Abstract
Background
Four influenza pandemics have occurred during the past 100 years, and new variants of influenza viruses will continue to emerge. The nasal mucosa acts as the primary site of exposure to influenza A virus (IAV) infection, but viral recognition and host immune responses in the nasal mucosa are still poorly understood.
Objectives
This study aimed to evaluate the utility of non-invasive nasopharyngeal swabs for longitudinal monitoring of mucosal immune responses in pigs experimentally challenged with two swine-adapted and one human-adapted IAV. By tracking antiviral immune responses from disease onset to recovery, we sought to assess the feasibility of this method for capturing dynamic changes in viral load and host responses across different IAV strains.
Methods
Forty-two IAV-negative pigs were divided into four groups and housed separately for infection studies. Viral and host RNA from nasopharyngeal swabs was analyzed using microfluidic qPCR, while statistical analysis was performed with a Bayesian approach in R. Additionally, immunohistochemical staining was used to assess MUC5AC expression in the nasal mucosa of infected pigs.
Results
RNA was successfully isolated from nasopharyngeal swabs, enabling gene expression analysis to monitor innate immune responses to IAV infection. A classical innate antiviral immune response was demonstrated after the three virus infections including expression of pattern recognition receptors (PRRs), transcription factors, interferons (IFNs), interferon-stimulated genes (ISGs), cytokines, and chemokines. The kinetics and magnitude of immune responses varied between infections, with notable downregulation of mucins following infection with the Danish swine-adapted isolate. Further, the Danish isolate induced a fast but transient IFN-mediated response concurrent with high expression of cytokines and chemokines, while the other swine-adapted Mexican isolate induced a prolonged immune response of ISGs, cytokines, and chemokines.
Conclusion
This study highlights the significance of highly translational nasopharyngeal swabs as a non-invasive method for assessing mucosal antiviral immune responses. Utilizing microfluidic mRNA analysis, we gained valuable insights into antiviral mucosal responses across 216 swab samples collected from viral inoculation through recovery in three distinct influenza virus infections.
Introduction
Influenza A virus (IAV) causes respiratory illness worldwide with annual seasonal epidemics and occasional global pandemics. The annual epidemics lead to approximately 3–5 million cases of severe illness, resulting in 290,000–650,000 deaths each year. 1 IAV has a high mutation rate and can infect many species, such as humans, pigs, poultry, ferrets, cats and horses.2–5 Even though the nasal mucosa acts as the primary site of exposure to IAV infection in mammals, viral recognition and host immune responses in the nasal mucosa are poorly understood. Pigs play a critical role in the transmission dynamics of influenza viruses, serving as a “mixing vessel” where genetic reassortment can occur due to their susceptibility to multiple influenza virus strains. 6 This poses a dual threat: not only do pigs facilitate the emergence of novel viruses with zoonotic potential, but they also contribute to significant economic losses in the agricultural industry. 7
IAV enters the respiratory airways through the nasal cavity, where the virus must penetrate the airway mucus layer to reach and infect the underlying respiratory epithelial cells. Infection is initiated by binding of the viral surface protein hemagglutinin to host cell receptors. The airway distribution of sialic acid α−2,6 influenza virus receptors is highly similar in humans and pigs, as is the airway mucin composition and distribution of mucus-secreting goblet cells. 8 The airway mucus of mammals consists of both secreted and transmembrane mucins, which are essential for binding and removal of IAV by mucociliary clearance. 9 If IAV does manage to reach and infect the underlying host cells, it is recognized by pattern recognition receptors (PRRs). Activation of the PRRs induces a signaling cascade resulting in the transcription and production of interferons (IFNs), pro-inflammatory cytokines, and chemokines.10–12 Secreted IFNs stimulate the expression of hundreds of IFN-stimulated genes (ISGs), which interfere with and restrict viral replication in the infected cell as well as in neighboring cells. Cytokines and chemokines recruit and activate immune cells,13,14 but they are also associated with the immunopathology observed during severe respiratory viral infections caused by IAV,15,16 SARS-CoV-2,17,18 and respiratory syncytial virus.19,20
Even though the nasal mucosa acts as the primary site of exposure to IAV infection, and the vast majority of all infections are assumed to be contained by the innate immune system, we still have limited knowledge regarding viral recognition and innate immune responses in the nasal mucosa. In this study, we demonstrated the utility of non-invasive nasopharyngeal swabs for tracking these responses from disease onset to recovery in pigs experimentally challenged with two swine-adapted and one human-adapted IAV. By including IAV strains well-adapted to the host and IAV originating from another host in the present study, we aimed to examine how viral adaptation shapes innate immune activation, offering valuable insights into the early mechanisms of immune recognition and control at the mucosal surface.
