Abstract
TLRs are PRRs that play a pivotal role in sensing exogenous pathogens and endogenous danger signals. Their role in the pathogenesis of inflammatory and immune-related diseases is gradually being unravelled. TLR2 and TLR4 are capable of sensing the oral microbial community, which is considered a potential trigger for Behçet’s disease (BD). This study aimed to investigate the expression and function of TLR2 and TLR4 in the oral mucosa of BD. A total of 87 patients was included: 55 BD, 24 healthy controls and eight recurrent aphthous stomatitis. Total RNA was purified from non-lesional oral mucosal brush biopsies and analysed for the presence of TLR2 and TLR4 mRNA, along with their splice variants. The response of peripheral blood mononuclear cells to classical TLR2 and TLR4 agonists was also investigated. TLR2b, TLR2d, TLR2e, TLR4.3 and TLR4.4 were significantly elevated in relapsed BD. A significant defect in the response to cognate agonists of TLR1/2 heterodimer and TLR4 was also observed in BD. The expression of unusual splice variants of TLR2 and TLR4 might explain the observed defect in these receptors’ function in BD.
Introduction
Behçet’s disease (BD) is one of the vasculitides of unknown aetiology that severely affects quality of life.1,2 Its most serious consequences are blindness, gastrointestinal bleeding and large vessel disease. 3 An oral bacterial and viral aetiology has been suggested,4–7 and the evidence points towards both genetic and environmental factors as contributors to the disease process. 8 Hypersensitivity or immune cross-reactivity to certain microbial molecules have also been investigated in relation to BD. 9
TLRs are PRRs recognising PAMPs, such as LPS, peptidoglycans, human and microbial heat shock proteins (HSPs), and viral antigens.10–12 TLR2 and TLR4 jointly are capable of identifying most members of the oral microbiome. These receptors are mainly expressed on cells of innate immunity; however, their expression has also been demonstrated on epithelial and endothelial cells. 13
Activation of TLRs triggers a cascade of protein kinases leading to the nuclear translocation of NF-κB and transcription of cytokine-encoding genes.13,14 TLR2 mRNA has five different splice variants (GenBank: AF424051.1, AF424049.1, AF424053.1, AF424052.1 and AF424050.1), while TLR4 mRNA is known to have three splice variants [National Center for Biotechnology Information (NCBI) reference sequences: NM_138554.4, NP_003257.1 and NM_138557.2].15,16
There are no differences in the putative open reading frames of the different TLR2 mRNA splice variants; however, differences in the untranslated sequence might influence mRNA stability and protein translation. 15 Furthermore, the presence of an early stop codon in TLR4 mRNA variants 3 and 4 makes them subject to nonsense decay mechanisms preventing the production of truncated protein (NCBI reference sequences: NP_003257.1 and NM_138557.2).16,17 Therefore, a defect in the nonsense decay mechanisms would be expected in cases of aberrant production of these truncated proteins. 17 As downstream signalling through the TLRs leads to cytokine induction it is also feasible that inappropriate signalling through faulty TLRs or the absence of properly functioning soluble receptors might prolong signalling, which—under normal circumstances—would be subject to negative feedback inhibition.18,19
Single-nucleotide polymorphism (SNP) analysis of TLR2 and TLR4 genes was not significantly associated with BD in Caucasian, Turkish or Tunisian cohorts.20–23 However, one SNP in TLR4 was associated with BD, and six others had effects on clinical features in a Japanese cohort. 24 Likewise, TLR4 haplotype TAGCGGTAA was considered a susceptibility gene for HLA-B51-positive BD and arthritis in a Korean cohort. 25
Investigation of TLR expression in BD PBMCs has demonstrated an elevated expression of TLR4 (mRNA and protein), but not TLR2, irrespective of disease activity, 26 and elevated TLR2 and TLR4 mRNA in active BD. 27 The expression of mRNA and protein of both receptors is suppressed by vitamin D, 28 and TLR2 is suppressed by HSP60 and LPS. 28 The total expression of TLR2 and TLR4 was also found to be elevated in broncho-alveolar lavage (BAL), and intestinal and buccal mucosa of BD.20,29,30 NOD2 expression showed correlation with TLR2 and TLR4 in BAL samples from BD patients. 30 However, to date, no study has investigated the differential expression and function of TLR2 and TLR4 splice variants in the oral mucosa of BD.
Materials and methods
Patient selection and ethical approval
All patients were diagnosed according to the International Study Group criteria, and informed consent was obtained. 31 The patient cohort was recruited from the Royal London Hospital and St. Thomas’ Hospital with appropriate local ethical approval. BD patients were stratified according to the overall disease activity, following a consultant-led assessment, into the following groups: (1) relapsed status (severe disease activity in more than two of the BD symptoms despite treatment); (2) minor activity status (disease activity in fewer than three of the BD symptoms); (3) inactive status (the absence of any symptoms at the time of sampling); (4) non-relapsed (minor activity BD group + inactive BD group). Healthy controls (HC) were recruited from laboratory staff with informed consent under the local ethically-approved study guidelines.
