Abstract
Pseudomonas aeruginosa infection can induce alveolar macrophage apoptosis and autophagy, which play a vital role in eliminating pathogens. These two processes are usually not independent. Recently, autophagy has been found to interact with apoptosis during pathogen infections. Nevertheless, the role of autophagy in P. aeruginosa-infected cell apoptosis is unclear. In this study, we explored the impact of P. aeruginosa infection on autophagy and apoptosis in RAW264.7 cells. The autophagy activator rapamycin was used to stimulate autophagy and explore the role of autophagy on apoptosis in P. aeruginosa-infected RAW264.7 cells. The results indicated that P. aeruginosa infection induced autophagy and apoptosis in RAW264.7 cells, and that rapamycin could suppress P. aeruginosa-induced apoptosis by regulating the expression of apoptosis-related proteins. In addition, rapamycin scavenged the cellular reactive oxygen species (ROS) and diminished p-JNK, p-ERK1/2 and p-p38 expression of MAPK pathways in RAW264.7 cells infected with P. aeruginosa. In conclusion, the promotion of autophagy decreased P. aeruginosa-induced ROS accumulation and further attenuated the apoptosis of RAW264.7 cells through MAPK pathway. These results provide novel insights into host–pathogen interactions and highlight a potential role of autophagy in eliminating P. aeruginosa.
Introduction
Pseudomonas aeruginosa, a Gram-negative bacterium, is commonly found in moist environments, and can cause acute and chronic infectious diseases. 1 Severe P. aeruginosa infections, such as urinary tract infections, ventilator-associated pneumonia and digestive tract infections, are determinants of mortality for hospitalized patients. 2 Notably, antibiotics are often used to prevent complications in P. aeruginosa-infected patients, an approach challenged by drug resistance. Thus, there is an urgent need to identify novel therapeutic targets for P. aeruginosa infection. According to previous studies, therapies based on autophagy and the innate immunity defence system of the host is an effective way to solve this problem. 3 , 4 Macrophages, being the host of P. aeruginosa and typical APCs of the innate immune system, play a substantial role in the development of P. aeruginosa infection. 5 Therefore, it is particularly essential to elucidate the interaction between macrophages and P. aeruginosa infection.
Macrophages can kill invading pathogens through phagocytosis and destruction of intracellular microorganisms. In addition to phagocytosis and intracellular killing, autophagy and apoptosis are also involved in innate immune response processes. 6 A process of cell self-digestion, autophagy is responsible for protein and organelle degradation to maintaining intracellular homeostasis. Thus, activation of autophagy in macrophages is generally regarded as a protective strategy against intracellular pathogens. However, autophagy dysfunction is associated with metabolic diseases, cancer, neurodegenerative diseases and microbial infection. 7 In this regard, autophagy is a double-edged sword, serving either pro-survival or pro-death functions. Different from autophagy, apoptosis is another type of programmed cell death that plays critical roles in maintenance of organism homeostasis by eliminating damaged cells. 8 However, several researchers uncovered that pathogens such as P. aeruginosa possessed the ability to interfere and evade the host immune system by inducing targeted cell apoptosis. 9 Accordingly, preventing excessive pathogen-induced cell apoptosis should be beneficial for eliminating bacteria. Interestingly, cellular contexts required a balanced interplay between autophagy and apoptosis, highlighting the relevance and impact of this crosstalk in normal development and in pathology. 10 For instance, etoposide, a general inducer of apoptosis, has been demonstrated to activate autophagy in mouse embryonic fibroblasts. 11 Moreover, autophagy also can activate caspase-dependent apoptosis. 12 Autophagy acts as a cellular cleanser, functioning to degrade misfolded proteins and attenuate endoplasmic reticulum stress, which further inhibits the activation of apoptosis. 13 These findings may imply that autophagy plays a modulatory role in apoptosis of host cells in response to a pathogen invasion. However, the mechanism of modulating the dynamic balance between autophagy and apoptosis during P. aeruginosa infection remains obscure. Thus, this study aimed to investigate whether autophagy is involved in regulating apoptosis in RAW264.7 cells infected with P. aeruginosa.
Materials and methods
Cell lines and P. aeruginosa strains
The macrophage mouse cell line RAW264.7 was purchased from the Shanghai Institute of Biochemistry and Cell Biology (Shanghai, China), cultured at 37°C and 5% CO2, in DMEM (Gibco, CA, USA) supplemented with 10% heated-inactivated FBS (Thermo Fisher) and 2 mM
P. aeruginosa strains were purchased from the American Type Culture Collection (ATCC, 9027) and cultured in LB broth.
Materials and reagents
The following reagents were used: Abs against LC3, ATG5, BAX, Bcl-2, caspase-8, and β-actin were purchased from Proteintech (CA, USA). Abs against JNK, phosphor-JNK, p38, phosphor-p38, ERK1/2, phosphor-ERK1/2, P62/SQSTM1 and cleaved caspase-3 were purchased from Cell Signaling Technology (Boston, USA). Abs against caspase-9 and poly (ADP-ribose) polymerases (PARP) were purchased from Abcam (Cambridge, United Kingdom). Goat anti-mouse IgG (H+L) cross-adsorbed secondary Ab Alexa Fluor 555 was purchased from Invitrogen (Massachusetts, USA). Immunofluorescence reagents were obtained from Zhongshan Golden Bridge Biotechnology (Beijing, China). An Annexin V/PI kit was purchased from KeyGen Biotech (Jiangsu, China). Reactive oxygen species (ROS) and JC-1 kits were purchased from Thermo Fisher Scientific (Massachusetts, USA). CytoID autophagic detection kit was purchased from Enzo Life Sciences. Rapamycin (RAPA), 3-Methyladenine (3-MA), N-acetylcysteine (NAC) and SB-202190 were purchased from MedChemExpress (New Jersey, USA).
Infection of the RAW264.7 macrophages with P. aeruginosa
The bacteria were cultured at 37°C (180 rpm/min) for 12 h and then centrifuged at 5000 g for 5 min at room temperature (20–25°C). The supernatant was discarded, and the cells were washed once with sterile saline (0.85% NaCl) and then re-suspended in saline to a concentration of approximately 2.0 × 1010 CFU/ml using a standard curve relating viable counts to the OD. The RAW264.7 cells were infected with P. aeruginosa at MOI of 20 under the indicated conditions before being harvested for analysis.