Material and methods
Preparation of virus inoculum
Three H1N1 viruses adapted to either the swine or human host were selected for inoculation. Two swine isolates were included, one belonging to the 1A.3.3.2 clade, A/Swine/Denmark/2017_10298/4_4p1/2017 (H1N1) (accession no. MT666901-MT666908), and will be referred to as the Danish isolate, and one belonging to the 1A.3.3 clade, A/Swine/Mexico/AVX-39/2012 (H1N1) (accession no. KU976900-KU976514), referred to as the Mexican isolate. A human-adapted isolate was included in the study as well, A/Denmark/238/2020 (H1N1) (human seasonal clade 1B.1; accession no. OQ062647-OQ062654). The Mexican isolate was kindly provided by Nacho Mena and Adolfo Garcia-Sastre (Mount Sinai School of Medicine). Virus stocks were prepared as described previously by passage in Madin–Darby canine kidney cells. 21
Characterization and quantification of the three IAVs
Full-genome sequencing and genetic characterization was performed previously on the inoculum virus. 21 HA gene nucleotide sequences were aligned with a selection of genetically related reference IAV sequences in CLC Main Workbench v24.0.1. 21 Reference IAV sequences were downloaded from publicly accessible sequence databases, GISAID EpiFlu™ (https://www.gisaid.org/) and NCBI GenBank (https://www.ncbi.nlm.nih.gov/genbank/), representing Northern hemisphere human seasonal vaccination IAV strains, IAVs circulating in Danish swine herds, and ancestral IAVs identified by Mena et al., 2016. 22 Swine-isolated HA sequences were classified into clades using the Swine H1 Clade Classification Tool hosted on the Bacterial and Viral Bioinformatics Resource Center (https://www.bv-brc.org/app/SubspeciesClassification). 23 An initial maximum likelihood phylogenetic tree was inferred from the HA gene sequence alignment using IQ-TREE v2.0.3 with the best-fitting substitution model (K3Pu + F + I), ultrafast bootstrap approximation = 1000 and number of iterations = 2500.24,25 The molecular clock phylogenetic tree was estimated from the topology of the maximum likelihood phylogenetic tree and sequence alignment by TreeTime v0.9.2 using a strict molecular clock model. 26 Trees were visualized in FigTree v1.4.4 and a time scale was fitted by reversing the scale axis and offsetting the time scale by the numerical collection date of the most recent isolate. The maximum-likelihood tree revealed the genetic differences of HA between the two swine isolates and the human isolate as they cluster into three different clades. 21 The Mexican swine isolate (A/swine/Mexico/AVX-39/2012) clusters into the swine H1 1A.3.3 clade and the Danish swine isolate, A/swine/Denmark/2017_10298_4_4p1/2017, clusters into a different swine clade namely H1 1A.3.3.2. The Human isolate clusters into the Human 6B.1 clade. To support the genetic characterization a pairwise comparisons of amino acid sequences of the viral proteins of the inoculum strains was performed in this study using CLC Main Workbench v22.0.2 (QIAGEN) to measure percentage identity. In addition, a sequence alignment of the NS1 sequences of the three inoculum strains was performed.
Severity of lung lesions was assessed by Kristensen et al. (2023) evaluating peribronchial and peribronchiolar infiltrates, bronchiolar luminal exudate, and alveolar infiltration, with a maximum score of 7. Briefly, peribronchial/peribronchiolar infiltrates were scored based on the percentage of affected sites as 0:none, 1:few (<10%), 2:many (10–50%), or 3: majority/all (>50%). Bronchiolar luminal exudate severity was categorized as 0:none, 1:minimal, or 2:heavy. Alveolar infiltration was similarly classified as 0:none, 1:minimal, or 3:heavy. 21 The most severe clinical signs and pathological changes were observed after infection with the Mexican isolate with pathological changes observed in all pigs at 3 dpi. Only 7/8 pigs and 6/8 pigs showed milder gross lesions after infection with the Danish and Human isolate, respectively. 21
The PCR assay used to detect IAV in nasopharyngeal swab samples was an in-house modified version of an RT-qPCR assay targeting the matrix gene (M-gene) using the SensiFast Probe No-ROX One-Step Mix kit (Meridian Bioscience, Cincinnati, Ohio) as described before. 21 Quantification was based on a 10-fold dilution series of the target sequence with known copy numbers. The total viral RNA load differed among the groups with the highest total viral RNA load after infection with the Danish isolate, peaking 4 days after infection (all control pigs were negative). 21
Only a few samples were positive after the peak in viral RNA 4 days after infection. Thus, in this study we determined the amount of infectious virus in the positive nasopharyngeal swab after day 4. The samples were homogenized, sterile filtered, and titrated in MDCK cells. The TCID50/ml was calculated using the Reed-Muench method. 27
Animal challenge
This study has been described in detail elsewhere. 21 In brief, 42 Danish Landrace Crossbred pigs (males and females, 7 weeks old), confirmed IAV negative and seronegative, were allocated into four groups by minimization (ARRIVE guidelines), with sex and size as nuisance variables, and housed in a separate isolation unit; groups 2–4 each included 12 pigs, and group 1 consisted of six pigs (Figure 1A). To simulate the natural route of IAV infection, pigs were inoculated intranasally by a MAD Nasal Intranasal Mucosal Atomization Device (Teleflex, Wayne, Pennsylvania). Pigs in groups 2–4 were inoculated in the right nostril with 3 ml of the swine isolates or the Human isolate with a titer of 107 TCID50/ml. Group 1 was mock inoculated with culture medium only (control). Nasopharyngeal swab samples were collected with FLOQSwabs (COPAN Diagnostics, Murrieta, California) from all animals just before inoculation (day 0) and at 1, 2, 3, 4, 7, 10, and 14 days post-infection (dpi) and stored in DNA/RNA Shield (Zymo Research, Irvine, California) at −20°C for immunological examination and in Sigma Virocult medium (Medical Wire, Corsham, UK) for viral quantification. Samples for viral quantification were kept at room temperature for 30 min then vortexed for 10 s before transfer to Eppendorf tubes and stored at −80°C until analyzed. All six pigs in group 1 (control) and eight pigs from each inoculated group (30 in total) were euthanized at 3 dpi, while the four remaining pigs from each inoculated group (12 in total) were euthanized at 14 dpi (Figure 1A). 21

A) pigs were acclimatized for seven days. On day 0, swab samples were collected before inoculation with either culture medium (control) or influenza A virus. Throughout the study, swab samples were collected (days 1–4, 7, 10, and 14 after inoculation) using FLOQSwabs. On day 3, all control pigs (n = 6) were euthanized together with eight pigs from the infected groups (n = 24). On day 14, the remaining pigs were euthanized (n = 12). B) Workflow to obtain transcriptional data included RNA extraction, a quality control step prior to reverse transcription. Pre-amplification was conducted using gene-specific primers prior to running high-throughput microfluidic qPCR. Created in BioRender.com.
