Abstract
In Yersinia pseudotuberculosis complex, the O-antigen of LPS is used for the serological characterization of strains, and 21 serotypes have been identified to date. The O-antigen biosynthesis gene cluster and corresponding O-antigen structure have been described for 18, leaving O:8, O:13 and O:14 unresolved. In this study, two O:8 isolates were examined. The O-antigen gene cluster sequence of strain 151 was near identical to serotype O:4a, though a frame-shift mutation was found in ddhD, while No. 6 was different to 151 and carried the O:1b gene cluster. Structural analysis revealed that No. 6 produced a deeply truncated LPS, suggesting a mutation within the waaF gene. Both ddhD and waaF were cloned and expressed in 151 and No. 6 strains, respectively, and it appeared that expression of ddhD gene in strain 151 restored the O-antigen on LPS, while waaF in No. 6 resulted in an LPS truncated less severely but still without the O-antigen, suggesting that other mutations occurred in this strain. Thus, both O:8 isolates were found to be spontaneous O-antigen-negative mutants derived from other validated serotypes, and we propose to remove this serotype from the O-serotyping scheme, as the O:8 serological specificity is not based on the O-antigen.
Introduction
LPS is a lipoglycan present on the cell surface of most Gram-negative bacteria. For many pathogens, including Yersinia pseudotuberculosis, the LPS is an important virulence determinant.1–3 The structure is comprised of three key segments: lipid A, which anchors the structure in the outer membrane; the core oligosaccharide, consisting of inner core and outer core sections; and, usually, O-specific polysaccharide (OPS; also known as the O-antigen). The OPS is composed of variable-length polymers of repeating oligosaccharide units (O units) that often vary in structure within a species. The different forms are classically differentiated using serology, and the O-serotyping scheme for Y. pseudotuberculosis currently defines 21 different O-serotypes. 4 Recently, it was proposed that this serotyping scheme can be applied to all members of the Y. pseudotuberculosis complex, which also includes Y. pestis, Y. similis and Y. wautersii.5,6
The majority of genes that are required for the synthesis, assembly and processing of O-unit structures in the Y. pseudotuberculosis complex are in a gene cluster flanked by conserved hemH and gsk genes. 7 The OPS gene clusters and corresponding O-unit structures of 18 serotypes have been determined. 8 Many Y. pseudotuberculosis OPS gene clusters contain a ddhDABC gene set encoding for a precursor common to most of the rare immunodominant 3,6-dideoxyhexose (3,6-DDH) sugars that are always attached as a side-branch of the O-unit structure. 9 Seven different 3,6-DDH sugars, five hexoses and two octoses branched at C-4 (yersinose A and B), have been found in the Y. pseudotuberculosis complex. 9 These sugars include paratose, tyvelose (Tyvp) and abequose, which are also found in Salmonella enterica. 10 The type of 3,6-DDH sugar produced is determined by one or two ddh-specific genes located immediately downstream of ddhDABC in the OPS gene cluster, being prt, prt and tyv, and abe, respectively, for paratose, Tyvp and abequose.
OPS gene clusters include wzx, wzy and wzz genes that encode proteins involved in the Wzx/Wzy-dependent processing pathway.
7
In this pathway, the OPS is made by polymerization of an oligosaccharide O unit, and in Y. pseudotuberculosis, O-unit synthesis is initiated by the transfer of N-acetyl- Representation of the main form of the carbohydrate backbone of the lipopolysaccharide from Y. pseudotuberculosis. Variations in the core region occur at different level: the external Kdo can be replaced by Ko, while Hep III can have either a 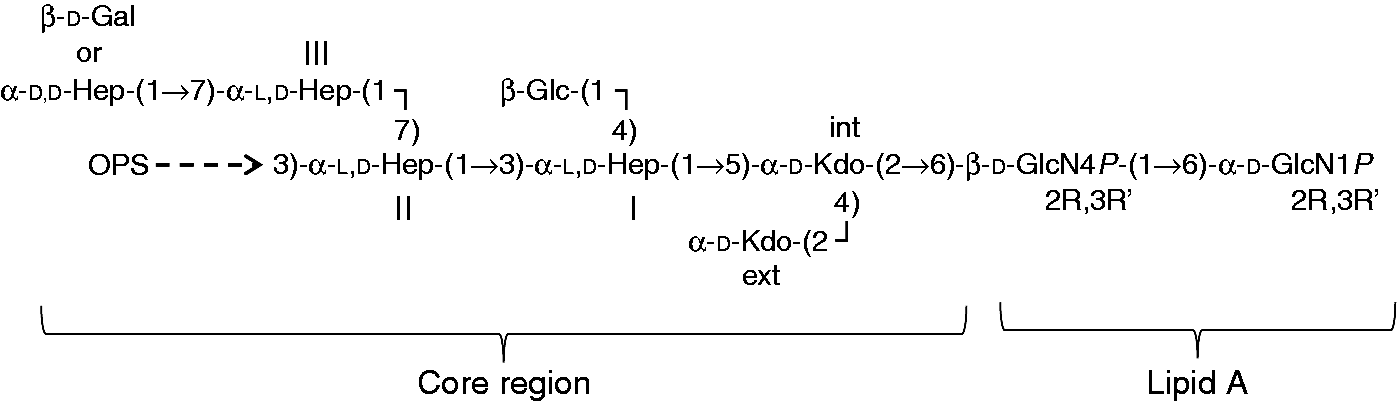
The core region contains a conserved hexasaccharide, constituted by two Kdo residues (named inner and external), three
The serological typing system for Y. pseudotuberculosis was developed in stages, and there have also been changes to the nomenclature. Six serogroups (I–VI) were established prior to 1984, as described by Tsubokura et al. 18 Two new ones, VII and VIII, were then proposed in 1984, 18 and the use of Roman numerals was replaced with an Arabic numbering system (O:1–O:8), with some groups having subtypes, for example O:1a and O:1b. 19 Serogroups O:9–O:15 were described in updates to the typing scheme.4,20 However, we now treat all as serotypes, as there is no further subdivision into serovars in Y. pseudotuberculosis.
The history of serotype O:8 is unusual. The first report of this serotype stated that the type strain (isolate 151) expressed only one epitope, O factor 20, that was unique to O:8, 18 but the 1993 revision of the scheme reported that the O:8 type strain has a rough phenotype, 20 that is, did not produce OPS, and this was not further investigated. In the following update, O:8 isolates were reported to react with O factor 11, 4 which had previously been described for serotypes O:2c and O:4a. 19 We now know that serotypes O:2c and O:4a have an O antigen with the same main chain, 9 which implies that epitope 11 is in the shared portion of the O-antigen structure. However, cross-reaction of epitope 11 with O:8 conflicts with the previously reported ‘rough’ phenotype. In this study, we examine the OPS gene clusters and LPS products by two O:8 isolates.
Materials and methods
Bacterial strains and growth
Bacterial strains and plasmids used in this study.