Western blot analysis
The cells were washed three times with PBS at 4°C, and the entire population was extracted and lysed with lysis buffer for 30 min on ice. Next, the lysates were centrifuged at 14000 g for 15 min at 4°C, and the precipitates and supernatants were collected. Then, the total protein (20–50 μg) was separated by SDS-PAGE, and the separated proteins were transferred by the wet method to PVDF membranes. Subsequently, the membranes were blocked with Superblock (Thermo Fisher Scientific, USA) for 2 h at room temperature. Next, the membranes were incubated with primary Abs overnight at 4°C and then with the appropriate secondary Abs for 2 h at room temperature. Finally, the proteins were visualized by an enhanced chemiluminescence (Advansta Inc., San Jose, CA, USA) system. The results were analysed by ImageJ software, and β-actin was used as the internal control.
Immunofluorescence detection
RAW264.7 cells were plated into 24-well plates at 1 × 105 cells/well, and adherent monolayers were infected with P. aeruginosa (MOI = 10). The cells were washed three times with ice-cold PBS and fixed with 4% paraformaldehyde for 20 min. The cells were then washed three times and permeabilized with TritonX-100 solution (0.01% in PBS). Next, the cells were re-washed, blocked with 5% normal goat serum for 30 min, and incubated with LC3 and caspase-3 Abs (1:50 dilution) for 12 h at 4°C. Finally, the cells were incubated with fluorescent secondary Abs for 1 h at room temperature, and nuclei were stained with DAPI.
Detection of apoptosis by flow cytometry
RAW264.7 cells were plated into six-well plates at 1× 106 cells/well, treated with rapamycin (100 nM) for 2 h and then infected with P. aeruginosa (MOI = 20) for 4 h. The apoptotic cells were measured by FACS using an Annexin V/PI kit (KeyGENBioTECH, Jiangsu, China). The cells were washed with PBS at 4°C and centrifuged for 5 min at 1000 r/min; the supernatant was discarded, and the cell pellet was suspended in 400 μl of 1 × binding buffer. Next, 5 μl of Annexin V-FITC and 5 μl of PI staining solution were added to the cells in each treatment group, after which the cells were gently mixed at incubated at room temperature in the dark for 20 min. The samples were analysed by flow cytometry (BD Biosciences, Franklin Lakes, NJ, USA) within 1 h.
Measurement of mitochondrial membrane potential
RAW264.7 cells were seeded into 6-well plates at 1 × 106 cells/well, pre-treated with rapamycin (100 nM) for 2 h or 3-methyladenine (3-MA, 5 mM) for 6 h and then treated with P. aeruginosa (MOI = 20) for 4 h. The mitochondrial membrane potential (MMP) was measured by FACS using a JC-1 kit (Thermo, USA). The JC-1 working solution was prepared by mixing 500 μl of PBS with 1 μl of JC-1 and mixing eddies. The cells were mixed with 500 μl of JC-1 working fluid, incubated at room temperature for 20 min and centrifuged for 5 min; the supernatant was discarded, and the cell pellet was re-suspended in PBS. Finally, the cells were detected by flow cytometry within 1 h. The results were determined by flow cytometry and a BD FACSCalibur system (BD Biosciences, Franklin Lakes, NJ, USA), and the data were analysed with Cell Quest software.
Detection of ROS by flow cytometry
Intracellular ROS levels were evaluated by flow cytometry. RAW264.7 cells were seeded in 6-well plates at a density of 1 × 106 cells/well, treated with rapamycin (100 nM) for 2 h and infected with P. aeruginosa for 4 h. The cells were then harvested and incubated with Cell ROX Orange Reagent (ThermoFisher Scientific, USA) for 20 mins. The cells were washed, harvested from the wells by scraping and analysed for intracellular fluorescence using a BD-FACS can instrument (BD, USA) and Cell Quest software.
Detection of autophagic vacuoles by flow cytometry
RAW264.7 cells were seeded in 6-well plates at a density of 1 × 106 cells/well, treated with rapamycin (100 nM) for 2 h or 3-MA (5 mM) for 6 h and infected with P. aeruginosa for 4 h. The formation of autophagic vacuoles was detected by CytoID detection kit (Enzo Life Sciences, USA). Briefly, CytoID working solution was prepared by mixing 500 μl of PBS with 1 μl of CytoID and mixing eddies. The cells were suspended with 500 μl CytoID working solution and incubated at room temperature for 30 min. The results were determined by flow cytometry.
Statistical analyses
Data statistical analysis was performed using GraphPad Prism version 8.0 (GraphPad Software, USA). The data are presented as the mean ± SD, and all data collected were obtained from at least three independent experiments for each condition. The data were statistically evaluated to compare differences between two groups by one-way ANOVA; *P < 0.05; **P < 0.01; ***P < 0.001.
Results
P. aeruginosa infection induced autophagy in RAW264.7 cells
RAW264.7 cells were infected with P. aeruginosa at various time and different MOI. As expected, P. aeruginosa infection increased the numbers of LC3-positive cells (Figure 1a). In addition, the expression of LC3 was in a time and dose-dependent manner (Figure 1b, c).