RNA extraction
Total RNA was extracted from nasopharyngeal swab samples with the Quick-RNA Microprep Kit (Zymo Research) according to the manufacturer's protocol. RNA purity and concentration were assessed using a NanoDrop One spectrophotometer (Thermo Fisher Scientific, Waltham, Massachusetts). RNA quality was determined using the Agilent RNA 6000 Nano Kit on the Bioanalyzer 2100 system (Agilent Technologies, Santa Clara, California) (Figure 1B).
cDNA synthesis and microfluidic qPCR
Using the QuantiTect Reverse Transcription Kit (QIAGEN, Hilden, Germany), 300 ng RNA was reverse transcribed into cDNA following the manufacturer's protocol. Two separate cDNA reactions were performed for each RNA sample (technical replicates) to validate the qPCR data (Figure 1B). The DNase treatment was performed by adding 1.5 µl Wipeout Buffer to 1 µL RNA and 11.5 µL RNase-free water. Reverse-transcription reactions were diluted 1:10 in TE-buffer (VWR International, Søborg, Denmark). All porcine primers used in this study were designed (design description in Supplementary Text S1) using Primer3 (v. 0.4.0) (https://bioinfo.ut.ee/primer3-0.4.0/) following an optimized protocol to generate high quality data from moderately degraded RNA (short amplicons and test of reproducibility of primer pairs). Sequences and qPCR efficiencies for all primers can be found in Supplemental Table S1. Target genes were pre-amplified using TaqMan PreAmp Master Mix (Applied Biosystems, Waltham, Massachusetts) as described previously 28 applying 20 pre-amplification cycles. Residual primers were digested by treatment with 4U of Exonuclease I (New England BioLabs, Ipswich, Massachusetts) by incubation at 37°C for 30 min, followed by 80°C for 15 min. Pre-amplified, exonuclease-treated cDNA samples were diluted 1:10 in TE Buffer. High-throughput microfluidic qPCR was carried out using 96.96 Dynamic Array IFC chips (Standard BioTools Inc., South San Francisco, California) running on the BioMark real-time platform (Standard BioTools Inc.). The following PCR protocol was used: Thermal mix (2 min at 50°C, 30 min at 70°C, and 10 min at 25°C) and Hot Start (2 min at 50°C and 10 min at 95°C), followed by 35 PCR cycles with a denaturation phase for 15 s at 95°C and annealing/elongation for 1 min at 60°C. The protocol ends with a Melting Phase (30 s at 60°C followed by 1°C increase every third second from 60°C to 95°C).
qPCR data analysis
Data were coded prior to analysis to complete the analysis blinded to treatment. Amplification and melting curves were visually inspected using the Fluidigm Real-Time PCR Analysis software (v4.8.1). Primer efficiencies were calculated for each assay from two independent 5-fold dilution series. Data were pre-processed in GenEx7 (MultiD, Gothenburg, Sweden), including interplate calibration, correction for PCR efficiency, evaluation of (using the algorithms geNorm 29 and NormFinder 30 ) and normalization to reference genes, and averaging of technical replicates. Samples were excluded if more than 15% of the technical replicates exceeded the accepted criteria (>1.5 Cq difference); less than 1% were excluded. A biologically relevant cut-off value of ±1 log2 fold change in gene expression compared to baseline (all data before inoculation and data from the control pigs) was used. The change in gene expression level was considered statistically significant with a posterior probability of >95%.
Statistical analysis
The data analysis was performed in the R programming environment (version 4.2.1 31 ) in RStudio (version 2022.12.0.353 32 ). The R-packages tidyverse (version 1.3.2 33 ) and bayesplot (version 1.9.0 34 ) were used for data manipulation and plotting. Statistical models were specified and fitted using Stan 35 and RStan (RStan v2.26.13 36 ). The R-package “loo” (version 2.5.1) was used to check model fit and for model comparison.37,38 Further, model fit assessment was done using bayesplot to do posterior predictive checking. 39
Using a Bayesian approach, the typical (average) expression levels for each combination of gene, treatment, and day was estimated, yielding a posterior distribution over the possible values of the parameters. For all Bayesian models (one model per gene), two independent Markov chain Monte Carlo chains for 10,000 iterations each were run, with 5000 iterations warm-up, resulting in 10,000 post-warmup samples for each parameter. Convergence of Markov chain Monte Carlo runs was monitored by checking that there were no divergent transitions during sampling and that the potential scale reduction factor (“R-hat”) was close to 1 at the end of the run for all model parameters. From the Markov chain Monte Carlo runs, all the quantities of interest were calculated (log2 fold change vs baseline for a given day). A comprehensive description of the statistical model and the model parameters can be found in Supplementary Text S2.
Immunohistochemical staining
Nasal mucosal tissues for immunohistochemical staining of MUC5AC were collected from eight additional pigs (same breed and similar age as the original study) four days after infection with the Danish isolate (n = 4) and the Human isolate (n = 4). The tissue specimens were fixed in Carnoy's Solution (Ampliqon, Odense M, Denmark) for a maximum of 24 h before exchange to 99% ethanol to preserve the mucus. The tissues were embedded in paraffin wax and sliced into 2–3 µm sections. After deparaffinization, immunohistochemical staining targeting MUC5AC was performed. Tissues were treated with citrate buffer in a heating cabinet at 90°C for 60 min and hereafter washed twice with TBS (pH 7.6) for 5 min. This washing step was performed after each step described below. Blocking for endogenous peroxidase was performed by adding 3% H2O2 to the sections for 10 min. Sections were blocked with UltraVision Protein Block (Epredia, Kalamazoo, Michigan) and an anti-MUC5AC antibody (diluted 1:320,000 in 1% BSA/TBS) (Invitrogen, Waltham, Massachusetts) and stored overnight at 4°C. A Primary Antibody Enhancer (Epredia) was added for 20 min. UltraVision Large Volume HRP-Polymer (Epredia) was added for 30 min, and the staining was visualised by adding AEC substrate (Vector laboratories, Newark, California) for 10 min. The sections were counterstained by Mayer's hematoxylin (VWR International). An isotype control (Agilent Technologies) (diluted to the same content of protein as the primary reagent in 1% BSA/TBS) was used as a negative control. Two images of each tissue with the highest amount of MUC5AC staining in the mucus and/or epithelium were selected as a region of interest in ImageJ. 40 Positive MUC5AC area (secreted and in the epithelium layer) was identified using the “split channels” function, choosing the blue channel, and setting a threshold of 90.
