Abstract
The health profile of Indigenous Australians is characterised by high rates of classic ‘lifestyle’ diseases. Potential roles of inflammation in pathophysiology of these diseases requires investigation. It is not clear if genetic regulation of inflammation in Indigenous Australians is similar to other populations. This study characterised frequencies of single nucleotide polymorphisms (SNPs) for eight cytokine genes for 100 individuals from a remote Indigenous Australian community and assessed novel genetic variants in four cytokine genes. We used a commercially-available allelic discrimination assay for SNP genotyping; re-sequencing was undertaken by standard Sanger sequencing methodologies for 26 samples. Frequencies of cytokine gene SNPs differed significantly from the Caucasian population (
Introduction
Indigenous Australians have a predicted life expectancy that is 17 yrs less than non-Indigenous Australians. 1 Both the incidence and effects of diseases associated with a modern lifestyle are greater in Indigenous populations than their non-indigenous counter-parts:2,3 rates of coronary heart disease are up to one and a half times higher; rates of type 2 diabetes (T2DM) are up to three times higher; and rates of end-stage renal disease are more than 10 times higher. 1 An increased incidence of lifestyle disease has also been reported in other Indigenous populations worldwide.4,5 Given the likely shared evolutionary path between different Indigenous populations, the involvement of biological factors in contributing to risk for lifestyle disease is possible. While it is recognised that reasons contributing to increased risk for lifestyle diseases in Indigenous populations are clearly multi-factorial (including obvious lifestyle and socio-economic factors), it remains surprising that the potential contribution of biological factors to disease risk in these populations has not been more extensively investigated.
Inflammatory events are increasingly accepted as contributing to the pathogenesis of a spectrum of interrelated classic lifestyle diseases, including T2DM, cardiovascular disease (CVD) and chronic kidney disease (CKD). Increased circulating levels of inflammatory cytokines, including IL-1β, TNF-α (both classic markers of inflammation) and IL-6 (a protein with a number of actions in the regulation of inflammation and metabolism), have been reported in individuals with T2DM compared with healthy controls.6–8 This is considered to be evidence of their possible role in disease pathogenesis. Increased concentrations of IL-69,10 and decreased concentrations of IL-10 (a key anti-inflammatory cytokine)11,12 have both been associated with an increased incidence of CVD in various clinical settings. Increased IL-6 concentrations 13 have also been associated with poorer prognosis and/or increased mortality in patients suffering CVD, further alluding to the possible role of inflammatory events in the mechanisms contributing to CVD and associated co-morbidities. Elevations in circulating concentrations of inflammatory markers have also been observed in patients with CKD.14,15 Collectively, such findings suggest involvement of inflammation in both the initiation and chronic progression of lifestyle diseases; therefore, the regulation of inflammatory mediators, including at the genetic level, is of particular interest and might act as a risk-modifying factor.
The potential contribution of single nucleotide polymorphisms (SNP), either in isolation or combination, on regulation of inflammation has been recognised previously by our group. The frequency of documented polymorphisms in genes encoding key pro- (IL-6, IL-1β) and anti-inflammatory (IL-10 and IL-1ra) cytokines was found to differ significantly between a cohort of Indigenous Australians compared with a Caucasian population of European descent.16–18 Other investigators have also reported differences in the distribution of documented SNPs in cytokine genes between (i) South African Zulu and Western European populations, 19 (ii) African American and Caucasian American populations,20,21 and (iii) Aboriginal Canadian and Caucasian populations. 22 These findings support the potential for common genetic traits within Indigenous populations worldwide.
Given the reported effects of gene polymorphisms on cytokine production, the genotype profile of Indigenous Australians (and other Indigenous populations) appears to favour a propensity for potent pro-inflammatory responses. While this might be beneficial in response to acute infection, failure to adequately regulate inflammatory responses following infection and/or inappropriate inflammation in response to non-infectious stimuli could result in dysregulation of these responses and contribute to risk for disease. 23 The potential for a heritable risk for excessive or dysregulated inflammation in Indigenous Australians formed part of the rationale for this project and might partly explain the increased rates of classic lifestyle diseases among Indigenous Australians. In this context, a series of cytokine genes was selected as a model for an initial examination of genetic variation in Indigenous Australians. The specific aims of the current study were to compare the frequency of selected cytokine gene polymorphisms in Indigenous and Caucasian controls, and to identify any additional (undocumented) genetic variants in selected cytokine genes.