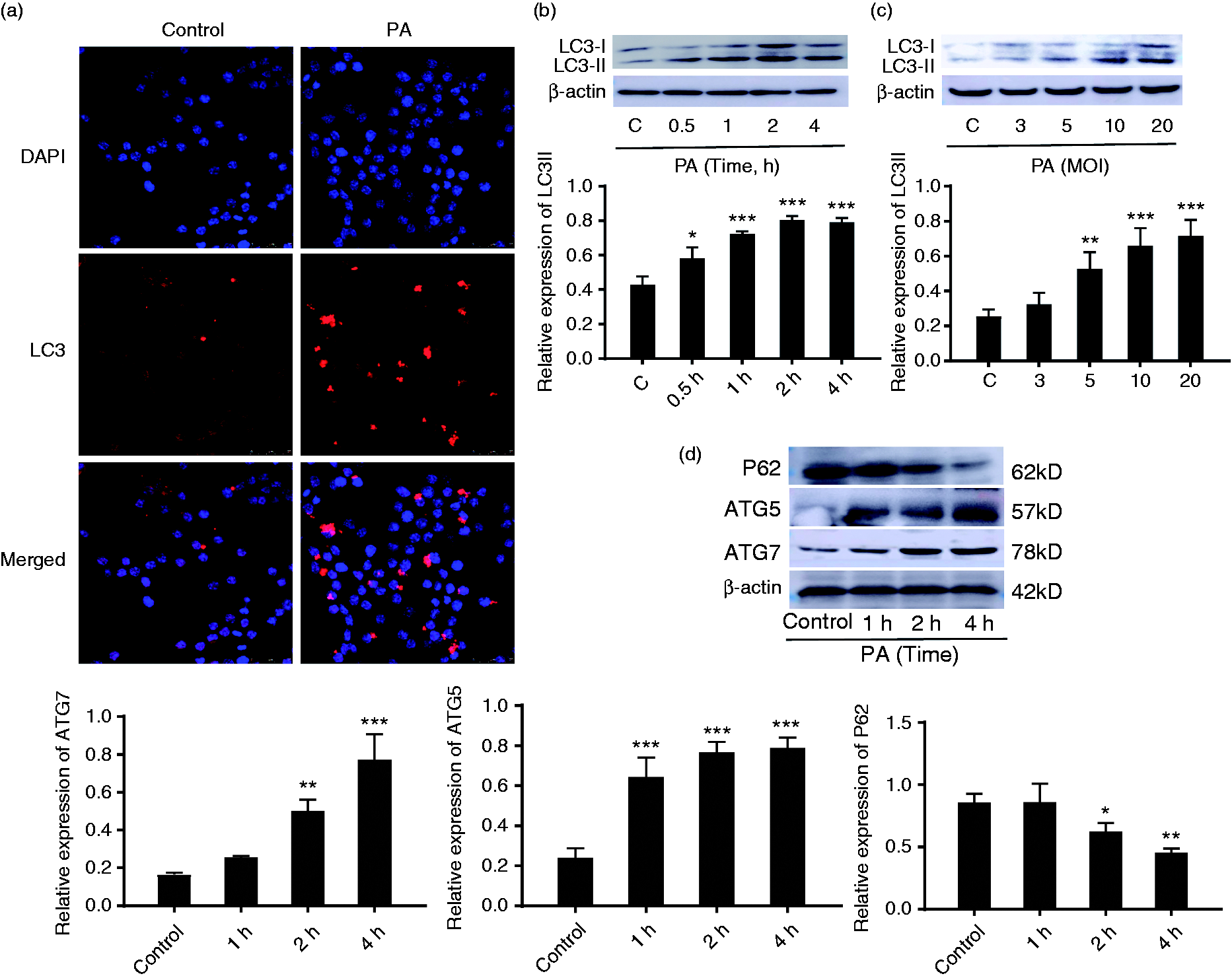
P. aeruginosa infection induced autophagy of RAW264.7 cells. (a) Immunofluorescence was used to detect intracellular LC3 puncta in RAW264.7 cells infected with P. aeruginosa (PA) for 2 h. (b,c) The expression of LC3II in RAW264.7 cells after infection with P. aeruginosa at different time and MOI were analysed by Western blot. (d) The expression of ATG5, ATG7 and P62 in RAW264.7 at different time points after P. aeruginosa infection. The protein ratio was calculated by ImageJ densitometry analysis. Data are shown as mean ± SEM, and one representative experiment from three independent experiments is shown. *P < 0.05; **P < 0.01; ***P < 0.001.
Importantly, autophagy-related (ATG) proteins known to regulate autophagy, such as ATG5 and ATG7, are involved in the extension and closure of autophagic vesicles. However, P62, a lysosomal substrate, is degraded when autophagosomes fuse with lysosomes. To further confirm P. aeruginosa-induced autophagy, Western blot was used to detect the expression of the autophagy-related proteins ATG5, ATG7 and P62 in RAW264.7 cells.
Compared with uninfected cells, the expression of ATG5 and ATG7 was significantly increased, accompanied with a decreasing P62 production in P. aeruginosa-infected RAW264.7 cells (Figure 1d). Furthermore, the formation of autophagic vacuoles was measured by flow cytometry. As shown in the Supplemental (Figure S1), P. aeruginosa infection increased the formation of autophagic vacuoles in RAW264.7 cells. These results unambiguously suggested that P. aeruginosa could induce RAW264.7 cells autophagy, which was consistent with the previous finding from other groups. 14
P. aeruginosa infection activated apoptosis in RAW264.7 cells
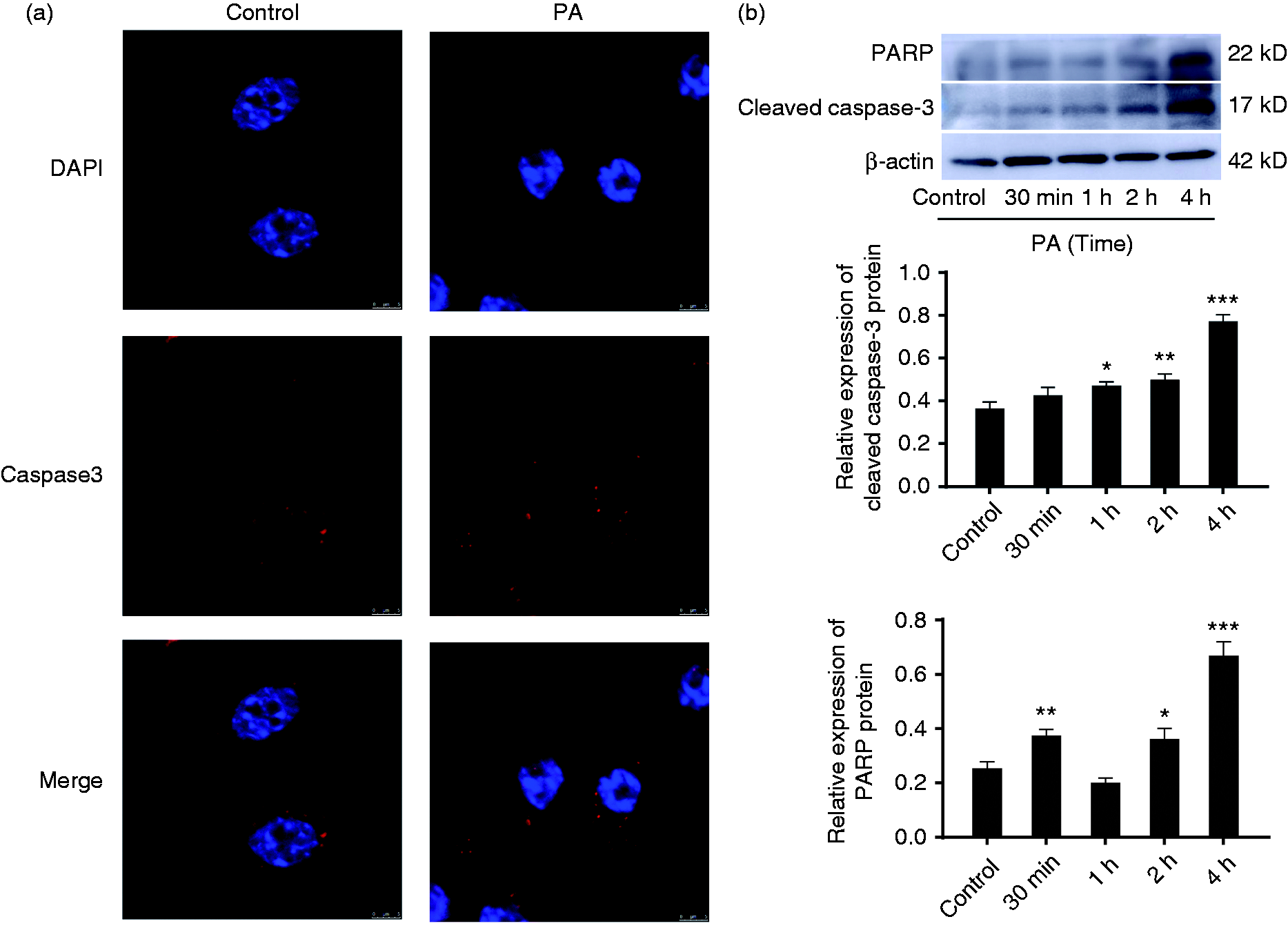
To confirm whether P. aeruginosa infection could induce apoptosis, RAW264.7 cells were infected with P. aeruginosa at a MOI of 20 in the indicated conditions. As shown in Figure 2a, P. aeruginosa infection increased the numbers of caspase-3-positive cells. In addition, the expression of cleaved caspase-3 and PARP detected by Western blot were up-regulated in a time-dependent manner in RAW264.7 cells upon P. aeruginosa infection (Figure 2b). The results indicated that P. aeruginosa infection could induce cell apoptosis.
