Abstract
TLRs represent one of the most important components of innate immunity. Currently, these receptors have been extensively studied in vertebrates and insects, but our knowledge for annelids is very limited. Therefore, the aim of our study was to identify earthworm TLR homologs by genome-wide screening, and to determine the expression of candidate genes as a response to Gram-positive and Gram-negative bacteria. Using a combination of deep pyrosequencing and quantitative PCR we found six candidate genes, for which all were expressed in
Introduction
Invertebrates have a simple immune system, recognizing harmful or pathogenic microorganisms mainly through innate PRRs.1–3 The PRRs recognize PAMPs associated with the invading microorganisms. PRRs are generally conserved throughout evolution, with TLRs being the most widespread. 4 These have an architecture consisting of extracellular leucine-rich repeats where the extracellular domains transmit the sensed signals through a cascade of adaptor molecules to activate a range of defense mechanisms. However, we still know very little about the role of TLRs in annelids,3,5 although they have been extensively studied for vertebrates—in which several families and sub-families have been identified 6 —and insects in which the TLRs were originally detected.7,8
Earthworms are oligochaetal annelids living in soil. 9 Soil represents a microbiologically hostile environment, with > 20,000 microbial genome equivalents. 10 Despite their simple immune system, earthworms utilize the soil environment as habitat. 11 TLRs could be an important component in doing so.
Therefore, the aim of this work was (i) to identify candidate TLRs in the earthworm
Our strategy was to first identify TLRs in the
We present results showing gene expression responses of the TLR candidates to bacterial exposure, supporting their role in earthworm immunology. 5
Materials and methods
Earthworm culture
The experiments were conducted with the earthworm species
Earthworm genome sequencing and analysis
