common mutations (MUT). Each clinical specimen (irrespective of smear microscopy status) and culture isolate were tested for resistance to first- and second-line drugs. All steps of LPA testing were performed following the manufacturer’s instructions and supervised by a staff member with appropriate skills in molecular assays. All staff involved in the molecular testing underwent training in conducting LPA procedures.
DNA isolation
Mycobacterial DNA extraction was done from 500 µL aliquot of each specimen using commercially available extraction kits, QIAamp DNA Blood Mini Kit (Qiagen, Hilden, Germany), and AID Smart Lysis Tool (AID/GenID GmbH, Straßberg, Germany) for culture isolates and decontaminated sputum samples, respectively.
Multiplex PCR amplification and reverse hybridization
A multiplex PCR was performed using DNA isolates with specific primers and nucleotide mixes for amplification in each diagnostic module. Following PCR, the respective biotinylated amplicons were identified by a hybridization reaction with oligonucleotide probes corresponding to a specific wild-type or/and mutation at nucleic acid positions and controls immobilized on a nitrocellulose membrane in a distinctive line format. Due to washing stage, the hybrids between amplicons and gene probes are only immobilized if the sequence of the probe is totally complementary to amplified DNA. The color reaction, as a result of binding of streptavidin-coupled alkaline phosphatase with hybrids, was then visually detected and interpreted.
Three commercially available LPA modules (strips) manufactured by AID Autoimmun Diagnostika GmbH to identify MTBC in clinical samples and its resistance to the main anti-TB drugs. Every module includes conjugate, amplification (glyceraldehyde-3-phosphate dehydrogenase gene for DNA quantity and quality), Mycobacterium universal, and MTBC controls. M. tuberculosis strain H37Rv and PCR-grade water were used as a positive and negative control samples, respectively.
The first module (MDR-TB) is used to detect WT and specific mutations associated with H (katG and inhA promoter), R (rpoB), and Pt/Et (inhA promoter) resistance. The second module (FQ-E) is used to detect WT and specific mutations associated with Lfx and Mfx (gyrA), and ethambutol (E) (embB) resistance. The third module (SLIs) is used to detect WT and specific mutations associated with aminoglycosides Am, Km, (rrs), S (rpsL and rrs), and polypeptide Cm (rrs) resistance.
Interpretation
Following the interpretation and reporting guide for laboratory staff and clinicians, 22 we used the term ‘Resistance not detected’ instead of ‘Susceptible,’ taking into consideration that even the presence of all WT probes cannot rule out resistance to the corresponding drug. Resistance was ‘inferred’ when one or more WT probes were not observed and none of the mutation probes in the corresponding region were developed. Resistance was ‘detected’ when any mutation probes identifying specific mutations conferring resistance to the drugs were bound with the amplicons (regardless of whether WT probes were developed). The test was considered to be invalid when the conjugate, Mycobacterium universal, or MTBC control did not produce any visible signal or there were issues with positive or/and negative control samples.
Clinical information and reference standard results were unavailable to the index test performers. However, routinely collected clinical information was available for the assessors of the reference standard, but we didn’t provide them with the results of index test.
Assessing impact of molecular methods on TB treatment
Routinely collected data on the treatment regimens administered to the patients were analyzed. After obtaining pDST results, we traced any changes in the treatment regimen based on Xpert testing. Then, we evaluated the feasibility and acceptability of a shorter RR/MDR-TB treatment with injectable agent in our cohort of patients and how initial LPA testing would contribute to the selection of appropriate treatment regimen if implemented.