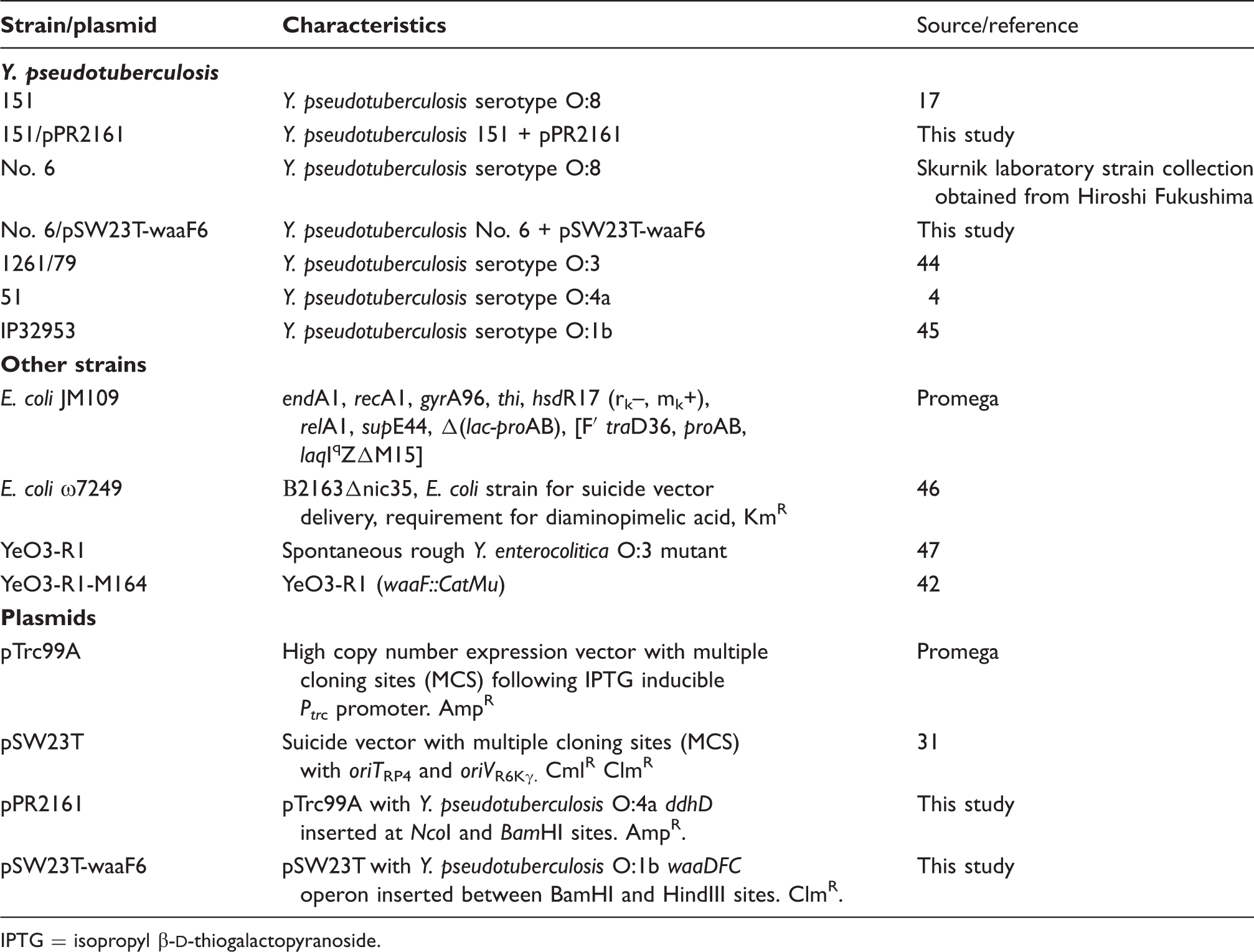
IPTG = isopropyl β-
Isolation of LPS
The bacteria (52.8 g and 11 g, strains No. 151 and 6, respectively) were washed successively with ethanol, acetone and ether. De-lipidated bacterial masses (9.96 g and 3.85 g, strains No. 151 and 6, respectively) were extracted with hot phenol/water (2.49 g and 430 mg crude LPS, 25% and 11.2% of bacterial dry mass, strains No. 151 and 6, respectively). 22 The obtained water phases were dialysed, then lyophilized and purified by enzymatic treatment, 23 then dialyzed and lyophilized. The samples were ultracentrifuged (150,000 g, 4℃, 4 h) three times, then lyophilized (yield of pure LPS: 4.3% and 6% for 151 and 6, respectively).
General and analytical methods
Chemical composition analyses of LPS included P, Kdo and GlcN determination, as well as methanolysis (0.5
Isolation of oligosaccharides 1, 2 and 3
LPS was dissolved in anhydrous hydrazine (25 mg/ml), stirred at 37℃ for 30 min, cooled, poured into ice-cold acetone (20 ml) and allowed to precipitate. The precipitate was then centrifuged (3000 g, 30 min, 4℃), washed twice with ice-cold acetone, dried, and then dissolved in water and lyophilized. The sample was N-deacylated with 4 M KOH as described,
27
and desalted by gel-permeation chromatography [Sephadex G-10 (GE Healthcare Europe GmbH, Freiburg, Germany) 50 × 1.5 cm, in 10 m
NMR spectroscopy
One-dimensional spectra were recorded with a Bruker DRX 600 spectrometer equipped with a z-gradients reverse probe, and chemical shifts were referred relative to internal acetone (δ1H 2.225, δ13C 31.5); two-dimensional (2D) NMR spectra of 1 and 2 were recorded in 0.5 ml 2H2O at 32℃; 2D NMR spectra of oligosaccharide 3 were acquired at 25℃.
For the homonuclear experiment, solvent-saturated double quantum-filtered correlation spectroscopy (DQF-COSY), total correlation spectroscopy (TOCSY), transverse rotating frame overhauser enhancement spectroscopy (TROESY) spectra and 512 free induction decays (FID)s of 2048 complex data points were collected, with 48 scans per FID and using standard manufacturer software. The spectral width was set to 10 ppm and the frequency carrier was placed at the residual HOD peak; mixing times of 100 and 300 ms were used for TOCSY and TROESY, respectively. For the heteronuclear single quantum coherence (HSQC) spectrum, 512 FIDS of 2048 complex points were acquired with 50 scans per FID; the GARP sequence was used for 13C decoupling during acquisition. As for heteronuclear multiple bond correlation (HMBC) and HSQC-TOCSY, the same resolution in the two dimensions was used, but scan number was doubled and tripled, respectively. Data processing and analysis was performed with the standard Bruker Topspin 3.0 program.
ESI-MS/IT-TOF spectrometry
Oligosaccharides 1, 2 and 3 were analyzed by ESI-MS analyses using an ESI-MS/IT-TOF instrument (Shimadzu Europe, Manchester, UK). Each sample was dissolved in aqueous solution at a concentration of 0.1 mg/ml, and 200-µl aliquots were injected directly in ESI at flow rate of 0.2 ml/min. The quadrupole was scanned in positive and negative mode over 400–2000 m/z range. An electrospray voltage of 1.90 kV and a nitrogen gas flow of 1.5 l/min were employed. Mass calibration and tuning were performed according to the manufacturer’s recommendations. The spectra were acquired using the LC MS Solution software; all mass values are reported as average masses, and deconvolution of mass spectra was performed using both Shimadzu deconvolution software and ESI prot software.
PCR, sequencing and bioinformatic analysis
Chromosomal DNA was extracted from representative isolates of O:1b and O:4a, and from isolates 151 and No. 6, using the ultra-pure method described previously. 28 The OPS gene cluster of No. 6 was typed by PCR using the genotyping scheme for Y. pseudotuberculosis. 29 The sequence of the 151 OPS gene cluster was obtained by a process of PCR walking from conserved hemH and gsk genes at either ends of the region, and also using primers designed from O:4a sequence. PCR products were sequenced using the AB3730xl sequencing platform carried out by the Australian Genome Research Facility (Sydney, Australia).
A consensus sequence was obtained for 151 by compiling sequence trace files, and bioinformatics analysis was performed as described previously. 28 Insertion sequences (IS) were identified and characterized using ISFinder (https://www-is.biotoul.fr). The final sequence for 151 was deposited into GenBank under accession number KM454907.