Samples
A total of 87 buccal mucosal brush biopsies (BMBB) from non-lesional oral mucosa from 55 BD (13 relapsed BD, 24 minor activity BD, 18 inactive BD) [female/male (F/M): 33/22, mean age (yrs): 43.3 ± 13], 24 HC [F/M: 14/10, mean age (yrs): 39.7 ± 13.8] and eight recurrent aphthous stomatitis (RAS) [F/M: 5/3, mean age (yrs): 46.5 ± 7] were collected using cytology brush (Flowgen, Nottingham, UK). 32 Fifty percent of RAS presented with oral ulcerations at the time of sampling. Total RNA and protein were purified simultaneously from the same BMBB samples according to the manufacturer’s protocol (Purescript-Gentra; Qiagen, Manchester, UK).
Expression of TLR2 and TLR4 mRNA by real-time PCR
Total RNA was transcribed to c-DNA using a high-capacity c-DNA reverse transcription kit according to the manufacturer’s instructions (Applied Biosystems, Warrington, UK). All samples were tested in a real-time (q) PCR experiment using TaqMan probes for TLR2 mRNA [probe for the common 5′ untranslated region (UTR) region of the five transcripts on the 2-3 exon boundary], TLR4 mRNA (probe for the common 5′UTR region of the three transcripts on the 1–2 exon boundary) (Applied Biosystems). The 18S rRNA acted as housekeeping gene for the relative expression analysis. Thirty-four positive samples (10 relapsed BD, 12 non-relapsed BD and 12 HC) were further investigated for expression of splice variants.
Expression of splice variants of TLR2 and TLR4 mRNA by qPCR
qPCR primers for GAPDH, TLR2 and TLR4 splice variants.

TLR protein expression: Slot-blot analysis
A total of 42 protein samples (14 relapsed BD, 14 non-relapsed BD and 14 HC) were quantified using the NanoDrop spectrophotometer (Labtech, Uckfield, East Sussex, UK). The Bio-Dot SF (Bio-Rad, Hertfordshire, UK) apparatus was assembled and used according to the manufacturer’s instructions. Purified protein in SDS buffer (25 µg/200 μl) was allowed to filter through the nitrocellulose membrane under vacuum. The membrane was washed with TBS and then incubated in blocking buffer, prepared according to standard protocols, for 1 h at 18–20℃ [room temperature (RT)]. The membranes were probed with specific Abs to TLR2 and TLR4 (TLR2 at 1 : 500 and TLR4 at 1 : 1000; eBioscience, Hatfield, Herts, UK and R&D Systems, Abingdon, Oxon, UK respectively) for 16–18 h (overnight) at 4℃. The membrane was then washed using 0.32 M TRIS-HCl pH 8.0 (1% v/v), 5 M sodium chloride (3% v/v) and 0.05% Tween 20 (wash buffer) for 60 min. Bound Ab was detected using HRP-conjugated anti-species Ab at 1 : 1000 for 1 h at RT. The membrane was developed using chemiluminescence (ECL plus, BioRad, Hemel Hempstead, UK), and the signal was quantified using densitometry analysis and expressed as relative expression compared with the negative control.
Western blot analysis
A total of 30 protein samples from the cohort investigated by slot-blot analysis (eight relapsed BD, 12 non-relapsed BD and 10 HC) was prepared in reducing buffer (1 µg/µl) and denatured at 100℃ for 5 min. Proteins were separated by SDS-PAGE in a 4–20% linear gradient ready cast gel (Invitrogen, Paisley, UK) at 150 V for 2 h. Separated protein samples were transferred to a polyvinylidene difluoride membrane (Immobilon membrane; Sigma-Aldrich, Poole, Dorset, UK) at 30 V for 1 h at 4℃.33,34 Proteins were blocked and probed with Ab as described above.
TLR functional study
A total of 46 blood samples (10 relapsed BD, 21 non-relapsed BD and 15 healthy individuals) was analysed. PBMC were isolated by density gradient centrifugation over Ficoll-Hypaque (1180 g for 35 min). Isolated cells were incubated at 37℃ overnight at 2 × 105 cells/ well. They were treated with specific TLR agonists: 0.1 µg Pam3CSK4 (TLR1/2 heterodimer), 108 heat killed preparation of Listeria monocytogenes (HKLM) (TLR2), 0.2 µg Escherichia coli K-12 LPS (TLR4) and 1 µg FSL-1 (TLR2/6 heterodimer; InvivoGen, San Diego, USA) for 16 h at 37℃. Phytohaemagglutinin (PHA) acted as a positive control; the negative control was unstimulated cells. The level of TNF-α in the culture supernatant was quantified by ELISA (R&D Systems). The stimulation index for each treatment was calculated by dividing the level of TNF-α produced after treatment by the spontaneous TNF-α secretion.
Statistical analysis
Pilot experiments were carried out and the results subjected to statistical power calculations in order to establish the numbers of samples required for robust statistical analysis (StatMate 2; GraphPad, La Jolla, CA, USA). Based on the average of the SD (SD average = 0.002207), a sample size of eight in each group has a 95% power to detect a difference between means of 0.0043 with a significance level (alpha) of 0.05 (two tailed). However, the maximum sample size achievable using the available resources was investigated.
