Abstract
Hyperbolicity is a global property of graphs that measures how close their structures are to trees in terms of their distances. It embeds multiple properties that facilitate solving several problems that found to be hard in the general graph form. In this paper, we investigate the hyperbolicity of graphs not only by considering Gromov’s notion of δ-hyperbolicity but also by analyzing its relationship to other graph’s parameters. This new perspective allows us to classify graphs with respect to their hyperbolicity and to show that many biological networks are hyperbolic. Then we introduce the eccentricity-based bending property which we exploit to identify the core vertices of a graph by proposing two models: the maximum-peak model and the minimum cover set model. In this extended version of the paper, we include some new theorems, as well as proofs of the theorems proposed in the conference paper. Also, we present the algorithms we used for each of the proposed core identification models, and we provide more analysis, explanations, and examples.
Keywords
Introduction
Using graph-theoretical tools for analyzing complex networks to characterize their structures has been the subject of much research. It aids identifying multiple key properties as well as explaining essential behaviors of those systems. A common structure that has been widely recognized in social networks as well as other network disciplines is the core–periphery structure. Multiple coefficients have been proposed to examine the existence of the core–periphery organization in a graph1,2; also, various methods have been introduced to identify the core of a graph.3–5
The core–periphery structure suggests partitioning the graph into two parts: the core which is dense and cohesive and the periphery which is sparse and disconnected. Vertices in the periphery part interact through a series of intermediate core vertices. This pattern of communication (where traffic tends to concentrate on a specific subset of the vertices) has been observed in trees where distant nodes communicate via the central node (or nodes) in the tree. δ-Hyperbolicity, which is a measure that shows how close a graph is to a tree, suggests that any shortest path (geodesic) between any pair of vertices bends (to some extent) toward the core of the graph. This phenomenon has been justified by the global curvature of the network which (in case of graphs) can be measured using hyperbolicity (sometimes called also the negative curvature). 6
There are multiple equivalent definitions for Gromov’s hyperbolicity. Let G = (V, E) be a graph with a distance metric d on V such that the distance between two vertices x and y is the length of a shortest path between x and y. A geodesic triangle
Multiple complex networks such as the Internet,8,9 data networks at the IP layer, 6 and social and biological networks10,11 show low δ-hyperbolicity (low hyperbolicity suggests a structure that is close to a tree structure 12 ). Also, it has been observed that networks with this property have highly connected cores. 6 Generally, the core of a graph is specified according to one or more centrality measures. For example, the degree centrality where the core of the graph is the set of vertices that have the highest number of connections, the betweenness centrality which considers the vertices that have the highest number of shortest paths passing through them as the core, and the eccentricity centrality in which the core has the subset of vertices that have the shortest distance to every other vertex.
The δ-hyperbolicity of graphs embeds multiple properties that facilitate solving several problems that found to be difficult in the general graph form. For example, diameter estimation, 13 distance and routing labeling, 14 and several routing problems.15,16 In this paper, we investigate implications of the δ-hyperbolicity of a network and exploit them for the purpose of partitioning the graph into core and periphery parts.
Our main contributions can be summarized as follows:
We study the hyperbolicity of several real-world biological networks and show that the hyperbolicity of almost all the networks in our datasets is small. This confirms the results in Albert et al.
11
However, unlike previous efforts, we analyze the relationship between the hyperbolicity and other global parameters of the graph. We find in most of our networks that the hyperbolicity is bounded by the logarithm of the diameter ( We formalize the notion of the eccentricity layering of a graph and employ it to introduce a new property that we find to be intrinsic to hyperbolic graphs: the eccentricity-based bending property. Unlike previous work, we investigate the essence of this bending in shortest paths by studying its relationship to the distance between vertex pairs. We exploit the eccentricity-based bending property, and based on it, propose two core–periphery separation models: the maximum-peak model and the minimum cover set model. We apply both models to our biological graph datasets. In contrast to what have been observed in Holme,
2
we find that biological networks exhibit a clear-cut core–periphery structure. Then we investigate the relationship between the hyperbolicity of a graph and the conciseness of its core.
