Abstract
Vitamin D supplementation is regarded as a novel approach to treat Alzheimer’s disease, but the underlying mechanism remains elusive. The cytokine IL-34 provides strong neuroprotective and survival signals in brain injury and neurodegeneration and could be an immunological mediator for the vitamin D-induced protection. The aim of this study was to investigate whether human IL-34 is up-regulated in neuronal cells by the hormonally active form of vitamin D, 1α,25-dihydroxyvitamin D3 [1α,25(OH)2D3]. We found that IL-34 was detectable in a variety of cell lines and its expression was strongly induced in SH-SY5Y neural cells in a dose- and time-dependent manner by 1α,25(OH)2D3 through the vitamin D receptor (VDR). Furthermore, we identified the core promoter of IL-34 gene and a VDR binding site (CGCCCT) that was required for 1α,25(OH)2D3-induced IL-34 expression. These findings suggest that the induction of IL-34 expression by 1α,25(OH)2D3 may constitute a mechanism that explains the protective function of vitamin D in Alzheimer’s disease.
Introduction
IL-34 is a homodimeric cytokine identified by functional screening of a representative set of extracellular proteins. 1 Its receptor is the already well-defined M-CSF receptor and it strongly promotes monocyte viability. 1 Although IL-34 and M-CSF share the same receptor Fms (the proto-oncogene c-fms products and receptor tyrosine kinase), they display different biological activities, signal activation kinetics and signal strength. 2 Recently, the receptor-type protein-tyrosine phosphatase (PTP-ζ), a cell-surface chondroitin sulfate proteoglycan that is highly expressed on glial and glioblastoma cells, has been shown to be another IL-34 receptor. 3 The binding of IL-34 to cell-surface PTP-ζ suppresses cellular activities such as proliferation, clonogenicity and motility in U251 glioblastoma cells. 3
IL-34 is constitutively expressed in various human tissues, including heart, brain, lung, liver, kidney, spleen, thymus, testes, ovary, small intestine, prostate and colon. 1 It is crucial for the development and maintenance of tissue-resident myeloid cells such as Langerhan’s cells, microglia and osteoclasts.4–8 The cellular origins of IL-34 are predominately keratinocytes, neurons and hepatocytes.4,5,9 The pro-inflammatory cytokines, IL-1β and TNF-α, induce IL-34 expression in osteoblasts and gingival fibroblasts.10–12 Additionally, toll-like receptor ligands (peptidoglycan, poly(I:C) and CpG) and TNF-α also induce IL-34 expression in lamina propria mononuclear cells. 13
The active form of vitamin D, calcitriol or 1α,25-dihydroxyvitamin D3 [1α,25(OH)2D3], mediates a variety of biological actions by binding to the cytoplasmic vitamin D receptor (VDR), a nuclear transcription factor, forming a complex with the accessory transcription factor, retinoid X receptor (RXR). This complex translocates to the nucleus and binds in a strongly cell-type-specific manner to vitamin D responsive elements of target genes initiating their expression and thus affecting basic cellular activities, including proliferation, apoptosis and DNA damage repair.14,15 In the murine IL-34 gene five putative VDR binding sites have been found 2 kb upstream of the transcription start site. 16 Both 1α,25(OH)2D3 and 2-methylene-19-nor-(20 S)-1α,25-dihydroxy-vitamin D3, a highly potent analog of 1α,25 (OH)2D3, up-regulate mouse IL-34 mRNA levels in osteoblastic cells in vitro. 16 However, it is unknown whether 1α,25 (OH)2D3 induces human IL-34 expression in vitro and in vivo.
A potential role of vitamin D in Alzheimer’s disease therapy has been discussed based on epidemiological data without strong biological underpinning.17,18 However, findings such as 1α,25(OH)2D3 stimulating amyloid-β phagocytosis by macrophages support the idea.19,20 Since IL-34 promotes a strong neuroprotective effect of microglia that attenuate oligomeric amyloid-β neurotoxicity, thus ameliorating Alzheimer’s disease,21,22 we investigated the regulation of IL-34 gene expression by 1α,25(OH)2D3 in a range of cell lines and, in particular, in SH-SY5Y neural cells. We found a significant up-regulation of IL-34 expression by 1α,25(OH)2D3 mediated by VDR binding to the core promoter.
Materials and methods
Cell culture and 1α,25(OH)2D3 stimulation
Human hepatocellular carcinoma cell line HepG2, human neuroblastoma cell line SH-SY5Y, human glioblastoma cell line A172, and human gastric cancer cell lines SGC-7901 and HGC-27 were maintained in DMEM supplemented with 10% FBS at 37℃ in 5% CO2. Human normal gastric epithelial cell line GES-1 was maintained in RPMI-1640 medium with 10% FBS. In stimulation experiments, SH-SY5Y and GES-1 cells were plated in six-well plates (4 × 105 cells/well) and grown to 80–90% confluency. Cells were treated with 10, 50, 100, 250 and 500 nM 1α,25(OH)2D3 (Sigma, St. Louis, MO, USA) and ethanol (control) for 48 h. Besides, SH-SY5Y and GES-1 cells (4 × 105 cells/dish) were plated in 35-mm dishes and then stimulated with 100 nM 1α,25(OH)2D3 for 0, 12, 24, 36, 48, 60 and 72 h. After stimulation, cells are harvested for quantitative RT-PCR (qRT-PCR) and Western blot analysis.