Data analysis
We calculated sensitivity, specificity, and predictive values for diagnostic accuracy assessments. The calculations were based on pDST results, where we used MGIT data as the preferred comparator whenever possible because it has CC defined for most of the drugs. Indeterminate LPA and pDST results were reported to estimate the impact on routine practice but were excluded from the analysis. We performed a blind interpretation of LPA testing results to minimize the effect of cognitive bias in the analysis. Missing data on the LPA and reference standard were also discarded from further evaluation. Although LPA results can take more than two values (resistance detected, inferred, and not detected), categorization of test results as positive or negative for sensitivity and specificity calculation was based on the presence of the mutation probe strip: binding of the amplicons to MUT probes was reported as resistance detected, and both, resistances inferred (no WT band developed) and identifying any of the WT probes without mutation bands at the same gene target were reported as resistance not detected. 22
For sensitivity, specificity, accuracy, and predictive values, the MedCalc Software Ltd. (Ostend, Belgium) Diagnostic test evaluation calculator 23 was used. The proportions were compared using Z Score Calculator for 2 Population Proportions. 24 Statistical analysis and graphics were performed with R Statistical Software [v4.2.3; R Core Team (15 March 2023)]. We used a calculator simulating sampling for the fixed pool size and assumed known test sensitivity and specificity for initial sample size estimation. Intended sample size estimation was calculated online (http://wnarifin.github.io), reflecting national surveillance data 25 of drug resistance to the targeted anti-TB medicines at 30%, expected overall LPA sensitivity of 0.8 and specificity of 0.9. The final estimated minimal sample size (5% dropout), ndrop = 123.
Results
Clinical evaluation of LPA modules
Eighty-four clinical specimens and 97 culture isolates from 126 active TB patients were tested in the study setting and included in the data analysis. The median age of patients was 43 years, interquartile range (IQR) 15 and 28 were assigned female at birth (22.2%). A flow diagram of participants who passed the screening for enrolment into the study and those who have withdrawn from the analysis is shown in Figure 1.

Number of patients and samples initially enrolled and finally assessed in the study.
