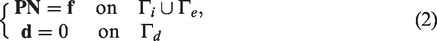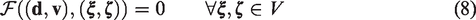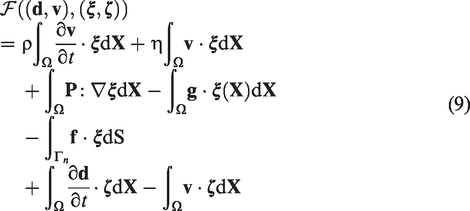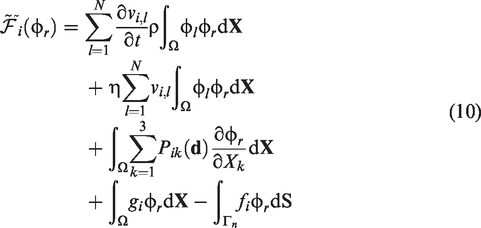Abstract
The blood vessels play a key role in the human circulatory system. As a tremendous amount of efforts have been devoted to develop mathematical models for investigating the elastic behaviors of human blood vessels, high performance numerical algorithms aiming at solving these models have attracted attention. In this work, we present an efficient finite element solver for an elastodynamic model which is commonly used for simulating soft tissues under external pressure loadings. In particular, the elastic material is assumed to satisfy the Saint–Venant–Kirchhoff law, the governing equation is spatially discretized by a finite element method, and a fully implicit backward difference method is used for the temporal discretization. The resulting nonlinear system is then solved by a Newton–Krylov–Schwarz method. It is the first time to apply the Newton–Krylov–Schwarz method to the Saint–Venant–Kirchhoff model for a patient-specific blood vessel. Numerical tests verify the efficiency of the proposed method and demonstrate its capability for bioengineering applications.
Introduction
In human circulatory system, the blood vessel network plays a key role that carries the blood from the heart to tissues over the entire body. Understanding the properties of blood vessels is essential to understand the health and disease of the vascular system, which in turn is necessary to patient evaluation and conduct therapeutic solutions in many clinical events, including surgery, angioplasty, tissue remodeling, and engineering. In particular, it has been recognized that the elastic motion of the vessel wall has significant effects on the hemodynamic quantities of interest. As such, the mechanical property of the blood vessel keeps being interested in many research areas. By this far, there have accumulated numerous studies on this topic, such as the stiffness, compliance, vessel diameter, and the internal and/or external pressures it may be subjected to.
During the past several decades, considerable efforts have been devoted to apply computational techniques to investigate the mechanical characteristics of blood vessels based on the continuum elasticity theory.1–4 On the other hand, as the underlying mathematical models have kept constantly evolving to more sophisticated levels and the problem size become unprecedentedly large, simulations on the vascular elasticity models have become very expansive tasks. Due to these reasons, developing high performance parallel numerical algorithms for blood vessel simulations has attracted more and more attention, see Quarteroni et al., 5 Takizawa et al., 6 Crosetto et al., 7 Balzani et al., 8 and the references therein. In this work, we present an efficient parallel numerical solver aiming at solving nonlinear partial differential equations raised in blood vessel deformational dynamics. More specifically, the mathematical model is built on the continuum hyperelasticity theory, and a nonlinear solver is proposed based on the finite element discretization, an implicit time stepping, and the Newton–Krylov–Schwarz method. 9 In this design, at each time step, we find the solution by successively solving the linearized discretized system. To achieve that, a Krylov subspace method is invoked at each Newton step. For improving the convergence and ensuring the scalability of whole method, a restricted additive Schwarz preconditioner 10 is used. Finally, an efficient program package is implemented based on the proposed algorithm, which is capable of carrying out blood vessel simulations. In the numerical experiments, we illustrate a patient-specific blood vessel under external surface loadings. These results not only demonstrate the efficiency and capability of our solver, but also provide a basic module for more sophisticated cardiac biomechanical applications.
The rest of the paper is organized as follows: the mathematical model and the discretization scheme are described in “Modeling” section. In “Methodology” section, we introduce the Newton–Krylov–Schwarz method for solving the induced nonlinear system. Finally, numerical experiments are demonstrated in “Numerical experiments” section for simulating a human blood vessel with patient-specific shape. The scalability results show the potential of the proposed method to efficiently solve large size problems with many processors.