This paper is organized as follows. The next section discusses some theoretical background on graph theory and presents the definition of δ-hyperbolic graphs. Related work on the core–periphery structure and graph centrality measures in networks in general and in biological networks in particular are also discussed. “Datasets” section describes the kinds of biological networks used in this paper and presents a summary of their parameters. In “δ-Hyperbolicity of networks” section, we measure, analyze, and classify the δ-hyperbolicity of the networks in our datasets. Our classification is based on how the hyperbolicity is evident in those graphs. We also discuss how other graph parameters factor in to this classification. Finally in “Core–periphery models based on δ-hyperbolicity” section, we propose our eccentricity-based bending property followed by two core–periphery separation models as one of this property’s implications.
Theoretical background and related work
Preliminaries on graph theory
A simple undirected graph G = (V, E) naturally defines a metric space (V, d) on its vertex set V. The distance d(u, v) is defined as the number of edges in a shortest path ρ(u, v) that connects two vertices u and v.
We define the size of the graph denoted as size(G) as the sum of the number of vertices and the number of edges in G, i.e.
The diameter of the graph diam(G) is the length of the longest shortest path between any two vertices u and v in the graph, i.e.
According to the distances between vertices in the graph, the center can be defined using the vertex eccentricities. The eccentricity of a vertex u is
δ-Hyperbolicity
In smooth geometry, hyperbolicity captures the notion of negative curvature which can be generalized as δ-hyperbolicity in more abstract concept of metric spaces including graphs. 17 The presence of hyperbolic networks in a big variety of applications attracted many researchers to investigate the negative curvature of different types of graphs. It turns out that many real-world networks are hyperbolic.6,8,9,11 Moreover, multiple applications of the hyperbolicity have been examined such as diameter estimation, 13 compact distance and routing labeling schemes and spanners,14,16,18 and several routing and traffic flow problems.6,15,16,19
The δ-hyperbolicity measure of a metric space was proposed by Gromov in 1987. 20 It measures how close the metric structure is to a tree structure. A connected graph G can be viewed as a metric space with the graph distance metric d. There are multiple equivalent definitions (up to constant factors 13 ) for Gromov’s hyperbolicity. In this paper, we use the following four-point condition definition.
Given a graph G = (V, E), x, y, u, and
A graph G = (V, E) is considered δ-hyperbolic for some nonnegative real number δ if for every set of four points x, y, u, and v, the larger two of the three sums
Trees are 0-hyperbolic (δ(G) = 0), cliques are also 0-hyperbolic, and chordal graphs are at most 1-hyperbolic. 21 On the other hand, a cycle with n vertices is approximately n/4-hyperbolic and an n × n grid is (n – 1)-hyperbolic. Generally, the smaller the value of δ the closer the graph is to a tree (metrically); this implies strong hyperbolicity. See Wu and Zhang 22 for a detailed discussion on tree-likeness and hyperbolicity.
Core–periphery and network centrality in complex networks
The notion of the core–periphery structure has a long history in social network analysis. It deals with identifying the part (or parts) of the network that represents the central part in terms of the network distance, the most congested part in terms of the network traffic, the highly connected part in terms of the vertices’ degrees, or any combination of the three. Borgatti and Everett 3 formalize the core–periphery structure by developing two families of core–periphery models: the discrete model where vertices belong to one of two classes (core and periphery) and the continuous model which includes three classes (or more) of vertices (core, semiperiphery, and periphery). Then they propose algorithms for detecting each model by finding the partition which maximizes the correlation between the data matrix and the pattern matrix.
Seidman 4 proposes the k-core decomposition as a tool to study the structural properties of large networks focusing on subsets of increasing degree centrality. It partitions the graph into subsets each of which is identified by removing vertices of degree smaller than k.