Quantitative RT-PCR analysis
Total RNA was isolated using Hipure Universal RNA Kit (Magen, Guangzhou, China) according to manufactures instructions and then reverse-transcribed into cDNA using HiScriptIIQ RT SuperMix for qPCR (Vazyme, Nanjing, China). qRT-PCR was performed using AceQ qPCR SYBR® Green Master Mix (Vazyme, Nanjing, China) and Stratagene Mx3000P real-time PCR system (Agilent Technologies, Santa Clara, CA, USA). The primers used were as follows: human IL-34, 5′-AAT CCG TGT TGT CCC TCT TG-3′ (sense) and 5′-CAG CAG GAG CAG TAC AGC AG -3′ (antisense); human GAPDH, 5′-CAG GAG GCA TTG CTG ATG AT -3′ (sense) and 5′- GAA GGC TGG GGC TCA TTT-3′ (antisense). The following PCR conditions were used: initial pre-degeneration at 95℃ for 5 min, followed by a 40-cycle reaction consisting of degeneration at 95℃ for 10 s and annealing and extension at 60℃ for 30 s. IL-34 expression levels were normalized using GAPDH mRNA levels. Relative gene expression levels were determined by the 2–ΔΔCt method.
Western blot analysis
Western blot experiments were performed as described previously. 23 Briefly, cells were lysed in RIPA lysis buffer and protein concentrations were determined using BCA kit (Beyotime, Shanghai, China). Proteins were separated by SDS-PAGE and transferred to PVDF membranes (Millipore, Billerica, MA, USA). After blocking, membranes were incubated with primary Abs specific for IL-34 (AF5265, 1:500 dilution; R&D Systems, Minneapolis, MN, USA), VDR (D151709; Sangon Biotech, Shanghai, China) or β-actin (TA-09; ZSGB-Biotech, Beijing, China) at 4℃ overnight. Subsequently, specific binding was revealed using secondary Abs (HRP-conjugated goat anti-rabbit IgG or HRP-conjugated rabbit anti-mouse IgG or HRP-conjugated rabbit anti-sheep IgG). All secondary Abs were obtained from ZSGB-Biotech. Signals were visualized by chemiluminescence using Super Signal West Femto (Thermo Scientific, Waltham, MA, USA) and images were captured using a digital imaging system (Tanon, Shanghai, China).
Plasmid construction
Genomic DNA was extracted from HepG2 cell line using the Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA). The DNA fragment containing the sequence ranging from position –506 to +207 relative to the transcription start site of IL-34 (ACCESSION: NM_152456 and NM_001172771) was amplified via PCR using the forward primer (5′-CGGGGTACCCTCCAGTCACCACCACAT-3′) and the reverse primer (5′-GAAGATCTTCAGAGGCAGAGGAAAGC-3′). PCR reactions were performed with DNA Ex Taq ® Polymerase (TaKaRa Bio, Shiga, Japan) using the following cycle conditions: one cycle 94℃ for 2 min; 30 cycles 94℃ for 30 s, 55℃ for 60 s, 72℃ for 60 s; one cycle 72℃ for 5 min. PCR products were cloned into pMD19 T-Vector (TaKaRa Bio) and subsequently sub-cloned into the promoterless luciferase reporter plasmid pGL3-Basic (Promega). The construct containing IL-34 –506/+207 region was designated pGL3-713. Reporter plasmids containing shorter fragments of the IL-34 promoter were constructed using pGL3-713 as the template and the following forward primers: pGL3-475 containing sequence –274/+207: 5′-CGG GGTACCTTGTCTAGGCCGAGTGTT-3′; pGL3-326 containing sequence –119/+207: 5′-CGGGGTACCGTTGAAGACTCCCTCCTACT-3′; pGL3-210 containing sequence –3/+207: 5′-CGGGGTACCCGGGAAGCACGGGAAGC-3′. The original reverse primer (pGL3-713) was used throughout. All IL-34 DNA sequences in luciferase reporter plasmids had been confirmed by DNA sequencing and were the same as the registered IL-34 sequence in NCBI and Ensembl databases.
Luciferase assay
For identification of IL-34 core promoter, SH-SY5Y and GES-1 cells were plated in 24-well plates (1 × 105 cells/well) and transfected with 0.4 µg firefly luciferase reporter plasmids pGL3-713, pGL3-475, pGL3-326 or pGL3-210 using Lipofectamine™ 2000 (Life Technologies, Carlsbad, CA, USA) according to the manufacturer's instructions. Forty-eight hours after transfection, cells were harvested in 100 µl of Passive Lysis Buffer (Promega) and luciferase assay was performed using the Dual Luciferase Assay System (Promega) according to the manufacturer’s direction. As an internal control, 10 ng pRL-TK plasmid (Promega), which expresses renilla luciferase constitutively, were co-transfected in each sample and firefly luciferase activities were normalized to renilla luciferase activities.