Modeling
We consider a nonlinear elastodynamic system for simulating a blood vessel with possible large elastic deformations due to external pressure loadings, such as the impact of blood flow on the interior surface or a squeeze pressure on the exterior surface. To clarify, we denote
In equation (1), the first Piola–Kirchhoff stress tensor
The Saint–Venant–Kirchhoff (SVK) model is usually regarded as one type of hyperelastic models,
11
for which category, there exists a strain energy density functional
In particular, for the SVK model, its energy density functional is in the form of
Despite its simplicity, the SVK model provides a basic module for many other elastic models which are more sophisticated and realistic, due to the fact that it captures some most essential features of soft materials.12,13 So far, the SVK model has been widely explored in many real-world applications, especially in studying the biomechanical characteristics of human body soft tissues, such as brain surgery, 14 liver, 15 respiratory system, 16 and so on.
Methodology
Spatial discretization
We discretize the partial differential system (1) spatially by using the standard finite element method. First, the variational form is found to be: find
Provided a N vertices tetrahedral mesh
On the other hand, when ζi is selected to be
Here,
Time discretization
To get a full discretization, we apply a backward differentiate formula to the time derivative terms in equations (10) and (11). Here, without loss of generality, we show the first-order case, i.e. the backward Euler method: by denoting
Thus, the nonlinear finite element equations (10) and (11) become
Provided that all information when
Here, we emphasize that the unknowns are the coefficients vectors:
A Newton–Krylov–Schwarz solver
At each time step tn, we use the Newton’s method to deal with the nonlinear algebraic equation system (12) and (13). That is, we start with an initial guess Find Update
where
The Krylov subspace methods are a family of iterative methods for solving algebraic linear systems, and they are highly desired for large number of unknowns with sparse coefficient matrices, such as the ones induced from partial differential equation systems, and play an essential role in modern supercomputings for solving versatile engineering problems.17–21 In this paper, we choose the GMRES method as the linear solver inside each Newton iteration. This method constructs a sequence of orthogonal vectors in a successively expanded Krylov spaces by minimizing the equation’s residual, such that they will eventually approach to the solution. A detailed introduction and justification of the GMRES method can be found in Saad.
22
In practice, applying a Krylov subspace method, such as GMRES, to solve a PDE problem often suffers a slow convergence and instability. This is mainly because the coefficient matrix of the induced linear system is ill-conditioned, which gets even worse when the problem size becomes larger. A common way to overcome this difficulty is to adopt a so-called “preconditioning.” For example, when a right preconditioner
Specifically, a Schwarz preconditioner
Correspondingly, two interpolation operators
In practice, the specific forms of these matrices may vary, depending on which domain decomposition strategy is adopted. In this paper, we choose the restricted additive Schwarz method, in which case
Based on the above settings,
Here
The whole algorithm, a sequence of Newton iterations embedded with Krylov subspace linear solvers each of which is accelerated by a Schwarz preconditioner, is usually referred to the Newton–Krylov–Schwarz (NKS) method.
Numerical experiments
In this section, we present some numerical results for investigating deformations of a blood vessel that undergoes external loadings. Figure 1(a) shows an unstructured finite element mesh of a segment of a patient-specific carotid artery vessel used in our tests, which is composed of 19,848 vertices and 75,454 cells and corresponds to 114,003 degree of freedoms. The main body of our program package is implemented based on the well-known scientific computing toolkit PETSc, and the mesh partitioning is performed by using ParMetis for parallel computing. In our tests, we use the time step size