Materials and methods
Sample material
The study adhered to the declaration of Helsinki. The study was undertaken with approval from local institutional ethics committees (University of Newcastle Human Research Ethics Committee and Hunter New England Health Research Ethics Committee). Written approval was also obtained from the traditional land owners of the community from which the Indigenous Australian samples were originally collected.
Genomic DNA previously extracted from whole blood samples collected from a community of isolated Indigenous Australians (
SNP genotyping
Commercially available real-time (RT) PCR allelic discrimination assays (PE Applied Biosystems, Foster City, CA, USA) were used for identification of the cytokine gene polymorphisms
All assays were undertaken in accordance with the manufacturer’s guidelines, with a total of 50 ng of DNA. Reactions were performed using an ABI Prism 7900HT Sequence Detection System (PE Applied Biosystems) under the following thermal cycling conditions: 40 cycles of 50℃ for 2 min, 95℃ for 10 min, 92℃ for 15 s and 60℃ for 1 min. Automatic classification of samples as homozygous (for either allele) or heterozygous was undertaken using the Sequence Detection Systems Software v2.3 for the 7900HT (PE Applied Biosystems).
DNA sequencing
Unidentified genetic variants in
Briefly, PCR primer pairs were designed using PerlPrimer v1.1.15, 24 and reaction conditions were optimised for each fragment. Fragment amplification was undertaken using a GeneAmp PCR System 9700 thermal cycler (Applied Biosystems), and standard denaturation and elongation conditions. For each fragment, PCR products were confirmed by visualisation on a 1.5% agarose gel stained with ethidium bromide. PCR products were prepared for use in respective sequencing reactions using an Agencourt Ampure PCR Purification System Kit (Beckman Coulter, Beverly, MA, USA) on a Biomek NXP Laboratory Automation Workstation (Beckman Coulter).
Cycle sequencing was completed using BigDye Terminator v3.1 Cycle Sequencing Kit (PE Applied Biosystems) and standard reaction conditions. After cycle sequencing, products were prepared for analysis using an Agencourt CleanSeQ Dye Remover Kit (Beckman Coulter) on a Biomek NXP Laboratory Automation Workstation (Beckman Coulter). Automated analysis of cycle sequencing products was undertaken with an ABI Prism 3730 DNA Analyser (PE Applied Biosystems) and sequence chromatographs saved for analysis of sequence variants.
On completion of automated sequence analysis, sequence chromatographs were visually inspected using Sequence Scanner Software v1.0 (Applied Biosystems) to confirm sequence quality. The sequence analysis software package Mutation Surveyor Version 3.23 (SoftGenetics, State College, PA, USA) was used to identify any sequence variants present in the samples relative to the GenBank reference sequence for
Data analysis
SNP genotyping data were analysed using the statistical software package Statistics/Data Analysis (STATA) Version 10.0 (Stata Corporation, College Station, TX, USA). A χ2 test or Fisher’s exact test, if appropriate, was used to assess the distribution of the SNPs. Statistical significance was accepted at
The frequency at which both previously documented and novel sequence variants were detected via DNA sequencing was recorded. Online tools were used to predict the potential effects of any novel variants on intron/exon splicing and transcription factor binding. ESE Finder v3.0 (Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, USA) was used to predict potential alterations in exonic splicing enhancer sites.25,26 FastSNP (Institute of Biomedical Sciences and Institute of Information Science, Taipei, Taiwan) 27 and TFSearch (Computational Biology Research Centre, National Institute of Advanced Industrial Science and Technology, Tsukuba, Japan) were used to predict potential alterations in transcription factor binding.