Figure 2. P. aeruginosa infection activated apoptosis in RAW264.7 cells. (a) Fluorescence microscopy was used to detect intracellular caspase-3 expression (red puncta) of RAW264.7 cells infected with P. aeruginosa (PA) for 2 h. (b) Time-dependent expression of cleaved caspase-3 and PARP in the RAW264.7 cells infected with P. aeruginosa (top panel). Western blot was used to analyse the expression of the proteins. The protein ratio was calculated using ImageJ densitometric analysis (bottom panel). The data are presented as mean ± SEM, and one representative experiment of three independent experiments is shown. *P < 0.05; **P < 0.01; ***P < 0.001 compared with the control group.
The promotion of autophagy inhibits P. aeruginosa-induced apoptosis in RAW264.7 cells
As both autophagy and apoptosis occurred in macrophages during P. aeruginosa infection, we hypothesized that apoptosis could be regulated by autophagy. To confirm the hypothesis, RAW264.7 cells were pre-treated with rapamycin or 3-MA following with P. aeruginosa infection for 4 h. The flow cytometry results showed P. aeruginosa infection significantly increased the apoptosis rate (Figure 3a, b). Meanwhile, compared with P. aeruginosa infection alone, rapamycin treatment effectively reduced the number of apoptotic cells. In contrast, 3-MA increased the number of apoptotic cells (Figure S2). The previously mentioned results indicated that the activation of autophagy attenuated the apoptosis rate of RAW264.7 cells infected with P. aeruginosa.
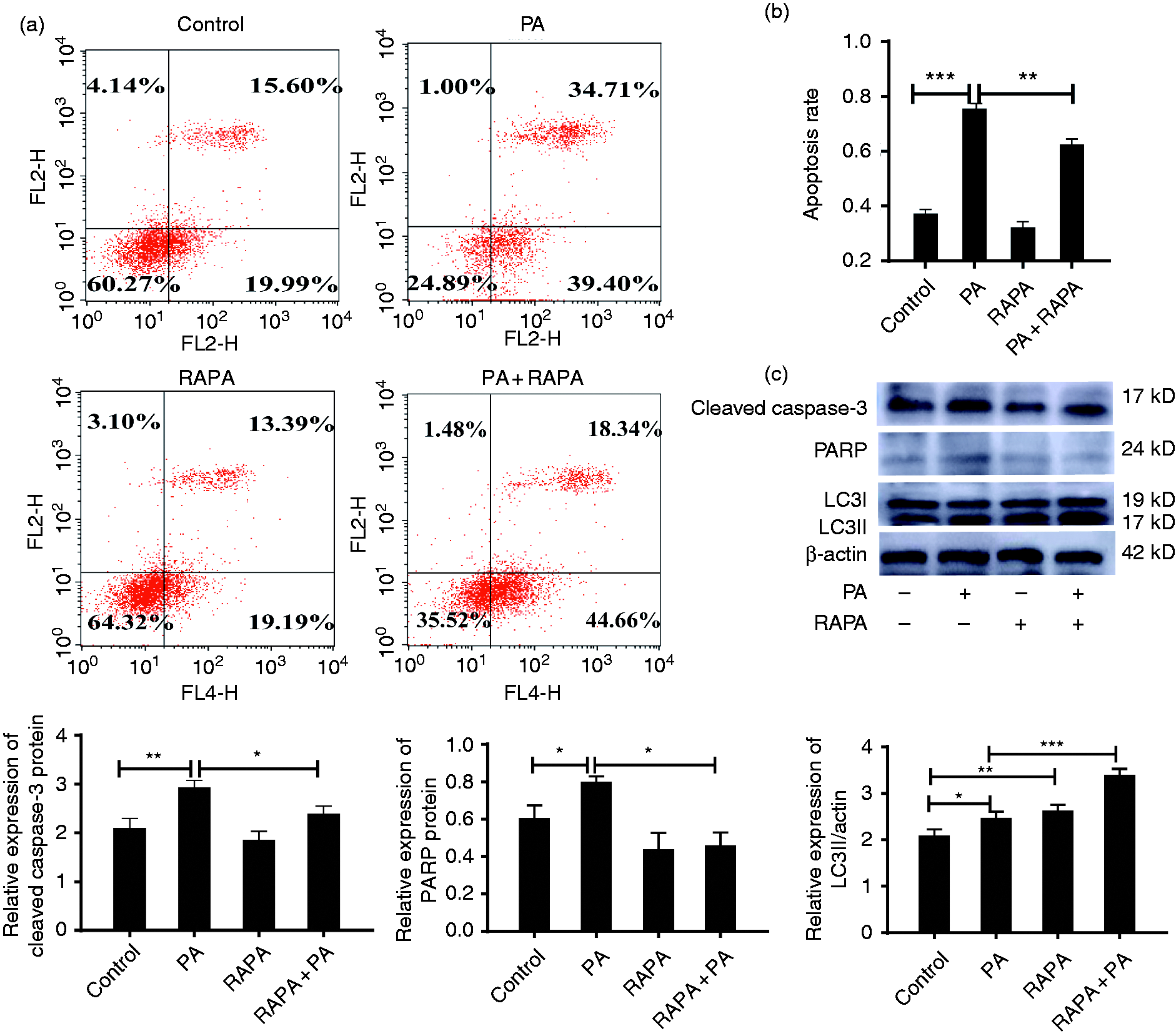
Autophagy prevents P. aeruginosa-induced apoptosis in RAW264.7 cells. (a) RAW264.7 cells were treated with rapamycin (RAPA) for 2 h and then infected with P. aeruginosa (PA) for 4 h. The apoptosis rates were determined by AnnexinV-FITC (AV) and PI staining. (b) Histogram of the RAW264.7 cell apoptosis rate as determined by the FACS assay. (c) The protein expression levels of PARP, cleaved caspase-3 and LC3 in RAW264.7 cells treated with P. aeruginosa or/and 10 μM rapamycin were determined by Western blot analysis. The data are represented as mean ± SD, and one representative experiment of three independent experiments is shown. *P < 0.05; **P < 0.01; ***P < 0.001 compared with the control group.