The results were analysed by Mann–Whitney U-test when comparing quantitative data of two groups, and the Kruskal–Wallis test when comparing more than two groups. Differences in the rate of expression of TLR2 and TLR4 were analysed using χ2 tests. Correlation studies were analysed using Spearman test (Prism; GraphPad).
Results
Characteristics of participants
Clinical characteristics of patients with BD.
mRNA expression of TLR2 and TLR4 is elevated in the oral mucosa of relapsed BD patients
There was a significant increase in BMBB expression of TLR2 mRNA in relapsed BD compared with HC (P = 0.0129). Similarly, there was an increase in BMBB expression of TLR2 mRNA in relapsed compared with inactive BD (P = 0.0260), relapsed compared with minor activity BD (P = 0.0396) and relapsed compared with non-relapsed BD (P = 0.0155). Likewise, there was an increase in the expression of TLR2 in relapsed BD compared to RAS (P = 0.0214). However, when HC were compared with the full cohort of BD (relapsed and non-relapsed) there was no significant difference (P = 0.3107). RAS expressed higher levels of TLR2 in comparison with HC; however, the difference was not statistically significant (Figure 1a).
Expression of TLR2 and TLR4 mRNA in oral mucosal brush biopsies. Data presented as mean and SEM. (a) The relative expression of TLR2 and TLR4 mRNA (BD: n = 55; relapsed BD: n = 13; non-relapsed BD: n = 42; inactive BD: n = 18; minor activity BD: n = 24; HC: n = 24; RAS: n = 8). *Comparison between relapsed BD and all the other groups was statistically significant (P < 0.05). **Comparison between relapsed BD and all the other groups was statistically significant (P < 0.01). (b) The relative expression of TLR2 splice variants a–e and TLR4 splice variants 1, 3 and 4 (BD: n = 22; relapsed BD: n = 10; non-relapsed BD: n = 12; HC: n = 12). *Comparison between relapsed BD and HC was statistically significant (P < 0.05). ***Comparison between relapsed BD and all the other groups was statistically significant (P < 0.001).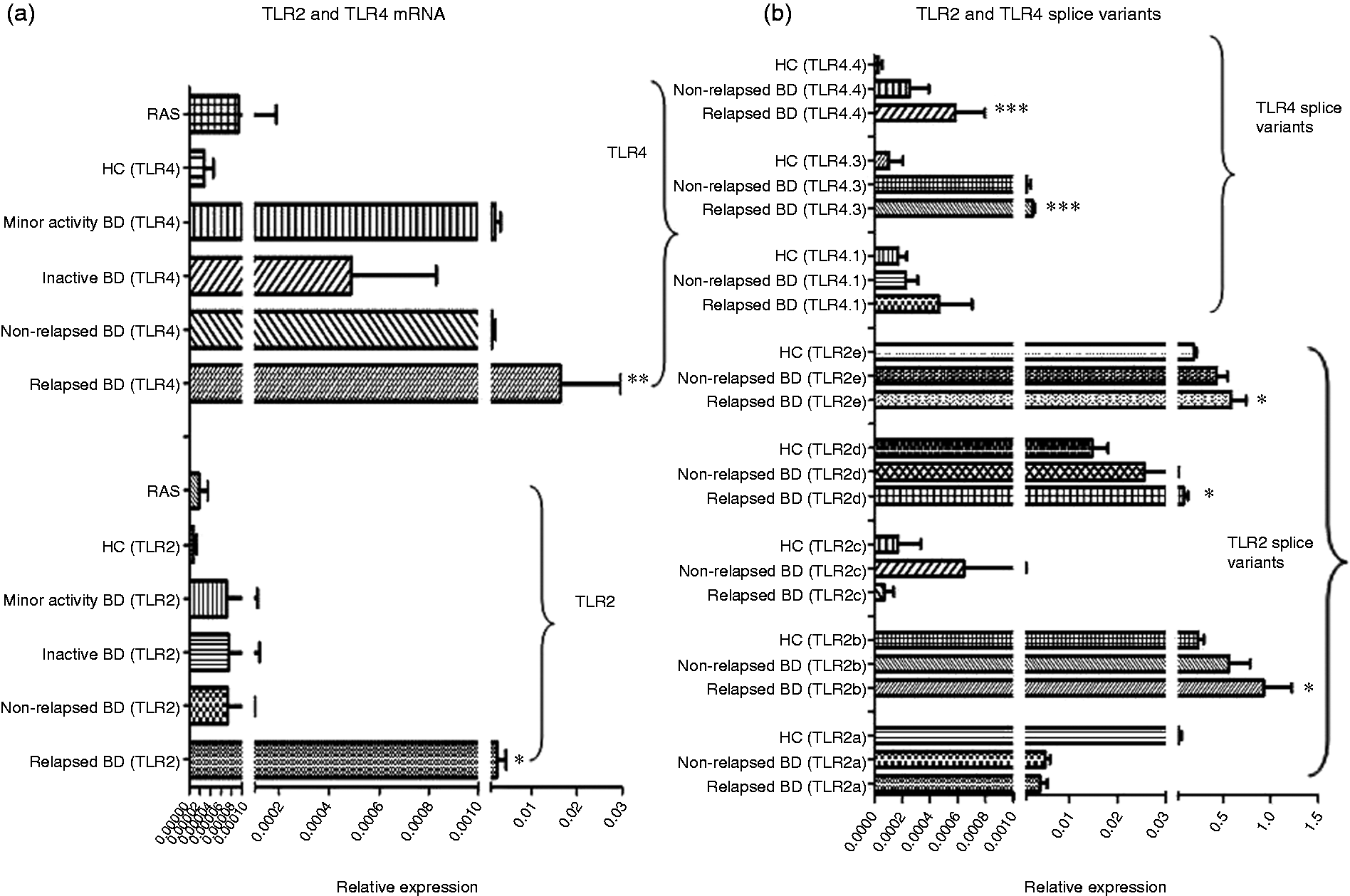
The expression of TLR4 (Figure 1a) showed a similar pattern, with a significant increase in expression of relapsed BD versus HC (P = 0.0019), relapsed compared with inactive BD (P = 0.0041), relapsed compared with minor activity BD (P = 0.0256), relapsed compared with non-relapsed BD (P = 0.0043) and relapsed BD compared to RAS (P = 0.0024). There was no significant difference in the expression of the HC and the whole of the BD cohort for TLR4 (P = 0.3107). RAS also expressed higher levels of TLR4 in comparison to HC; however, this was not statistically significant. The level of expression of TLR4 appeared higher than TLR2, but, again, was not significant (Figure 1a).