In stimulation experiments, SH-SY5Y cells were co-transfected with three plamsids containing 713-bp, 475-bp and 326-bp fragments (pGL3-713, pGL3-475 and pGL3-326, respectively) or pGL3-326 and siRNAs (VDR-siRNA or control siRNA) or pGL3-326 and its derivative plasmids without each VDR binding site. After transfection, SH-SY5Y cells were treated with 100 nM 1α,25(OH)2D3 for 48 h. Then cells were harvested for luciferase assay or Western blot analysis.
RNA interference
RNA interference against VDR was performed using known siRNA which had been verified in previous studies.24,25 The following VDR siRNA was used: 5′-GGAGUUCAUUCUGACAGAUTT-3′(sense) and 5′-AUCUGUCAGAAUGAACUCCTT-3′ (antisense). A nonspecific siRNA was used as the control. All of these siRNAs were synthesized by Shanghai GenePharma (Shanghai, China). SH-SY5Y cells were co-transfected pGL3-326 with VDR siRNA or control siRNA using Lipofectamine™ 2000 (Life Technologies) according to the manufacturer's instructions. After further stimulation with 100 nM 1α,25(OH)2D3 for 48 h, cells were harvested for Western blot and luciferase assay.
Site-directed mutagenesis
The DNA sequence containing IL-34 promoter region was analyzed by the online available software Patch (http://gene-regulation.com/cgi-bin/pub/programs/patch/bin/patch.cgi). Three possible VDR binding sites in –119/–3 region were predicted. pGL3-326 was used as the template to construct three pGL3-326 mutants which lack each predicted VDR bind site, according to the instructions of the Fast Mutagenesis Kit (Tiangen Biotech, Beijing, China). The following primers were used:
pGL3-326-M1 (CTCCCT deletion): Forward: 5′-CGATAGGTACCGTTGAAGACCTACTTCACAGTCCAAGG- -3′ Reverse: 5′- CCTTGGACTGTGAAGTAGGTCTTCAACGGTACCTATCG–3′ pGL3-326-M2 (TGCCCC deletion): Forward: 5′-GCGGGGGCCACGTGGCCCGCCCCTGCTGGCTTCG–3′; Reverse: 5′-CGAAGCCAGCAGGGGCGGGCCACGTGGCCCCCGC–3′ pGL3-326-M3 (CGCCCT deletion): Forward: 5′-GCAGTGGCCTGCTTTCCCTCTCATCAGACGGGAAGCCTG–3′ Reverse: 5′-CAGGCTTCCCGTCTGATGAGAGGGAAAGCAGGCCACTGC–3′
Statistical analysis
Data are presented as mean ± SD. The representative results from at least three experiments are shown. The results are compared by GraphPad Prism 5.0 using Student's t-test or Chi-square test between two groups. A value of P < 0.05 was considered significant.
Results
IL-34 is detectable in a variety of cell lines
We analyzed the expression status of the IL-34 gene in different cell lines, using Western blot. The results showed that IL-34 was expressed at low levels in the human liver cancer cell line HepG2 and at relatively high levels in the human gastric epithelial cell line GES-1, the human gastric cancer cell lines SGC-7901 and HGC-27 (Figure 1a). IL-34 was also expressed in human neuroblastoma cell line SH-SY5Y and human glioblastoma cell line A172 with a stronger expression in A172 than SH-SY5Y cells (Figure 1a).