Subsequently, to further provide definitive evidence, we detected the expression of cleaved caspase-3, PARP and LC3II in RAW264.7 cells by Western blot. Figure 3c shows that P. aeruginosa infection alone resulted in a marked increase the expression of PARP and cleaved caspase-3 in RAW264.7 cells. In contrast, rapamycin exhibited a capacity to suppress, while 3-MA promoted, P. aeruginosa-induced PARP and cleaved caspase-3 expression (Figure S3). Taken together, these results demonstrated that autophagy could reduce P. aeruginosa-induced RAW264.7 cells apoptosis.
Autophagy alleviated P. aeruginosa-induced apoptosis via the intrinsic apoptotic pathway
According to whether caspase-8 or caspase-9 is activated, apoptosis is divided into extrinsic and intrinsic pathways, respectively. 15 We detected caspase-8 and caspase-9 expression to explore which pathway was involved in the process. The results showed that P. aeruginosa infection dramatically up-regulated the expression of caspase-8 and caspase-9 in RAW264.7 cells (Figure 4a). Interestingly, rapamycin decreased the P. aeruginosa-induced caspase-9 expression, but not that of caspase-8. These results indicated that autophagy alleviated P. aeruginosa-induced apoptosis through the intrinsic apoptotic pathway.
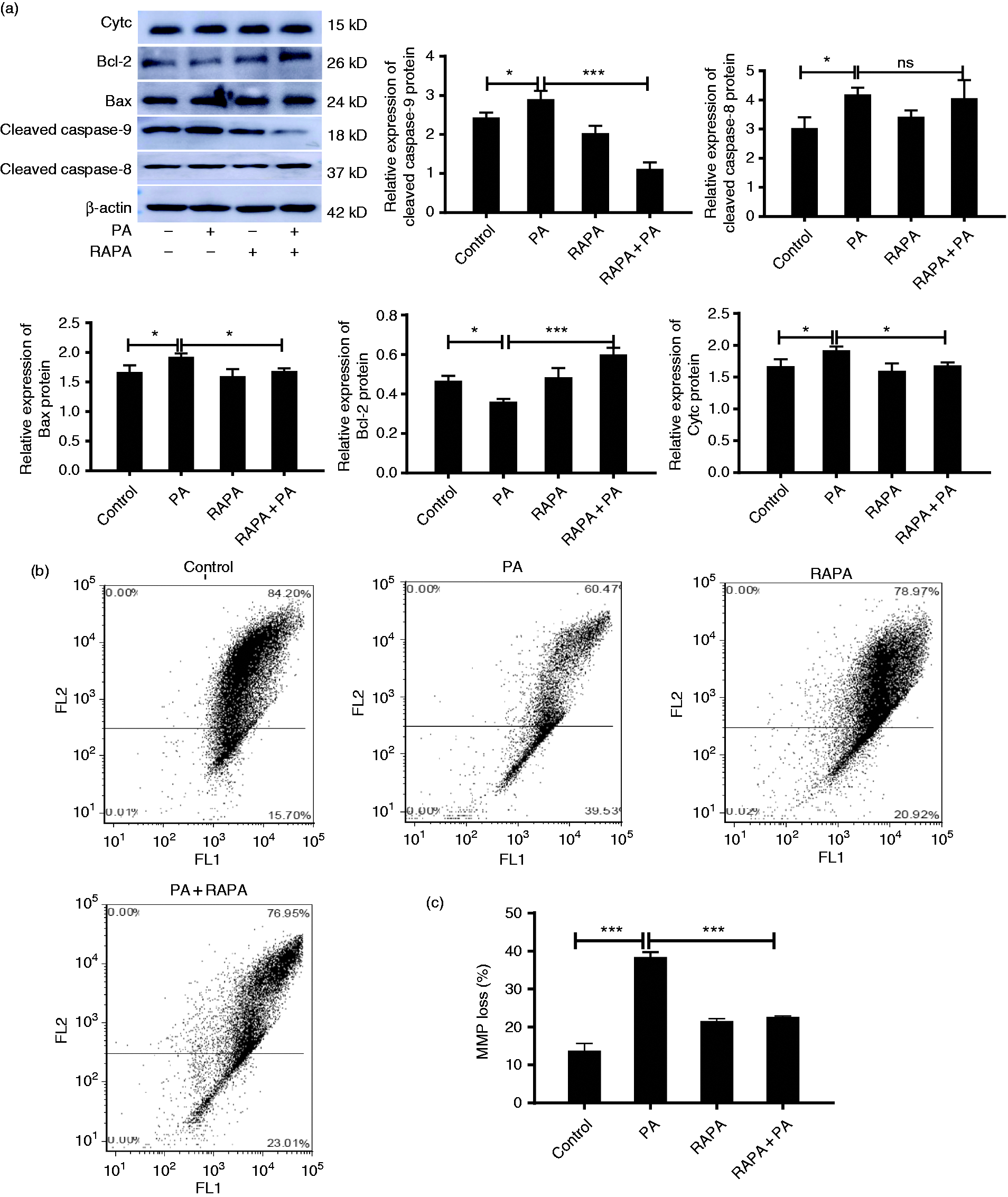
Autophagy alleviated P. aeruginosa-induced intrinsic apoptosis. (a) RAW264.7 cells were treated with P. aeruginosa (PA) or/and 10 μM rapamycin (RAPA). The expression of cleaved caspase-8, cleaved caspase-9, cytc, Bax and Bcl-2 were measured by Western blot. (b) Mitochondrial membrane potential (MMP) was determined by flow cytometry. (c) Quantitative analysis of MMP in RAW264.7 cells by the FACS assay. The data are shown as mean ± SEM of three independent experiments. NS, no significance; *P < 0.05; **P < 0.01; ***P < 0.001.