Examining the ocular activity showed that ocular active BD expressed statistically higher levels of TLR2 and TLR4 in their BMBB than ocular inactive BD (P = 0.0389 and P = 0.0497 respectively). However, the number of the ocular active patients was very small (n = 4). Furthermore, orally-active BD expressed higher levels of TLR2 and TLR4 in their BMBB in comparison with orally inactive BD, but the differences were not statistically significant. The other systemic activities (skin, genital, rheumatological, etc.) and treatment protocol did not significantly affect the expression of TLR2 and TLR4 mRNA (data not shown).
TLR splice variants show differential mRNA expression in BMBB of BD patients compared with HC
TLR2a and TLR2c expression was broadly similar in all groups. The expression levels of TLR2b, TLR2d and TLR2e were higher in relapsed BD than HC (P = 0.032, P = 0.0272 and P = 0.0134 respectively). There was also a trend for non-relapsed BD to have higher levels of expression of these splice variants compared with HC, but this observed trend was not statistically significant (P = 0.2366, P = 0.6236 and P = 0.0690 respectively) (Figure 1b).
The relative expression of TLR4 variant 1, 3 and 4 mRNA was also measured in the same cohorts. All investigated groups had broadly similar levels of TLR4.1 expression. Relapsed BD expressed higher levels of TLR4.3 compared with non-relapsed BD, and HC (P = 0.0127 and P = 0.0002 respectively). Likewise, relapsed BD expressed higher levels of TLR4.4 compared with non-relapsed BD and HC (P = 0.0373 and P = 0.0127 respectively) (Figure 1b).
The rate of positivity was high for relapsed BD compared with all the other groups [TLR2: 76.9% (10/13) and TLR4: 100% (13/13)] (Figure 2a). There were no statistical significant differences in positivity rate between HC, non-relapsed BD, minor activity BD, inactive BD and RAS (Figure 2a). TLR2 variant c was low in all the investigated groups (Figure 2b). Expression levels of TLR4 splice variants 3 and 4 were infrequent in HC (1/12: 8.3%) and non-relapsed BD [3/12 (25%) and 4/12 (33.3%) respectively] compared with relapsed BD (9/10, 90%) (Figure 2c).