Holme
2
introduces a coefficient that measures if a network has a clear core–periphery structure based on the closeness centrality (defined below) and the basic definition of clusters. He also shows that the core’s neighborhood (for increasing radius) is highly dense (with respect to the number of edges). Unlike our result, he concludes that biological networks do not have a strong core–periphery organization. In Chung and Lu,
23
the authors show that for some families of random graphs with expected degrees there is a core to which almost all vertices are at distance less than or equal to
In the study of communication networks, the core is usually identified by the small dense part of the network that carries out most of the traffic under shortest-path routing.6,25,26 Narayan and Saniee 6 show that the load scales as O(n2) with the number of vertices n at the core of the network. The asymptotic traffic flow in hyperbolic graphs has been studied in Baryshnikov and Tucci. 25 It shows that a vertex v belongs to the core if there exists a finite radius r such that the amount of the traffic that passes through the ball centered at v and with radius r behaves asymptotically as θ(n2) as the number of vertices n grows to infinity. The existence of the core in large networks such as the Internet motivates researchers to embed the Internet distance metric in a hyperbolic space for distance estimation.9,27
The notion behind centrality is identifying key vertices in a graph that are considered high contributors. There are multiple centrality measures in the literature; some of which are: the betweenness centrality (similar to Newman, 28 we will call it the shortest-path betweenness centrality) and the eccentricity centrality.29–31
The shortest-path betweenness measure is a widely used concept in social networks. It expresses how much effect each vertex (or actor) has in the communication of the network assuming that all traffic follows shortest paths between every two vertices (taking into account that one or more shortest paths may exist between any given pair).
Given a connected finite graph G = (V, E), the shortest-path betweenness of a vertex
The eccentricity centrality suggests that the center of the graph includes the vertex (or vertices) that has the shortest distance to all other vertices (minimum eccentricity). For a given vertex u, the eccentricity centrality of u is
The eccentricity centrality is sometimes referred to in literature as the closeness centrality. To avoid any ambiguity, we point out that the closeness centrality considers the center as the subset of vertices with the minimum total distance to all other vertices. In other words, the closeness centrality
Biological networks and the core–periphery structure
It has been found in several fields that looking at the overall system reveals more about the functionality of its components as opposed to inspecting its individual elements. Therefore, various types of large-scale biological networks have been constructed to capture the different kinds of interactions between their components. For instance, protein–protein interaction (PPI) networks have been used to identify the function of individual proteins as well as the purpose behind some unknown interactions.
33
A lot of research efforts were directed to discover some topological properties of the biological networks. It has been shown that some protein structures and PPI networks34,35 and a number of metabolic networks36–38 exhibit the small-world property (a graph is small world when its diameter is bounded by the logarithm of its size (
Structure analyses of some biological networks have detected the presence of hierarchal, modular, and core–periphery organization structures. The core–periphery model of several types of biological networks has been studied thoroughly in the literature. Da et al. 1 propose a parameter that detects the existence of a core–periphery structure in a metabolic network based on the closeness centrality of metabolites and the network connectivity. In Junker and Schreiber, 40 the yeast protein interaction network was analyzed based on the betweenness centrality. They conclude that the high betweenness low connectivity proteins may be working as connectors between separate modules. Using mathematical tools that have been used to analyze sociological networks, the authors in 36 study recognizing the central metabolites in a metabolic pathway network. In Luo et al., 41 the authors demonstrate a systematic exploration of the core–periphery model in protein interaction networks that depends on the connectivity of the vertices. They also classify the peripheral vertices based on their structural relationship with the core. In Ma and Zeng, 42 the authors identify the central metabolites using degree centrality and closeness centrality. They also show the relationship between the average path length and the closeness centralization index of metabolic network.
Datasets
There are various forms of biological networks that have different characteristics according to their origins and to their construction methodologies. Generally, the vertices in a biological network represent biomolecules such as proteins, genes, or metabolites, and the edges represent a chemical, physical, or functional interaction between the connected vertices. Each of the networks used in this work belongs to one of the following types:
Protein interaction (PI) networks: Generally, in a PI network, the vertices represent different proteins and the edges represent the connections between the interacting proteins. PI networks have been described as small-world and scale-free networks.
40
Neural networks: They contain neurons (vertices) which are connected together through synapsis (edges). Neurons have a high tendency to form clusters based on their spatial location. Neural networks are small-world networks.
40
Metabolic networks: Metabolic networks are represented by metabolites (vertices) and biochemical reactions (directed edges). Usually, metabolites are small molecules (for example amino acids); however, they also can be macromolecules. Metabolic networks show the small-world property and they have a high clustering coefficient. Also, they follow the power-law degree distribution.
40
Transcription networks: Networks in which vertices are genes and edges represent different interactions (interrelationships) between genes. A transcription network is one type of the gene regulatory networks.