IL-34 is detectable in a variety of cell lines and 1α,25(OH)2D3 induces IL-34 expression in SH-SY5Y cells. (a) The expression level of IL-34 in HepG2, SH-SY5Y, A172, GES-1, SGC-7901 and HGC-27 cell lines was determined by Western blot. β-Actin was used as a loading control. (b) SH-SY5Y cells were treated with 0 nM (ethanol control), 10 nM and 100 nM 1α,25(OH)2D3 for 48 h, then cells were harvested for qRT-PCR. (c) SH-SY5Y cells were stimulated with 100 nM 1α,25(OH)2D3 for 0 h, 24 h, 48 h and 72 h, then cells were harvested for qRT-PCR. (d) SH-SY5Y cells were treated with increasing concentrations of 1α,25(OH)2D3 [blank (ethanol), 10 nM, 50 nM, 100 nM, 250 nM, or 500 nM] for 48 h. IL-34 expression was analyzed by Western blot. β-Actin was used as a loading control. (e) SH-SY5Y cells were incubated with 100 nM 1α,25(OH)2D3 for the indicated time intervals. IL-34 expression was analyzed by Western blot. β-Actin was used as a loading control. For qRT-PCR results, mean values (±SD) of a representative experiment performed in triplicate are shown. *P < 0.05; **P < 0.01.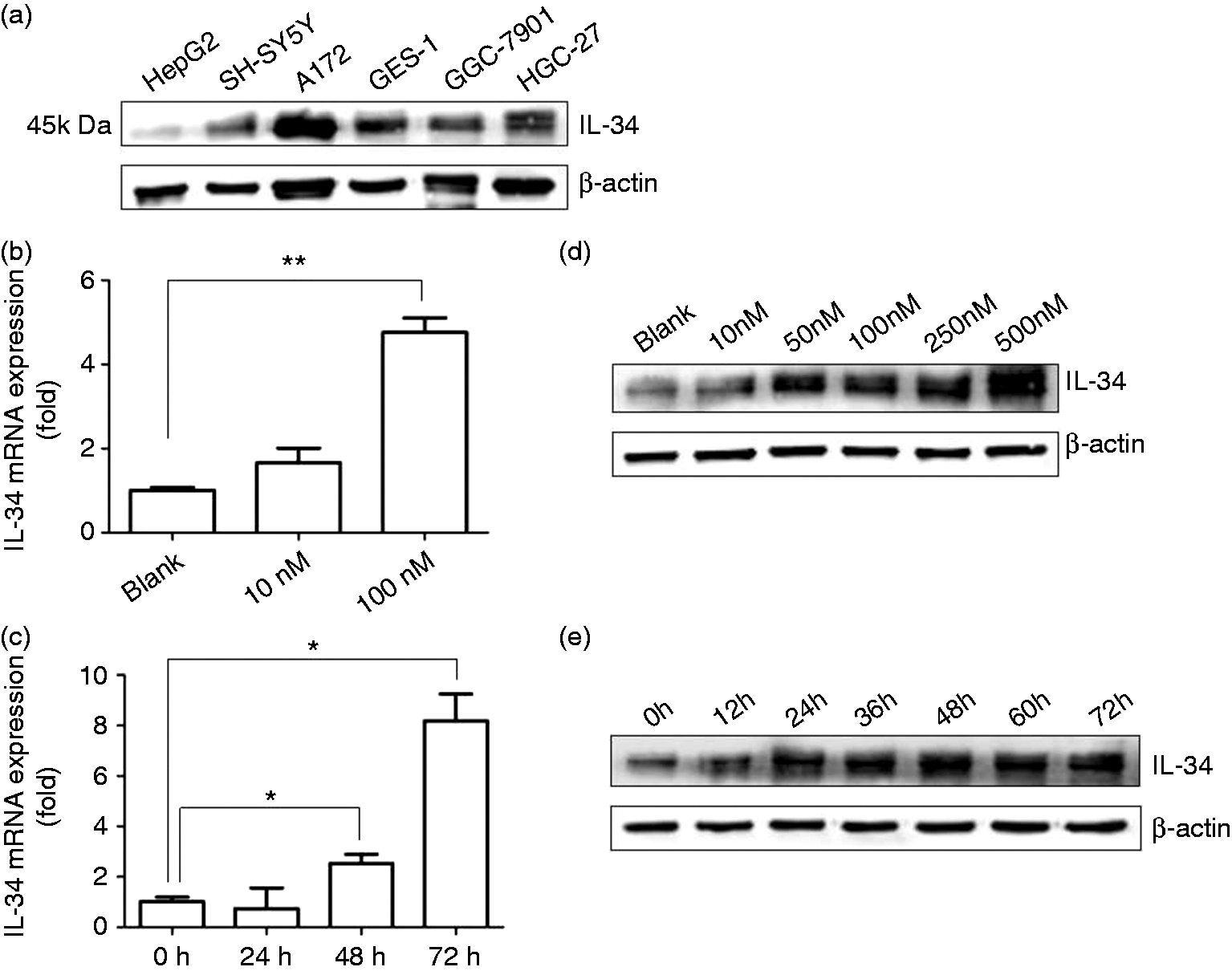
IL-34 expression is up-regulated by 1α,25(OH)2D3 in SH-SY5Y cells
Since neurons are main sources of IL-34 in brain,4,5 we examined whether 1α,25(OH)2D3, the active form of vitamin D, induced IL-34 expression in SH-SY5Y cells. IL-34 mRNA and protein levels were, indeed, up-regulated by 1α,25(OH)2D3 in a dose-dependent manner in SH-SY5Y cells (Figure 1b, d). Ten nM 1α,25(OH)2D3 induced IL-34 mRNA and protein expression partially but not significantly (Figure 1b, d). Supplementation with > 50 nM 1α,25(OH)2D3 induced IL-34 protein expression to a high level in SH-SY5Y cells (Figure 1d). Additionally, 1α,25(OH)2D3 up-regulated IL-34 mRNA and protein expression in SH-SY5Y cells in a time-dependent manner (Figure 1c, e). Treatment with 100 nM 1α,25(OH)2D3 for over 24 h induced IL-34 protein expression to a high level in SH-SY5Y cells (Figure 1c, e). In order to make the results more stable, we chose the dose of 100 nM 1α,25(OH)2D3 and the 48-h treatment time to do further experiments.
The core promoter of IL-34 gene is located within the region –274/–3 relative to the transcription start site
Next, we investigated the transcriptional regulation of IL-34 in SH-SY5Y cells. To identify the IL-34 core promoter, we cloned a 713-bp fragment of the IL-34 promoter region (–506 to +207 relative to the transcription start site) (ACCESSION: NM_152456 and NM_001172771) into pGL3-Basic luciferase reporter plasmid (pGL3-713). Additionally, we generated three truncated versions pGL3-475 (containing sequence –274/+207), pGL3-326 (containing sequence –119/+207) and pGL3-210 (containing sequence –3/+207) (Figure 2a). All these plasmids were transiently transfected into SH-SY5Y cells. Subsequent luciferase assays revealed that the 475-bp fragment had the highest transcriptional activity. Compared with the 475-bp fragment, a drop in the luciferase activity was observed for the 326-bp fragment. The luciferase activity of 210-bp fragment was further reduced to a level that was comparable with that of pGL3-basic negative control (Figure 2b). Similar results were also obtained in the normal gastric epithelial cell line GES-1 (Figure 2c). These results indicate that the core promoter of the IL-34 gene was located within the region –274/–3 relative to the transcription start site.