LPA specificity (%) in detecting RR-TB by testing clinical specimens was 80.0 (95% CI 51.9–95.7), while sensitivity (%) was only 64.7 (95% CI 50.1–77.6). Then, to design the treatment regimen for patients with R-susceptible TB, it is essential to provide upfront detection of resistance to H (Hr-TB). In our study, the sensitivity to identify resistance to H by testing clinical specimens was 80.4 (95% CI 66.9–90.2), and the specificity – 80.0 (95% CI 51.9–95.7). The rifapentine (P) based regimen for drug-susceptible TB, both RR-TB and Hr-TB treatment regimens, requires FQ in the scheme. Thus such testing of clinical samples is a high priority for the management of TB. The best specificity was obtained for Lfx 93.0 (95% CI 80.9–98.5). Nevertheless, the sensitivity of LPA in the detection of FQ resistance was low and varied from 8.7% (Lfx) to 12.5% for low-level resistance Mfx0.25 (critical concentration value of 0.25; Mfx could be considered as an effective drug only at a high dose up to 800 mg in shorter regimen if concomitant high-level resistance was not documented). Interestingly, using culture isolates instead of clinical samples improves LPA sensitivity in identifying resistance to Lfx – 6.7-fold increase (from 8.7% to 58.3%) and 5.7-fold increase to Mfx0.25 (from 12.5% to 71.4%). Estimates of diagnostic accuracy in comparison with pDST are represented in Table 1.
Results of LPA testing of clinical samples and ICs in comparison with culture-based drug susceptibility tests.
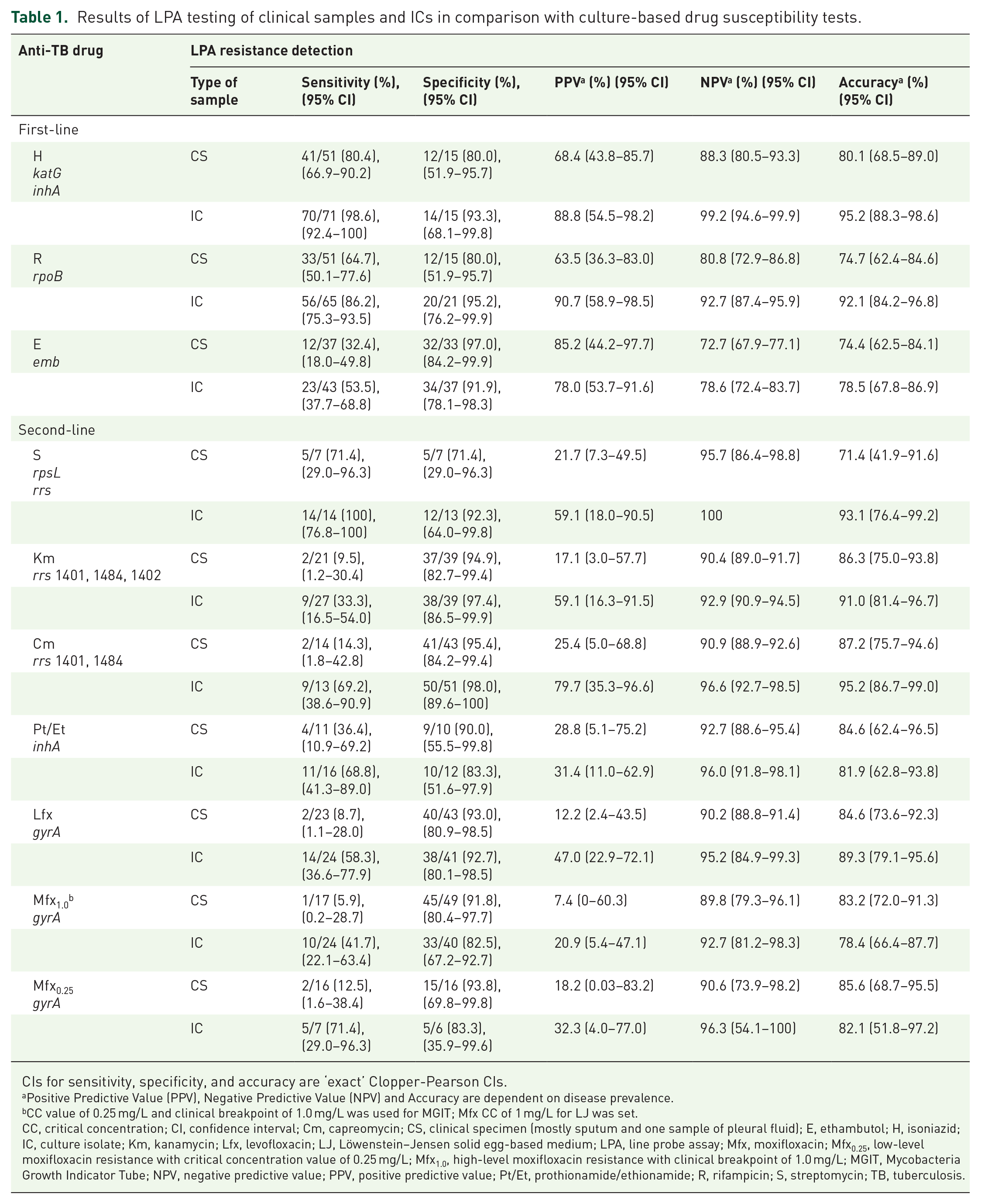
CIs for sensitivity, specificity, and accuracy are ‘exact’ Clopper-Pearson CIs.
Positive Predictive Value (PPV), Negative Predictive Value (NPV) and Accuracy are dependent on disease prevalence.
CC value of 0.25 mg/L and clinical breakpoint of 1.0 mg/L was used for MGIT; Mfx CC of 1 mg/L for LJ was set.
CC, critical concentration; CI, confidence interval; Cm, capreomycin; CS, clinical specimen (mostly sputum and one sample of pleural fluid); E, ethambutol; H, isoniazid; IC, culture isolate; Km, kanamycin; Lfx, levofloxacin; LJ, Löwenstein–Jensen solid egg-based medium; LPA, line probe assay; Mfx, moxifloxacin; Mfx0.25, low-level moxifloxacin resistance with critical concentration value of 0.25 mg/L; Mfx1.0, high-level moxifloxacin resistance with clinical breakpoint of 1.0 mg/L; MGIT, Mycobacteria Growth Indicator Tube; NPV, negative predictive value; PPV, positive predictive value; Pt/Et, prothionamide/ethionamide; R, rifampicin; S, streptomycin; TB, tuberculosis.
Within-study comparison of LPA accuracy in clinical samples versus culture isolates
For a direct within-study comparison of LPA between clinical specimens and culture isolates, we prospectively evaluated available testing results of both samples from 45 patients. Accuracy with corresponding 95% confidence intervals (CI) of LPA (pDST as a comparator test) was higher when performed in isolated strains than in clinical samples, except for FQ (Figure 2). However, a statistically significant difference was observed only for H – 100 (95% CI 89.7–100) versus 75.6 (95% CI 57.9–88.6) two-tailed z test, p = 0.002. However, we found significantly higher sensitivity for H, R, and Lfx analyzing culture isolates (p < 0.05). More detailed information is presented in Table 2.