Results
Characterization of the three influenza A virus strains
Based on data from Kristensen et al. (2023), viral shedding peaked at 4 dpi after all three infections with the highest viral shedding after infection with the Danish isolate (2.10 × 109 copies/ml), followed by the Mexican isolate (1.61 × 109 copies/ml) and lastly, the Human isolate (6.25 × 108 copies/ml) (all control pigs were negative). Only few animals were IAV positive from 7 dpi and onwards. 21 To investigate if the infection was cleared at this point, the positive samples were tested for infectious virus by TCID50, and they were all negative (data not shown). Furthermore, a pairwise comparisons of all viral gene sequences were performed to display genetic differences beyond HA, which could have an impact on the immunological response. The comparison resulted in high amino acid identities (>95%) between the Human and the Danish strains for all gene segments but, HA (92.6%) and NA (91.7%), and the NS1 protein (94.5%). Higher variations were seen when the Human and Danish strains were compared to the Mexican strain (<95% amino acid identities in HA, NA, NS1, NS2/NEP, M2, and PAX). The lowest similarity was found in the NS1 gene sequence (Human isolate: 86.2% and Danish isolate: 89.4%). Overall, data from Kristensen et al. (2023) and additional analysis from this study demonstrate distinct viral shedding patterns, pathogenicity, and genetic relationships among the isolates, with the highest viral RNA load after infection with the Danish isolate and the most severe clinical outcomes after infection with the Mexican isolate.
High-quality RNA isolated from swab samples
RNA extracted from the nasopharyngeal specimens were of good quantities (mean of 112.5 ng/µL [SEM = 2.8]) and quality, 41 with a mean RNA integrity number of 6.8 [SEM = 0.1] and mean 260/280 and 260/230 absorbance ratios of 2.0 and 1.9, respectively. High reproducibility was seen for cDNA replicates produced from the same RNA sample using all included primer assays optimized for medium to low RNA quality. 42 Different primer pairs targeting the same transcript were highly and significantly correlated (p-values < 0.00001, Pearson correlation coefficient ranging from 0.72–0.99).
Gene expression analysis can be performed on non-invasive nasopharyngeal swab samples to monitor innate immune responses following IAV infection and clearance
Table 1 shows the complexity and duration of the innate immune response to viral infection, as indicated by the total number of statistically significant upregulated innate antiviral genes (±1 log2 fold change compared to baseline, posterior probability >95%) at the seven time points after infection. Gene expression levels after all three infections compared to baseline are summarised in Supplemental Table S2.
The total number of statistically significant upregulated innate antiviral genes (+1 log2 fold compared to baseline, posterior probability >95%) at the seven time points following infection underpins the complexity and duration of the innate immune response.
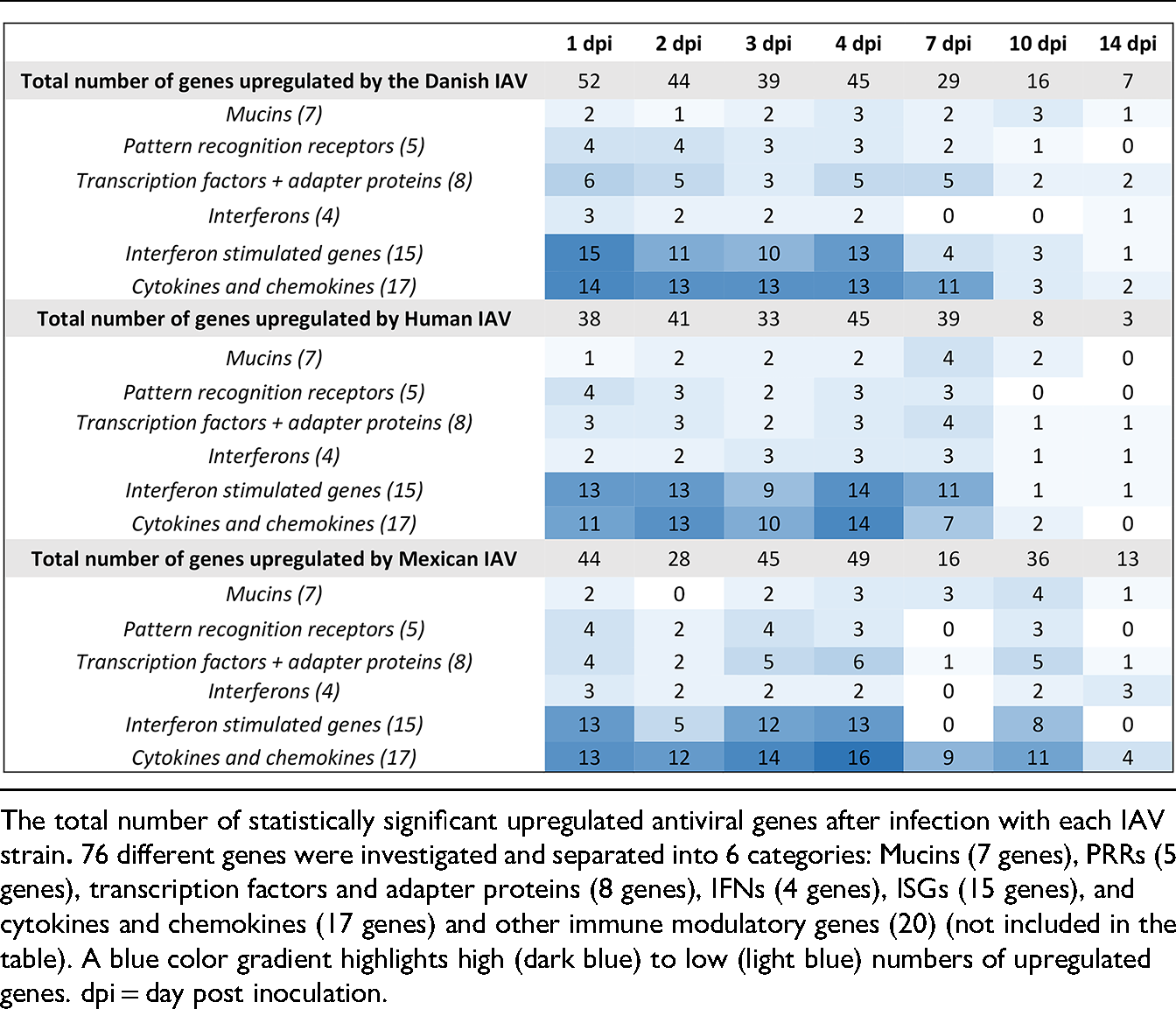