Positivity rate of TLR2 and TLR4 mRNA in oral mucosal brush biopsies. (a) TLR2 and TLR4 mRNA positivity rate in the non-lesional oral mucosa (BD: n = 55; relapsed BD: n = 13; non-relapsed BD: n = 42; inactive BD: n = 18; minor activity BD: n = 24; HC: n = 24; RAS = 8). (b) TLR2 mRNA splice variants positivity rate (BD: n = 22; relapsed BD: n = 10; non-relapsed BD: n = 12; HC: n = 12). (c) TLR4 mRNA splice variants positivity rate (BD: n = 22; relapsed BD: n = 10; non-relapsed BD: n = 12; HC: n = 12).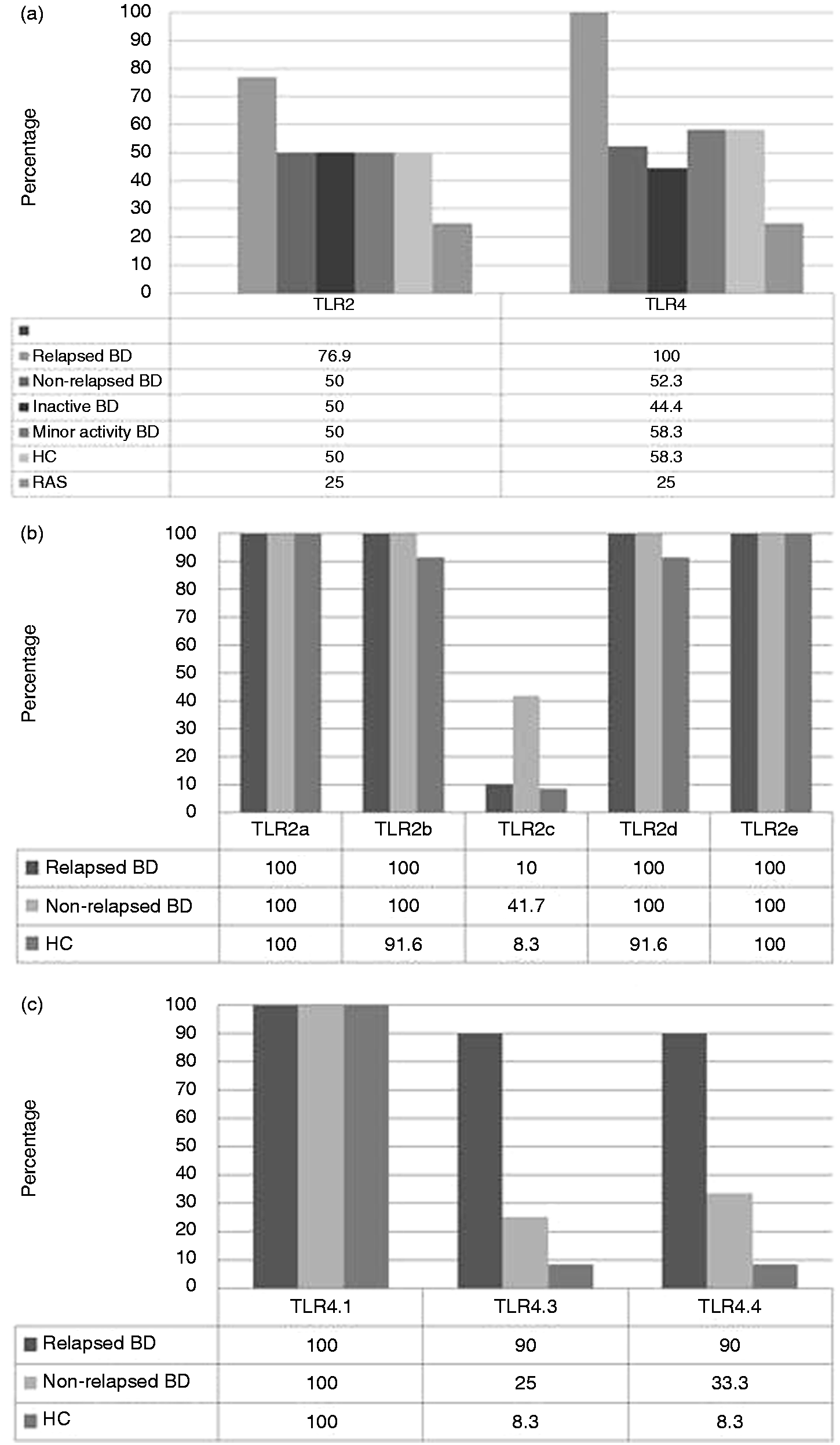
Descriptive statistical analysis including mean, median, SD, SEM, and percentages of TLR2 and TLR4 mRNA and their splice variants is presented in Supplementary Table S1.
TLR2 and TLR4 mRNA correlation study
There was positive correlation between TLR2b and TLR2e in all the investigated groups (relapsed BD: P = 0.0001 and r = 0.9758; non-relapsed BD: P = 0.0139 and r = 0.6853; HC: P = 0.0004 and r = 0.8531). Additionally, TLR2b positively correlated with TLR2d in non-relapsed BD (P = 0.0220 and r = 0.6503), and TLR4.1 positively correlated TLR4.4 in relapsed BD (P = 0.0330 and r = 0.6727) (Supplementary Figure S1).
Total protein for TLR2 and TLR4 are similar in BD and HC
There was no significant difference in the expression of the TLR2 protein between non-relapsed BD, relapsed BD and HC (P = 0.2937), or between the whole BD cohort (relapsed and non-relapsed) and HC (P = 0.3158). Similarly, there was no statistically significant difference in the TLR4 protein between relapsed BD, non-relapsed BD and HC (P = 0.4673), or between the whole BD cohort and HC (P = 0.4329) (Figure 3a).