Luciferase activities of IL-34 promoter constructs in SH-SY5Y and GES-1 cell lines. (a) Four DNA fragments containing the indicated regions of the IL-34 gene were amplified and cloned into pGL3-basic luciferase reporter plasmids. Numbering is relative to the transcription start site (TSS). (b) SH-SY5Y and (c) GES-1 cells were transfected with the IL-34 promoter constructs. After 48 h, cells were harvested and IL-34 promoter activities were determined using luciferase assay. Firefly luciferase activity was normalized using renilla luciferase activity. Mean values (±SD) of a representative experiment performed in triplicate are shown. *P < 0.05; **P < 0.01.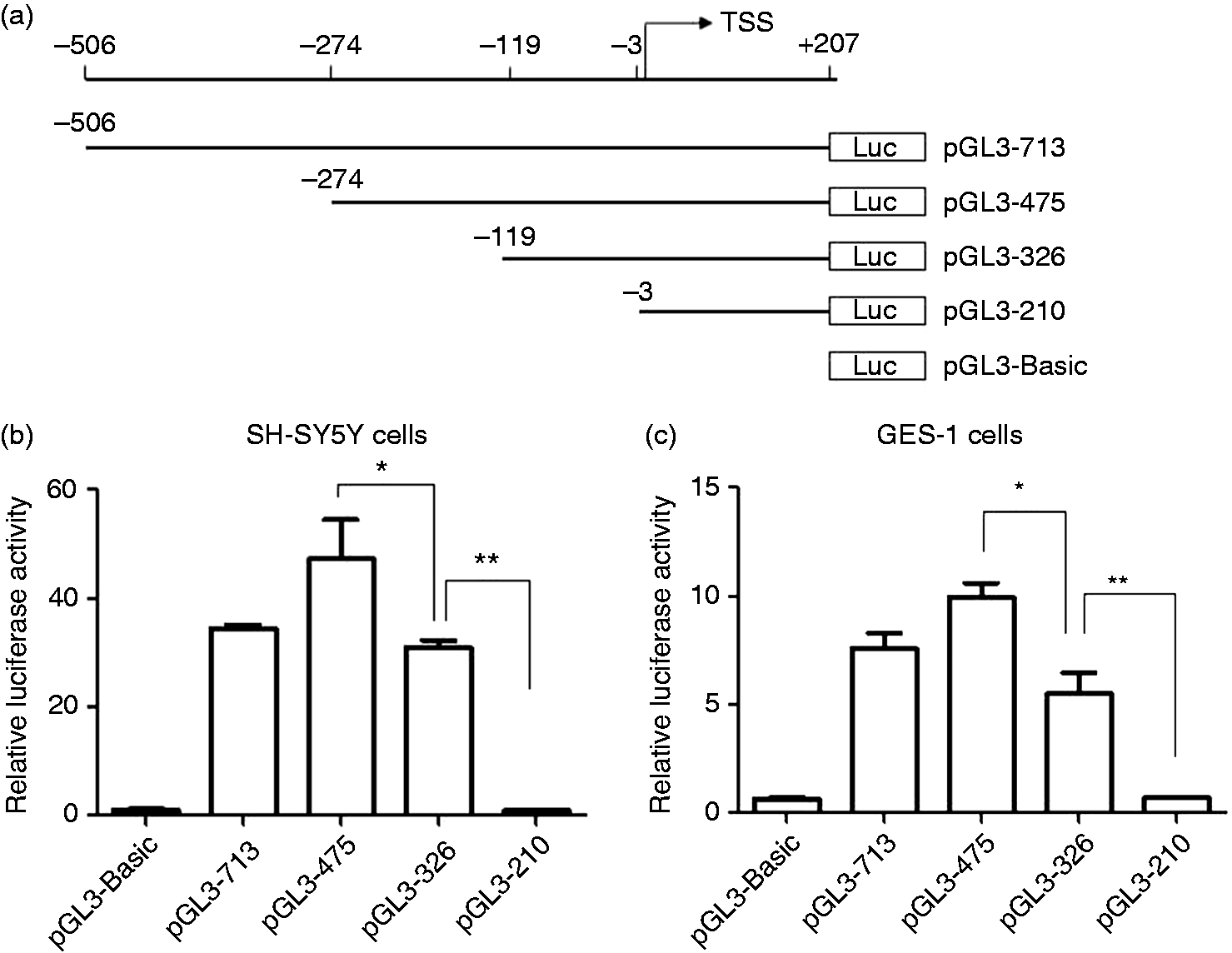
1α,25(OH)2D3 induces IL-34 expression through VDR in SH-SY5Y cells
In order to detail how 1α,25(OH)2D3 interacts with the IL-34 core promoter and induces IL-34 expression in neuronal cells, we transfected pGL3-713, pGL3-475 and pGL3-326 into SH-SY5Y cells and treated transfected cells with 1α,25(OH)2D3. It was found that 1α,25(OH)2D3 increased luciferase activities of all these IL-34 constructs in SH-SY5Y cells, suggesting that active VDR binding sites were present in all fragments and VDR binding sites in pGL3-326 was enough for this induction (Figure 3a). To test the specificity of the interaction we employed Western blot analysis of SH-SY5Y cells which were co-transfected with pGL3-326 and VDR siRNA and further stimulated with 1α,25(OH)2D3 for 48 h. It was found that 1α,25(OH)2D3 induced IL-34 expression under treatment with control siRNA (or no siRNA) and VDR siRNA transfection specifically down-regulated VDR expression in SH-SY5Y cells (Figure 3b). Consistent with a previous report,
26
1α,25(OH)2D3 slightly induce VDR expression in SH-SY5Y cells, partially attenuating the silencing effect of VDR siRNA (Figure 3b). However, 1α,25(OH)2D3 induced more IL-34 expression under treatment with control siRNA and no siRNA than under treatment with VDR siRNA (Figure 3b). Luciferase assay also showed VDR siRNA blocked the increase of pGL-326 luciferase activities induced by 1α,25(OH)2D3 in SH-SY5Y cells (Figure 3c). Together, these results suggested that IL-34 expression was induced by 1α,25(OH)2D3 through VDR in SH-SY5Y cells.