If IAV successfully reaches and infects host cells, the virus is detected by pattern recognition receptors (PRRs). PRR activation triggers several signaling cascades that leads to the transcription and production of interferons (IFNs), pro-inflammatory cytokines, and chemokines. Secreted IFNs then induce the expression of hundreds of IFN-stimulated genes (ISGs), which inhibit viral replication in the infected cell. A classical innate antiviral immune response was observed in the nasal mucosa specimens after infection with all three IAVs. Expression of 76 genes centrally involved in the antiviral innate immune response was investigated, and statistically significant upregulation in the expression levels of mucins, PRRs, transcription factors and adapter proteins, IFNs, ISGs, cytokines, and chemokines were seen at one or more time points after all three infections compared to baseline (Table 1 and Supplemental Table S2).
Figure 2 summarizes and compares the innate mucosal expression profiles 1 day after infection with three viruses. A rapid increase of important viral PRRs, that detect viral RNA and activate downstream signaling pathways, including DDX58 (RIG-I), IFIH1 (MDA5), TLR7, and TLR8, as well as downstream transcription factors (IRF7, IRF9, and STAT1), IFNs, ISGs, and cytokines and chemokines, were seen in the nasopharyngeal swab samples already at 1 dpi in response to all three IAV strains (Figure 2). Many of the innate factors were continuously expressed from day 1 to 4. Furthermore, differences between the infection strains are also evident with significant upregulation of MYD88 (adaptor protein for TLR signaling) and IFNG solely after infection with the two swine isolates, while the key transcription factor for inflammatory cytokines, NFKB1 expression was exclusive after infection with the Danish isolate one day after infection (Figure 2).

To demonstrate how a classical antiviral immune response can be measured and followed daily in non-invasive nasopharyngeal swab samples, the gene expression levels of a selection of central PPRs and ISGs after infection with the Danish isolate are shown in Figure 3. A strong and long-lasting response of DDX58 (RIG-I) was demonstrated with highly increased expression during the initial 4 days of infection, which still remained elevated at 7 and 10 dpi (Figure 3). In contrast, the TLR7 response was limited to 1 and 2 dpi (Figure 3). Similar patterns were observed for other PRRs, with a strong and long-lasting response of IFIH1 (MDA5) and a shortened response of TLR8, respectively (Supplemental Table S2).

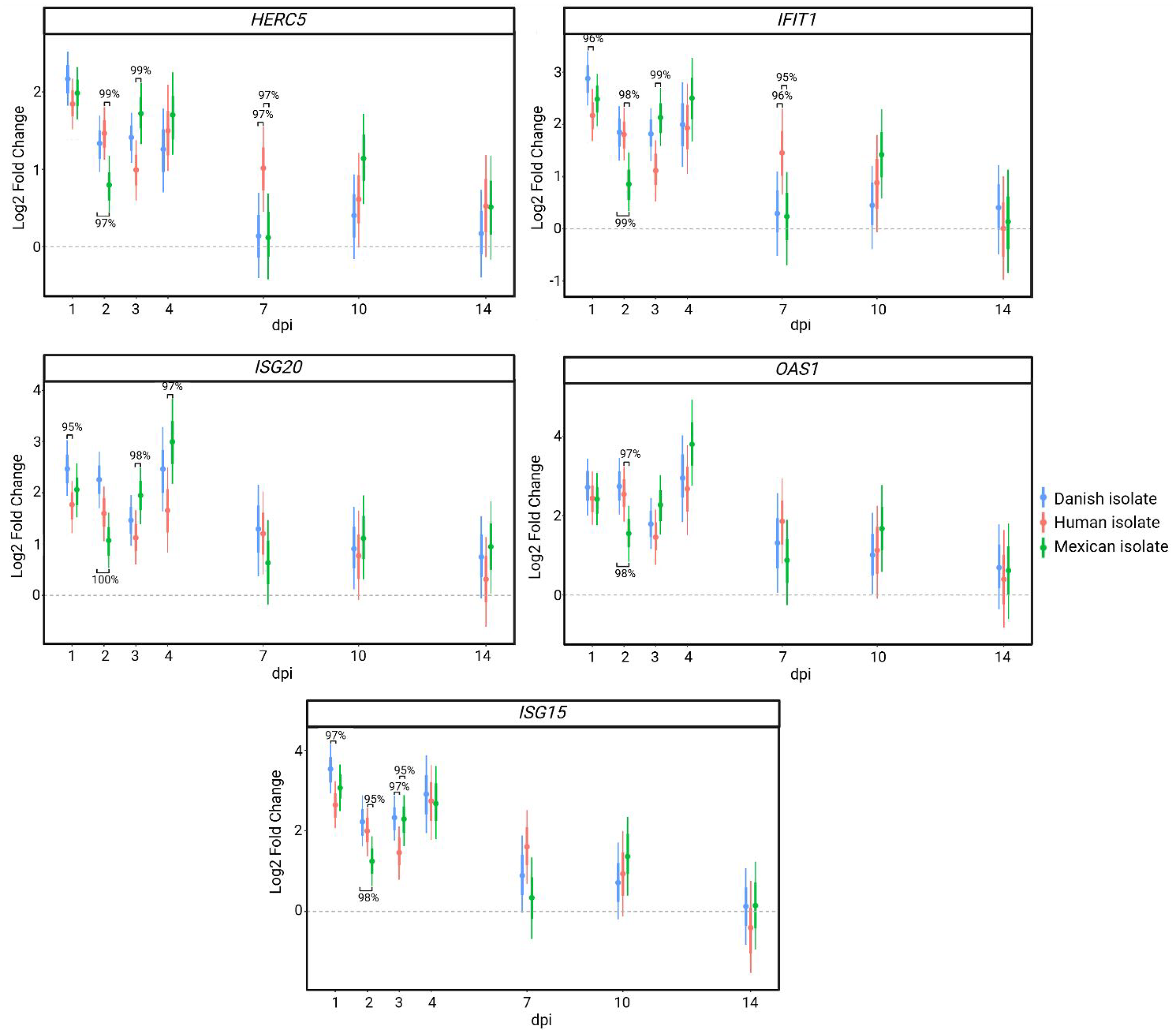
The ISGs showed varying expression patterns as well. Induction of the antiviral ubiquitin-like modifier, ISG15 expression was strong and transient response with significantly increased levels at 1 to 4 dpi compared to baseline, while the interferon-stimulated GTPase MX1 showed a strong initial response followed by a sustained albeit diminished response compared to ISG15. Lastly, some ISGs displayed fast and moderate expression levels after infection with the Danish isolate as depicted by EIF2AK2 (PKR) (Figure 3). Members of the same ISG families, such as OAS, which activates the RNase L pathway for viral RNA degradation, and IFIT, which inhibits viral RNA translation, did not necessarily show the same expression patterns. IFIT1 and OASL demonstrated strong and brief responses like ISG15, while IFIT3, ISG20 and OAS1 had strong and longer lasting expression levels after infection with the Danish isolate (Supplemental Table S2), underlying the complexity of an antiviral immune response against IAV. In summary, an antiviral innate immune response can be monitored daily using non-invasive nasopharyngeal swab samples where differences in kinetics and magnitude of the innate factors can be investigated throughout the infection until clearance.