The anterior part of a single earthworm was subjected to genome sequencing. Total DNA was isolated using the Qiagen tissue kit following the manufacturer’s recommendations (Qiagen, Hilden, Germany). The genomic DNA was prepared according to the manufacturer’s instructions for 454 sequencing of total DNA and subsequently sequenced using the Genome Sequencer FLX Titanium System (Roche, Basel, Switzerland). The genome sequencing was done at the Norwegian Sequencing Center (www.sequencing.uio.no). The sequences were deposited in the National Center for Biotechnology Information (NCBI) database with the ID SRS523963 under the BioSample accession SRP034887.
TLR homologs were identified by BLAST searches in a local database generated from the pyrosequencing reads by the CLC Genomic Workbench (CLC bio). Candidate genes used for the searches were TLR homologs identified from the
PCR primers
Primers and probes used in this study.
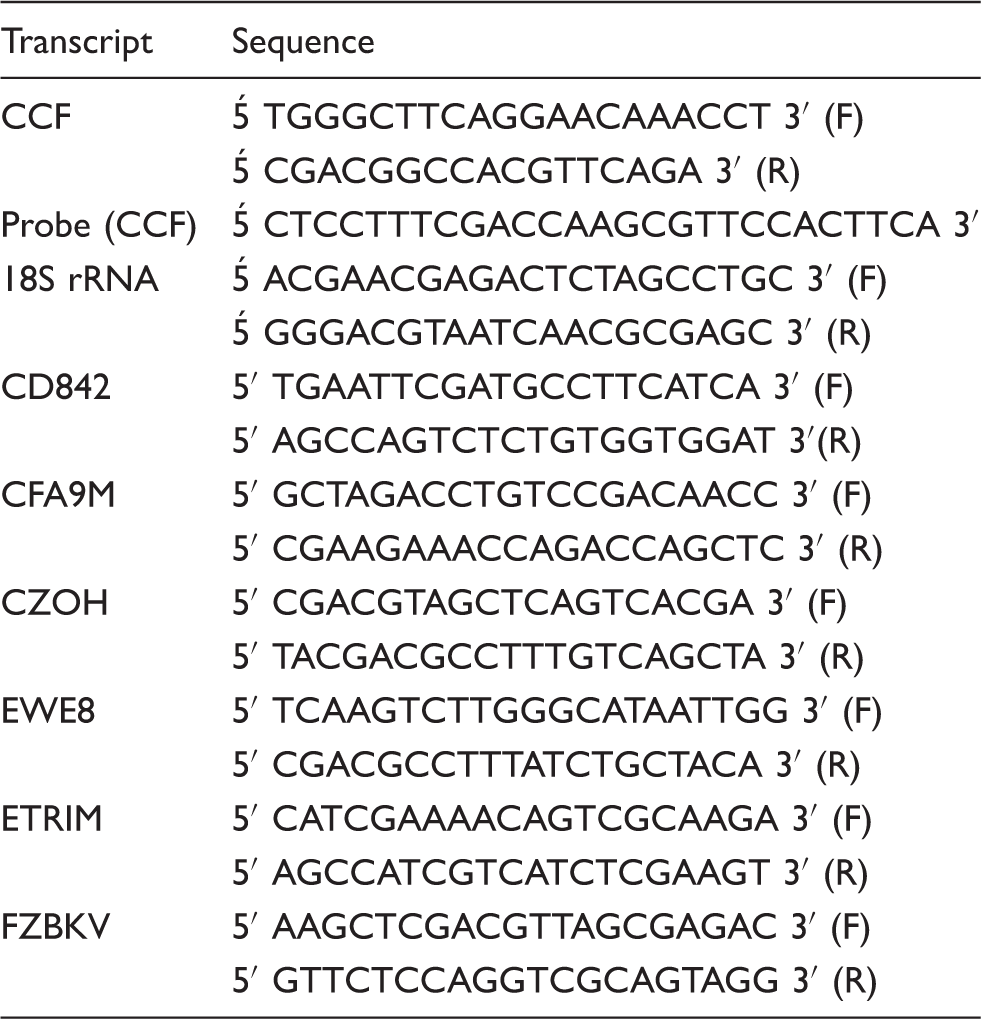
Gene expression experiments
Three main gene expression experiments were conducted: (i) the effect of bacterial exposure, (ii) the effect of starvation and (iii) longitudinal investigation of individual differences. The longitudinal nature of the experiments was both to identify potential confounding gene expression responses, and to investigate individual differences in expression patterns as these could influence results interpretation.
The effect of bacterial exposure was determined by soaking the earthworms in cultures containing 109 CFU/ml of
Immune cells were harvested after extraction of immune cells by electro stimulation of earthworms at 5–10 V for 5 s in a modified
RNA isolation was performed using Qiagen RNeasy Mini Kit on freshly harvested immune cells. Homogenized lysed samples were stored at −80℃ until all samples in the experiment were collected as described by the manufacturer. Frozen lysates was thawed at 37℃ and protocol from the manufacturer was followed for isolation of RNA.
Further, the isolated RNA was treated with DNase1, Amplification Grade (Cat. no. 18068-015; Invitrogen, Carlsbad, CA, USA) before used as template for complementary DNA (cDNA) using reverse transcriptase. cDNA was synthesized using RNase OUT, Ribonuclease Inhibitor (Recombinant), (Cat. no.10777-019; Invitrogen) and SuperScript III Reverse Transcriptase (Cat. no. 18080-044, Invitrogen). The manufacturer’s protocol was followed. A control containing reagents, but no template, was included in the experiments as a negative control.