Subsequently, we examined intrinsic apoptotic pathway-related proteins (Figure 4a). As expected, P. aeruginosa remarkably increased the expression of the pro-apoptotic proteins Bax and cytochrome C (cytc) in RAW264.7 cells and down-regulated the expression of the anti-apoptotic protein Bcl-2. In contrast, rapamycin reversed the associated protein expression induced by P. aeruginosa infection. Furthermore, loss of the integrity of the MMP was observed by flow cytometry assay based on JC-1 staining. As shown in Figure 4b and 4c, P. aeruginosa infection moderately decreased the MMP of RAW264.7 cells, while rapamycin partially reversed the loss of MMP induced by P. aeruginosa infection. Together, these data unambiguously indicated that autophagy could partially alleviate mitochondrial-mediated intrinsic apoptosis of RAW264.7 cells promoted by P. aeruginosa infection.
Autophagy attenuated P. aeruginosa-induced apoptosis via ROS-dependent MAPK pathway
Autophagy can depress the production of ROS which play key roles in mediating the intrinsic apoptotic pathway. 10 Therefore, we hypothesized that the decreased apoptosis rate induced by autophagy is achieved via scavenging cellular ROS. To confirm the hypothesis, we explored the effect of autophagy on the potential of -mediated ROS production to determine the biological significance of autophagy in P. aeruginosa infection. We first used flow cytometry to detect ROS levels in RAW264.7 treated with rapamycin for 2 h and then infected with P. aeruginosa for 2 h. The results are shown in Figure 5a and 5b. P. aeruginosa infection significantly accumulated ROS production while rapamycin partial reversed P. aeruginosa-induced ROS accumulation in RAW264.7 cells. Collectively, these data implied that autophagy attenuates the ROS production induced by P. aeruginosa infection.
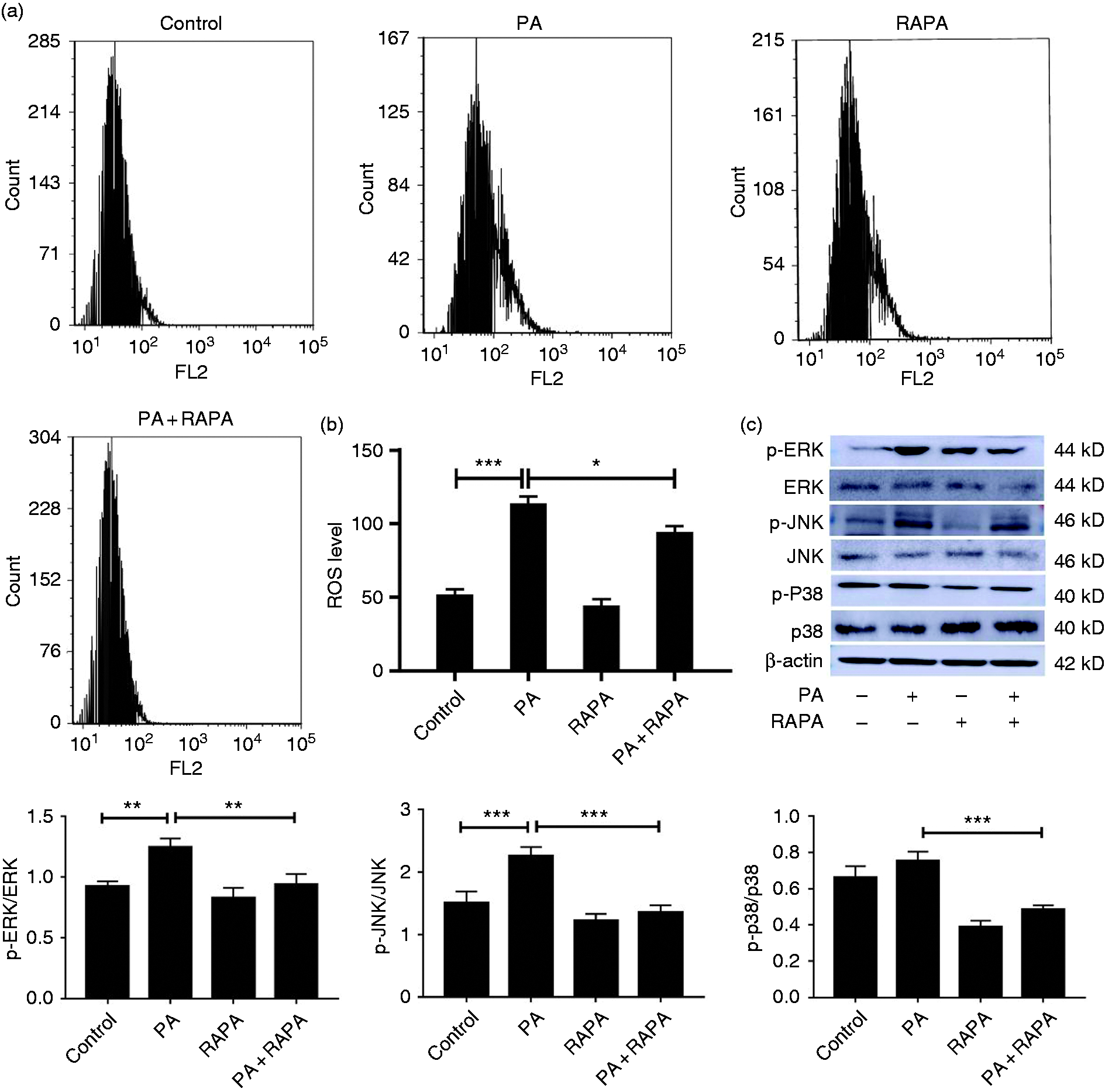
Autophagy blocked P. aeruginosa-induced activation of ROS-MAPK pathways in RAW264.7 cells. (a) ROS accumulation was determined by flow cytometry in RAW264.7 cells which were treated with rapamycin (RAPA) for 2 h, followed by exposure to P. aeruginosa (PA) for 4 h. (b) Quantitative analysis of ROS levels in RAW264.7 cells by FACS assay. (c) The expression of p-JNK, p-ERK1/2 and p-p38 was analysed by Western blot after exposure to rapamycin and P. aeruginosa alone or combined. The data are shown as mean ± SEM of three independent experiments. NS, no significance; *P < 0.05; **P < 0.01; ***P < 0.001.