43
We analyze the PI networks of budding yeast, 44 Escherichia coli, 45 yeast, 46 Saccharomyces cerevisiae, 47 and Helicobacter pylori. 48 Also, we analyze two different brain area networks of the Macaque monkey49,50 and the metabolic networks of the Escherichia coli 51 and the Caenorhabditis elegans. 52 Finally, we analyze the yeast transcription network. 43
Graph datasets and their parameters: number of vertices |V |; number of edges |E|; graph’s size
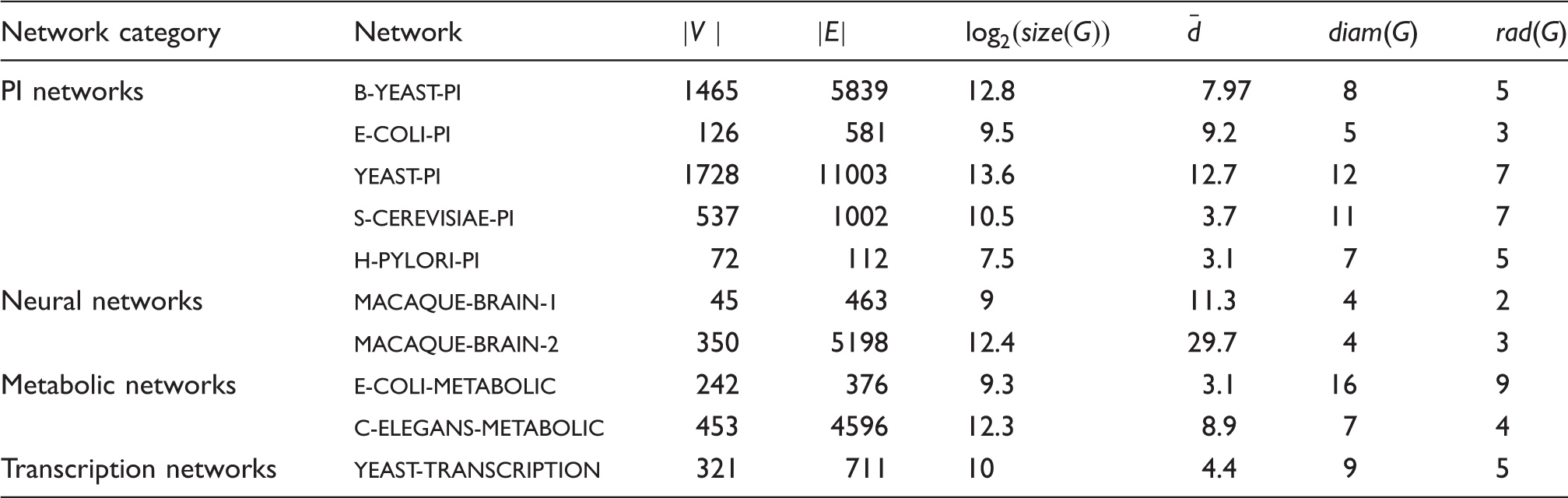
δ-Hyperbolicity of networks
For the purpose of investigating the hyperbolicity of networks, it seems natural to analyze and classify the graphs based on their hyperbolicity. The classification should reflect how strong (evident) the tree-likeness is in the graph’s structure.
In the following subsections not only we measure the hyperbolicity of each of the networks in our graph datasets, but also we relate this value to other graph’s parameters. Upon this analysis, we provide our hyperbolicity-based classification of the graphs.