Cloning and complementation of ddhD
The ddhD gene was amplified from chromosomal DNA from strain 51 (O:4a) using high-fidelity PCR (described previously
19
) with primers that incorporated flanking NcoI (5’) and BamHI (3’) restriction sites into the product. The product and the pTrc99A expression vector (obtained from Promega, Madison, WI, USA)
30
were digested with NcoI and BamHI, and then ligated together according to the manufacturers instructions (New England Biolabs, Ipswich, MA, USA). The sample was cloned into Escherichia coli K-12 JM109, for selection of positive constructs. A positive transformant was selected, and the constructed plasmid purified and confirmed by PCR and sequencing prior to electroporation into strain 151. Electroporation was conducted using a BioRad gene pulser with the following conditions: 25 μFD, 25 kV, 200 Ohm. Expression of the ddhD gene in pTrc99A was induced by supplementing cultures with 1 mM isopropyl β-
Cloning of waaDFC operon and complementation of the waaF mutation in strain No. 6
The 3407-bp DNA fragment carrying the waaDFC operon was amplified by PCR using genomic DNA of Y. pseudotuberculosis strain IP32953 as the template, and primers waaF6-F (5’-GCGaagcttGCTGATAAAAGGGGTGCTTG-3’) and waaF6-R (5’- GCGggatcCGTTAGAGCTGTTGGGTGCT-3’), which contained the BamHI and HindIII restriction sites for cloning. The 1.8-kb suicide vector pSW23T, 31 and the waaDFC fragment were digested with restriction enzymes BamHI and HindIII. The linearized pSW23T DNA was dephosphorylated by a 20-min FastAP alkaline phosphatase (Fermentas, Waltham, MA, USA) treatment at 37℃. The DNA fragments were ligated for 1 h at room temperature (RT) and 16 h at 4 ℃. After ligation the DNA was ethanol-precipitated, solubilized into 10 µl water and electroporated into E. coli ω7249 cells. The transformants were plated on LA plates containing 0.3 mM diaminopimelic acid (DAP), 20 µg/ml chloramphenicol and 100 µg/ml kanamycin. The obtained colonies were checked for the presence of the insert PCR using insert primers waaF6-F and waaF6-R. One of the PCR-positive colonies was selected and the constructed plasmid was named as pSW23T-waaF6. The plasmid was mobilized from the E. coli strain ω7249/pSW23T-waaF6 into Y. pseudotuberculosis strain No. 6 by conjugation. After 16 h mating on a LA plate supplemented with DAP the mating mix was collected into PBS, washed three times with PBS and aliquots plated on selective Yersinia agar plates supplemented with chloramphenicol. Chloramphenicol-resistant Yersinia transconjugants were recovered and checked for waaF complementation by DOC-PAGE.
SDS- or DOC-PAGE analysis and Western blotting
The bacteria were grown for 16–20 h at RT in 5 ml LB medium. The exact OD600 of the cultures was measured and 1 ml of the cultures was centrifuged. For DOC-PAGE, the pellets were resuspended in deoxycholate (DOC) lysis buffer [2% DOC, 4% 2-mercaptoethanol, 10% glycerol and 0.002% bromophenol blue in 1 M Tris-HCl buffer (pH 6.8)] in a volume adjusted according to density of the culture (100 µl/OD600 = 1). For SDS-PAGE, the pellets treated as described previously. 13 LPS phenotypes of proteinase K-treated whole-cell lysates were analyzed by silver-stained SDS or DOC-PAGE,13,32,33 and by Western blotting using rabbit anti-O:1b antiserum. The samples were transferred after DOC-PAGE from the gel onto a PVDF membrane by semidry blotter (Panther Semidry Electroblotter; Owl Separation Systems, Thermo Scientific, Waltham, MA, USA) at 12 V, 22℃, for 2 h. After blocking 12–16 h at 4℃ in 5% skimmed milk/1 × Tris-buffered saline (TBS) Tween20 buffer (0.15 M NaCl, 10 mM Tris-HCl, 0.05% Tween20, pH 7.5), the membrane was incubated for 1 h at 22℃ in a rolling tube with 1:1000–1:2000 diluted rabbit antisera in 5% skimmed milk/1 × TBS Tween20 buffer. To detect the bound Abs, the membrane was incubated for 1 h at 22℃ in a 1:2000-diluted HRP-conjugated secondary swine anti-rabbit Abs (DAKO, Glostrup, Denmark) in 5% skimmed milk/1 × TBS Tween20 buffer. After washing, the secondary Abs were detected with the enhanced chemiluminescence solution (0.1 M Tris-HCl, 12.5 mM luminol, 2 mM coumaric acid, 0.03% H2O2) exposed to X-ray film (Kodak BioMax MR, Sigma-Aldrich, Helsinki, Finland).
Results and discussion
Purification and visualization of LPS from O:8 isolates
LPS was purified from Y. pseudotuberculosis O:8 isolates 151 and No. 6 using the hot phenol/water extraction procedure, and the sample was purified by enzymatic treatment, dialysis and ultracentrifugation, then lyophilized to remove contaminants (see ‘Materials and methods’).
Visualization of the LPS profiles revealed that the O:3 control has a single band of LPS (Figure 2). Consistent with our previous observation, the O:3 OPS chain could not be visualized as the periodate treatment in the staining process requires vicinal hydroxyl groups, which are rare in the O:3 structure.
19
However, the O:3 band is as expected for lipid A with core oligosaccharide and one O unit. Both O:8 isolates produced bands with faster migration and presumably a lower molecular mass than the O:3 control. The LPS band of 151 runs faster than that of O:3, which is consistent with the previous serological classification that reported the isolate as rough.
20
However, the LPS band produced by No. 6 runs faster than that of 151 (Figure 2), indicating that the isolates are not the same, and the LPS is presumably significantly shorter.
LPS profiles of Y. pseudotuberculosis serotype O:8 and O:3 isolates analyzed by DOC-PAGE.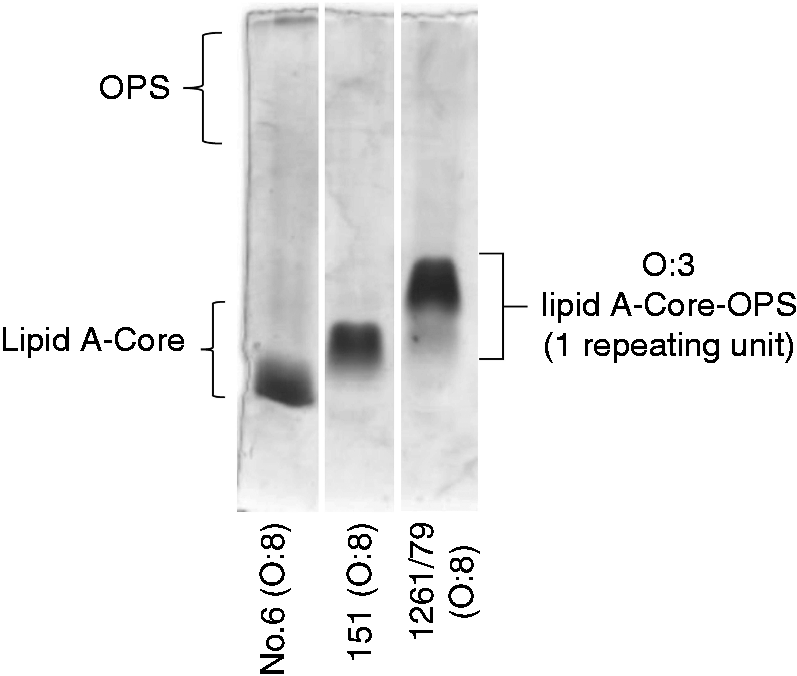
Compositional analyses of LPS molecules from O:8 isolates
To determine the differences between the LPS of the two O:8 isolates, chemical analysis was performed on purified LPS. The content of neutral sugars [Glc, Gal,
Genetic analysis of the two strains provided insight into the mutations responsible for the phenotype observed.