Amplifications were conducted in a 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) using 20 µl reaction volumes containing 1 unit Dynazyme II HotStart DNA polymerase (Thermo Scientific, Waltham, MA, USA), 1 × Dynazyme TM II Hot start reaction buffer (Thermo Scientific), 400 µmol dNTPs (Thermo Scientific), 0.2 × Rox (Life Technologies, Invitrogen) 0.2 µmol probe (Biomers), 0.4 µmol of each primer (Biomers). All amplifications were conducted using 40 cycles with denaturation at 95℃ for 30 s and elongation at 60℃ for 1 min. The primers and probes used are listed in Table 1. LinRegPCR was used to estimate the amplification efficiency,
21
while gene expression was measured relative to the 18S rRNA gene. Statistical testing of the measured expression levels of the identified homologs was done using the Mann–Whitney
Results
Identification of TLR homologs
The genome sequencing resulted in a total of 556,684 sequencing reads with an average length of 325 base pairs (bp). The sequences represent 180 million bp. The genome size of
TLR homologs identified in
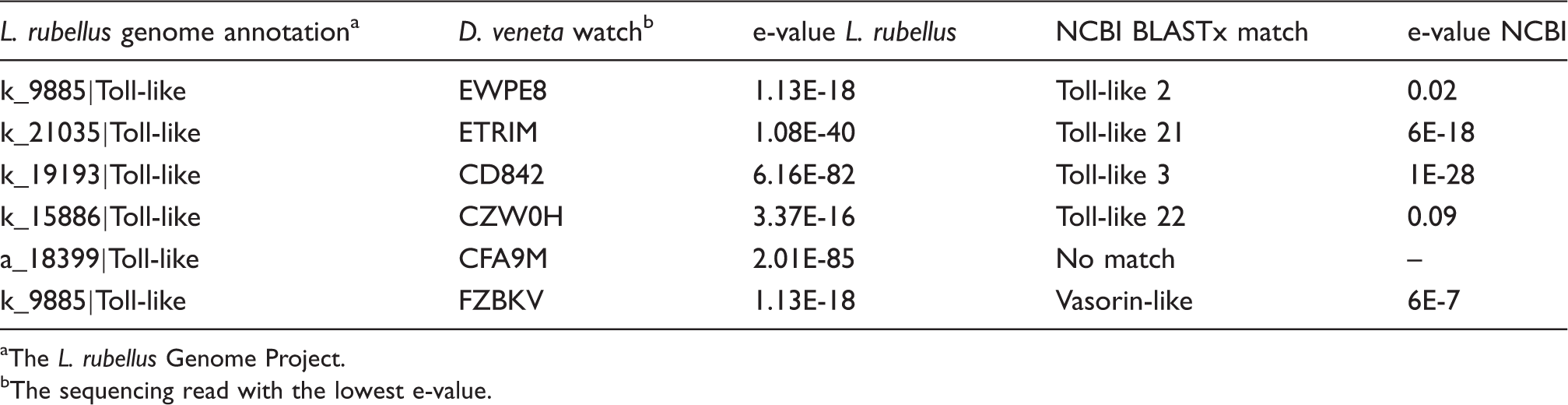

The
The sequencing read with the lowest e-value.
Gene expression of TLR homologs