Comparative accuracy (%) of line probe assay for clinical samples and culture isolates from the same cohort of patients.
Comparison of LPA tests for detection of resistance to anti-TB drugs in clinical samples and culture isolates, with phenotypic drug susceptibility testing as reference standard.

We used z test to compare two proportions. Comparative performance of LPA was analyzed between 2 smear microscopy negative (SM−) and 32 smear-positive (SM+) clinical samples versus 34 culture isolates in the same patients for H, R, Km; 11 SM+ clinical samples versus 11 culture isolates for Pt/Et; 2 smear microscopy negative (SM−) and 31 smear-positive (SM+) clinical samples versus 33 culture isolates for Cm; 2 smear microscopy negative (SM−) and 29 smear-positive (SM+) clinical samples versus 31 culture isolates for E; 1 SM− and 6 SM+ clinical samples versus 7 culture isolates for S, 1 SM− and 27 SM+ clinical samples versus 28 culture isolates for Lfx and Mfx1.0; 1 SM− and 9 SM+ clinical samples versus 10 culture isolates for Mfx0.25.
CI, confidence interval; Cm, capreomycin; E, ethambutol; H, isoniazid; Km, kanamycin; Lfx, levofloxacin; LPA, line probe assay; Mfx0.25, low-level moxifloxacin resistance with critical concentration value of 0.25 mg/L; Mfx1.0, high-level moxifloxacin resistance with clinical breakpoint of 1.0 mg/L; Pt/Et, prothionamide/ethionamide; R, rifampicin;. S, streptomycin; TB, tuberculosis.
Second-line gDST in R-susceptible TB cases
Of particular interest is the rate of resistance to second-line drugs in patients with TB and documented susceptibility to R because these data are not routinely collected. As shown in Table 3, in 6/17 (35.3%) clinical samples and 14/35 (40.0%) culture isolates from patients without the rpoB mutation detected by Xpert, LPA identified mutations in other regions associated with resistance to some anti-TB drugs, including rifampicin.
Mutations detected by line probe assay in clinical samples and culture isolates from patients with GeneXpert positive for Mycobacterium tuberculosis, but negative for rifampicin detection.

Potential alternatives that might be considered in settings with no access to molecular detection of FQ resistance
In our study, 66 clinical samples were tested by LPA with FQ resistance documented by pDST. Among those specimens with no detected mutations associated with resistance to FQ, over 90% (60/66) of the amplicons did not bind to WT probes, inferring resistance. In comparison, specific mutations were detected in 7.6% (5/66) of samples (Table 4). According to our data, based on 79 culture isolates with LPA testing, a combination of rpoB, katG, and inhA mutations was significantly associated with FQ resistance detected by pDST χ(1) = 7.4 (Pearson Chi-Square test with one degree of freedom), p = 0.007. Individuals with such mutations are 3.8 times more likely to have FQ resistance than remaining patients [odds ratio 3.8 (95% CI 1.4–10.1)].
Distribution of line probe assay (AID Autoimmun Diagnostika GmbH) results among clinical samples and culture isolates.

Cm, capreomycin; E, ethambutol; H, isoniazid; Km, kanamycin; Lfx, levofloxacin; Mfx0.25, low-level moxifloxacin resistance with critical concentration value of 0.25 mg/L; Mfx1.0, high-level moxifloxacin resistance with clinical breakpoint of 1.0 mg/L; MUT, mutation probe; pDST, phenotypic drug susceptibility test; Pt/Et, prothionamide/ethionamide; R, rifampicin; S, streptomycin; WT, wild-type probe.