Kinetics and magnitude of the immune response differed between the three infections
A rapid onset of the immune response upon infection with the Danish isolate, including high expression of PRRs, transcription factors, interferons, ISGs, cytokines, and chemokines was seen at 1 dpi, followed by a decrease in expression until 14 dpi with the exception of 4 dpi where the expression levels were re-induced, though not to the same magnitude as at 1 dpi (Table 2 and Figure 4). The highest number of antiviral genes upregulated in the nasopharyngeal samples was seen after infection with the Danish isolate at 1 dpi (n = 52) (Table 1). The highest number of upregulated genes after infection with both the Mexican isolate (n = 49) and the Human isolate (n = 45) was seen at 4 dpi (Table 1). This later peak in number of antiviral genes and generally also expression levels after infection with the Mexican isolate and the Human isolate was represented in all parts of the antiviral innate immune pathway from PRRs to IFNs and downstream expression of ISGs. Expression levels of pro- and anti-inflammatory cytokines (IL10, IL12B, IL17A, IL1A, IL1B, IL1RN) peaked at 2 dpi followed by a re-induction at 4 dpi after infection with the Danish isolate (Table 2). Expression of cytokines after infection with the human strain had a similar expression pattern, though the response was dampened compared to the infection with the swine viruses (Table 2).
Significant expression levels of cytokines after all three infections compared to baseline, depicted as log2 fold change, from 1 to 14 dpi.

In contrast to the two other strains, infection with the Mexican isolate resulted in a prolonged duration of increased immune gene expression. At 10 dpi substantially more genes were upregulated (Table 1) with high expression levels after infection with the Mexican isolate compared to the other isolates (Supplemental Table S2). PRRs, TLR8, IFIH1 (MDA5), and DDX58 (RIG-I), and especially the cytokines CXCL8 (IL8), IL10, IL12B, IL18, IL1A, IL1B, IL1RN, IL6, and TNF were highly expressed at 10 dpi (Table 2). ISGs such as OAS1, IFIT1, ISG15, ISG20, and HERC5 were found to be differentially expressed at 10 dpi compared to baseline solely in response to the Mexican isolate (Figure 4). Inflammatory and anti-inflammatory genes such as IL1A, IL1RN, IL17A, and TNF were still significantly increased at day 14 after infection with the Mexican strain, demonstrating a substantially prolonged response compared to the two other isolates (Table 2). Thus, the expression pattern of the immune response after infection with the Mexican isolate differed from the two other IAV infections with a prolonged response with high expression levels of inflammatory genes, anti-inflammatory genes and ISGs past day 7. Overall, these findings highlight the value of non-invasive nasopharyngeal swabs for tracking differences in kinetics and magnitude of innate factors throughout the infection until viral clearance. The upregulation of antiviral genes and strain-specific immune signaling differences underscore host response complexity and highlight the potential of gene expression profiling in differentiating infection dynamics among viral strains.