IL-34 expression is up-regulated by 1α,25(OH)2D3 through VDR in SH-SY5Y cells. (a) SH-SY5Y cells were transfected with pGL3-713, pGL3-475 and pGL3-326, respectively, and treated with 100 nM 1α,25(OH)2D3 for 48 h. IL-34 promoter activities were analyzed in luciferase assay. Firefly luciferase activity was normalized using renilla luciferase activity. Mean values (±SD) of a representative experiment performed in triplicate are shown. (b, c) SH-SY5Y cells were co-transfected with pGL3-326 and VDR-siRNA or control siRNA. After further stimulation with 100 nM 1α,25(OH)2D3 for 48 h, VDR and IL-34 expression were analyzed in (b) Western blot and luciferase activities of pGL3-326 were determined in (c) luciferase assay. Firefly luciferase activity was normalized using renilla luciferase activity. The value of cells transfected with control siRNA without 1α,25(OH)2D3 stimulation is set as 1. Mean values (±SD) of a representative experiment performed in triplicate are shown. *P < 0.05; **P < 0.01.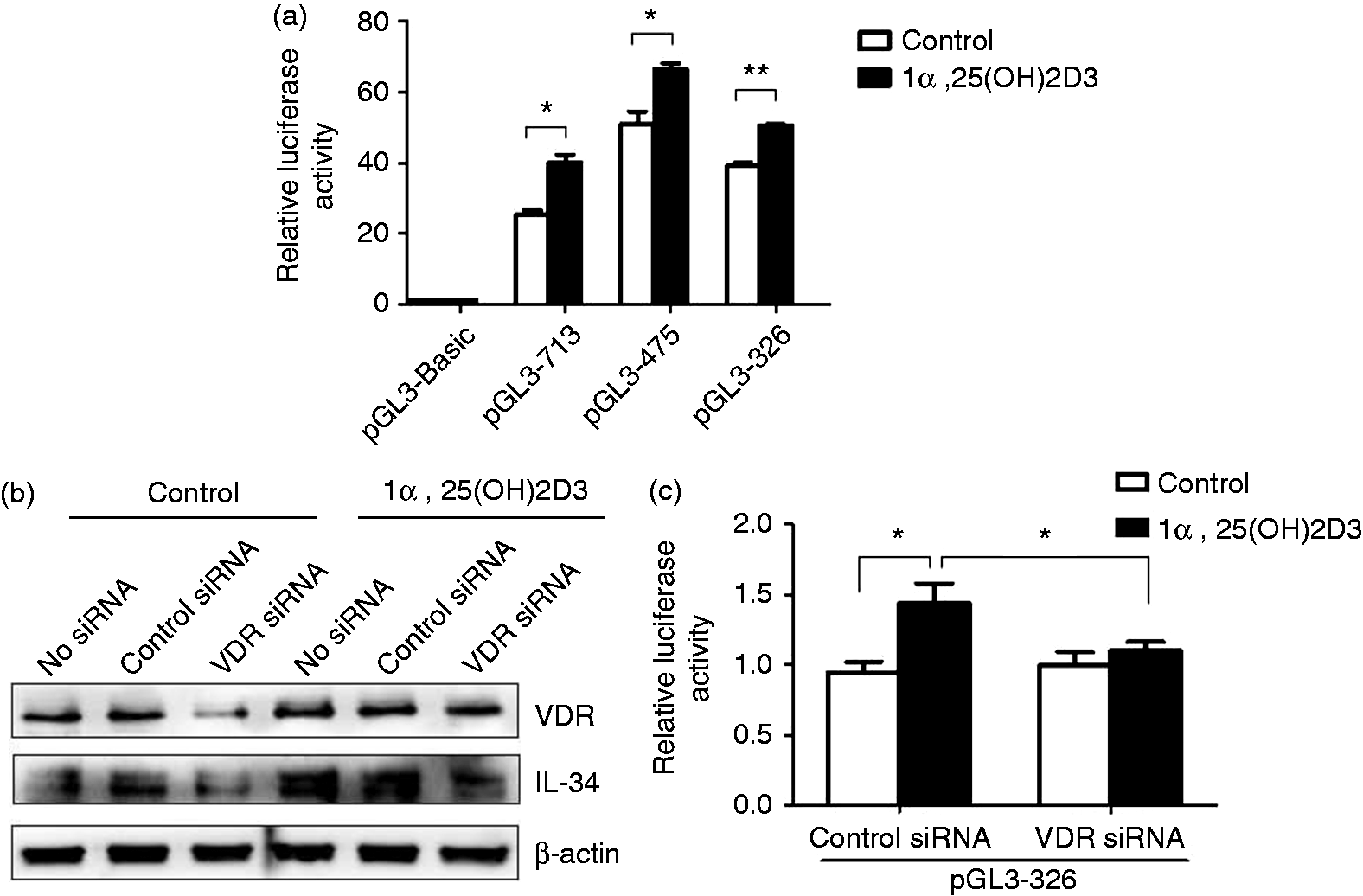
Identification of the VDR binding site that is responsible for the induction of IL-34 expression by 1α,25(OH)2D3
It was predicted that there were four human VDR binding sites in the 326-bp core promoter region (–119/+207 region) of the IL-34 gene. Three were located in the –119/–3 region and one in the first exon of the IL-34 gene. To assess the individual role of these VDR binding sites three predicted VDR binding sites in the –119/–3 region were deleted using site-directed mutagenesis (pGL3-326-M1: CTCCCT-defective; pGL3-326-M2: TGCCCC-defective; pGL3-326-M3: CGCCCT-defective) (Figure 4a). The deletion mutants were transfected into SH-SY5Y cells, and the promoter activity in response to 1α,25(OH)2D3 stimulation was determined using luciferase assay. All these deletions decreased the luciferase activities of pGL3-326 slightly but not significantly (Figure 4b). The M1 (CTCCCT) deletion led to the loss of 1α,25(OH)2D3-induced luciferase activity while M2 (TGCCCC) and M3 (TGCCCC) deletions had no effects on the increase of luciferase activities induced by 1α,25(OH)2D3 (Figure 4b). These results suggest strongly that M1 (CTCCCT) is the VDR binding site required for the induction of IL-34 expression by 1α,25(OH)2D3 in SH-SY5Y cells.