We found relatively large gene expression effects for the TLR gene homologs ETRIM and FZBKV for the earthworms externally exposed to Expression of TLR gene homologs as response to bacterial exposure. Earthworms surface exposed to about 109 CFU/ml were analyzed 2–4 d after exposure. Error bars represent SEM between individual earthworms. **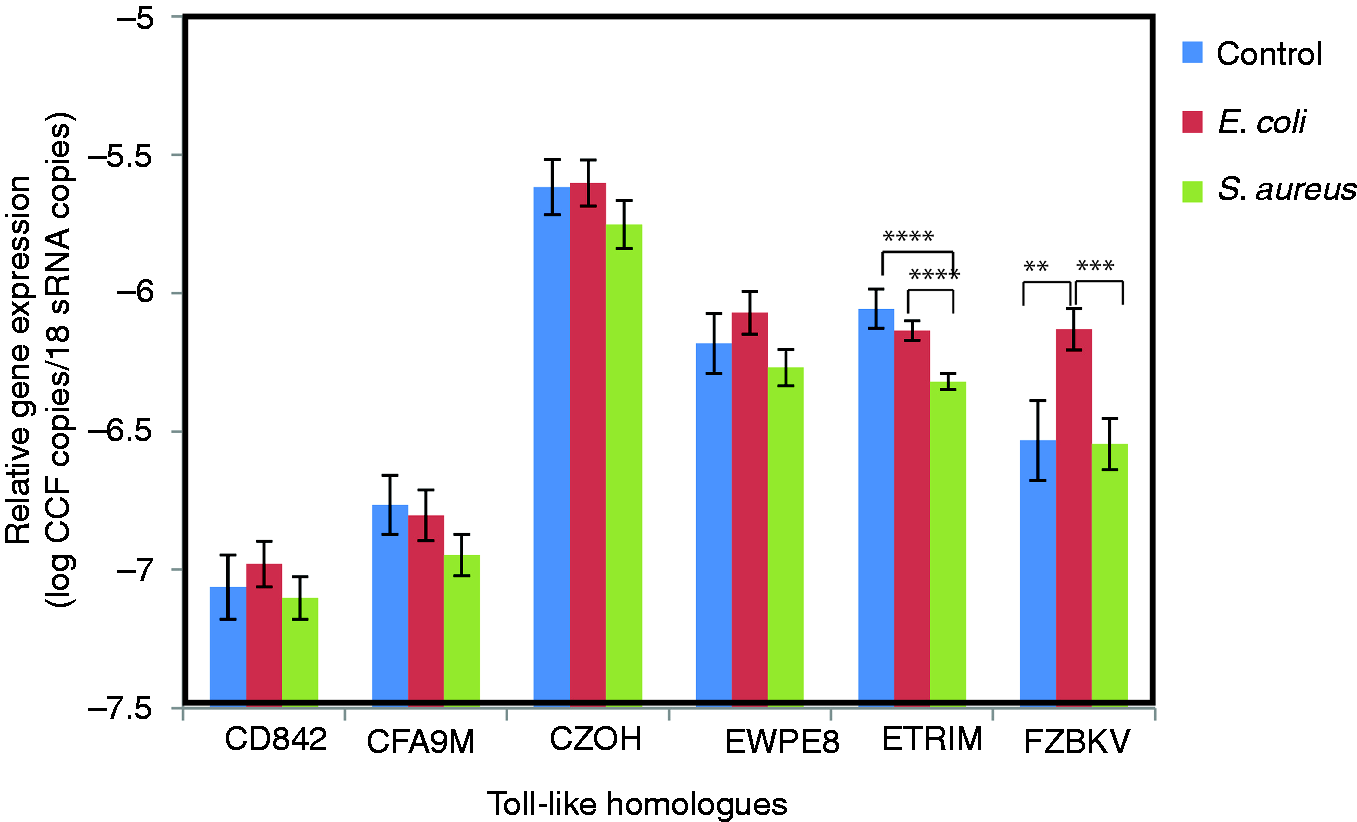
As all experiments were done on starved earthworms we conducted an experiment in which we tested the gene expression effect of starvation over 5 d. We found no effect of starvation for any of the genes investigated (Supplementary Table 1). Finally, we tested whether there were individual differences in the gene expression by analyzing the immune cell expression of the TLR homologs over a period of 3 mo. The testing was repeated three times for 12 earthworms. We did not detect any major individual differences in the expression patterns for any of the TLR homologs (Supplementary Table 2).
CCF gene expression
We could not detect any systematic differences in CCF expression for any of the experimental conditions tested (results not shown). The average CCF expression was about 5 log lower than that of 18S rRNA. What was apparent, however, were stable individual differences in the gene expression patterns over the 28 d investigated, where one earthworm showed a more than 10-fold higher expression than the others (Figure 2).