According to a previous study, autophagy may alter the activity of MAPKs, including JNK, ERK1/2 and p38, and affect cell proliferation, survival and apoptosis under various conditions. 16 Therefore, to further verify whether autophagy prevents P. aeruginosa-induced RAW264.7 cell apoptosis by MAPK pathway, we detected p-JNK, p-ERK1/2 and p-p38 protein expression of RAW264.7 cells treated with rapamycin for 2 h or 3-MA for 6 h and then exposed to P. aeruginosa for 4 h. The results showed that rapamycin potently suppressed the activation of p-JNK, p-ERK1/2 and p-p38 production in RAW264.7 cells infected with P. aeruginosa (Figure 5c). However, 3-MA promoted the activation of p-JNK and p-p38 (Figure S3). To further address our conclusion, NAC, which is a ROS scavenger, was employed to block ROS accumulation. As shown in Figure S4-a, NAC depressed the activation of MAPK pathways and abolished the 3-MA-induced MAPK pathway activation in P. aeruginosa-infected cells. What is more, we also employed SB-202190 to block p38 phosphorylation. The expression of apoptosis-related factors were detected by Western blot (Figure S4-b). The results showed that the inhibition of MAPK pathways by SB-202190 decreased the expression of apoptosis and alleviated 3-MA induced apoptosis in P. aeruginosa-infected cells. Taken together, these results clearly suggested that activated autophagy attenuated P. aeruginosa-induced cells apoptosis by scavenging the cellular ROS and blocking MAPK pathway.
Discussion
P. aeruginosa is a highly infectious bacterium which can cause host cell death, tissue and organ damage, and destroy the body’s immune mechanisms. 17 Exploring the interaction between pathogenic bacteria and host cells is very important for a comprehensive understanding of the pathogenic mechanism of P. aeruginosa infection. In addition, this is increasingly critical in developing strategies to prevent P. aeruginosa infection. 18
Accumulating evidence has shown that autophagy is considered an essential pathway for eliminating intracellular pathogenic bacteria infections.19–22 Moreover, apoptosis is also involved in the process for bacterial infection. However, cell fate is neither determined by autophagy nor by apoptosis. To date, a thorough comprehension of multifaceted relationship between autophagy and apoptosis has been established in many diseases. 23 , 24 A previous study reported that autophagy plays a protective role in high Glc-induced apoptosis in adipose tissue-derived stem cells. 25 On the contrary, once apoptosis is blocked, autophagy can induce cell death. Meanwhile, elimination of damaged mitochondria reduces ROS accumulation, creating a feedback loop that causes inhibition of autophagy. 26 Thus, exploring the role of autophagy in apoptosis will contribute to understanding the pathogenesis of P. aeruginosa. With this in mind, we initially hypothesized that autophagy could be involved in regulating the apoptosis induced by P. aeruginosa infection.
We first determined the impact of P. aeruginosa infection on autophagy. The expression of LC3 in RAW264.7 cells was detected by Western blot and immunofluorescence. The results indicated that P. aeruginosa infection increased the numbers of LC3-positive cells and expression of LC3II as well in RAW264.7 cells. Consistent with the LC3II result, ATG7 and ATG5 expression was elevated in a time- and dose-dependent manner by P. aeruginosa infection followed by a decreased P62 expression. This implied that P. aeruginosa infection induced autophagy in RAW264.7 cells. Subsequently, we confirmed that P. aeruginosa infection induced RAW264.7 cell apoptosis with an increased expression of cleaved caspase-3 and PARP. Thus, consistent with other findings, P. aeruginosa infection can induce cell autophagy and apoptosis. 14 , 27
In an effort to evaluate the role of autophagy on apoptosis, we used rapamycin or 3-MA to regulate autophagy. Flow cytometry results showed that rapamycin effectively reduced the apoptosis rate of RAW264.7 cells upon P. aeruginosa infection followed by decreasing cleaved caspase-3 and PARP expression. However, 3-MA exhibited a totally reversed function. These findings were consistent with a previous model, in which the authors determined that rapamycin protects cell death by activating autophagy, inhibiting oxidative stress, and mitochondria apoptosis. 28
The intrinsic and extrinsic pathways are considered to be two different apoptosis pathways. One characteristic of intrinsic apoptosis is mitochondrial dysfunction, which occurs upon the release of cytc from the mitochondrion into the cytoplasm. These molecules lead to the self-splicing of caspase-9, which triggers caspase-9 activation, and enhances the catalytic properties of other caspases, such as caspase-3 maturation. This process is tightly regulated by Bcl-2 family proteins, which function in the mitochondrial membrane. 29 According to our study, rapamycin decreased the P. aeruginosa-induced expression of the caspase-9, Bax and cytc proteins, but increased the expression of Bcl-2. Furthermore, the activation of mitochondrial apoptosis was also associated with a change in MMP. A flow cytometry assay also showed that rapamycin partially reversed the loss of MMP induced by P. aeruginosa infection. However, rapamycin did not change the expression of the extrinsic apoptotic pathway-related protein caspase-8 in RAW264.7 cells infected with P. aeruginosa. Collectively, our results indicated that autophagy inhibited P. aeruginosa-induced apoptosis in RAW264.7 cells via the intrinsic apoptotic pathway, and not by the extrinsic apoptotic pathway.
Increasing evidence has demonstrated that during pathogen infection, ROS-induced oxidative stress could cause cell injury through mitochondrial dysfunction and reduced activity of antioxidant enzymes.30–32 Multiple studies have revealed that exposure to P. aeruginosa can initiate apoptotic cell death via excessive ROS generation. 33 , 34 Given this evidence, we investigated the effect of rapamycin treatment on P. aeruginosa-induced ROS accumulation. The flow cytometry results showed that rapamycin effectively diminished ROS production in response to P. aeruginosa infection, indicating that autophagy inhibits P. aeruginosa-induced ROS production. As mentioned previously, ROS have been shown to induce apoptosis, therefore, decreased ROS accumulation may be the potential mechanism by which autophagy regulates apoptosis during P. aeruginosa infection.
The MAPK pathway, including the critical members of JNK, p38 and ERK, is a pivotal cellular signalling pathway implicated in cell survival and apoptosis. 35 Mechanistically, activation of the MAPK pathway is tightly related to ROS-induced apoptosis. 36 Moreover, P. aeruginosa infection can activate all three MAPK members, further leading to apoptosis through ROS accumulation. 37 In addition, many lines of evidence confirmed that rapamycin can alter the activity of MAPKs, which further affects cell proliferation, survival and apoptosis.38–40 We proposed that the autophagy-mediated regulation of MAPK signalling pathway and scavenging ROS should be critical for inhibiting P. aeruginosa-induced apoptosis.
