Abstract
Innate immunity coordinates LPS detection via TLR4 on polymorphonuclear leukocytes (PMNs) to elicit responses to many Gram-negative bacteria. In this study, we describe the effects of five subtypes of LPS [isolated from Escherichia coli B4, Pseudomonas aeruginosa PAO1, multidrug-resistant P. aeruginosa (MDRP), Acinetobacter baumannii and multidrug-resistant A. baumannii (MDRA)] on gene expression in PMNs. LPS isolated from B4, PAO1, and A. baumannii did not significantly alter TLR2 expression. However, LPS from MDRP and MDRA caused a 0.6-fold decrease and 2.7-fold increase, respectively, in TLR2 expression. Similarly, TLR4 expression was not significantly altered by LPS isolated from B4, PAO1 and A. baumannii but was down-regulated by LPS isolated from MDRP and MDRA by 0.1- and 0.6-fold, respectively. All LPS subtypes, excluding PAO1, down-regulated CD14 expression in PMNs. However, all five LPS subtypes up-regulated TNFA, IL1B, IL6, IL10 and TREM1 expression in a concentration-dependent manner, with the most substantial responses observed following exposure to LPS from MDRP and MDRA. These different effects on the gene expression in PMNs may depend on variation in LPS structural modifications related to acquired drug resistance, such as acylation and/or glycosylation.
Introduction
The human innate immune system identifies bacterial components via PRRs. 1 LPS present in the outer membrane of Gram-negative bacteria is often a diphosphorylated, polar macromolecule that contains hydrophobic elements within its lipid A core structure, as well as hydrophilic elements in its repeating polysaccharide surface. This molecule is recognized by several extracellular and cell surface proteins including the LPS-binding protein, CD14, MD-2 and TLR4, which are expressed by polymorphonuclear leukocytes (PMNs) and macrophages. The abundance of proteins involved in LPS detection reflects the requirement for specific protein–LPS and protein–protein interactions. 1 TLR4 induces both innate and adaptive immune responses to LPS. 2 This system triggers abundant inflammatory responses that enable clearance of bacterial infections; however, improper regulation can also result in immune system malfunction.
Studies on LPS structure during the course of infection have shown that LPS can be modified in response to environmental stimuli. 3 Lipid A acylation patterns have been shown to be altered in LPS isolated from Pseudomonas aeruginosa strains infecting the lung of patients with cystic fibrosis, and O-antigen composition has similarly been shown to vary with environmental conditions.4,5 However, gene expression analysis in PMNs stimulated by LPS from different Gram-negative bacteria strains has not been reported. In this study, we selected clinically important opportunistic organisms—P. aeruginosa, Acinetobacter baumannii (A.b) and their respective, multidrug-resistant strains—to elucidate their effects on PMN gene expressions.
Materials and methods
Reagents
Endotoxin-free Hanks’ balanced salt solution (HBSS), PBS and deionized water were purchased from Life Technology Japan (Tokyo, Japan). All reagents were checked and confirmed to be endotoxin free, DNase free, RNase free and pyrogen free by Life Technology Japan. Furthermore, all reagents and solutions included in the commercial kits in this study were confirmed to be free from contamination by endotoxin testing, which was performed in compliance with the following guidelines: ANSI/AAMI ST72: 2011; Bacterial endotoxins-Test methods, routine monitoring, and alternative to batch testing; USP <85> Bacterial Endotoxin Test. 6
LPSs
LPSs isolated from Escherichia coli O111:B4 (B4), P. aeruginosa (PAO1), multidrug-resistant P. aeruginosa (MDRP), A.b and multidrug-resistant A.b (MDRA) were purchased from Wako Pure Chemical Industries (Tokyo, Japan). LPS was prepared by phenol extraction. 7 The phenol extract contained < 1% protein. LPSs contained endotoxin levels of not less than 500,000 EU (endotoxin units)/mg.
PMN preparation
Human PMNs were isolated from healthy volunteers’ peripheral blood. 8 Briefly, 20 ml whole blood was mixed with 4.5% dextran solution and allowed to stand for 40 min at room temperature. The leukocyte-rich plasma was centrifuged at 400 g on an endotoxin-free Ficoll-Paque Plus gradient (Amersham Bioscience, Milwaukee, WI, USA) for 20 min. Hypotonic (0.2%) saline was used to lyse erythrocytes and osmolality was restored by hypertonic (1.6%) saline. PMNs were only used in experiments if the viability was 99% or more (as assessed by trypan blue exclusion) and the purity was at least 95% (as assessed by morphological analysis). PMNs were adjusted to a final concentration of 1 × 107 cells/ml in endotoxin-free HBSS. We obtained blood from nine volunteer donors [six men and three women, aged 29–58 yr (mean age 39 yr)]. Each donor donated blood more than once and the mRNA expression levels of 11 genes (one was the reference gene, ACTB) were averaged. Healthy individuals were confirmed, at the start of the study, through consent forms and interviews to be physically healthy, non-smokers who were not taking any medication and were informed about the study purpose; consent was obtained from all participants. The protocol was approved by the ethical review committee at the Teikyo University School of Medicine (No. 07-104).
PMN stimulation with LPS
LPS, resuspended in 100% DMSO (endotoxin-free), was inoculated into PMN suspension (5 × 106 cells/ml) at 10, 100, or 500 ng/ml. The control group in this study was untreated PMNs (5 × 106 cells/ml) without any added LPS. The mixtures were incubated with controls for 10 and 30 min in an environment containing 5% CO2. Following stimulation, PMNs were collected and washed with cold endotoxin-free PBS.