Hyperbolicity of biological networks
We measure δ-hyperbolicity on each bi-connected component of each network in the datasets presented in “Datasets” section using Gromov’s four-point condition. For each network, we identify a bi-connected component with the maximum value of δ since the hyperbolicity of a graph equals the maximum hyperbolicity of its bi-connected components.8,22,53
Graph datasets and their parameters: number of vertices |V |; number of edges |E|; diameter diam(G); radius rad(G); hyperbolicity δ(G); and the average hyperbolicity
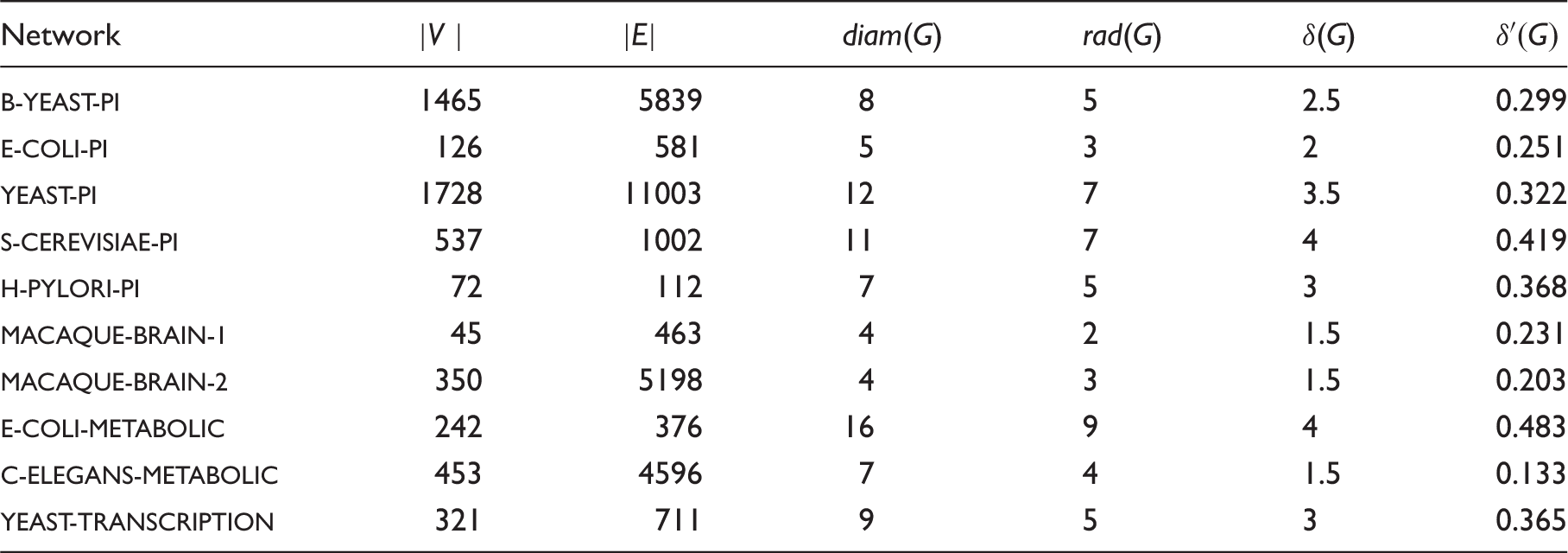

Distribution of quadruples over different values of δ.

Distribution of quadruples over different values of δ (three datasets): (a)
This observation makes it equally important to calculate the value of the average delta
Analysis and discussion
Our goal is to categorize graphs with respect to their hyperbolicity into three classes: strongly-hyperbolic graphs, hyperbolic graphs, and nonhyperbolic graphs.
Taking into account that trees are 0-hyperbolic, generally, the smaller the value of δ(G), the closer the graph’s structure to a tree. However, studying the tree-like structure of graphs based solely on the value of the hyperbolicity may not be sufficient to characterize their closeness to a tree topology for two reasons. First, the hyperbolicity is a relative measure. For example, for a given graph G = (V, E), a value of δ(G) = 10 can be seen as too large when
Since finite graphs will always have a finite value for δ such that the four-point condition is true, it is natural to think that the nonhyperbolic class includes only infinite graphs. However, in this study, we only consider finite graphs; accordingly, a nonhyperbolic graph in our sense is a graph with too large δ with respect to the logarithm of the graph’s size, i.e. when it violates the following condition:
Recall that
Multiple previous works have analyzed the relationship between the hyperbolicity and the diameter. In fact, the graph’s diameter represents an upper bound for δ.
54,55
For any graph G with diameter diam(G) and hyperbolicity
Lemma 1
Moreover, the authors in Kennedy et al. 17 (using the slim triangles condition for hyperbolicity) and in Jonckheere et al. 56 (using the four-point condition) argue that the hyperbolicity of the graph is “actually” present when the value of δ is much smaller than the graph’s diameter. They specify that for a graph to be hyperbolic the value of δ/diam(G) must asymptotically scale to zero. In this work, our goal is to ensure that the low value for the hyperbolicity is not a consequence of the graph’s small diameter.
Interestingly, for most of the networks in our graph datasets, we find that Strongly-hyperbolic if it exhibits (1) Hyperbolic when it violates either (1) or (2); Nonhyperbolic in all other cases.