Selection of treatment regimen based on LPA results
To assess the clinical utility of LPA implementation in selecting treatment regimens (either standardized/short or individualized/long) in routine practice, we analyzed a cohort of all RR/MDR-TB patients hospitalized at our clinic in 2019. From 156 patients with RR/MDR-TB, only 35 (22%) patients met the criteria for prescribing a shorter treatment regimen with an injectable agent (4–6 Km/Am-Mfx-Pt/Et-Cfz-Z-Hhd-E/5 Mfx-Cfz-Z-E), such as no exposure to one or more second-line medicines in the regimen for >1 month; no suspected ineffectiveness to compound in the shorter MDR-TB regimen; no pregnancy; no bilateral disseminated lesions on the chest X-ray, or severe extrapulmonary TB (meningeal TB or TB spondylitis). All patients started the shorter regimen pending the results of the pDST.
Due to the detection of additional resistance by pDST (exclusion criteria from shorter treatment), 22/35 (63%) patients with RR-TB were transferred to individualized long-term treatment schemes, another 5/35 (14%) patients were excluded due to documented intolerance or toxicity associated with one or more agents in the shorter RR-TB regimen. Therefore, only 8/35 (23%) patients completed an initial shorter treatment regimen with an injectable agent, demonstrating a low suitability rate of 5% (8/156) of cases in the RR-TB cohort. For these 35 patients with RR-TB enrolled in the study, for 7 patients, LPA testing was performed at the Genetic Research Laboratory (with 7 clinical specimens and 6 culture isolates available for the LPA testing). We found that LPA could correctly exclude the six RR-TB patients transferred to long-term treatment from a shorter regimen, while the remaining single patient with RR-TB was correctly selected for shorter treatment (no further resistance detected by LPA). In five culture isolates and one sputum sample from those excluded patients, at least a mutation in the katG gene was identified, indicating that a high dose of H is likely to be ineffective. Regarding results in clinical samples, four out of seven (57.1%) LPA results were consistent with pDST results. In contrast, two samples had indeterminate results (missing the chance to detect resistance to H). WT bands were detected in regions corresponding to the resistance to H, E, and Pt on pDST in one clinical sample. In three patients who had been excluded from a shorter treatment regimen, one sample showed inferred mutations to H and Pt. In one specimen showed inferred resistance to H, and embB M306V mutations were detected, and one sample showed katG, embB M306V mutations, and inferred resistance to FQ. Finally, the patient with RR-TB who continued the shorter treatment had WT results (in the regions corresponding to other drugs) detected by LPA and no additional resistance detected in pDST.
We have also calculated TAT for first- and second-line pDST in days from sputum collection to reporting of results. All LPA results were available during the second part of the next working day after sample shipment to the laboratory (max TAT was 2 days). In our cohort of patients, the TAT of pDST for H was 30.5 (IQR 36) days, for R – 30.5 (IQR 37) days, for E – 30.0 (IQR 36) days, for S – 30.0 (IQR 30) days, for Km and Cm – 38.5 (IQR 48) days, for Lfx – 37.5 (IQR 48) days, for Mfx – 38.0 (IQR 48) days, and for Pt/Et – 71.0 (IQR 49) days. Implementation of LPA has shown a 15-fold (30/2) to 36-fold (71/2) reduction in the time needed for DST. It could also reduce the triage time for suitable ward sharing by infection control requirements and time under exposure to anti-TB drugs with undetected resistance.
Discussion
This study provides evidence of high diagnostic accuracy of LPA (AID Autoimmun Diagnostika GmbH) for detecting resistance to core anti-TB drugs such as H (80.1%, 95% CI 68.5–89.0), R (74.7%, 95% CI 62.4–84.6), and FQ (e.g. Mfx0.25 – 85.6%, 95% CI 68.7–95.5) in clinical samples. We found that initial LPA testing in sputum specimens could correctly guide clinicians in choosing between longer and shorter TB treatment regimens in 57.1% of RR-TB cases. However, the WHO recommends minimal requirements to target product profiles for the next generation of the gDST at peripheral centers, such as aggregated sensitivity of at least 88% for all clinically relevant mutations and specificity ⩾98%, 26 which could only be obtained for culture isolates tested in the diagnostic module that identifies resistance to R, H, and Pt/Et. A within-study comparison of LPA confirmed (two-tailed z test) that using culture isolates was associated with higher resistance detection accuracy than sputum samples (except for FQ). Furthermore, a combination of rpoB, katG, and inhA mutations increased the probability of FQ resistance by 3.8 times. It can be considered an ‘off-label’ alternative for the treatment decision-making process in resource-limited settings (e.g. irregular supply of reagents/consumables for molecular or culture-based DST for FQ) or when gDST to FQ has suboptimal sensitivity.