RNA preparation
Human PMNs (5 × 106 cells/ml) were washed with 1 ml of cold endotoxin-free PBS. Total RNA was extracted using the RNeasy Plus Mini Kit (QIAGEN, Germany) according to the manufacturer’s instructions. The quantity and quality of total RNA samples were determined using the Agilent 2100 Bioanalyzer (Agilent Technologies, Waldbronn, Germany).
Complimentary DNA synthesis
Total RNA was reverse-transcribed to cDNA by using SuperScript VILO cDNA synthesis kit (Invitrogen Life Technologies, Carlsbad, CA, USA). Briefly, 1 µg of total RNA was incubated with 2.5 μM oligo (dT)20, 50 ng of random hexamers and 200 U SuperScript III RT enzyme in a 20-μl reaction volume at 42℃ for 60 min, followed by 50℃ for 20 min. The reactions were terminated by heating the solution at 85℃ for 5 min.
Quantitative real-time PCR (qPCR) analysis
Primer sets for quantitative polymerase chain reaction.

F: forward primer; R: reverse primer; bp: base pairs.
Briefly, in this study, we employed the absolute quantification method for qPCR analysis. The copy numbers of each target gene were divided by the copy number of the same gene in untreated PMNs. All copy numbers of the target gene were adjusted relative to the number of copies of the housekeeping gene ACTB.
Statistical analysis
All P-values were determined by the nonparametric unpaired or paired t-tests (two-tailed) using Excel 2011 (Microsoft, Tokyo, Japan). We considered P < 0.05 to be significant and the degree of significance is indicated as **P < 0.01.
Results
We did not find any significant differences in gene expression levels of the 11 genes tested in PMNs between healthy men and women stimulated with LPS under the same conditions (data not shown).
PRRs in LPS-stimulated PMNs
TLR2 gene expression levels in B4, PAO1 and A.b LPS-stimulated PMNs were not significantly altered. MDRP LPS treatment, however, down-regulated TLR2 expression in PMNs 0.6-fold, while MDRA LPS stimulation up-regulated it 2.7-fold (Figure 1A).
Gene expression analysis of PRRs in LPS-stimulated PMNs. mRNA expression levels of (A) TLR2, (B) TLR4,and (C) CD14 can be seen. LPS was administered at 0, 10, 100 and 500 ng/ml. All reaction tubes were incubated at 37℃ for 10 min. mRNA expression levels observed for 0 ng/ml LPS were considered as baseline and designated as 1.0. The fold-change is indicated as a ratio. *P < 0.05; **P < 0.01 compared with healthy subjects. Error bars represent SEM. Data are representative of at least three separate experiments.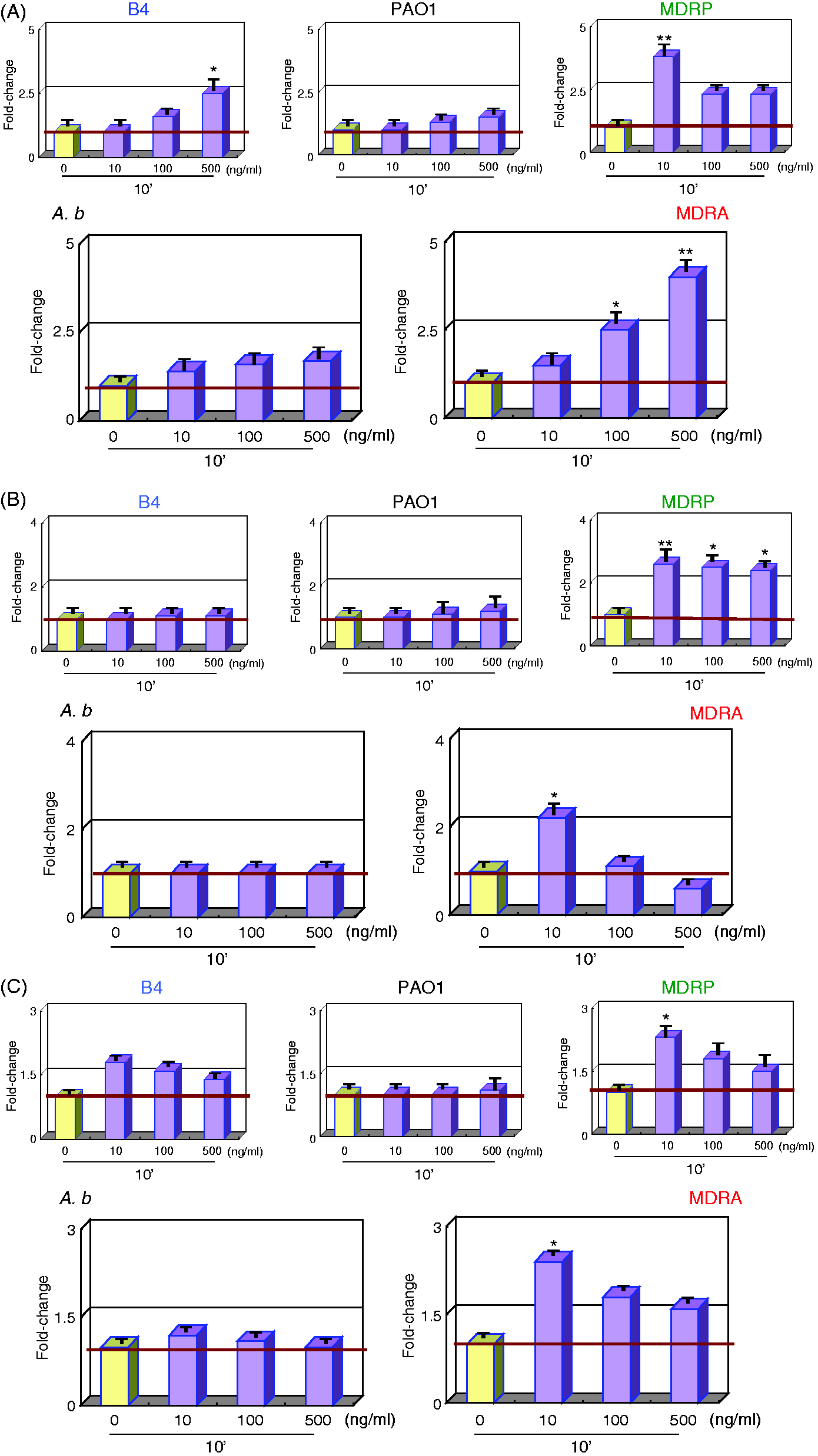
Similarly, B4, PAO1 and A.b LPS did not significantly alter TLR4 expression in PMNs. In comparison, MDRA and MDRP LPS-stimulated PMNs expressed 0.1- and 0.6-fold lesser levels of TLR4 transcript, respectively (Figure 1B).