For each of the variants identified by sequencing, the genotype frequencies for the Northern and Western European ancestry population (CEU), the Yoruba Nigerian population (YRI) and the Han Chinese population of Beijing (CHB) were obtained from the NCBI SNP Database to act as reference populations. Distribution of reported genotype frequencies for each of the reference populations was compared using a χ2 test. Further comparisons were made with the genotype distributions noted in the sample of Indigenous Australians to determine if there were any similarities in the genotype frequencies between the groups. Statistical significance was accepted at
Results
SNP genotyping
Genotype frequencies for Indigenous Australians (
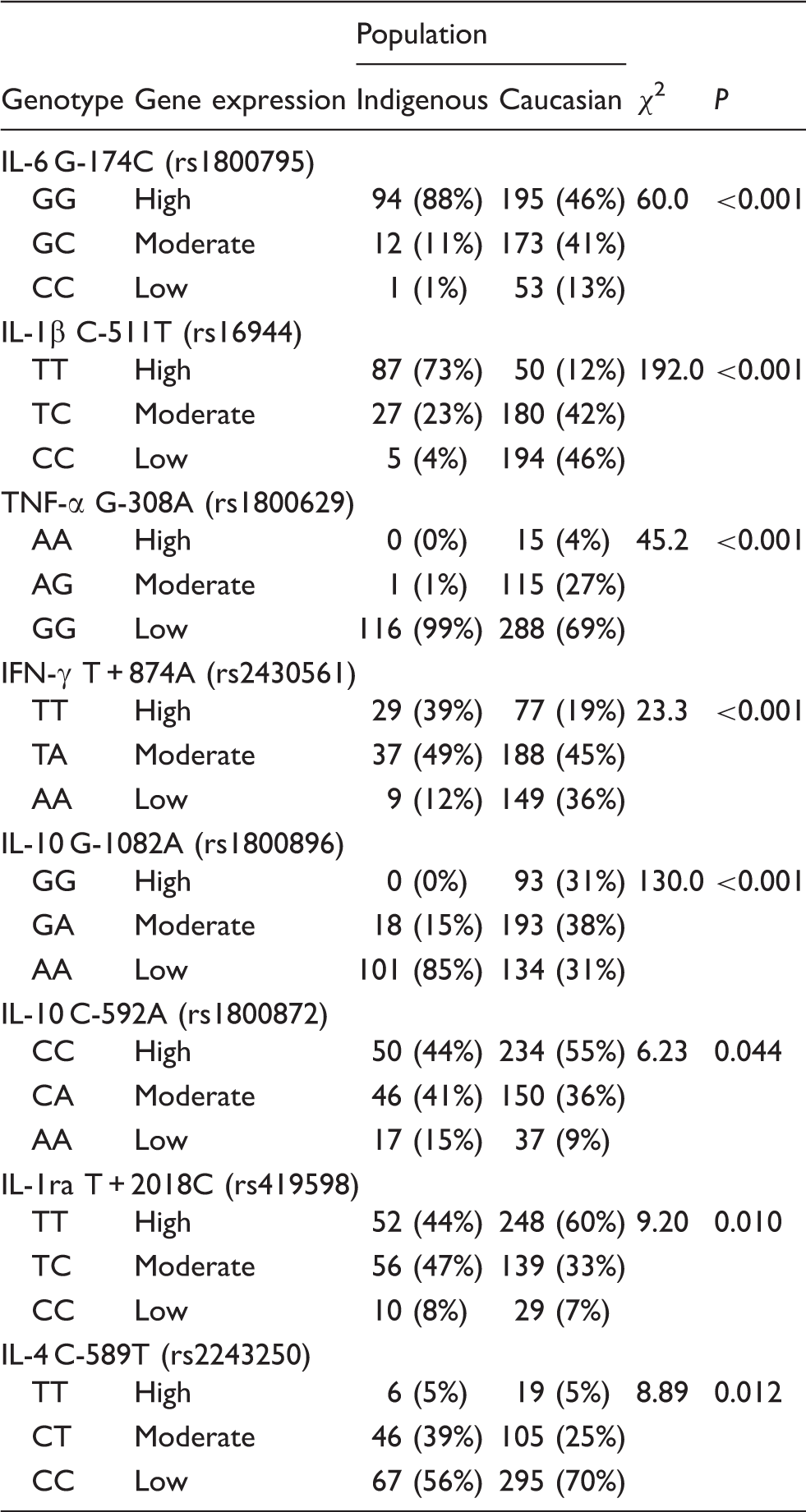

DNA sequencing
TNF-α and IL-1β
Only a small number of novel variants were identified for
IL-6
Genotype frequencies for documented and novel SNPs in the gene encoding IL-6. Predicted effects on splicing enhancer sites (SES) and transcription factor (TF) binding are indicated. For documented SNPs, HapMap frequencies from the CEPH population of CEU, YRI and CHB are also included.