Identification of the VDR binding site, which is necessary for the induction of IL-34 expression by 1α,25(OH)2D3 in SH-SY5Y cells. (a) Three pGL3-326 deletion mutants were constructed (each lacked one predicted VDR binding sites in –119/–3 region). Numbering is relative to the transcription start site (TSS). (b) SH-SY5Y cells were transfected with pGL3-326 or pGL-326 deletion mutants and further treated with 100 nM 1α,25(OH)2D3 for 48 h. Then cells were harvested for luciferase assay. Firefly luciferase activity was normalized using renilla luciferase activity. The value of cells transfected with pGL3-326 without 1α,25(OH)2D3 stimulation is set as 1. Mean values (±SD) of a representative experiment performed in triplicate are shown. *P < 0.05; **P < 0.01.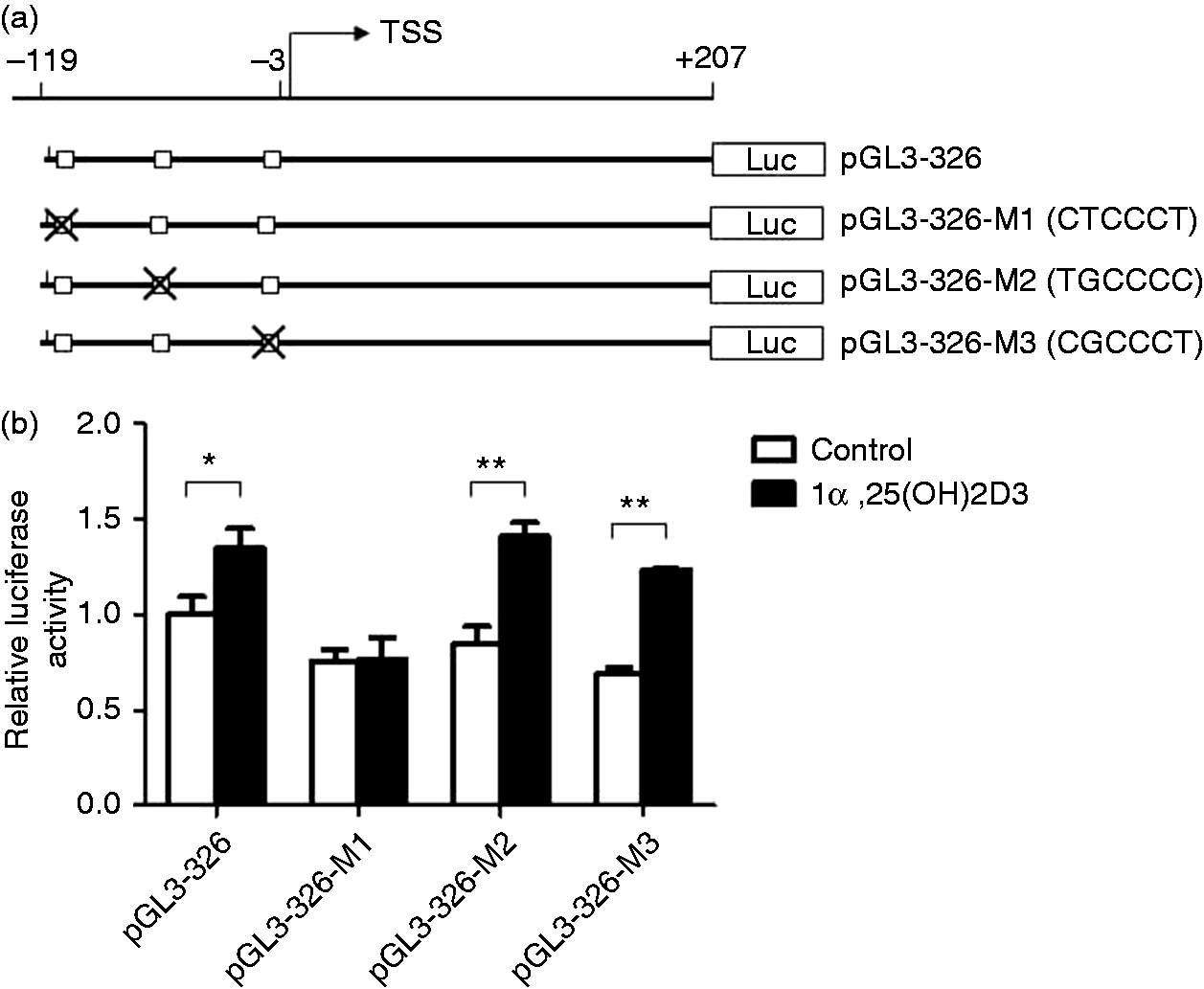
Discussion
Vitamin D plays a beneficial role in Alzheimer’s disease and its supplementation has been suggested to treat patients with Alzheimer’s disease. However, the underlying mechanism remains elusive.17,18,27,28 The cytokine IL-34 initiates strong neuroprotective and survival signals in brain injury and neurodegeneration. 29 This study now discovers that 1α,25(OH)2D3 , the active form of vitamin D, induces IL-34 expression in cultured human neural cells. Results from this study suggest that IL-34 is a vitamin D target gene and its up-regulation by 1α,25(OH)2D3 provides a mechanistic explanation of the beneficial effects of vitamin D for patients with Alzheimer’s disease.
The IL-34 promoter region was predicted to be the –224/+26 region relative to the transcription start site by using online software PromoterScan (https://www-bimas.cit.nih.gov/molbio/proscan/). Our luciferase assay results showed that the region –274/–3 relative to the transcription start site was IL-34 core promoter region, which was similar to the predicted promoter region. The identified promoter region of IL-34 contained no TATA box and CCAAT box, indicating that the IL-34 core promoter lacked a classical sequence structure. By using online software GPMiner (http://gpminer.mbc.nctu.edu.tw/), it is predicted that there are four GC boxes (–10/–5, –55/–60, –78/–84, –127/–133) within a 475-bp fragment. The GC box (–127/–133) may be potentially responsible for the decline of luciferase activities of the 326-bp fragment vs. the 475-bp fragment (Figure 2b, c). CTCCCT was a predicted VDR and RXR binding site in the upstream region of the IL-34 gene. Our results suggested that 1α,25(OH)2D3 induced IL-34 expression through the binding of VDR to this site in SH-SY5Y cells (Figures 3 and 4).