PMNs stimulated by all LPS subtypes, excluding PAO1, down-regulated CD14 expression. MDRA and MDRP LPS elicited the most significant repression, which caused an average 0.3-fold reduction in CD14 transcript levels (Figure 1C).
Inflammatory cytokines in LPS-stimulated PMNs
All LPS subtypes up-regulated TNFA, IL1B and IL6 expression in PMNs in a dose-dependent manner. Moreover, the transcript levels of these inflammatory cytokines from MDRA and MDRP LPS-stimulated PMNs were higher than those in PMNs stimulated by A.b and PAO1 LPS (Figures 2A–C) and induced an average fold change of 3.2—7.5.
Gene expression analysis of inflammatory cytokines in LPS-stimulated PMNs. mRNA expression levels of (A) TNFA, (B) IL1B and (C) IL6 can be seen. Sample treatments and data analyses were as outlined in Figure 1.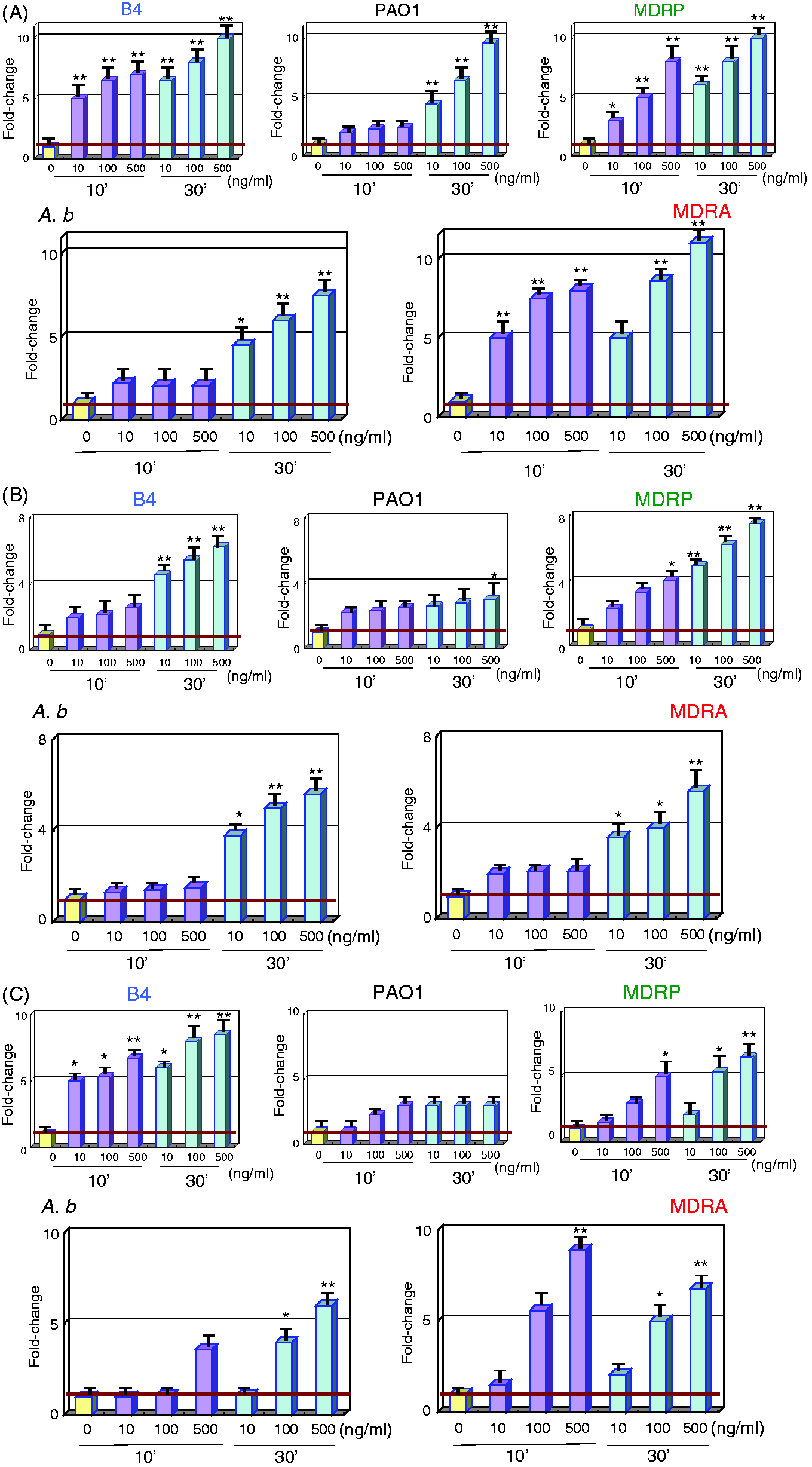
Chemokine receptors in LPS-stimulated PMNs
IL8Rs expression levels in LPS-treated PMNs were not significantly altered (Figure 3A, B). However, MDRA and MDRP LPS stimulated higher levels of ITGAM expression in PMNs compared with A.b and PAO1 LPS (Figure 3B). MDRA and MDRP LPS-stimulated PMNs up-regulated ITGAM expression by 1.5-and 1.7-fold, respectively.