Immuno-blot analyses of the expression of TLR2 and TLR4 proteins. (a) Slot-blot analysis of total TLR2 and TLR4. Data presented as mean and SEM. (b) Characterisation of TLR2 and TLR4 molecular forms by Western blot (WB).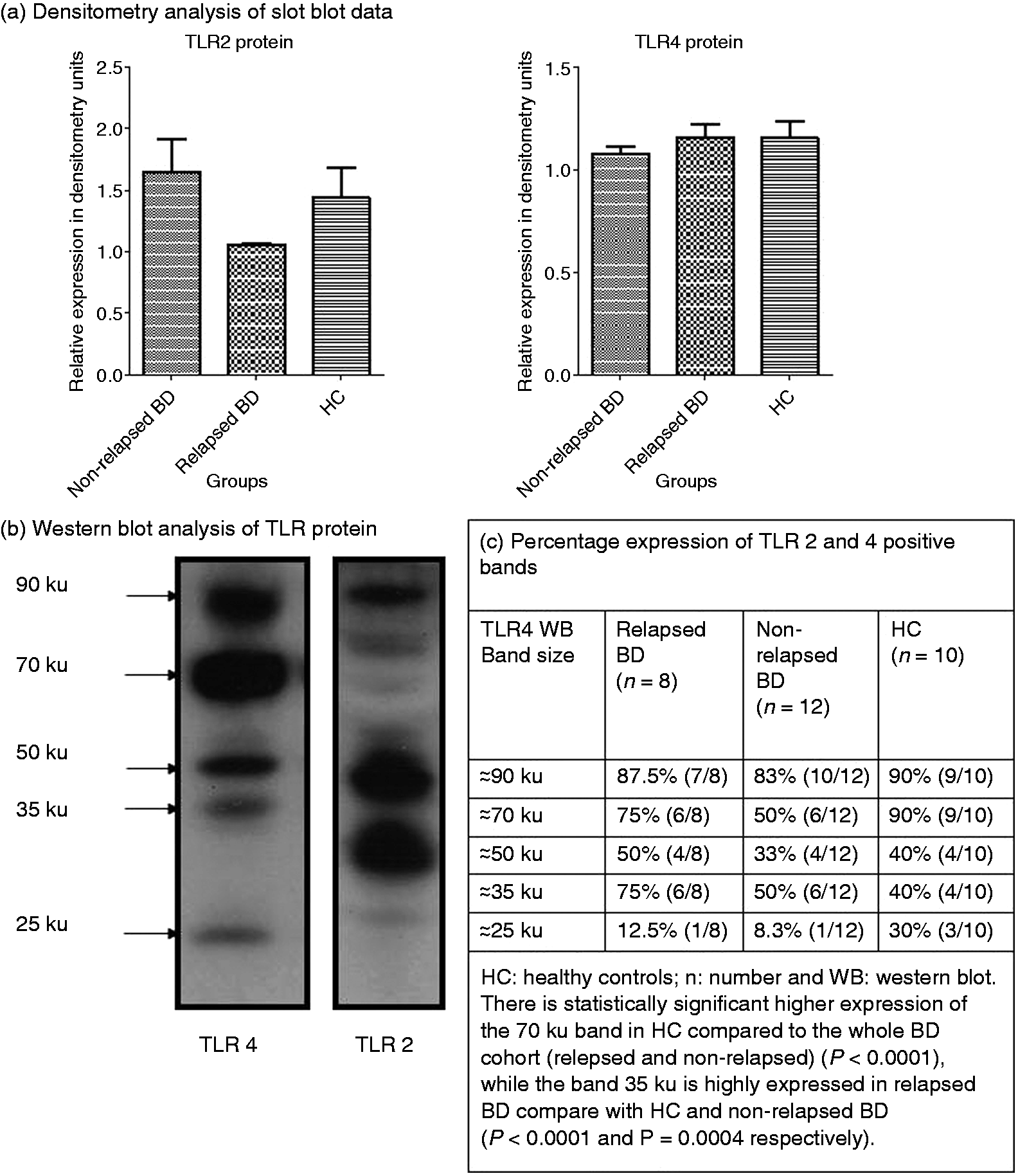
TLR2 and TLR4 show heterogeneity in Western-blot analysis
Western blot analysis of protein lysates showed heterogeneous bands with 90% of HC showing a band at ∼ 90 ku (Figure 3b). The 70-ku band was also strong in 90% of HC, while the 50-ku band appeared in 50% of relapsed BD. Additionally, 75% of relapsed BD had a band of 35 ku and 12.5% had a band of 25 ku. There was significantly higher expression of the 70-ku band in HC in comparison with whole BD cohort (relapsed and non-relapsed) (Figure 3c: P < 0.0001), while the 35-ku band was expressed to a significant level in relapsed BD compared with HC and non-relapsed BD (P < 0.0001 and P = 0.0004 respectively) (Figure 3c). TLR2 protein expression also appeared to be somewhat heterogeneous despite a constant amount of protein loading. Three significant bands were observed for TLR2 at about 90 ku, 50 ku and 30 ku (Figure 3c). These observations suggest that either incomplete sequences of the TLR proteins are produced or that intracellular degradation of the TLR had occurred.
TLR2 and TLR4 function is dampened in BD
The spontaneous TNF-α secretion was elevated in relapsed BD PBMC supernatant; however, this difference was not statistically significant (Figure 4a). All three groups (relapsed BD, non-relapsed BD and HC) responded equally to PHA treatment. After treatment with the different TLRs agonists, PBMC from the whole BD cohort (relapsed and non-relapsed) failed to respond to TLR4 and TLR1/2 agonists efficiently compared with HC (P = 0.0093 and P = 0.0500 respectively) (Figure 4b).