Isolate 151 appears to have arisen from serotype O:4a
The OPS gene cluster of isolate 151 was sequenced (GenBank accession number KM454907) and found to have a gene cluster closely related to that of serotype O:4a, (GenBank accession number KJ504355).
7
However, in comparison to O:4a, the remnant ISYps7 element in the 151 gene cluster is ∼270 bp shorter at the end adjacent to wzy (Figure 3; Table S1), and the ddhD gene of strain 151 includes a frame-shift mutation 297 bp downstream of the start codon, which produces a premature stop codon 16 bp further downstream (Figure 3).
Comparison of the OPS gene clusters of O:8 and O:4a. All genes are oriented in the forward direction, and gene names are shown above or below. Genes that encode processing enzymes are gray, and bold horizontal lines indicate genes that encode glycosyltransferases. IS remnants are shown in black, and the vertical arrow indicates the position of a short repeat. Nucleotide sequence percentage identity levels are shown. The segment deleted in the O:8 ddhD gene is shown below. Figure is drawn to scale from GenBank accession numbers KJ504355 and KM454907.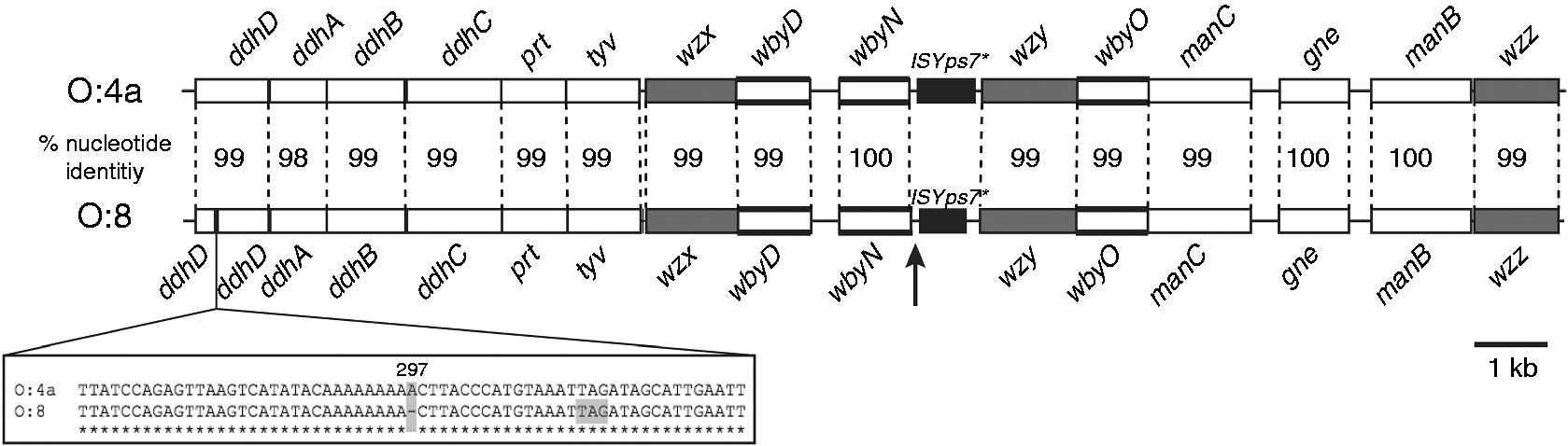
Serotype O:4a is known to produce smooth LPS that includes an O-unit main-chain structure containing
We used the O:4a ddhD gene to complement the mutation in 151, and observed that while production of long-chain O antigen was restored, the length was much less than that of O:4a LPS (Figure 4). Also, the lipid A core component of 151, with or without the cloned ddhD gene, ran faster than that of the O:4a wild type. The presence of sub-stoichiometric amounts of Complementation of ddhD in strain 151. Concentrated LPS was loaded into wells. Track 1 shows an SDS-PAGE gel of O:4a LPS, and tracks 2 and 3 LPS from O:8 strain 151 without and with complementation by a cloned ddhD gene.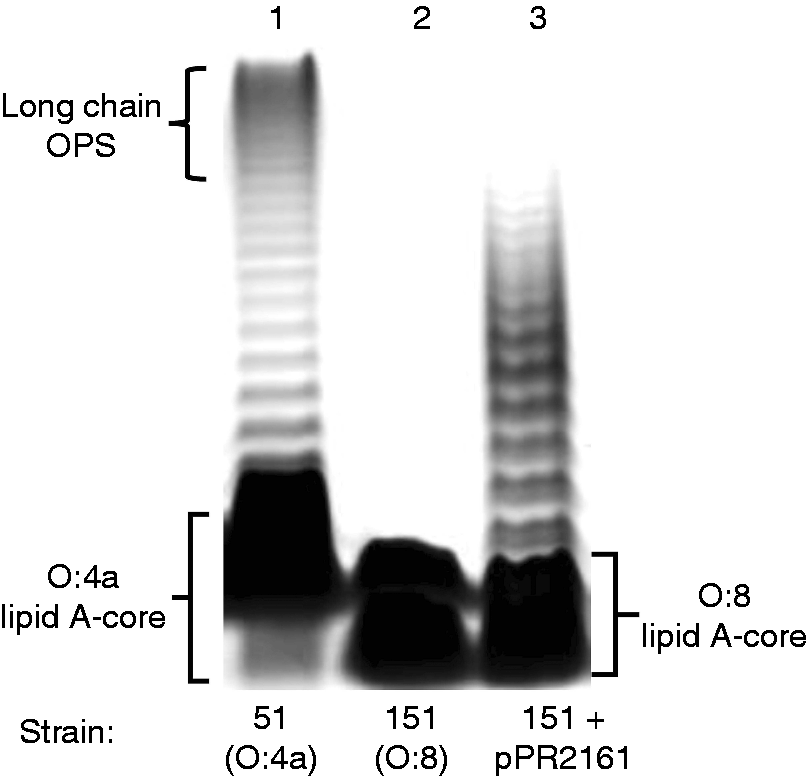
Isolate No. 6 appears to have arisen from serotype O:1b
PCR genotyping, using the scheme described previously, 29 revealed that isolate No. 6 carried the O:1b gene cluster (data not shown). However, even though No. 6 still harbours the epitope 20, its LPS is deeply truncated and smaller than that observed for 151 (Figure 2), indicating the isolate probably has a mutation in the gene cluster responsible for core oligosaccharide synthesis.
Thus, structural analysis was performed on the LPS recovered from No. 6 because it contained already the structural elements of epitope 20, and to determine the extent of the LPS truncation to provide insights into the possible type of genetic mutation.