Gene expression analysis of chemokine receptors in LPS-stimulated PMNs. mRNA expression levels of (A) IL8Rs and (B) ITGAM can be seen. Sample treatments and data analyses were as outlined in Figure 1.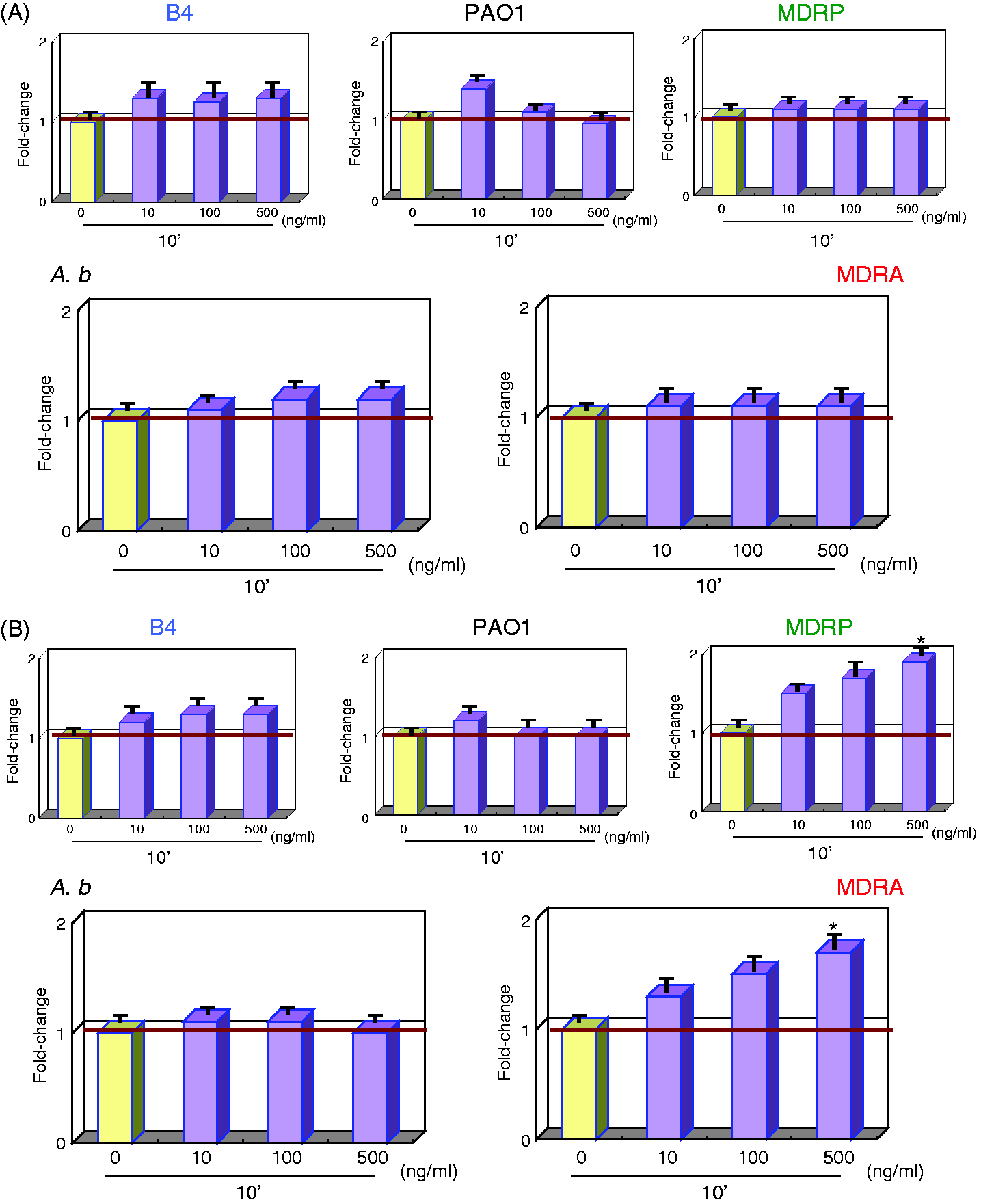
Anti-inflammatory cytokine in LPS-stimulated PMNs
All LPS subtypes up-regulated IL10 expression in a dose-dependent manner. In particular, IL10 expression in MDRA LPS-stimulated PMNs was higher than that in A.b-stimulated PMNs (Figure 4). On average, IL10 expression was up-regulated 3.8-fold in response to LPS.