Initial exclusion of FQ resistance is a decision-making step for the recently recommended shorter regimen for drug-susceptible TB (2 months isoniazid (H)-rifapentine (P)-pyrazinamide (Z)-moxifloxacin (Mfx) followed by 2 months HP-Mfx) 27 and drug-resistant TB schemes, including Hr-TB with 6 months RZE-Lfx and RR/MDR-TB with 6-month BPaLM/BPaL regimens. 28 However, the low sensitivity of this assay for identifying FQ resistance could be associated either with using only the gyrA locus as a quinolone-resistance determining region or with a prevalence of unknown mutations in dominant M. tuberculosis strain types circulating among our cohort of patients (atypical mutations outside of the interrogated regions of the test). Moreover, systematic errors of LPA are possible due to synonymous and non-synonymous mutations in the gyrA region of some M. tuberculosis isolates, preventing the binding of WT and mutant probes. 29 In another commercial LPA evaluation study, 63% of sputum sample results were classified as resistant based only on the lack of hybridization with the WT probe associated with a double gyrA mutation, 80Ala and 90Gly, detected by gyrA sequencing. 30
Although culture-based DST methods remain the gold standard for identifying drug resistance, these tests require a consolidated approach to select optimal CC, allowing minimization of false-susceptible and false-resistant results with low-resistant strains containing a gyrA mutation. 31 Taking into account the diagnostic importance of pDST on solid media in resource-limited settings, we have also included an interim CC for Mfx of 1 mg/L on Löwenstein–Jensen solid egg-based medium (LJ) in our analysis. However, it could lead to misclassification of gyrA mutants of M. tuberculosis as a susceptible strain at this concentration due to disparities in the validation of this CC with other commercial LPAs. 32 Predominantly M/XDR M. tuberculosis Beijing clades circulating in our region could contribute to individually acquired mutations associated with additional drug resistance. 33 After that, a better understanding of the genetic diversities of M. tuberculosis strains can help to adapt diagnostic algorithms according to each country’s actual needs.
To avoid pseudo-outbreaks or minimize underestimation of the risk of resistance, TB clinical management teams should follow evidence-based practices in the interpretation of gDST results. Conceptually, the WT strain is a drug-sensitive variant, but absence of the WT probe is not equal to the detection of the mutation. However, in an earlier expert report 34 on this issue, the sensitivity and specificity calculation methodology could have been better described, or a better approach used when an absent WT probe has been considered documented resistance, which is discordant with current recommendations. 22 LPA-based detection of resistance to anti-TB drugs is focused on the hotspot regions due to limited space on the strip. However, the results of such testing give a wider pattern for interpretation: mutation detected (MUT probe band developed conferring resistance), heteroresistance (WT and MUT band coexisting at the same loci), all WT probes are developed without mutation band (resistance not detected) and no WT band developed (resistance inferred). Nonetheless, discrepancies between LPA and pDST are better explained through clinical implications or additionally needed diagnostic actions instead of considering the data as inconsistent results. 10 We represented the rate of uninterpretable LPA results as a proportion of all tests performed by each module irrespectively of pDST result availability because, for some drugs such as S, Pt/Et, and FQ, we have performed many more LPA tests than there are available results of pDST for comparison (e.g. patients with susceptible TB had results of LPA but rarely pDST was done for second-line drugs). The rate of inconclusive results was significantly higher while analyzing clinical samples (p < 0.05 for all modules). To explain conceptual differences in determining resistance between phenotypical and molecular-based DST, we described a detailed distribution of the results (including indeterminate) for both methods in Tables 4 and 5.
Phenotypic drug susceptibility test results for each anti-TB drug evaluated in the study.

Contamination and uninterpretable results.
Cm, capreomycin; E, ethambutol; H, isoniazid; Km, kanamycin; Lfx, levofloxacin; LJ, Löwenstein–Jensen solid egg-based medium; Mfx0.25, low-level moxifloxacin resistance with critical concentration value of 0.25 mg/L; Mfx1.0, high-level moxifloxacin resistance with clinical breakpoint of 1.0 mg/L; MGIT, Mycobacteria Growth Indicator Tube; Pt/Et, prothionamide/ethionamide; R, rifampicin; S, streptomycin; TB, tuberculosis.