(a) The unstructured tetrahedral mesh on the blood vessel used in our finite element simulations (19,848 vertices, 75,454 cells); (b) the time-variant pattern of the magnitude of the surface loading
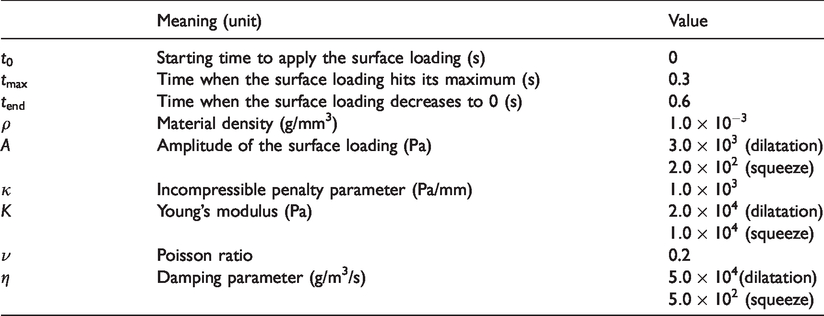
Model parameters and their standard values used in the numerical tests.

Boundary settings
In our simulations, the two ends of the vessel are fixed, on which a homogeneous Dirichlet boundary condition is applied. To deform the vessel, we either apply a dilatation force on the interior surface or squeeze it on the exterior surface, respectively. This force is dynamically loaded to ensure that its current direction is always normal to the surface no matter how the shape is changed. More specifically, in equation (9), we set
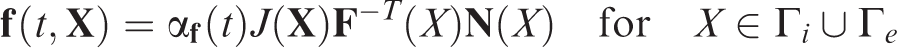

Incompressibility and damping
It is usually regarded that many soft materials, including human and animal tissues, are incompressible. For taking such an effect into consideration, we add an extra term in the strain energy functional equation (7)


Similar to the dynamic loading term, it also induces an extra contribution into the Jacobian of the Newton’s equation.
Another practical setting is about the damping parameter η. In physics, damping is understood as an effect to prevent oscillatory motions of a dynamic system. Numerically, based on our experiences, introducing the damping term with a properly selected η usually improves the stability of the Newton solver.
In the following, we show the numerical results obtained from these two sets of experiment.
Vessel dilatation
In this test, a loading is applied over the whole interior surface (except for the parts close to two ends) to expand the vessel. We first tested five cases with different choices of Young’s modulus K (

(a)–(e) Vessel dilatation for different choices of Young’s modulus; (f) a cross-section view on a vertical plane for different Young’s modulus (Unit: Pascal): (a) K = 1 × 104; (b) K = 1.5 × 104; (c) K = 2 × 104; (d) K = 2.5 × 104; (e) K = 3 × 104; (f) slice cuts.
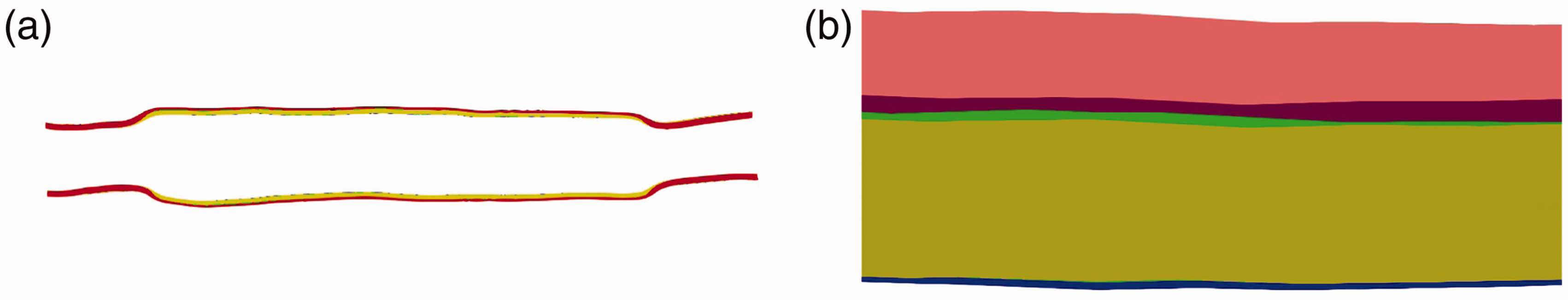
(a) The slice cuts across the middle plane of the vessel in dilatation for different Poisson’s ratio ν: 0.1 (blue), 0.2 (green), 0.3 (yellow), and 0.4 (red); (b) a close look on the top boundary area.
Visualization of the numerical results clearly shows that as the Young’s modulus K gets larger, the smaller deformation of the vessel can be expanded. On the other hand, when the Poisson ratio ν changes, the results keep almost the same, and the most significant difference that can be observed is the thickness of the deformed vessel boundary layer. The observations agree with the common understanding of these model parameters.
Vessel squeeze
We repeated our tests for applying a squeezing force on the exterior surface and collected the results in Figure 4. In these tests, the force is applied on a top and bottom areas near the center of the vessel, which is dynamically loaded to ensure the normality to the exterior surface. Similarly, we also tested different Poisson ratios ν, in which case, the Young’s modulus K is fixed at