Gene expression analysis of the anti-inflammatory cytokine IL10 in LPS-stimulated PMNs. mRNA expression levels of IL10 can be seen. Sample treatments and data analyses were as outlined in Figure 1.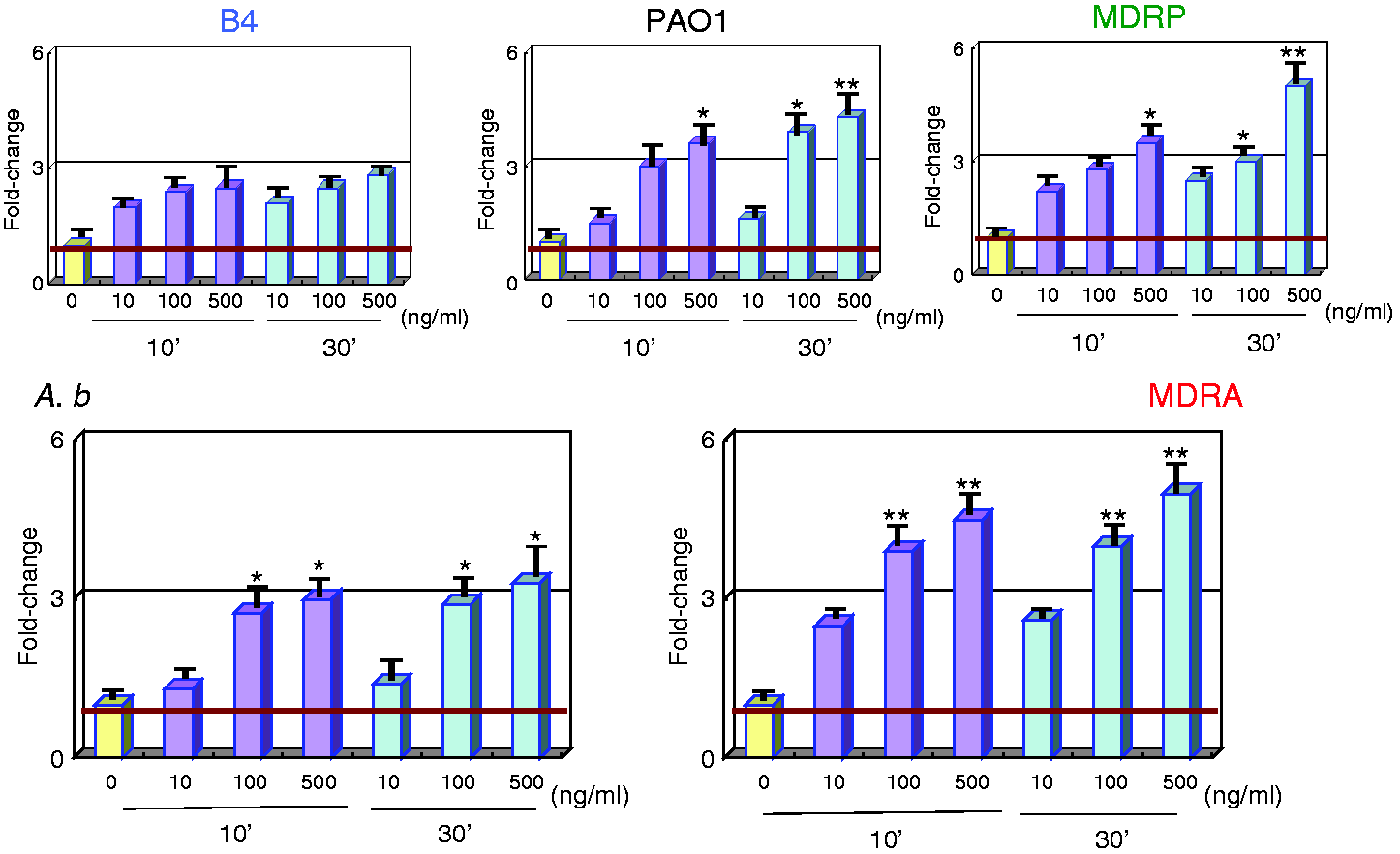
Pro-inflammatory myeloid cell receptor in LPS-stimulated PMNs
All LPS subtypes, excluding A.b, induced up-regulation of TREM1 in a dose-dependent manner. TREM1 levels in MDRA and MDRP LPS-stimulated PMNs were higher than those in A.b and PAO1 LPS-stimulated PMNs (Figure 5). LPS from these multidrug-resistant subtypes up-regulated TREM1 expression 2.7- and 2.2-fold, respectively.