Our findings show that in over 35% of clinical samples from patients without the rpoB mutation detected by Xpert, LPA identified mutations in other regions that could be associated with resistance to anti-TB drugs. In areas with a high prevalence of drug resistance and limited capacity for single-occupancy patient-care rooms, like Ukraine, it is essential to avoid the prescription of standardized treatment regimens for RR-TB patients based on XpertMTB/RIF testing only, as it may lead to nosocomial transmission of mutant M. tuberculosis strains. On the other hand, the current approach, where only patients with Xpert-detected rpoB mutations remain eligible for second-line pDST testing, could contribute to an underestimation of resistance due to selection bias.
Reprioritization of resources due to COVID-19 (including exploitation of the Xpert platform to perform molecular detection of SARS-CoV-2) throughout Ukraine might have worsened the drug-resistant TB delay during the pandemic’s peak. In a recent cross-sectional multicenter study, Penn-Nicholson et al. 35 found that detection of H, FQ, Et, and SLIs resistance by the Xpert MTB/XDR assay and by LPA had equivalent accuracy. Black et al., 36 in a retrospective laboratory-based study, concluded that discordance between XpertMTB/RIF and other commercial LPA (MTBDRplus) results could be associated with false detection of RIF-resistance by Xpert (22%), heteroresistance of samples (16%) and some errors in either transcription (7%) or interpretation of LPA results (7%). In our study, lower sensitivity of LPA on clinical specimens to detect R resistance (64.7 versus 80.4) compared to H could be explained by a relatively more comprehensive range of genetic loci associated with resistance to H than to R. 37 However, in our cohort of patients, using XpertMTB/RIF instead of pDST as a reference standard test, detection of RR-TB by LPA was associated with even lower sensitivity 61.1 (95% CI 46.9–74.1) versus 64.7 (95% CI 50.1–77.6), p = 0.704 and specificity 75.0 (95% CI 42.8–94.5) versus 80.0 (51.9–95.7), p = 0.757, respectively. It could also be influenced by the lower limit of detection for XpertMTB/RIF (112.6 CFU/mL) 11 compared with LPA (10,000 CFU/mL). 38 On the other hand, low TB bacillary load in clinical samples could increase the probability of false R resistance detection by Xpert assay, while CC used in pDST can misclassify strains with specific R resistance-conferring mutations located at codon 526, 511Pro, 516Tyr, 533Pro, and 572Phe of the rpoB gene. 39
About half of the samples tested for R and S have heteroresistance in our cohort. There are three ways heteroresistance can be explained; first, it is seen in coexisting susceptible and resistant bacterial cells due to naturally developing spontaneous mutations in the initial population. The second situation is when a patient is infected/re-infected with two different M. tuberculosis strains or acquires resistance during exposure to ineffective treatment. Finally, it might also be caused by different limits of detection for WT and MUT probe reactions for R and S compared with the rest of probes. 40 As a result of the investigation of over 31,000 MTBC genomes, Vargas Jr et al. 41 demonstrated that the presence of loss-of-function mutations in the coding region of eis can lead to systematic diagnostic errors of many molecular assays to detect resistance to SLIs.
A low proportion of patients (5%) in the RR-TB cohort who completed the shorter regimen with an injectable agent (known as the ‘Bangladesh’ regimen) was consistent with estimations of other studies.42,43 Our findings could apply to a new 9-month entirely oral regimen containing Bdq (instead of an injectable agent) (4–6 Bdq-Mfx/Lfx-Pt/Et-Cfz-Z-Hhd-E/5Mfx-Cfz-Z-E) for RR/MDR-TB. However, in our study, adding a single new component (Bdq) is unlikely to be sufficient to cover a high rate of resistance (63%) and intolerance (14%) to other documented components of this regimen. Moreover, using Bdq in these circumstances may create an additional risk of acquiring resistance to this important drug. Considering the abovementioned precautions, we suggest replacing Pt/Et with 2 months of Lzd (600 mg daily) whenever possible during the initial phase of the short regimen for an RR/MDR-TB cohort of patients in Ukraine. High dose H (10–15 mg/kg per day in adults) is likely effective if susceptibility is confirmed or only specific inhA promoter mutations are detected (without katG mutations). 44 However, this approach neglects the prediction of resistance to Pt/Et, making prescribing high dose H controversial. Due to high probability of preXDR-TB in patients with documented combination of katG, rpoB, and inhA mutations, 6-month BPaL regimen (without Mfx) could be more suitable in such population living in our geographic area.