Downregulation of several mucins after infection with the Danish isolate
Several mucins were found to be downregulated in the nasal mucosal swabs upon viral infection. MUC5AC was downregulated at 2 dpi in response to all three isolates. Notably, the Danish isolate uniquely caused downregulation of both secreted and transmembrane mucins at various time points post-infection. The main secreted mucins, MUC5AC and MUC5B, were downregulated in the early period of infection, where the transmembrane mucin MUC12 was significantly downregulated at 2, 3, and 7 dpi compared to baseline (Figure 5). In order to support the data on reduced MUC5AC expression after infection with the Danish isolate, nasal mucosal tissues were collected from additionally eight pigs (same breed and similar age as the original study) four days after infection with identical isolates, the Danish isolate (n = 4) and the Human isolate (n = 4). The tissues were fixed in Carnoy's solution to preserve mucus and were stained for MUC5AC. Two areas of each of the stained slides were analyzed for the percentage of area staining positive for MUC5AC in the epithelial and mucus layer to determine the amount of MUC5AC production after each infection (Figure 6). Although the MUC5AC positive area was smallest after infection with the Danish isolate, the difference was not statistically significant (data not shown).


Discussion
The innate immune response plays a key role in the host defense against invading pathogens, and despite extensive efforts to elucidate the role of innate immunity in response to IAV and other respiratory infections, viral recognition and innate responses in the nasal mucosa remain poorly understood. Particularly, the contribution of host factors in either facilitating or inhibiting viral infection at the site of first contact remains to be investigated. The use of nasopharyngeal swabs as a non-invasive sampling method to investigate immune responses in the nasal mucosa to viral infections has only been demonstrated to a limited extent. Nasopharyngeal swabs have been used to study immune responses against SARS-CoV-243,44 and respiratory syncytial virus (RSV)45–47 in humans, but to the best of our knowledge, only protein expression of a few cytokines have been studied in nasopharyngeal swab samples collected from pigs upon infection with influenza A virus. 48 In the present study, we demonstrate that factors of the entire innate pathway can be measured and explored in nasopharyngeal swabs throughout the infection and recovery phases in pigs infected with highly host-adapted IAV strains and IAV originating from another host.
A classical innate antiviral immune response was demonstrated with expression of PRRs, transcription factors and adapter proteins, IFNs, ISGs, and cytokines at one or more time points after infection with the two swine-adapted (Danish and Mexican) and the human-adapted isolate. These innate immune responses are consistent with what has been reported in human studies on IAV infection both in vivo and in vitro,13,49–51 suggesting that both the sampling approach and the use of pigs may yield valuable innate immune data relevant to human IAV infections. We demonstrate that good RNA quality and highly reproducible data can be obtained by the nasopharyngeal swab sampling method, which proves its applicability for continuous monitoring of the local host response during infection, allowing in-depth investigations of the temporal dynamics of the mucosal antiviral innate immune response.
Differences in kinetics and magnitude of the antiviral innate immune responses were identified both during each individual IAV infection and between the infections. After infection with the swine-adapted Danish strain, we observed transient early increased expression of both TLR7 and TLR8 whereas the cytoplasmic PRRs DDX58 (RIG-I) and IFIH1 (MDA5) were continually upregulated after infection also after the viral RNA load had decreased at 7 and 10 dpi. Similarly, other studies have found simultaneous and continued activation of all PRRs in human peripheral blood of symptomatic individuals several hours after H3N2 IAV infection 52 and in mouse alveolar type II epithelial cells until 3 days after infection. 53 But the long-term differences observed several days post IAV infection in this study have not been described previously. TLR7 and TLR8 recognize ssRNA genomes in endosomes without requiring viral replication whereas cytoplasmic RIG-I also recognizes triphosphorylated ssRNA, which is generated during viral replication. 54 Thus, the initial short-term response of TLRs seen after infection with the Danish isolate, as well as the other isolates (Supplementary Table S2), might be more important early in the infection prior to viral replication to trigger and activate an early immune response, while the long-term RIG-I activation is important to induce a more long-lasting antiviral environment mainly in response to replicating viruses. A prompt onset of the innate immune response with peak expression of interferons, notably IFNB1 and IFNL1 (IL29), was seen at 1 dpi after infection with the Danish isolate compared to a delayed IFN response peaking at 4 dpi after the two other infections. Lower expression of IFNs was been seen in vitro during the first 12 h after infection of porcine tracheal epithelial cells with a swine-adapted IAV compared to a human-adapted IAV. 55 The presence of a fully functional protective mucus layer including sialic acid-coated decoy receptors in vivo may impact the infection dynamics, as suggested in the present study. A decreased expression of both secreted and transmembrane mucins (MUC5AC, MUC5B, and MUC12) was observed in the nasopharyngeal specimens at several time points after infection with the Danish isolate, potentially allowing this virus to reach and infect epithelial cells more rapidly, thereby inducing an innate immune response with an earlier onset than the two other IAVs. Increased mucin expression in relation to H1N1 and H3N2 IAV infection has been reported in vivo in mouse lungs56,57 and in a few in vitro studies in human epithelial cells,58,59 but the ability to measure mucin expression dynamics over time in nasopharyngeal swabs has to our knowledge not been demonstrated before. In addition, a thorough in vivo analysis of mucin expression after challenge with IAV or other important respiratory viruses (corona viruses and RSV) has to our knowledge never been performed, positioning the results presented here as an important foundation for future studies of mucosal virus-host interaction in both animals and humans.