Gene expression analysis of the pro-inflammatory myeloid cell receptor TREM1 in LPS-stimulated PMNs. mRNA expression levels of TREM1 can be seen. Sample treatments and data analyses were as outlined in Figure 1.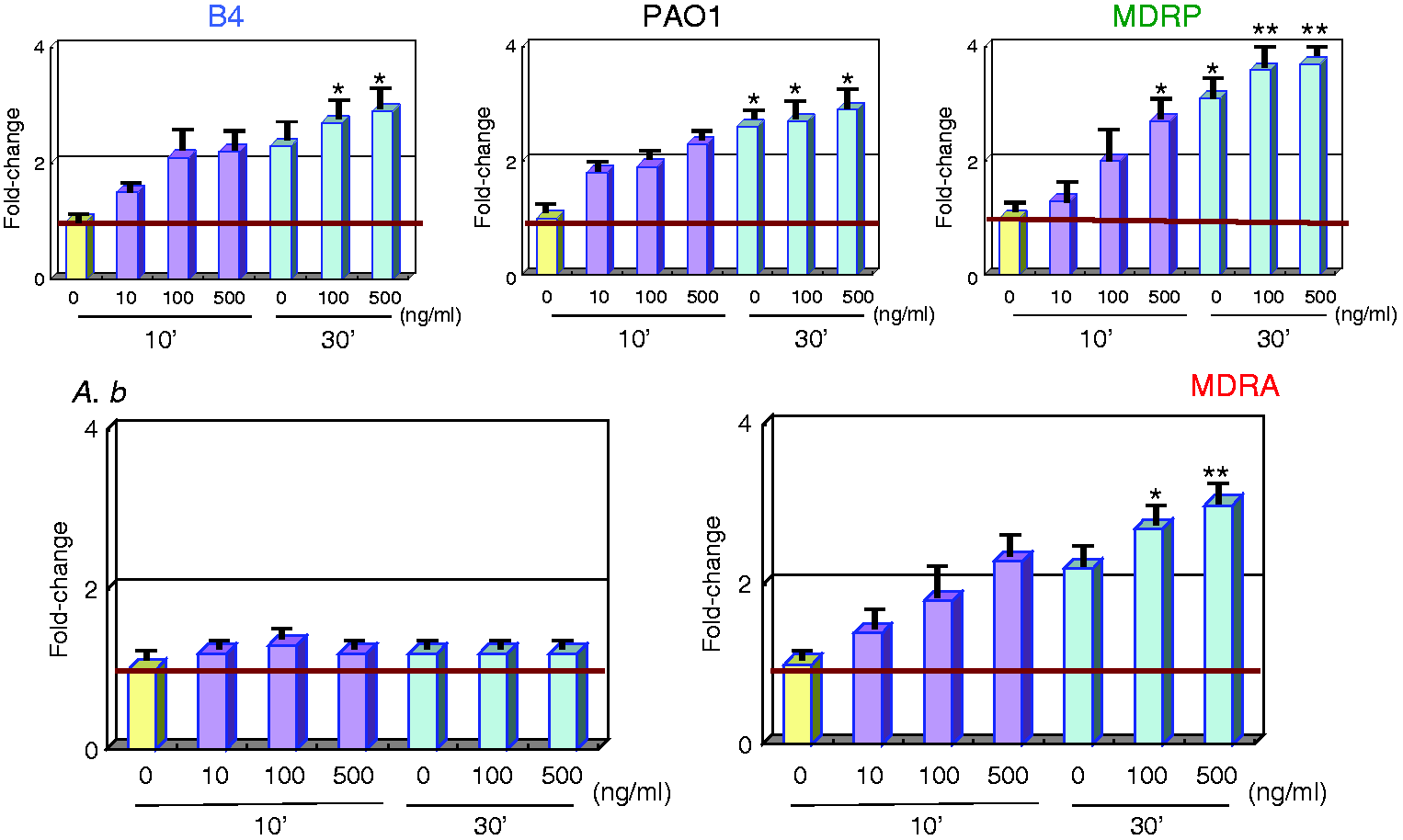
Discussion
Even at a minute dose, LPS can induce endotoxemia, which causes increased levels of circulating pro-inflammatory cytokines, exacerbated hypothermia and an often-fatal outcome. It can also activate non-transcriptional cellular responses, including autophagy, endocytosis, phagocytosis, and oxidative burst.11–13 In human cells, three receptors for LPS have been identified: (1) soluble or membrane-bound CD14–MD2–TLR4 complex, (2) ITGAM (β2 integrins) and (3) scavenger receptors for lipid molecules. Soluble and membrane-bound CD14 greatly potentiates the host response to small quantities of LPS and other microbial mediators. 14
MD2–TLR4 best detects bisphosphorylated diglucosamine to which six saturated fatty acyl chains with lengths of 12, 14 or 16 carbons are attached; this structure is called ‘hexa-acyl LPS’.
15
Escherichia coli (E. coli) and A. b. produce LPS with predominantly hexa-acyl lipid A structures. However, P. aeruginosa produces LPS with predominantly penta-acyl lipid A structures.
15
Lipid A is recognized as the most toxic and/or inflammatory region of LPS, although the polysaccharide portion of the molecule also has potent immunomodulatory and immunostimulatory properties.