NMR spectroscopy of oligosaccharide phosphates isolated from No. 6
Mild hydrazinolysis of the LPS mixture from isolate No. 6 (57 mg) led to the O-deacylated products (42.2 mg; 76%) that were further subjected to strong alkaline treatment in order to remove the amide-linked acyl residues. From this mixture (13.44 mg; 24% of the LPS), oligosaccharides
The complete assignment of 1H and
13
C resonances of each oligosaccharide was achieved (Tables 2–4) by combining the information obtained from DQF-COSY, TOCSY, TROESY, HSQC, HMBC and HSQC-TOCSY NMR experiments, and started on oligosaccharide 1H NMR spectra of 600-MHz 1H and 150-MHz 13C chemical shifts of 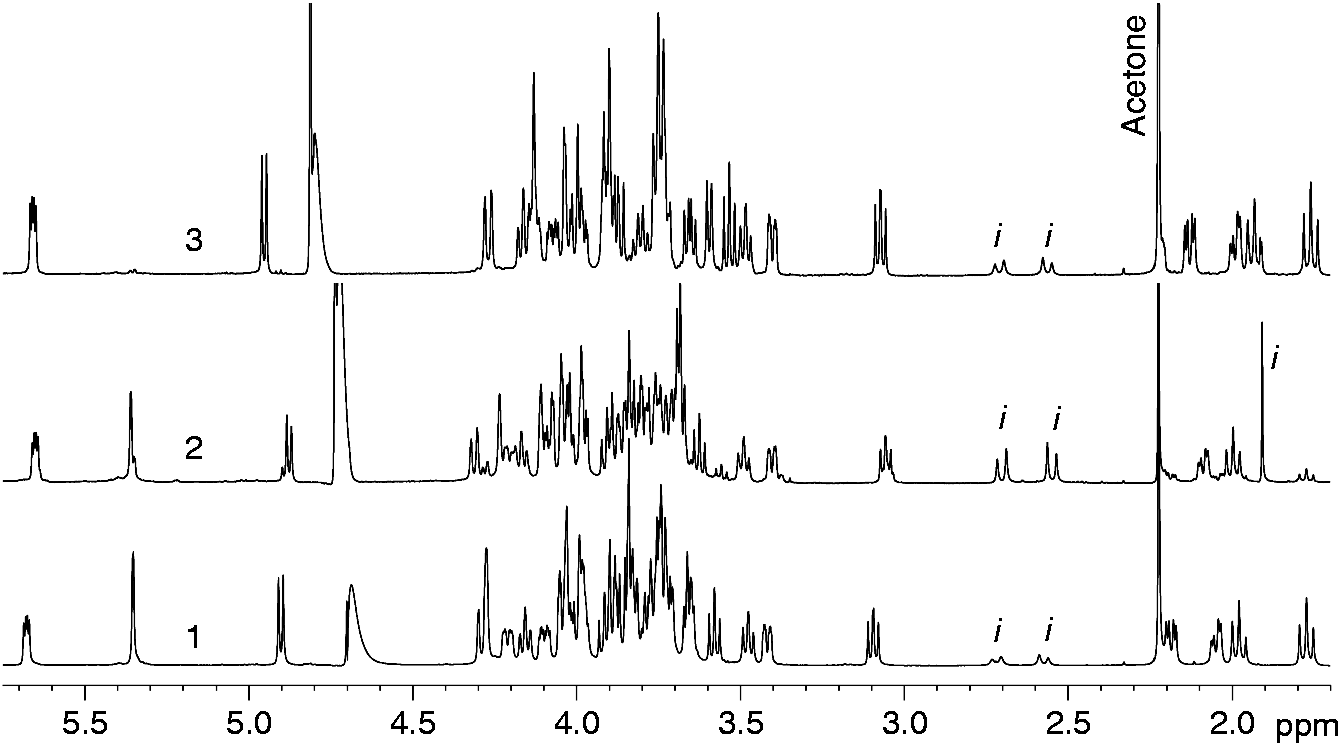
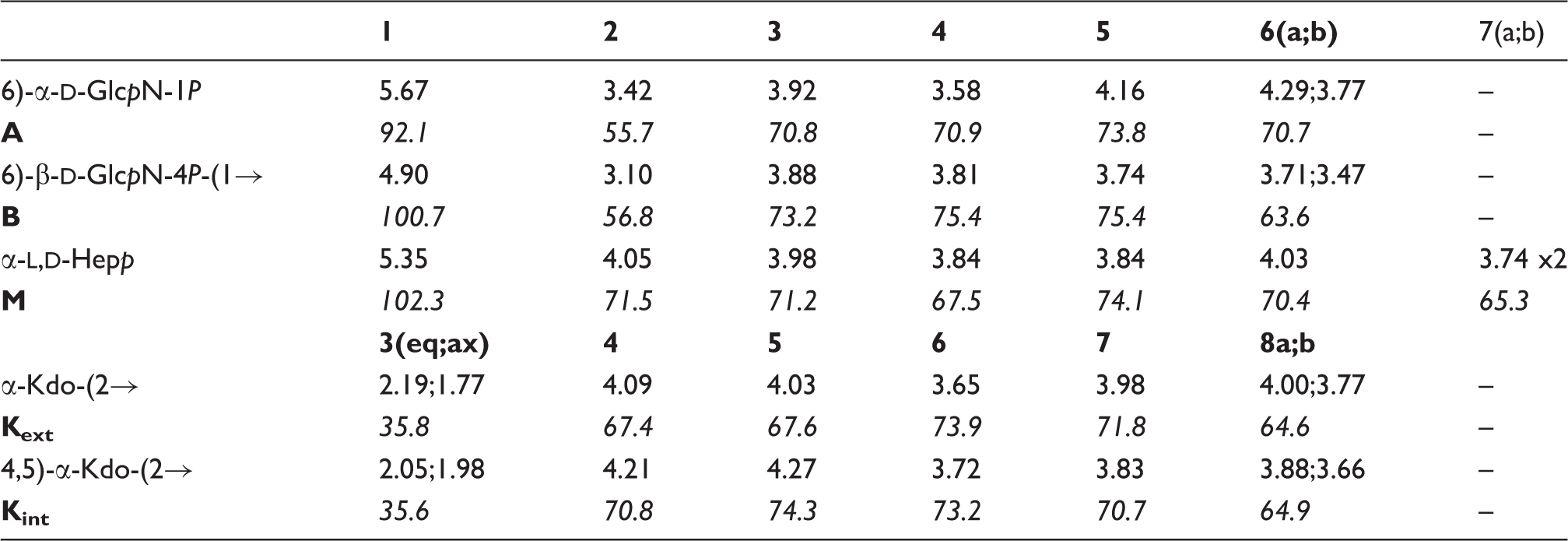
The 1H-NMR spectrum of Structures of 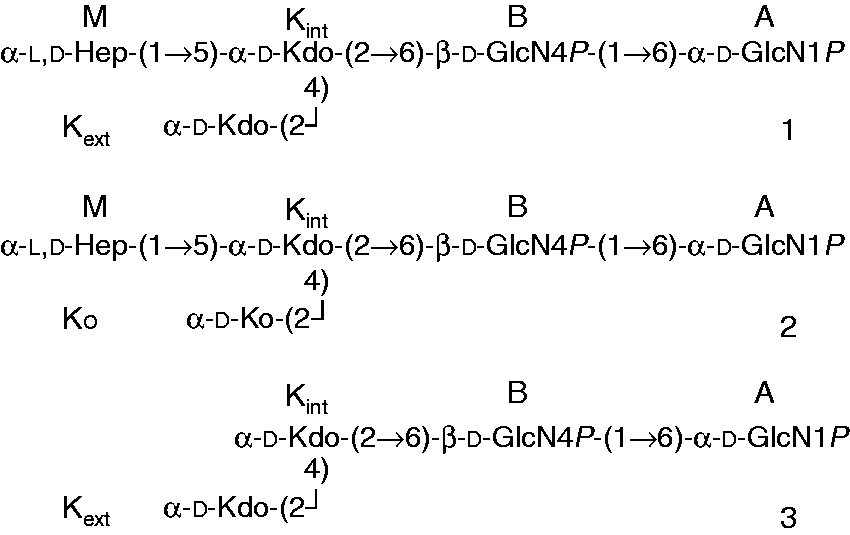
600-MHz 1H and 150-MHz
13
C chemical shifts of
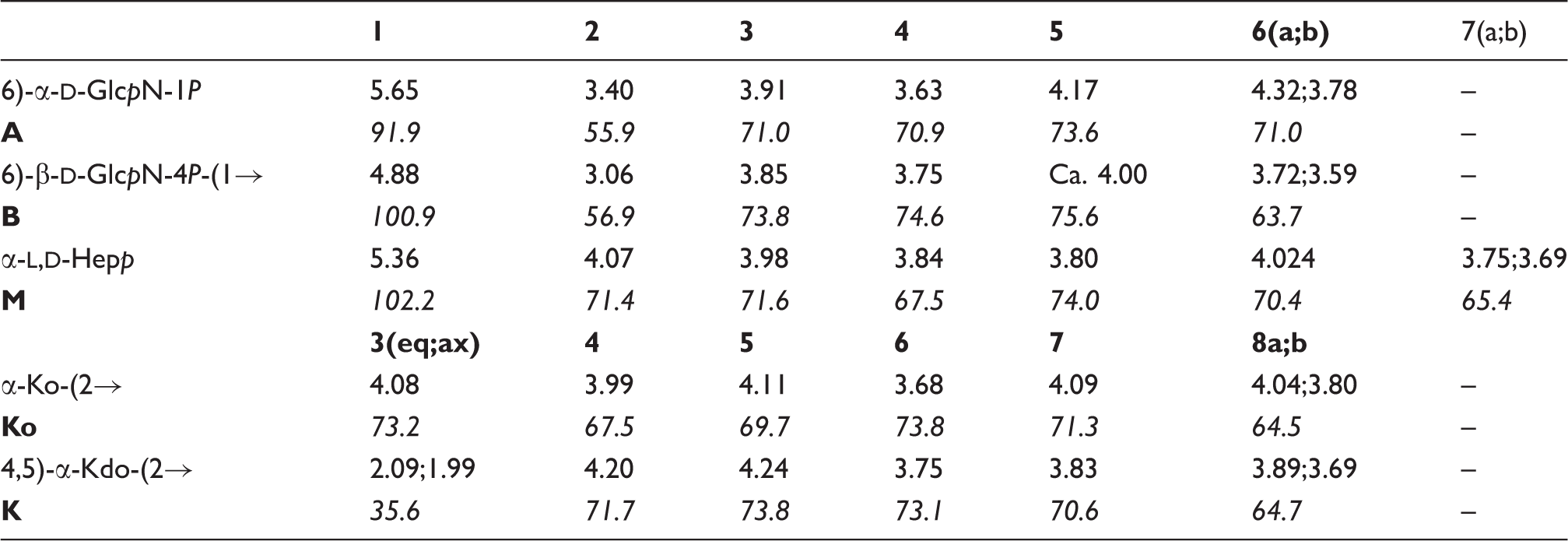
600-MHz 1H and 150-MHz 13C chemical shifts of
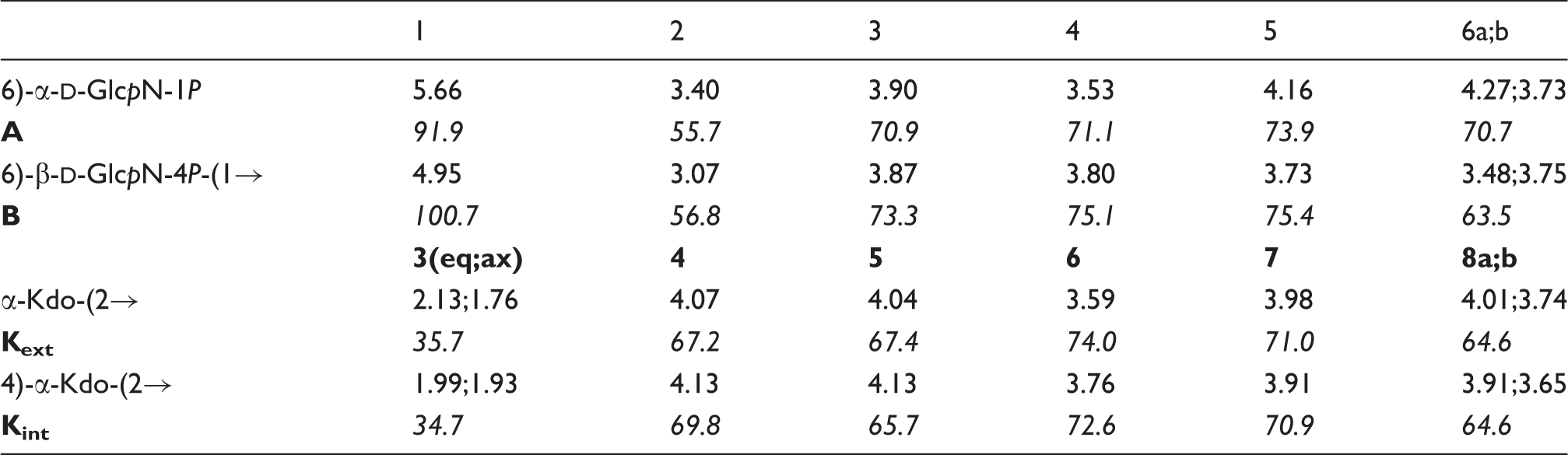
Consistent with this, H-3eq of Expansion of HSQC (black) and HSQC-TOCSY (grey) spectrum of 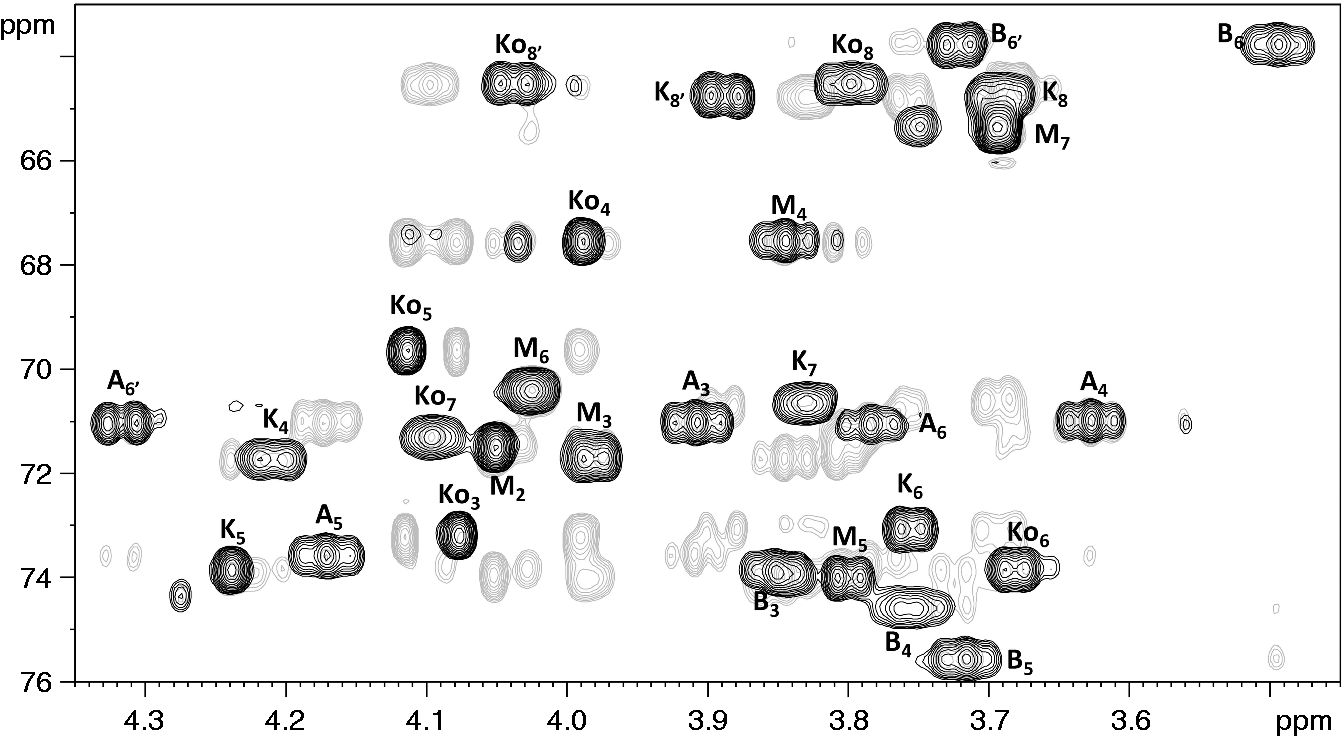