Note that in the case when δ is small but greater than
As Table 1 shows, all networks in the datasets, with the exception of networks Classification of the graph datasets based on their hyperbolicity.
The ratio of δ to logarithm of the size of the graph

We also compared the value of δ to
Core–periphery models based on δ-hyperbolicity
It was suggested by Baryshnikov and Tucci 25 and Narayan and Saniee 6 that the highly congested cores in many communication networks can be due to their hyperbolicity or negative curvature. Those cores are represented by vertices that belong to most shortest paths (shortest-path betweenness centrality) and (or) have minimum distances to all other vertices (eccentricity centrality). It was also observed that the negative curvature causes most of the shortest paths to bend making the peak of the arc formed by a shortest path to pass through a vertex in the core. In this section, we formalize this notion of bending in shortest paths by introducing an important property that is intrinsic to δ-hyperbolic graphs (the eccentricity-based bending property). Then we use the implication of this property to aid the partitioning of a graph into core and periphery parts by proposing two models: the maximum-peak model and the minimum cover set model. we further apply our models to the biological networks in our datasets. In contrast to what have been observed in Holme, 2 we show that biological networks do exhibit a core–periphery structure.
Eccentricity layering of a graph
The eccentricity layering of a graph Eccentricity layering of a graph. Darker vertices belong to lower layers.
Any vertex Distribution of vertices over different layers of the graph’s eccentricity layering. Distribution of vertices over different layers with respect to the graph’s eccentricity layering. rad(G) is the graph’s radius; |V | is the number of vertices of each graph.

In this work, we use layer 0 to indicate the central layer, layer 1 to indicate the layer that directly succeeds the central layer, etc. Therefore, when we say lower layers, we mean layers toward the central layer and the opposite for higher layers.
Eccentricity-based bending property of δ-hyperbolic networks
Let G = (V, E) be a δ-hyperbolic graph,
Let G be a δ-hyperbolic graph and
We use this property to establish the following few interesting statements.
Let G be a δ-hyperbolic graph and x, y, s be arbitrary vertices of G. If
Let w be a middle vertex of a shortest (x, y)-path and let
Let G be a δ-hyperbolic graph and x, y be arbitrary vertices of G. If
Let w be a middle vertex of a shortest (x, y)-path given by Proposition 1 and s be a vertex such that Lemma 2 [14]
Proposition 1
Proof
Proposition 2
Proof
Denote by
Let G be a δ-hyperbolic graph and y be its arbitrary vertex. Then,
Consider an arbitrary vertex Proposition 3
Proof
We define the bend in shortest paths between two distinct vertices u and v with ∀ Definition
We say that shortest paths between u and v bend if and only if a vertex z with
Now we analyze how vertex pairs behave in terms of their bending toward the center of the graph C(G). Specifically, we investigate the effect of the distance between a pair of vertices on the bend of the shortest paths between them. Our findings in this context are summarized in the following two statements.
Despite their distances, most vertex pairs bend. Moreover, among those bending vertex pairs, the majority is represented by those that are sufficiently far from each other. There is a direct relation between the distance among vertex pairs and how close to the center a shortest path between them bends.
Motivation and empirical evaluation of (A)
In light of Proposition 2, we investigate how vertex pairs of various distances act with respect to the eccentricity-based bending property. Given two vertices u and v, we know from Proposition 2 that when
Interestingly, we noticed the bend in the majority of shortest paths with lengths A case in which the shortest path between a diametral pair x, y does not bend toward the graph’s center The effect of the distance k between vertex pairs on the bending property. Out of all vertex pairs with distance at least k, we show the percentage of those that bend for three of the networks in our graph datasets.