16
In genetic analysis, the activation of a two-component system (TCS), PmrA/B, is triggered by environmental stimuli and specific mutations within the TCS that result in their constitutive activation and subsequent overexpression of LPS-modifying genes.17,18 Activation of the PmrA/B TCS leads to the up-regulation of the pmrCAB and arnBCADTEF-pmrE operons, which mediate the synthesis and transfer of pEtN and 4-amino-4-deoxy-
Modification of lipid A with pEtN occurs as a result of mutations in the pmrA/B TCS in A. b. Mutations in pmrA and/or pmrB have been found to induce the constitutive expression of pmrA (the cognate response regulator) and the subsequent autoregulation of the promoter region of the pmrCAB operon, which leads to the modification of the 4′- or 1-phosphate group of lipid A with pEtN. Mutations have been reported in different domains of both PmrA and PmrB.21,22 Additionally, pmrB appears to be the most commonly mutated gene in A. b. 23 Moreover, glycosylation of lipid A with galactosamine (hexosamine) at the 1-phosphate group has been recently reported in resistant strains. 24
Unlike A. b, P. aeruginosa has both the pmrA/B and phoP/Q TCSs, each of which can separately regulate the arnBCADTEF operon. 25 Several studies have shown that P. aeruginosa can develop resistance to polymyxins via the constitutive modification of its LPS with Arap4N, which is stimulated by the pmrA/B and phoP/Q TCSs.25,26 The synthesis and transfer of Arap4N to the lipid A are accomplished by the arnBCADTEF operon in P. aeruginosa. These genetic alterations are known to lead to the modification of lipid A with Arap4N. For P. aeruginosa, only one study has reported a mutation in pmrA that may be responsible for resistance; 27 the other mutations that have been described are mainly localized to PmrB or generally distributed to cognate regulators of TCSs. In terms of lipid A modifications in resistant bacteria, there appears to be an absolute relationship between colistin resistance and Arap4N addition and a significant relationship between colistin resistance and the loss of secondary lauroyl chain hydroxylation. Moreover, there is wide structural variation in lipid A in polymyxin-resistant P. aeruginosa in terms of the presence of either penta- or hexa-acylated lipid A. 28
Once TLR4 binds to LPS, two possible pathways of cellular activation can occur through either the myeloid differentiation factor MyD88 or the TLR domain adaptor-inducing IFN-β (TRIF) pathway.29,30 The cascade of inflammatory cytokines and other inflammatory mediators following LPS exposure contributes to generalized inflammation, procoagulant activity, tissue injury and septic shock.31,32 Similarly, TLR4 activation in response to LPS signaling can also lead to LPS toxicity and plays a role in septic shock. 33
Core genes in innate immunity, such as PRRs, inflammatory cytokines, chemokine receptors, anti-inflammatory cytokines and the pro-inflammatory myeloid cell receptor were selected, 34 and quantitative real-time RT-PCR was performed to validate their expression. Specifically, we evaluated TLR2, which recognizes bacterial lipoproteins; TLR4 and CD14, which recognize LPS from Gram-negative rods; TNFA, IL6 and IL1B, which were induced by activation of PRRs (for IL1B, this occurred via the inflammasome); IL8 (a strong chemoattractant of PMNs) and ITGAM, which are recognized by IL8Rs and mediate PMN migration; IL10, an anti-inflammatory and resolving cytokine; and TREM1, an amplifier of inflammation and useful biomarker of bacterial infections.
The mRNA expression levels of TLR2, TLR4, CD14, IL8Rs and ITGAM after 30 min of LPS stimulation were not significant or demonstrated a similar pattern as that observed after 10 min of LPS stimulation (data not shown). However, the mRNA levels of TNFA, IL1B, IL6, IL10 and TREM1 after 10 and 30 min LPS stimulation were significantly increased in a dose-dependent manner. The results of TLR2, TLR4 and CD14 expression did not indicate a dose-dependent effect (Figure 1). We speculate that this was mainly attributed to an overdose of LPS. PRRs recognize specific PAMPs and are sensitive because they guard the entrance to the signal-transduction cascade. 14 The minimum concentration of LPS for recognition may therefore be < 10 ng/ml (data not shown). In our study, gene expression levels of PRRs (TLR2, TLR4 and CD14), inflammatory cytokines (TNFA, IL1B and IL6), chemokine receptors (IL8Rs and ITGAM), an anti-inflammatory receptor (IL10), and a pro-inflammatory myeloid cell receptor (TREM1) in LPS-stimulated PMNs were altered between multidrug-resistant organisms (MDROs) and non-MDROs. In MDROs, various modifications (i.e. acylation and glycosylation) of lipid A can occur. MD2–TLR4 best detects hexa-acyl LPS. E. coli and A. b produce LPS with predominantly hexa-acyl lipid A structures. However, P. aeruginosa produces LPS with predominantly penta-acyl lipid A structures. 15
In MDROs, such as MDRA and MDRP, the presence of mutations in pmrA and/or pmrB can cause modification in the lipid A structure. In MDRA, hexa-acyl lipid A may be predicted to be modified by glycosylation, whereas penta-acyl lipid A in MDRP may be characterized as either penta- or hexa-acylated lipid A. 28
Average fold change of gene expression levels in LPS-stimulated PMNs related to untreated controls. Positive numbers indicate up-regulation and negative values represent down-regulation.