ESI-MS/IT-TOF analysis
ESI-MS spectra of ESI-MS spectra in negative mode of ESI-MS MS/MS spectra of the fully de-lipidated oligosaccharides 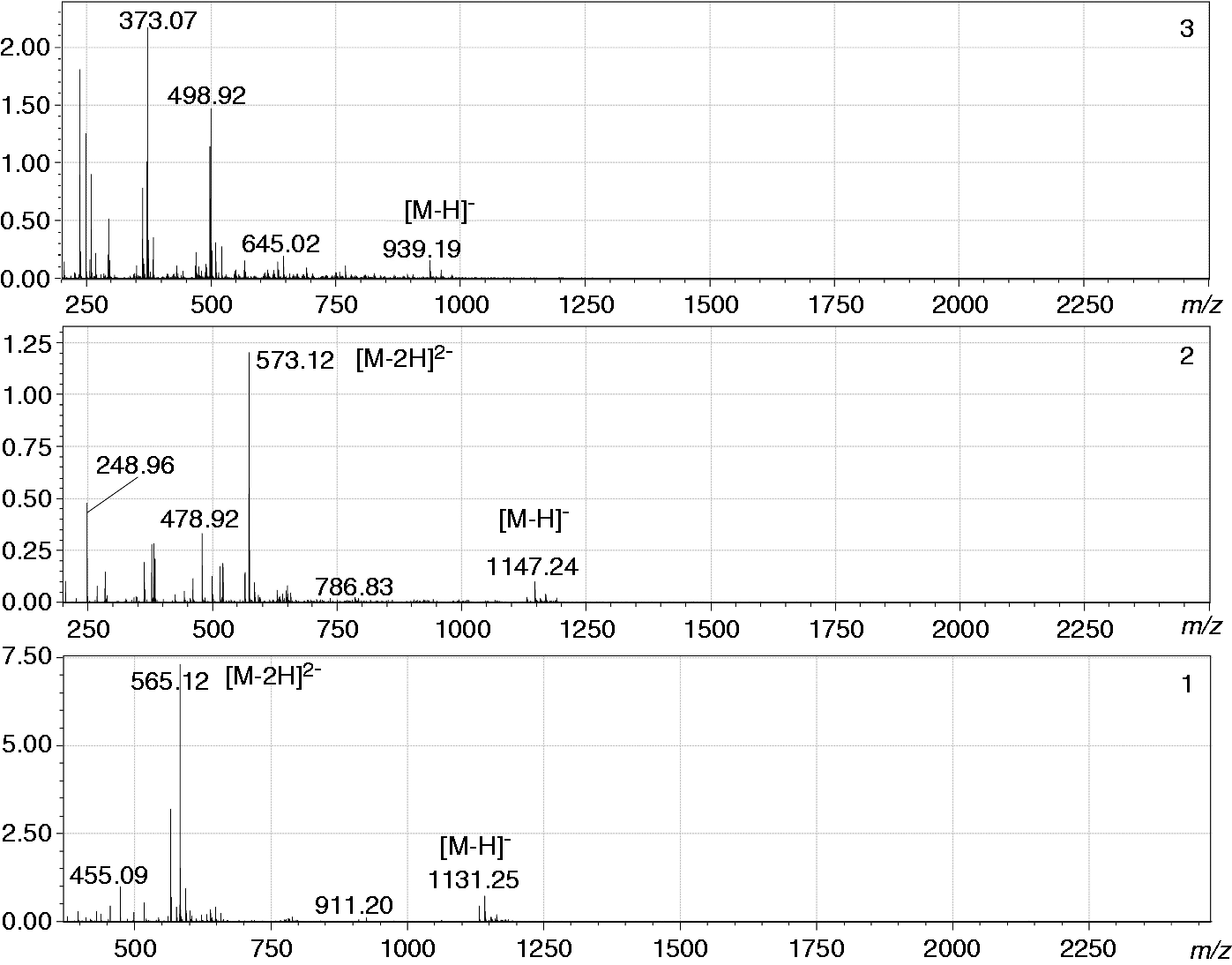
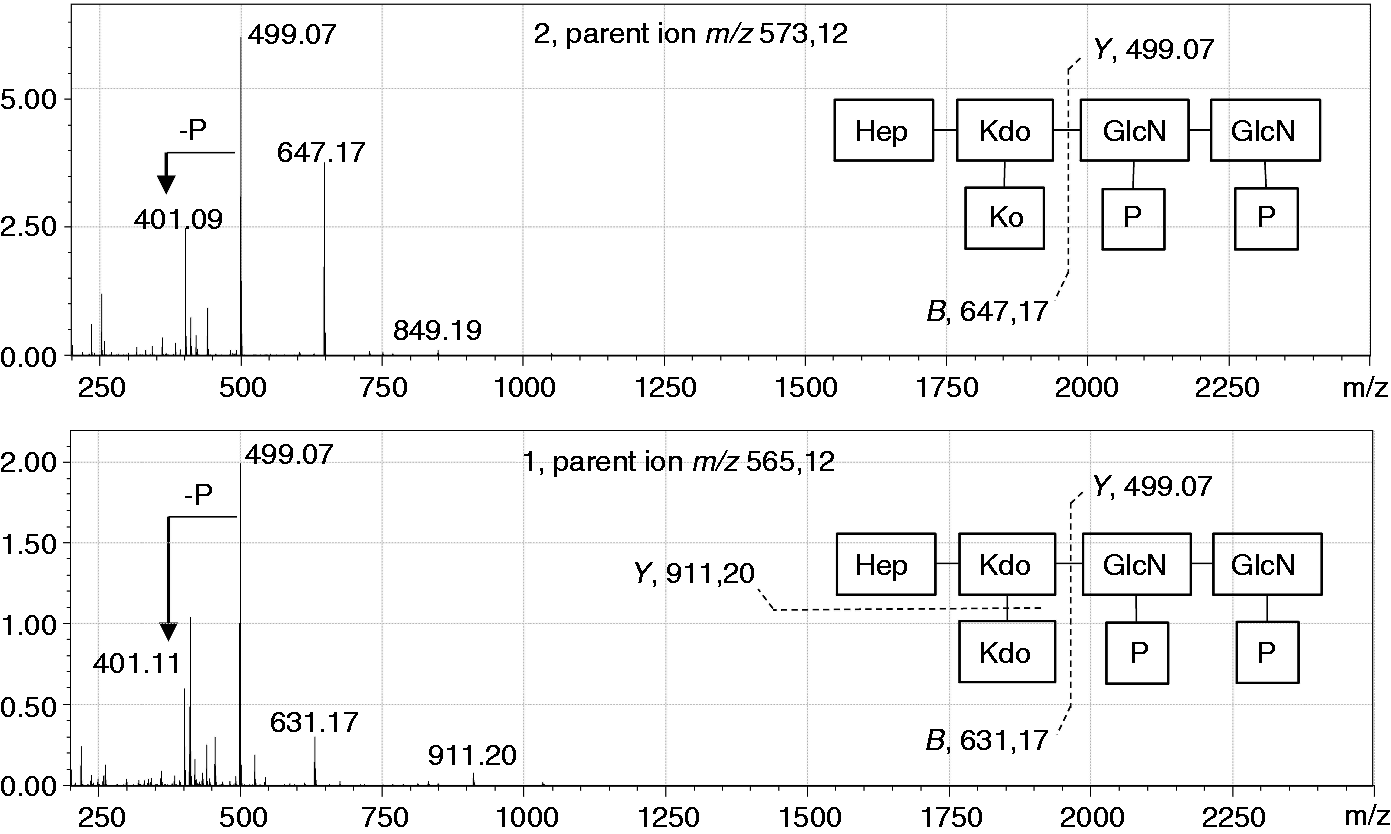
The MS/MS spectrum (negative ion mode) of
LPS truncation in No. 6 is the result of a mutation in waaF
The core oligosaccharides of Enterobacteriaceae species usually include three Hepp residues (Hep I, Hep II and Hep III) and three Hexp residues (Hex I, Hex II and Hex III).