n/a: not applicable.
IL-10
Genotype frequencies for documented and novel SNPs in the gene encoding IL-10. Predicted effects on splicing enhancer sites (SES) and transcription factor (TF) binding are indicated. For documented SNPs, HapMap frequencies from the CEPH population of CEU, YRI and CHB are also included.

n/a: not applicable.
Comparison of genotype frequencies with reference populations
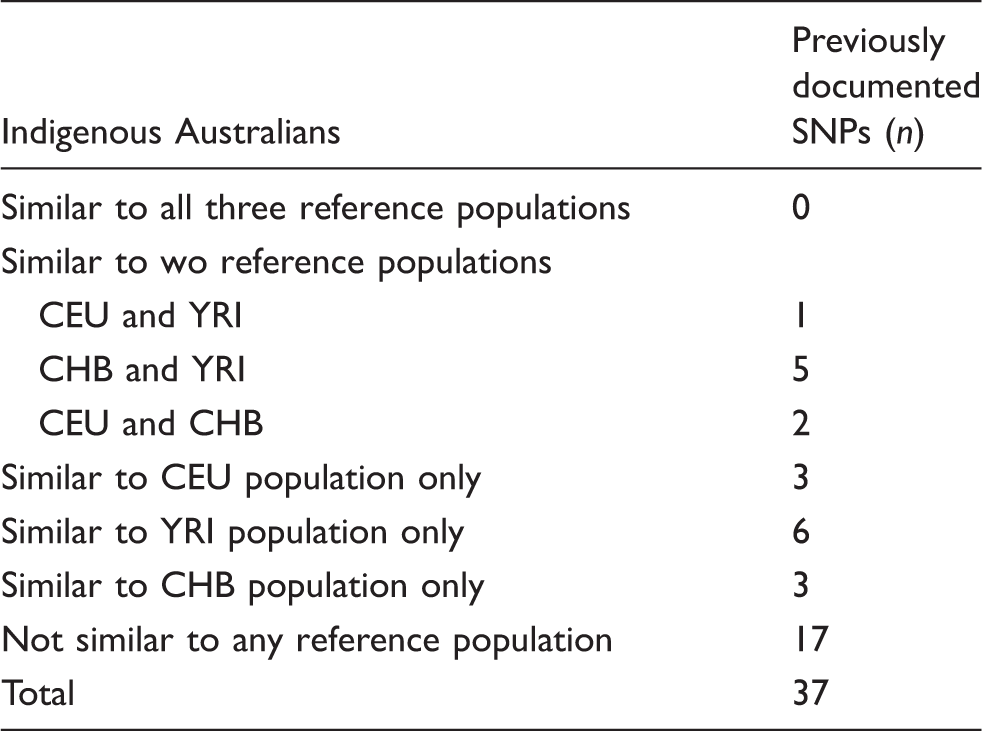
Numbers of previously documented SNPs also identified in Indigenous Australians where the distribution of genotype frequencies resemble one or more of the three reference populations: CEU, YRI and CHB.
Discussion
The current study was undertaken to investigate the potential regulation of cytokine genes in Indigenous Australians, particularly in the context of inflammatory control as a risk-modifying factor for classic lifestyle disease. Findings from the current study confirm genotype frequencies for a set of known cytokine gene polymorphisms are markedly different in Indigenous Australians than Caucasian controls. In addition, we were able to identify the presence of a number of additional, and previously unreported, sequence variants in several key cytokine genes using a subset of the Indigenous sample. Comparison of genotype frequencies for previously reported SNPs between Indigenous Australians and three HapMap populations (CEU, YRI and CHB) revealed unique patterns of distribution in the Indigenous Australians. Together, these findings support a need for further examination of genetic variation in Indigenous Australians.