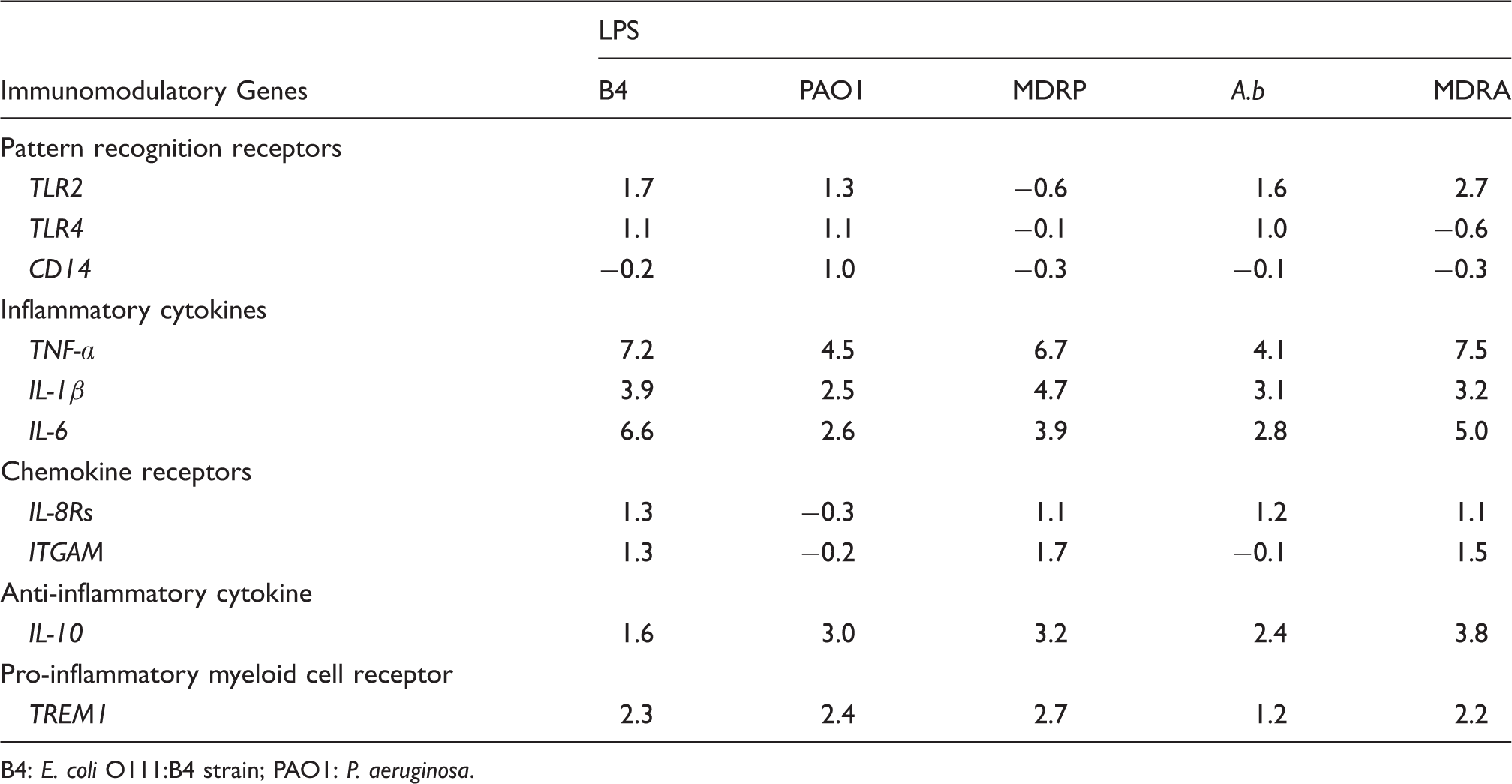
B4: E. coli O111:B4 strain; PAO1: P. aeruginosa.
These results suggest that two common single-nucleotide polymorphisms in TLR2 and TLR4—Arg753Gln and Asp299Gly, respectively—are associated with shorter time to onset of severe sepsis or septic shock in patients admitted to the intensive care unit. 37 Additionally, it is possible that gene-specific epigenetic reprogramming occurs in endotoxin-tolerant murine macrophages. For example, TLR4-dependent processes remodel chromatin at distinct classes of genes such that pro-inflammatory genes are silenced, while anti-inflammatory and antimicrobial genes are activated. 38
It is also possible that these LPS subtypes contain different biochemical modifications, and are thus structurally distinct. Indeed, gene expression levels in PMNs in response to LPS from drug-sensitive strains (PAO1 or A.b) different from that to LPS from their respective drug-resistant strains (MDRP or MDRA). Several mechanisms of resistance to polymyxins such as colistin and other antimicrobial peptides have been reported; most of these pathways alter LPS synthesis to decrease the affinity between the antimicrobial agent and its target. 39 For example, two mechanisms of colistin resistance have been reported in A.b. The first involves inactivation of one of three genes required for lipid A biosynthesis—lpxA, lpxD, or lpxC—which results in total loss of LPS.40,41 In a second mechanism, lipid A modification mediated by the addition of phosphoethanolamine results in loss of colistin affinity and subsequent resistance; these modifications are regulated by the PmrA/B TCS.22,42
LPS and lipid A from A. b are implicated in diverse biological activities such as toxicity, pyrogenicity, mitogenicity and inflammation.43,44 Particularly, Fernandez et al. reported that biofilm formation and swarming motility of MDRP led to multiresistant states. 45 Pelletier et al. predicted that Acinetobacter harbors lptA, which encodes pEtN transferase at the 4′ position of lipid A. 24 Mutation and pEtN addition to lipid A resulted in an increased inflammatory response via TLR4. These findings may also affect the virulence and cytotoxicity of MDRP and MDRA.
Furthermore, A.b is naturally competent for DNA uptake and demonstrates high rates of natural transformation. 46 Genomic studies have reported that some A.b strains have ≥ 28 genomic islands encoding antibiotic resistance determinants. 47
Among PAO1 variants, psrA mutants exhibit supersusceptibility to polymyxin B and indocilin because of outer membrane permeabilization. Multiple pathways are implicated in this increased sensitivity, including virulence factors such as type III secretion systems, biofilm formation, and adhesion or motility. 48
In addition, pure LPS is likely to exhibit more hydrophobic qualities, resulting in rapid binding to available host membranes 3 . Furthermore, the physical size of LPS complexes directly influences CD14-dependent internalization of these complexes. 49 Future studies about the genetic and cellular mechanisms that govern LPS-stimulated effects on gene expression are necessary to appreciate fully this fascinating recognition of PAMPs by PRRs.
Footnotes
Acknowledgements
We thank Ms. C Miyazaki for providing technical assistance.
Funding
Financial support for this study was provided by a grant-in-aid from the Ministry of Education, Culture, Sports, Science and Technology of Japan (24591490).
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