41
However, while the three truncated LPS forms obtained from isolate No. 6 contained a normal lipid A component substituted with Kdo-Ko moieties, two of the forms had only a single Complementation of loss of WaaF function in strain No. 6 by the waaDFC operon. (A) Silver staining of LPS samples separated in DOC-PAGE. (B) Western blotting of the same samples using rabbit antiserum specific for Y. pseudotuberculosis serotype O:1b. The LPS samples loaded to the lanes are identified between the panels: No. 6/pWaaF6 = seven independent clones of the No. 6 strain complemented with waaDFC operon using plasmid pSW23T-waaF6; M164 = Y. enterocolitica strain YeO3-R1-M164, a waaF::CatMu mutant;
42
IP32953 = Y. pseudotuberculosis O:1b strain IP32953;
45
YeO3-R1 = Y. enterocolitica O:3 spontaneous rough mutant;
47
No. 6 = Y. pseudotuberculosis strain No. 6.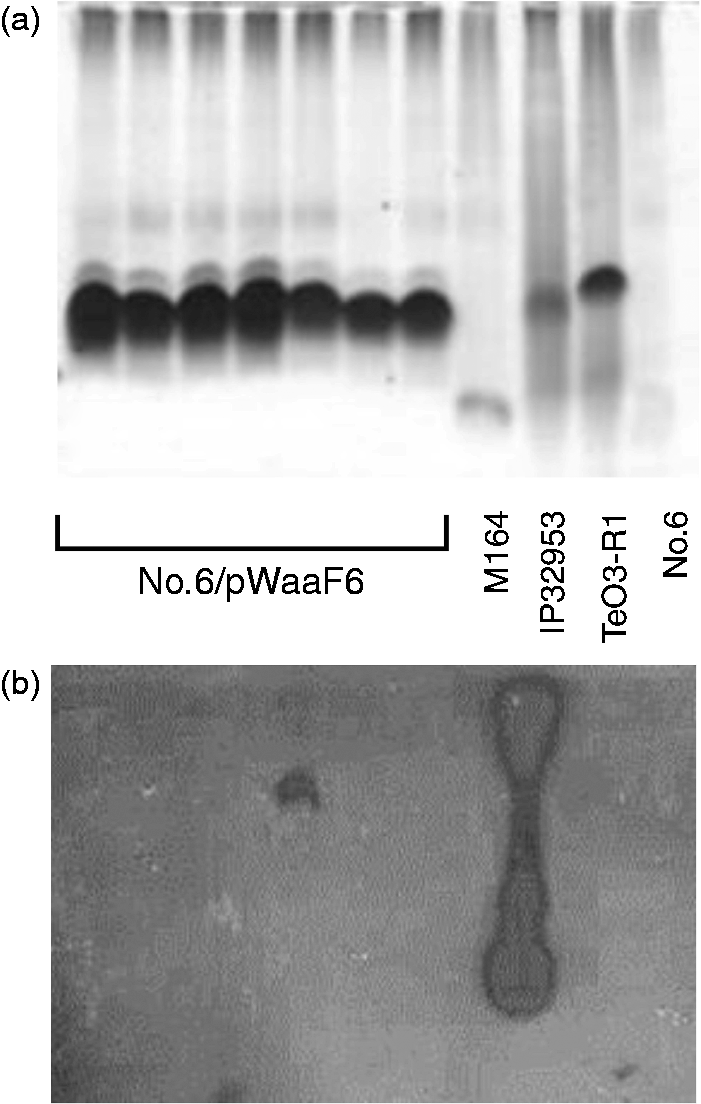
Sequencing of the waaF gene in No. 6 revealed a 53-bp deletion in the middle of the gene causing a frame shift and truncation of the gene product by about 160 C-terminal amino acids. As the waaF gene is a middle gene in a three-gene waa operon, we cloned the entire operon from serotype O:1b (isolate IP32953) to create plasmid pSW23T-waaF6, and used it to complement isolate No. 6. Complementation resulted in the restoration of the LPS core band to a size equivalent to that of serotype O:1b (Figure 10A). However, the complemented strain failed to show the O:1b-specific O antigen in Western blotting, indicating that strain No. 6 may additionally carry a second mutation in the O antigen gene cluster (Figure 10B). The nature of the second mutation was not addressed, as the deletion in waaF was sufficient to cause the truncation of LPS, and explained the origin of this O:8 strain.
Final comments
The two O:8 strains studied have different origins as shown by the O-antigen genes present, and we propose that epitope 20 is on a structure that is masked by O antigen, as both strains are rough mutants.
The two O:8 strains studied both have rough LPS, as reported previously for type strain 151. It is well established that under normal laboratory conditions many bacteria, including Y. pseudotuberculosis, accumulate point mutations in the OPS gene cluster that change the colony morphology from smooth to rough. In a wild type bacterium, the cell surface includes a heterogeneous population of LPS molecules with some containing lipid A and core oligosaccharide only, while others include either OPS or a single O unit. Thus, when the serotyping scheme for Y. pseudotuberculosis was developed, the antisera typically contained anti-core Abs that were removed when the antisera were cross-absorbed with bacteria of other serotypes.
In the last update to the scheme, the anti-O:8 antiserum was reported to also react with O:2c and O:4a bacteria, 4 and our analysis of the 151 strain that was used for this showed that it had an O:4a gene cluster and that a mutation in the ddhD gene of that gene cluster conferred the rough phenotype. The No. 6 strain is a more typical rough strain with a mutation in waaF that blocks completion of the inner core.
Here we have provided evidence that the two Y. pseudotuberculosis serotype O:8 isolates assessed in this study, are spontaneous mutants of other validated serotypes. It is interesting that both appear to have separate mutations affecting the O antigen and the LPS core. For 151 the mutation in ddhD would make the strain rough but did not explain the data obtained later on the core composition, which must be due to a separate mutation, probably in waaQ. For No. 6, the LPS composition data showed that the core was defective and so the O-antigen gene cluster was not sequenced. But later data showed that restoration of the core structure did not rescue O-antigen expression, probably because there is a mutation in the O-antigen gene cluster. The presence of two mutations in both strains is unlikely to be a coincidence. However, it is not surprising as mutants making incomplete O antigen are often deleterious and leads to further mutations. This phenomenon is not well understood, and falls outside of the scope of this study. However, these findings do show that serotype O:8 is not due to presence of a specific form of O antigen, and epitope 20 should be removed from the serotyping scheme, bringing the total number to 20 serotypes for the Y. pseudotuberculosis complex. The fact that the ancestral strains were of different serotypes indicates that it is being rough that exposes epitope 20, which is present but not exposed in the ancestral strains. In these circumstances there is no reason to further explore the nature of the mutations in the 151 core or the No 6 O-antigen gene cluster, but the nature of epitope 20 is a topic for future research.
Footnotes
Acknowledgements
The authors thank Sylvia Düpow (RCB) for technical assistance. JK was supported by an Australian Postgraduate Award (APA).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