Our study has some limitations. We did not have the possibility to analyze the discordance between genotypic and pDST by whole genome sequencing of the M. tuberculosis isolates. TAT for pDST results depends on the time to culture positivity, and it is significantly influenced by media (MGIT versus LJ) and bacterial load in sputum specimens, which should be considered for the pragmatic evaluation of LPA benefits. Critical concentrations of some anti-TB drugs have not been updated for many years, affecting the design and interpretation of molecular assays. LPA is a moderate-complexity molecular DST associated with some technical issues, such as open plate format with possible cross-contamination and compromised interpretation if either all bands or none are developed. 22 The high proportion 91/126 (72.2%) of RR/MDR-TB patients in the study cohort corresponded to national hospitalization practices with more prevalent initial ambulatory treatment for drug-susceptible TB but, on the other hand, it could potentially impact the accuracy estimates for individual drug resistance detection by LPA. To minimize this effect while using diagnostic test evaluation calculator, we set up the prevalence rate of resistance to first-line medicines (H, R, and E) at 35% and the rate of resistance to second-line drugs (S, Km, Cm, Lfx, Mfx, Pt/Et) – at 10%, representing national surveillance data. 25 The association between katG, rpoB and inhA mutations and FQ resistance may be setting-specific and therefore the findings should not be extrapolated beyond studied populations without local molecular epidemiology data analysis.
However, our study has several important strengths. As far as we know, our findings were based on the first direct within-study comparison of LPA results between clinical samples and culture isolates from the same cohort of patients. Another strength of our study is the development of local capacity for the implementation of LPA technology in the Genetic Research Laboratory in Vinnytsia, obtaining results in only 1–2 working days, in comparison with the standard procedure of shipping the samples to the reference laboratory located in Kyiv (at 300 km). Another advantage of our research implementation was enrolling patients into the study regardless of smear microscopy status, where one in five had SM-negative results. Herein, the inconclusive rate of the investigated LPA among SM-negative clinical samples varied from 20.0% (3/15) for FQ-E to 43.8% (7/16) for the MDR-TB module in comparison with over 60% results that could not be interpreted in another commercially available assay. 45
Conclusions
In summary, accuracy of LPA varied from 71.4% (S) to 87.2% (Cm) in clinical specimens and from 78.4% (Mfx1.0) to 95.2% (H, Cm) in the corresponding culture isolates depending on the anti-TB drugs studied and the type of sample used. Implementing LPA in high M/XDR-TB burden settings can facilitate clinicians providing an optimal distribution of available wards between patients with different profiles of resistance and quick antimicrobial stewardship in the clinical management of TB.
Supplemental Material
sj-docx-1-tar-10.1177_17534666241249841 – Supplemental material for Impact of line probe assay-based molecular testing on individualized treatment in patients with rifampicin-resistant tuberculosis: data from the prospective INNOVA4TB cohort study in Ukraine
Supplemental material, sj-docx-1-tar-10.1177_17534666241249841 for Impact of line probe assay-based molecular testing on individualized treatment in patients with rifampicin-resistant tuberculosis: data from the prospective INNOVA4TB cohort study in Ukraine by Andrii Dudnyk, Matthias Hempel, Oksana Lytvyniuk, Halyna Liudkevych, Volodymyr Matsera, Tetiana Nikitchenko, Svitlana Blyzniuk, Barbara Molina-Moya, Rosemarie Preyer and José Domínguez in Therapeutic Advances in Respiratory Disease
Footnotes
Acknowledgements
We thank the patients included in the study for their cooperation and consent to participate. We also thank microbiology laboratory personnel at the Regional Phthisiopulmonology Center and the Genetic Research Laboratory technician for their contributions to this study.
Declarations
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