(a)–(e) Vessel squeezing for different choices of Young’s modulus; (f) the central slice cut show for different Young’s modulus cases (Unit: Pascal): (a) K = 1 × 104; (b) K = 1.5 × 104; (c) K = 2 × 104; (d) K = 2.5 × 104; (e) K = 3 × 104; (f) a cross-section view.

(a) The slice cuts across the middle plane of the vessel in pressure for different Poisson’s ratio ν: 0.1 (blue), 0.2 (green), 0.3 (yellow), and 0.4 (red); (b) a close look on the top depressed area.
Solver performance
We present some performances of our finite element solver for the vessel dilatation and squeeze simulations. In the following tables, we denote
We first test the model parameters: Young’s modulus K and Poisson ratio ν, respectively. As the results are shown in Table 2, the numerical solver is able to complete all simulations very efficiently and robustly. For each case, both the nonlinear and the linear solver terminates with a small number of iterations.
Solver performances on the vessel dilatation and squeeze tests.

This table presents the average Newton iterations per time step, the average GMRES iterations per Newton step, and the total computing time, for each vessel dilatation or squeeze case, respectively. All data are counted on the first 30 time steps (when it reaches the maximum pressure loading) for each case of simulations.
Based on the same mesh, we also studied the impact on the solver performance brought by the fill-in level of the ILU subdomain solver. In particular, we repeat the dilatation and the squeeze simulations with the ILU fill-in level to be 0, 1, 2, and 3, respectively. The problem is solved by using four processors, and the results are collected in Table 3. It can be seen that the higher ILU fill-in level can reduce the GMRES iteration numbers, but may cost more total computational time. According to the results by these tests, to set the fill-in level 1 may be the best choice for most of the cases.
Sensitivity of solver performance on the ILU fill-in level.

This table presents the average Newton iterations per time step, the average GMRES iterations per Newton step, and the total computing time cost, for each vessel dilatation or squeeze case, respectively. All data are counted on the first 30 time steps for each case of simulations.
In addition, we demonstrate the strong scalability of our numerical solver. The mesh used here has the same geometry as the previous case, with 129,298 vertices and 604,383 cells, and the computing time for each case is counted for the first 30 steps. For two jobs, which take time of T1 and T2, by using n1 and n2 processors, respectively, we define

We test on 16, 24, 32, 48, 64, and 96 CPU processors and compute the parallel scalability based on the 16-processor case. The results can be found in Table 4. It can be seen that our numerical solver has a very good efficiency, and the nonlinear and linear solver are stable in terms of the Newton and GMRES iteration numbers. This indicates that the proposed algorithm has a great potential for larger-sized elastodynamic problem with more processors. We plan to investigate this topic further in the future.
Strong scalability performance of our numerical solver for blood vessel simulations.

The average Newton iterations per time step, the average GMRES iterations per Newton step, and the computing time are reported for each case. The speed-up and parallel efficiency are the computed based on the computing time for the first 30 time steps.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Shenzhen research projects: JCYJ20160331193229720, JCYJ20170307165328836, and JSGG20170824154458183 and NSFC funding 61531166003.