Individual differences in CCF gene expression. Longitudinal analyses were conducted for 12 earthworms after 0, 14 and 28 d. Error bars represent SEM of three measurements for each earthworm.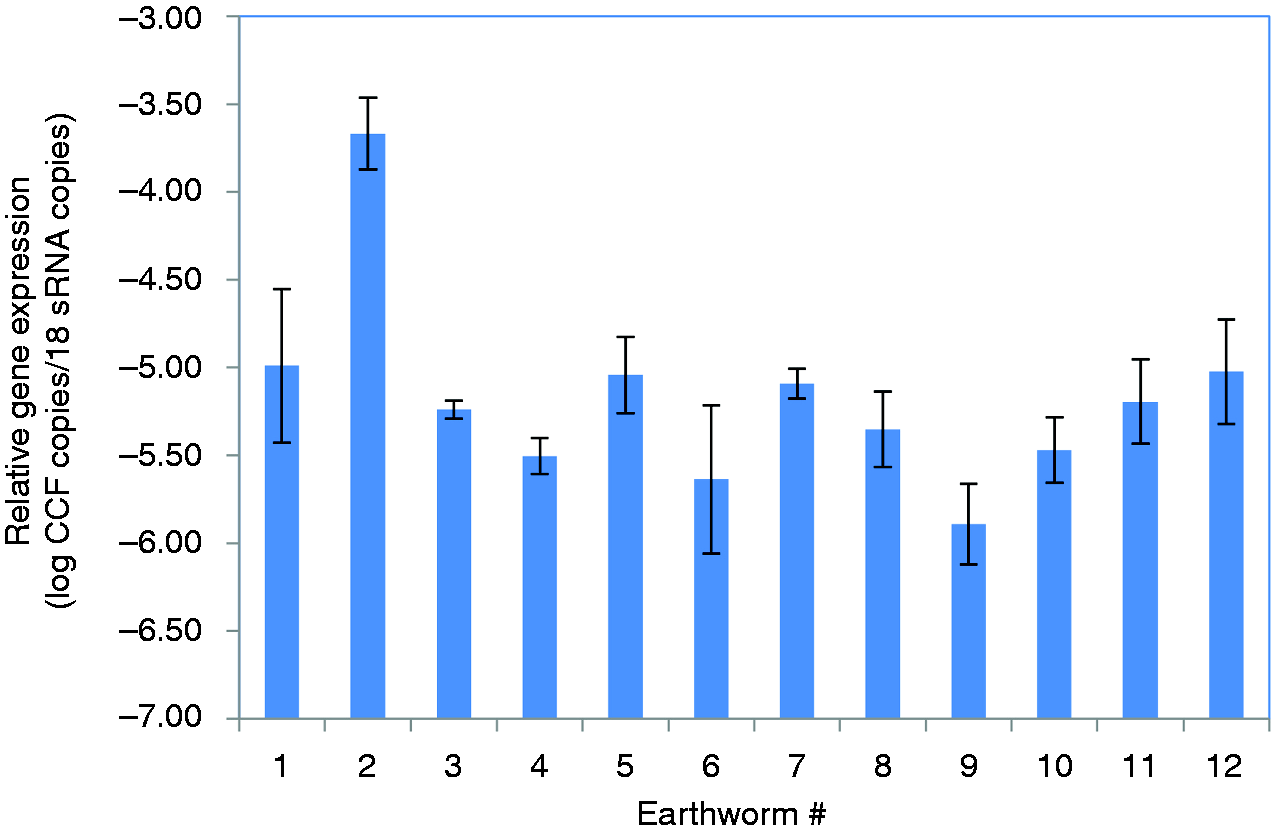
Discussion
The most prominent and systematic gene expression responses were the down-regulation of the TLR 21 homolog (ETRIM) after earthworm exposure to
TLR 21 has recently been identified in a range of non-mammalian vertebrates.
23
The receptor recognizes CpG oligonucleotides with bacterial origin, and activation requires acidification of endosomes.
Vasorin expression, however, has been shown to be connected to tissue damage.
26
For vasorin homolog expression, the
It has previously been suggested that TLRs play little or no role in annelid immunology.
3
The TLR homolog gene expression response detected by us, however, suggests that TLRs also have a role in annelids. This has also been confirmed by another study identifying a TLR homolog in the earthworm species
CCF in
In conclusion, we have shown gene expression responses of TLR homologs in earthworms after microbial exposure and individual variation in CCF expression. These results represent new knowledge about the innate immunology of earthworms.
Footnotes
Funding
This work was financed by Hedmark Sparebank and Hedmark University College.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