TLR2 and TLR4 functional study. Data presented as mean and SEM. (a) Spontaneous TNF-α secretion by PBMC. There is a trend of increased spontaneous TNF-α secretion by relapsed BD PBMC; however, the difference is not statistically significant. (b) TNF-α secretion upon stimulation with PHA and TLR agonists. There is a statistically significant defect in the function of TLR4 and TLR2/1 heterodimers.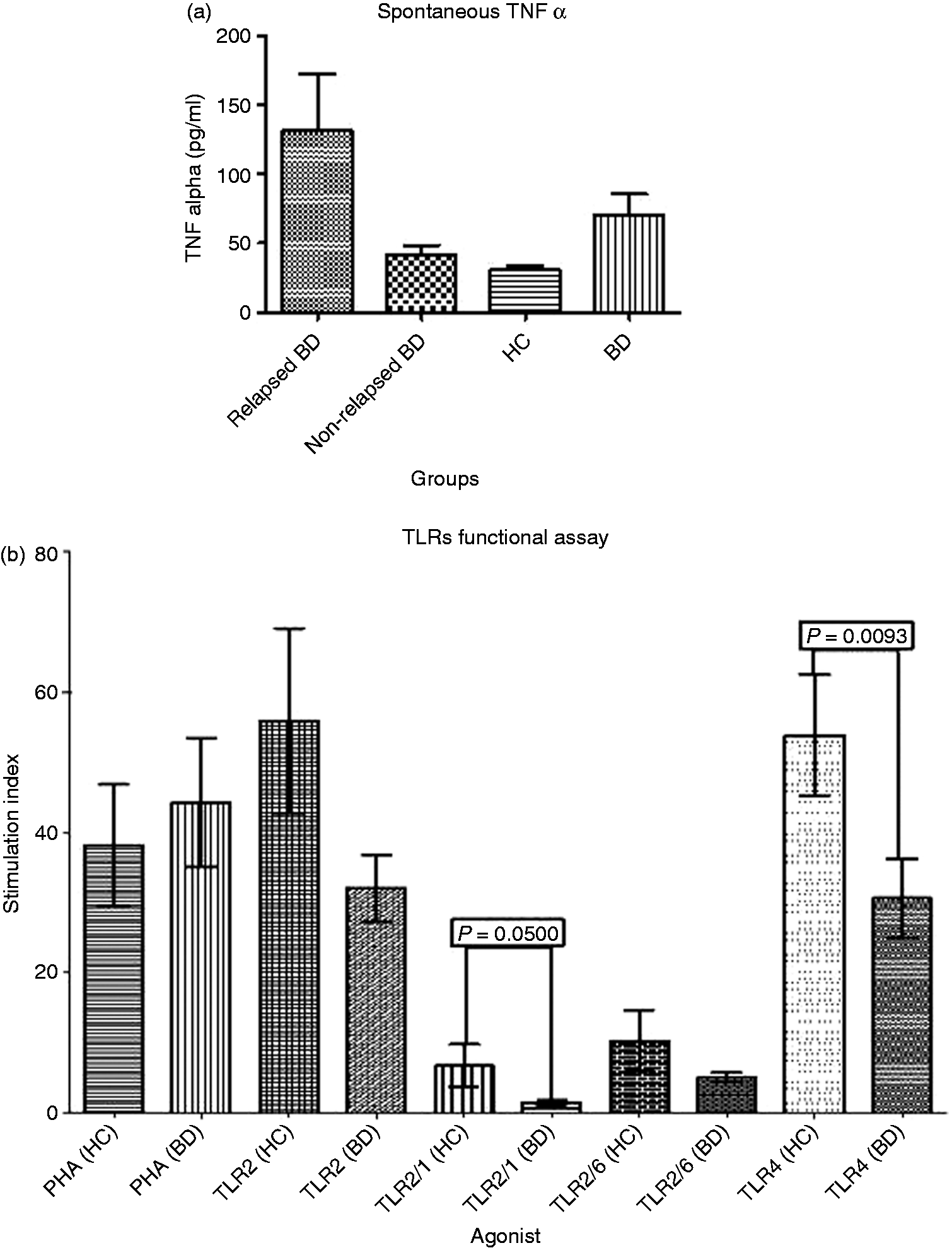
Discussion
Enhanced inflammatory reactions are a recognised characteristic pathological feature of BD exemplified by high serum pro-inflammatory cytokine levels. 35 Exogenous and endogenous antigens binding to TLRs induce a signalling cascade, which results in the nuclear translocation of NF-κB and transcription of pro-inflammatory cytokine-encoding genes. 12 Evidence from genome wide association study (GWAS) and SNP study suggest that elements of the innate immune system might be compromised in BD.24,25,30,36–38 Furthermore, an oral bacterial and viral aetiology was suggested in the literature.4–7 Interestingly, TLR2 and TLR4 are capable of sensing most of the oral microbiome. Therefore, the aim of the study was to investigate the differential expression of TLR2, TLR4 and their splice variants in relation to BD patients’ oral mucosa.
The oral mucosa is considered an important site of BD disease activity as most patients experience recurrent episodes of oral ulceration, and more than 80% of patients will present first with oral ulcers. 39 There is evidence that dental and periodontal therapies are associated with initial oral flare-up, but help in controlling the number and frequency of these flare-ups in the long term. 40 Furthermore, there are clinical observations suggesting that good control of the oral manifestations of the disease ameliorates the systemic symptoms (unpublished data). Homeostatic relationships between commensals and host responses have become an important area of study in a variety of disease conditions, including autoimmune encephalomyelitis, rheumatic arthritis and inflammatory bowel diseases.41,42 It is therefore important to understand the ability of BD patients to maintain their oral homeostasis.