IL-34 may be produced from many tissues and cells.1,4,5,9 However, the regulation of its expression is largely unknown. This study firstly identified the IL-34 core promoter and provided the data that 1α,25(OH)2D3 induced IL-34 expression in SH-SY5Y neural cells. We also treated GES-1 cells with 1α,25(OH)2D3 for different times, and in different doses. It was found that 1α,25(OH)2D3 did not increase IL-34 mRNA and protein levels in GES-1 cells (data not shown). Other cells are needed to determine whether IL-34 expression induced by 1α, 25(OH)2D3 is specific for neuronal cells.
In the NCBI database, there are three human IL-34 mRNA variants. Variants 1 (NM_152456.2) and 2 (NM_001172771.1) use one promoter, whereas variant 3 (NM_001172772.1) uses an alternative promoter. In the Ensembl database, there are six transcripts (splice variants). The RefSeqs of IL-34-201 and 202 are the same as those in NCBI database. IL-34-203 and 204 are derived from an Ensembl automatic analysis pipeline and have no Refseq yet. IL-34-205 and 206 do not encode proteins. We focus on the identification of IL-34-202 promoter because IL-34-202 is the firstly identified and major mRNA variant. 1 However, it is possible that the alternative promoter also initiates the transcription of other IL-34 mRNA variants. Further investigation is needed to identify the alternative promoter of the IL-34 gene.
In summary, we found that 1α,25(OH)2D3 induced IL-34 expression in SH-SY5Y neural cells. Further studies are required to explore whether 1α,25(OH)2D3 induces IL-34 in primary human neurons in vitro and in vivo, and if it indeed mediates a protective function in Alzheimer’s disease.
Footnotes
Declaration of Conflicting Interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Natural Science Foundation of China (No. 81372576), Anhui Provincial Natural Science Foundation (No.1508085MH158), Postdoctoral Science Foundation of Anhui Province (No. 2015B066) and National Undergraduate Training Programs for Innovation and Entrepreneurship-China (No. 201610366006).