The two swine isolates induced the fastest and highest magnitude of innate immune factor upregulation, which are consistent with the viral RNA load measured in the same animals. Association between a higher viral load and higher levels of ISGs (MX1, MX2, ISG15, and OAS1) has likewise been documented in Covid-19 positive patients. 60 In general, the magnitude of the inflammatory immune response towards the Human isolate was lower compared to the swine isolates. Lower cytokine expression and viral load after infection with the human IAV agree with the absence of transcription factor NFKB1 upregulation during this infection.61–64 Contrary to the infection with the Human isolate, infection with the Mexican swine isolate induced a substantial and prolonged mucosal immune response with high expression of PRRs, transcription factors, cytokines, and ISGs early after infection (1 to 4 dpi) and again at 10 dpi. Interferons (IFNA1, IFNB1, and IFNL1 (IL29)), cytokines (IL17A, IL1A, IL1RN, and TNF), and other pro- and anti-inflammatory proteins, such as S100A7, SELL, PTGS2 (COX2) and TNFAIP3, were even differentially expressed at 14 dpi (Supplementary Table S2) when the infection was cleared in all pigs. The viral protein NS1 may play a key role in modulating the innate immune response, and was the only viral protein with amino acid identity below 90% when comparing the Mexican isolate to the other strains (Human: 86.2%, Danish: 89.4%) (Supplemental Figure S1). NS1 limits host IFN production through two main mechanisms: i) RIG-I inhibition and ii) CPSF4/CPSF30 inhibition, essential for host pre-mRNA processing. The NS1 region (aa 175–210) is critical for CPSF4/CPSF30 binding, with mutations at D189 and V194 impairing host gene inhibition. The Mexican NS1 gene carries substitutions at both positions, along with four additional changes (aa 202, 206, 207, 209) within the CPSF30 binding site. These mutations may weaken CPSF4/CPSF30 binding, reducing immune evasion and contributing to the prolonged immune response observed after infection with the Mexican isolate (Supplemental Figure S2). It may be speculated that the prolonged activation of PRRs, cytokines, and inflammatory genes after infection with the Mexican strain contribute to the increased severity of pathological lesions associated with this strain. 21 Indeed, over-stimulation of multiple viral sensing pathways and their downstream innate factors has been linked to the development of clinical signs and symptoms in humans experimentally infected with H3N2 IAV compared to asymptomatic individuals. 52
Conclusion
Our study highlights the utility of nasopharyngeal swabs as a non-invasive method for monitoring the innate immune response during viral infections. While primarily based on microfluidic mRNA analysis, our approach provides valuable insights into mucosal immunity and antiviral responses. Future studies incorporating complementary protein-level assessments would further strengthen these findings. Importantly, our work underscores the potential of nasopharyngeal swabs as a powerful yet underutilized tool for investigating mucosal immune responses, offering a translationally relevant approach for studying respiratory viral infections.
Supplemental Material
sj-docx-1-ini-10.1177_17534259251331385 - Supplemental material for Tracking mucosal innate immune responses to three influenza A virus strains in a highly translational pig model using nasopharyngeal swabs
Supplemental material, sj-docx-1-ini-10.1177_17534259251331385 for Tracking mucosal innate immune responses to three influenza A virus strains in a highly translational pig model using nasopharyngeal swabs by Helena A Laybourn, Charlotte Kristensen, Anders G Pedersen, Louise Brogaard, Sophie George, Betina L Henriksen, Chrysillis H Polhaus, Ramona Trebbien, Lars E Larsen and Kerstin Skovgaard in Innate Immunity
Supplemental Material
sj-xlsx-2-ini-10.1177_17534259251331385 - Supplemental material for Tracking mucosal innate immune responses to three influenza A virus strains in a highly translational pig model using nasopharyngeal swabs
Supplemental material, sj-xlsx-2-ini-10.1177_17534259251331385 for Tracking mucosal innate immune responses to three influenza A virus strains in a highly translational pig model using nasopharyngeal swabs by Helena A Laybourn, Charlotte Kristensen, Anders G Pedersen, Louise Brogaard, Sophie George, Betina L Henriksen, Chrysillis H Polhaus, Ramona Trebbien, Lars E Larsen and Kerstin Skovgaard in Innate Immunity
Supplemental Material
sj-docx-3-ini-10.1177_17534259251331385 - Supplemental material for Tracking mucosal innate immune responses to three influenza A virus strains in a highly translational pig model using nasopharyngeal swabs
Supplemental material, sj-docx-3-ini-10.1177_17534259251331385 for Tracking mucosal innate immune responses to three influenza A virus strains in a highly translational pig model using nasopharyngeal swabs by Helena A Laybourn, Charlotte Kristensen, Anders G Pedersen, Louise Brogaard, Sophie George, Betina L Henriksen, Chrysillis H Polhaus, Ramona Trebbien, Lars E Larsen and Kerstin Skovgaard in Innate Immunity
Supplemental Material
sj-docx-4-ini-10.1177_17534259251331385 - Supplemental material for Tracking mucosal innate immune responses to three influenza A virus strains in a highly translational pig model using nasopharyngeal swabs
Supplemental material, sj-docx-4-ini-10.1177_17534259251331385 for Tracking mucosal innate immune responses to three influenza A virus strains in a highly translational pig model using nasopharyngeal swabs by Helena A Laybourn, Charlotte Kristensen, Anders G Pedersen, Louise Brogaard, Sophie George, Betina L Henriksen, Chrysillis H Polhaus, Ramona Trebbien, Lars E Larsen and Kerstin Skovgaard in Innate Immunity
Footnotes
Acknowledgements
Karin Tarp is kindly thanked for her technical assistance.
ORCID iDs
Ethical considerations
The animal experiment was performed under biosafety level 2 conditions and in accordance with an animal study protocol approved by The Danish Animal Experimentation Council (protocol no. 2020-15-0201-00502).
Author contributions/CRediT
HAL: Data curation, Formal Analysis, Visualization, Investigation, Writing – original draft, Writing – review & editing. CK: Formal Analysis, Visualization, Investigation, Writing – review & editing. AGP: Formal Analysis, Software, Writing – review & editing. LB: Formal Analysis, Writing – review & editing. SG: Formal Analysis, Visualization, Writing – review & editing. BLH: Formal Analysis, Writing – review & editing. CHP: Formal Analysis, Writing – review & editing. RT: Conceptualization, Methodology, Resources, Writing – review & editing. LEL: Conceptualization, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Writing – review & editing. KS: Conceptualization, Investigation, Methodology, Project administration, Resources, Supervision, Writing – review & editing.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work presented in this study is part of the FluZooMark project supported by Novo Nordisk Fonden (grant number NNF19OC0056326).
Conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