Finally, we examined the expression of p-JNK, p-ERK1/2 and p-p38 by Western blot analysis. The result strongly confirmed that the activation of autophagy blocked the MAPK signalling pathway by down-regulating JNK, ERK1/2 and p38 expression in P. aeruginosa-infected RAW264.7 cells. In contrast, the inhibition of autophagy by 3-MA activated them. The previously mentioned findings were further confirmed by ROS scavenger and p38 MAP kinase inhibitor. The results showed that ROS scavengers abolished 3-MA-induced MAPK pathways activation and the inhibition of MAPK pathways alleviated 3-MA induced apoptosis in P. aeruginosa-infected cells. Conclusively, we defined a mechanism by which autophagy activated by rapamycin suppressed P. aeruginosa-induced oxidative damage and apoptosis in RAW264.7 cells.
As illustrated in Figure 6, P. aeruginosa infection could induce RAW 264.7 cells apoptosis and autophagy. Rapamycin could eliminate ROS accumulation, which will further suppress endogenous apoptosis through MAPK pathways in RAW264.7 cells infected with P. aeruginosa. This study suggested that autophagy plays a protective role in modulating apoptosis during P. aeruginosa infection. It also may provide a novel insight in further understanding the molecular mechanism of P. aeruginosa infection.
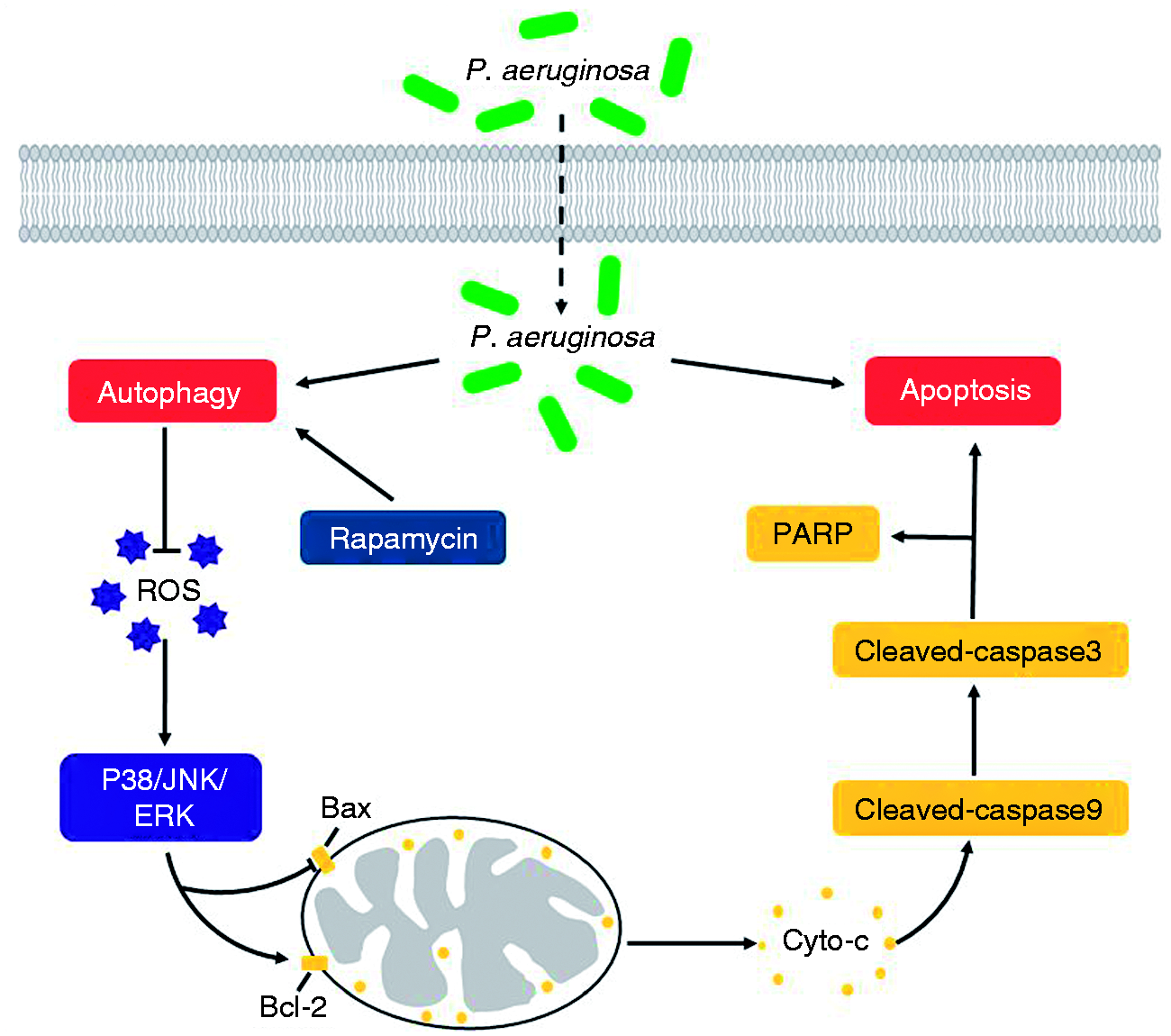
Proposed model for suppressed P. aeruginosa-induced apoptosis in RAW264.7 cells by activation of autophagy. The invasion of P. aeruginosa markedly increases abundance of apoptosis and autophagy in RAW264.7 cells. Rapamycin stimulates autophagy effectively and eliminates the accumulation of ROS which would reduce cellular damage caused by oxidative. Then the activated MAPK signalling components and their down-stream target gene further regulate apoptosis associated proteins Bax, Bcl-2, Cytc, caspase-9, caspase-3 and PARP production. Consequently, the activated autophagy suppresses P. aeruginosa-induced apoptosis in RAW264.7 cells.
Supplemental Material
sj-pdf-1-ini-10.1177_1753425920952156 - Supplemental material for Autophagy plays a protective role during Pseudomonas aeruginosa-induced apoptosis via ROS–MAPK pathway
Supplemental material, sj-pdf-1-ini-10.1177_1753425920952156 for Autophagy plays a protective role during Pseudomonas aeruginosa-induced apoptosis via ROS–MAPK pathway by Lu Han, Qinmei Ma, Jialin Yu, Zhaoqian Gong, Chenjie Ma, Yanan Xu, Guangcun Deng and Xiaoling Wu in Innate Immunity
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the key research and development program of Ningxia [Grant Numbers 2018BFH03017, 2017BN04]; the project for cultivation of scientific and innovative talent [Grant Number KJT2017002] and Innovative program for postgraduate students of Ningxia University [Grant Number GIP2019-13]; The project of building first-class discipline of western higher education [Biology, NXYLXK2017B05].
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