The possibility that genetic evolution can influence risk for disease has been discussed previously. In a review by Tishkoff and Williams, 28 the prevalence of complex lifestyle diseases was highlighted in populations with an African ancestry living outside of Africa. While the role of a Western lifestyle cannot be ignored in the context of this discussion, the possibility that hereditary factors influence the type and magnitude of responses to common environmental risk factors must be similarly acknowledged. 28 A large number of studies have already examined possible associations between genetic variants and risk for lifestyle diseases (see reviews by Lango and Weedon 29 , Stolerman and Florez, 30 and Daley and Cargill 31 ). The relevance of any findings for ethnic minorities or Indigenous populations is uncertain, and the need to verify reported associations in specific population groups remains. Findings from the current study suggest a unique cytokine SNP profile in a sample of Indigenous Australians compared with Caucasian controls, which could contribute to risk for lifestyle disease in modern environments. The additional genetic variants identified in the four genes subjected to DNA sequencing are likely to be replicated across the genome, and such genetic variation could prove important in further understanding the differing health profiles between Indigenous and non-Indigenous populations. As such, the findings reported here provide compelling evidence supporting the need to examine further the extent of genetic variation in Indigenous Australians.
The DNA samples used in the current study were obtained from an isolated Indigenous community from central Australia. The traditional nature of the community was such that there had not been any significant admixture with the non-Indigenous population. In this respect, it was considered that this particular cohort was likely to reflect closely an ancestral genome and, as such, offer unique insight into the genetic architecture of Indigenous Australians. Comparison of the genotype frequencies presented here, with those listed for CEU, YRI and CHB populations, failed to reveal any clear patterns of similarity. It was initially anticipated that the genotype frequencies in the Indigenous Australians would more closely resemble the YRI population based on a shared genetic ancestry; however, the recent article from Rasmussen et al. 32 describing the complete sequencing of the Aboriginal genome would not support this assumption. While similarity to the YRI population was obvious for a number of the detected SNPs (16%), for an even greater proportion of SNPs (∼ 46%), the genotype frequencies for Indigenous Australians did not resemble any of the reference populations. A divergent evolutionary path in Indigenous Australians in response to the unique selective pressures of the Australian environment is one explanation for the unique genotype profiles observed. These observations support the need for additional targeted studies in specific minority populations to catalogue the extent of genetic variation relative to the human reference sequence and profile potential effects on disease risk in specific population groups.
Any functional effects of the novel variants detected in the current study were not investigated directly. Implications of these findings on protein function remains theoretical and based primarily on results from
The potential contribution of genetic variation on individual susceptibility to a particular disease is becoming an important focus of investigation. In the current study, the frequencies of a set of cytokine gene polymorphisms were found to be markedly different in Indigenous Australians compared with Caucasian controls. In addition, a series of cytokine genes was used as a model for assessing genetic variation in the context of risk for lifestyle disease in a subset of the available Indigenous sample. A number of additional variants were observed in the cytokine genes examined relative to the human reference sequence. This finding suggests that the selective pressures encountered by Indigenous populations might have contributed to the presence of additional genetic variants. Accordingly, the extent of genetic variation in Indigenous populations needs to be catalogued such that better informed assessment of the potential for a heritable risk for lifestyle (and other) diseases in these population groups can be made.
Footnotes
Funding
The project was supported by an Early Career Researcher Grant (to AJC) from the University of Newcastle (Newcastle, Australia).
Acknowledgements
We acknowledge the valuable contribution of the Indigenous participants over an extended period of time. Prior to commencement of the recent study phase, the research team engaged in a renewed process of consultation to inform community members of the research findings and to reaffirm consent for continued use of the blood samples for immune gene studies. We would also like to acknowledge the support and input of the late Dr June Roberts-Thomson in the initiation of this study. We gratefully acknowledge the assistance of Professor Barry Boettcher in corresponding with the Indigenous community elders. Buffy Coats for non-Indigenous Australians were obtained from the Australian Red Cross Blood Service.