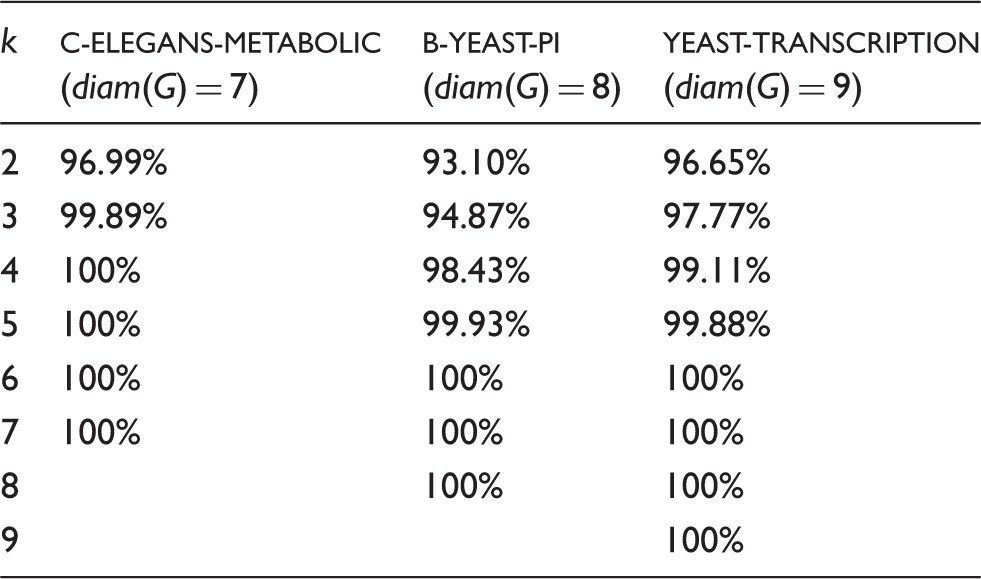
To quantify the distances at which the bend in vertex pairs happens in each of the networks in our datasets, we define two parameters: the absolute curvity and the effective curvity.
The absolute curvity

The sixth and the last columns in Table 6 present the absolute curvity and the effective curvity of each graph as a function of δ so we can compare it with the upper bound Decide the lowest layer μk that all vertex pairs with distance 1. 2. max ← -1 3. 4. 5. 6. max ←bend(x,y) 7. 8. 9. 10. μk ← max 11. print μk 12. Algorithm 1
Motivation and empirical evaluation of (B)
Here we examine the impact of the distance on the level to which vertex pairs bend. Let k be the distance between two vertices such that
This allows us to look at how the bends of the vertex pairs behave with respect to different distances. The algorithm used to find every μk is shown in Algorithm 1. A summary of the results is shown in Figure 7.
μk values for each network in the graph datasets.
As expected, we found a direct relation between the distance of vertex pairs and their bend. For example, Figure 7 shows that in network
In fact, this observation is a direct implication of Proposition 1 and Proposition 2. For every pair of vertices u and v that are sufficiently far from each other, there is a vertex z in the middle of
Also, we know from (A) that the distance among vertex pairs does not need to be very large for a vertex pair to bend. For example, most vertex pairs with distance as small as two from each other bend.
Core–periphery identification using the eccentricity-based bending property
A well-defined center of a graph is a good starting point for locating its core. According to the pattern of data exchange discussed earlier (in which a shortest path between distant vertices bends toward graph’s center), we choose to identify the core of the graph using the eccentricity centrality measure (see “Core–periphery and network centrality in complex networks” section).
Even though the center C(G) contains all vertices that are closer to other vertices, this subset is not sufficient to carry out the communication between every pair of vertices
Obviously, not all graphs exhibit a core–periphery structure. Also, graphs follow this structure with different extents with respect to the quality of their cores. We identify a good graph’s core as the one that (1) includes a small number of layers with respect to the eccentricity layering; and (2) has size (with respect to the number of vertices) that is small compared to the total number of vertices in the graph. The core should also contain vertices that participate in the majority of interactions among other vertices.
In the following two subsections, we discuss two models that exploit the eccentricity-based bending property to partition the vertices in a graph G into two sets: the core core(G) and the periphery.
Model I. The maximum-peak model
Given a δ-hyperbolic graph G = (V, E) along with its eccentricity layering
In light of the eccentricity-based bending property, each bend(x, y) for a pair of vertices x and y represents a peak for shortest paths connecting x and y. In this model, we are locating the index of the lowest layer p over all layers that vertex pairs bend to. Index p represents the separation point where the layers of the graph can be partitioned to a core and a periphery. See Figure 8 for an illustration.
A simplified illustration of the eccentricity layering of a graph (with four layers) and the maximum-peak model. 
After identifying all peaks, the core will include all vertices starting at
Again, to avoid the impact that outlier vertices may impose (as discussed in “Eccentricity-based bending property of δ-hyperbolic networks” section), we define two types of the separation index p: the absolute separation index
The algorithm used to decide the effective separation index Decide the effective separation index The cores of the graph datasets based on the maximum-peak model. |V | is the number of vertices; Algorithm 2

2.