We demonstrated that TLR2 and TLR4 expression increases in relapsed BD BMBB, mirroring the results of a previous study, which indicated elevated expression of TLR2 and TLR4 in BD PBMC, 27 and highlighting the importance of these markers in monitoring the disease activity. Given the higher levels of TLR2 and TLR4 in relapsed disease, one of their cognate ligands may play an important role in the pathogenesis of BD. However, it is also possible that multiple antigens contribute to the activity of the disease with some of them acting as shared ligands for both TLR2 and TLR4.
This study is the first to investigate the relative expression of the different splice variants of TLR2 and TLR4 in BD. These splice variants have different capacities for cell surface expression. TLR2b is known to transcribe a full-length protein. Variants c, d and e are thought to be unstable mRNAs and may not be translated completely. Interestingly, TLR variants b, d and e were highly expressed in relapsed BD. The elevated TLR2e is of great importance, as this may reflect the monocyte infiltration in the non-lesional oral mucosa with the possibility that a certain endogenous and/or exogenous antigen challenges the mucosa during the relapsed phase of the disease.
The up-regulation of the unusual mRNA splice variants of TLR4 may lead to incomplete protein translation. TLR4.3 was significantly elevated in relapsed BD; this splice variant has an extra exon (compared to splice variant 1), which results in a frameshift and an early stop codon. The translated protein from this variant is expected to be significantly truncated and a candidate for nonsense decay. Interestingly, TLR4.4 was also elevated in relapsed BD and lacks an exon (compared with TLR4.1), resulting in a frame-shift and an early stop codon. The exact protein product and function of these splice variants is an inviting point for further research. The truncated proteins might be unable to respond to negative feedback inhibition, resulting in prolonged induction of pro-inflammatory cytokines.
Interestingly, the correlation study pointed toward a positive correlation between TLR2b (known to translate to full length protein) and TLR2e (known to be expressed by fresh monocytes) in all investigated groups. It is documented in the literature that TLR activation can induce chemokine production and leukocyte recruitment. 43 However, contradictory observations were also documented pointing towards inhibition of leukocyte chemotaxis following TLR activation by high concentrations of their agonists. 44
The functional study was carried out to assess the ability of the TLRs to respond to their classical cognate ligands. The dampened response of BD PBMC TLR4 upon stimulation with its agonist is one of the novelties of this study, which could be explained by the higher levels of TLR4 variants 3 and 4 in relapsed BD. Also, the decrease in the BD PBMC response upon treatment with TLR1/2 agonist might highlight a defect in the heterodimer formation between these two receptors. Heterodimer formation between TLR1 and TLR2, and between TLR2 and TLR6 significantly increase the repertoire of these PRRs, and the formation of faulty heterodimers may lead to insights into their functions. 45 Inappropriate signalling through faulty TLRs might also lead to prolong signalling, which, under normal circumstances, would be subject to negative feedback inhibition. This might further explain the over-production of the pro-inflammatory cytokines characteristic of BD.
Examination of our slot-blot data suggested that protein expression of TLR2 and TLR4 was not raised in relapsed BD compared with inactive BD. This was discordant with the elevated levels of TLR2 and 4 mRNA. It is not unprecedented for protein and mRNA levels of the same gene/protein to show a poor correlation in the same tissue. 46 However, this may reflect a defect and the translation of unusual splice variants.
In Western blot analysis, we consistently observed multiple bands, despite the use of protease inhibitor cocktail tablets (Roche Applied Science, Burgess Hill, Sussex, UK) to remove the potential for ex vivo protein degradation. Jaresova et al. 16 reported the expression of four TLR4 splice variants in human APC. Moreover, Koch et al. 47 demonstrated the presence of about five bands at 130, 110, 72, 50 and 26 ku by Western blot. The band at 130 ku is a glycosylated TLR4 isoform, expressed at the cell surface, and the band at 110 ku is a partially glycosylated protein, unable to translocate to the surface. However, the bands at lower molecular masses may represent translation products of the TLR4 splice variants which contain early stop codons.47,48 These splice variants should be a candidate for the nonsense decay mechanism, preventing them from translation into truncated protein. The abundance of a 35-ku band of TLR4 in relapsed BD patients may indicate either a defect in the nonsense decay mechanism or enhanced proteolysis in this cohort.
We conclude that expression of TLR2 and TLR4 mRNA was elevated in relapsed BD and, furthermore, that a shift toward expression of unusual splice variants might explain the observed defect in TLR2/1 heterodimer and TLR4 response to their cognate ligands indicating functional abnormalities in the innate immune response to oral microbes.
The exact protein product of the TLR2 and TLR4 mRNA splice variants is not completely understood. The correlation between the different TLR2 and TLR4 splice variants and the bands observed in Western blot was not possible in matched samples. This, in turn, invites further research to investigate the function of these splices in health and disease.
Footnotes
Acknowledgements
We are grateful for Dr Duncan Clark, Dr Graham Wallace and Professor Judith Breuer for reviewing the initial manuscript and highlighting areas requiring further work.
Conflict of interest
The authors do not have any potential conflicts of interest to declare.
Funding
This work was funded by the Research Centre for Clinical and Diagnostic Oral Sciences. Institute of Dentistry, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, London, UK.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