3.
4.
5. cnt + + and mark pair(x,y) as counted
6.
7.
8. break and return
9.
10.
11.
12.
The results in Table 7 show (as expected) a big difference in the sizes (with respect to the number of vertices) of the absolute core and the effective core in the majority of the networks in the datasets. This implies that the identification of the absolute separation index was highly affected by a small percent of vertices. Closer analysis to the effective core set
Recall that the core according to this model is defined by the layers and not by the vertices which may result in the addition of some vertices to the core that do not have real contribution (they were added only for their location in the graph’s eccentricity layering). This issue can be resolved by the identification of the core according to the minimum cover set model presented in the following subsection.
Model II. The minimum cover set model
Consider a graph G = (V, E) with the eccentricity layering The eccentricity ecc(v) (see “Eccentricity layering of a graph” section). Vertices with smaller eccentricities have higher priority to be in the graph’s core. The distance-to-center for a vertex v, denoted as f(v), which expresses the distance between v and its closest vertex from the center C(G), i.e. The betweenness b(v). The definition of the betweenness b(v) of v is close to the definition of the classic shortest-path betweenness centrality. It measures how many pairs of distant vertices x and y have v in one of their shortest paths (versus counting all shortest paths in the classic definition (see “Core–periphery and network centrality in complex networks” section)). It quantifies the participation of a vertex v in the traffic flow process, and we define it as:
Our goal in this model is to identify the smallest subset of vertices that participate in all traffic throughout the network. The algorithm for this model comprises two stages. First, in a priority list T we lexicographically sort the vertices according to the three attributes: ecc(v), f(v), and b(v). T now has the vertices in the order that they should be considered to become part of the core. The goal is to ensure that for each pair of vertices Decide the core of each graph based on the minimum cover set model. T is a priority list in which vertices are ordered based on: ecc(v), f(v), and b(v). 1. C 2. 3. v ← next vertex in T 4. 5. C 6. 7. 8. return CAlgorithm 3
The second stage starts with a vertex v at the head of T being removed from T and added to an initially empty set
The cores of the graph datasets based on the minimum cover set model. |V | is the number of vertices; δ(G) is the hyperbolicity;

Close analysis of Table 8 shows that each produced absolute core The percentage of the uncovered vertex pairs after the orderly addition of vertices to the core set 
To keep only vertices that are considered higher contributors (cover a large number of vertex pairs) we define the effective core set
Because hyperbolic graphs adhere to the property of having shortest paths that bend to the core of the graph, it was natural to think that hyperbolic graphs with lower δ(G) should have even smaller number of vertices in the core. A quick comparison between the
Concluding remarks
The structure of several biological networks has been often described as a chain-like or tree-like topology in molecular biology. 11 This motivates investigating if those networks also admit tree-like structures based on different aspects such as their distances. In “Hyperbolicity of biological networks” section, we observed that most biological networks appear to have low hyperbolicity suggesting their closeness to a tree structure with respect to their distances. 10
In the tree structure, the communication among distant vertices is carried out through the center of the tree which is one or two vertices with the smallest eccentricities. The center in this case represents the core of the network while the rest of the vertices belong to the periphery. Since strongly-hyperbolic graphs have a structure that is closer to a tree structure, this motivates the following hypothesis: does hyperbolicity indicate the existence of a sharper core–periphery dichotomy? In other words, do strongly-hyperbolic graphs have more concise cores compared to (weak) hyperbolic graphs?
Before we answer this question we will analyze the networks in our datasets. In “δ-Hyperbolicity of networks” section, we differentiated between two types of hyperbolicity that appear in our graphs. First, the low hyperbolicity that is caused by a small diameter or graph size; second, the low hyperbolicity with the value of δ(G) smaller than or equal to the value of
Summary of the graph datasets’ parameters and cores: size of the graph size(G), diameter diam(G), hyperbolicity δ(G), average hyperbolicity

We also observed two patterns in strongly-hyperbolic networks named groups 1 and 2 in Table 9. The networks in the first group (the
Footnotes
Acknowledgment
Results of this paper were partially presented at CompleNet 2015.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
