Abstract
A new Cd(II) complex, [Cd(H4pbidc)(H2O)]
n
(
Keywords

A new one-dimensional Cd(II) complex, [Cd(H4pbidc)(H2O)]
n
(
Introduction
Rational design and synthesis of metal-organic complexes (discrete or polymeric) have attracted considerable interest owing to their structural diversity and significant potential applications in catalysis, ion exchange, chemical sensing and drug delivery.1–6 Although many factors need to be considered in the construction of the complexes, careful choice of the organic ligands has been paid more attention in crystal engineering than ever.7–10 It is well known that imidazole and its derivatives, such as imidazole-4,5-dicarboxylic acid, 2-(hydroxymethyl)-1H-imidazole-4,5-dicarboxylic acid, 2-phenyl-1H-imidazole-4,5-dicarboxylic acid, 2-(pyridin-4-yl)-1H-imidazole-4,5-dicarboxylic acid, 2,2′-(1,2-phenylene)bis(1H-imidazole-4,5-dicarboxylic acid), 2,2′-(ethane-1,2-diyl)bis(1H-imidazole-4,5-dicarboxylic acid), are widely used in the preparation of new coordination complexes since they contain N-atom donors, as well as O-atom donors, and have a rich variety of coordination modes which can lead to complexes with intriguing structures and interesting properties.11–17 Among the reported ligands, 2,2′-(propane-1,3-diyl)bis(1H-imidazole-4,5-dicarboxylic acid) (H6pbidc) is a good linker since it has 12 potential donors and can coordinate to almost all soft and hard metal ions with various coordination modes, resulting in final products with diverse structures. To the best of our knowledge, only four mononuclear complexes based on H6pbidc ligand have been reported.18–21 In order to further enrich the categories and numbers of complexes based on this ligand, we continue the use of H6pbidc as a ligand to self-assemble with CdCl2·2.5H2O and have successfully obtained a new one-dimensional (1D) complex [Cd(H4pbidc)(H2O)]
n
. In addition to the determination of the single-crystal structure of complex
Experimental
General information and materials
BSA was purchased from Aokxing biotechnology Co, Ltd, Beijing, and was used without further purification. Tris and dimethyl sulfoxide (DMSO) were obtained from Sinopsin group chemical reagent Co, Ltd. Other chemicals were of analytical reagent grade and were used without further purification. Double-distilled water was used throughout the experiment. The element analyses (C, H and N) were obtained by a FLASH EA 1112 elemental analyzer (Thermo Fisher Scientific Company, Waltham, MA, USA). The infrared (IR) data were recorded on a Nicolet iS50 spectrophotometer (Thermo Scientific, Waltham, MA, USA) with KBr pellets in the range 400–4000 cm−1. All fluorescence measurements were carried out on an F-7000 spectrofluorimeter (Hitachi, Ltd., Chiyoda-ku, Tokyo, Japan) equipped with a thermostat bath and 1.0-cm quartz cells. Fluorescence quenching spectra of BSA were recorded from 285 to 450 nm at an excitation wavelength of 280 nm. The width of the excitation and emission slit was set to 5.0 and 5.0 nm, respectively. All quenching experiments were performed at 298, 308 and 313 K, for BSA concentration of 1.0 × 10−6 mol L−1, and for complex
Synthesis of [Cd(H4pbidc)(H2O)]
n
(1 )
A mixture of CdCl2·2.5H2O (0.05 mmol), H6pbidc (0.05 mmol), methanol (2 mL) and distilled water (4 mL) was sealed in a 25-mL Teflonlined stainless steel container and heated at 413 K for 72 h. After the mixture had been allowed to cool to room temperature at a rate of 5 K h−1, colourless crystals of [Cd(H4pbidc)(H2O)]
n
(

The reaction for the synthesis of complex
Preparation of stock solutions
BSA (relative molecular weight 65,000) was dissolved in Tris-HCl buffer (pH = 7.36, containing 0.1 mol L−1 Tris, 0.1 mol L−1 NaCl) to prepare a stock solution of 5.00 × 10−5 mol L−1 and was kept refrigerated at 273—277 K. A sample of 5.00 × 10−3 mol L–1 stock solution of complex
Single-crystal structure determination
Single-crystal X-ray diffraction data collection for complex
Crystal data and structure refinement for complex

Selected bond lengths (Å) and angles (o) for complex

Symmetry transformations used to generate equivalent atoms: #1 −x + 1, −y + 1, −z + 1; #2 −x, −y + 1, −z + 1.
Hydrogen bonds for complex

Symmetry transformations used to generate equivalent atoms: #3 x, y, z − 1; #4 x, y + 1, z; #5 x, y − 1, z; #6 x, y, z + 1.
Results and discussion
Crystal structure of [Cd(H4pbidc)(H2O)]
n
(1 )
The asymmetric unit of complex
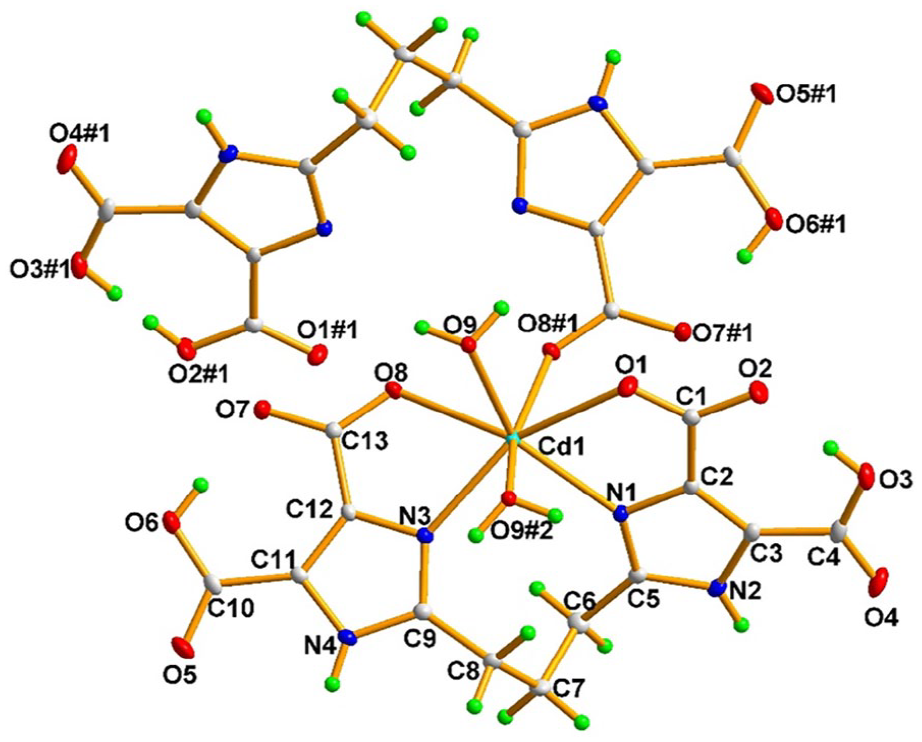
Coordination environment of the Cd(II) ion in complex
As depicted in Figure 2, as well as two imidazole nitrogen atoms and two carboxylato oxygen atoms of the same H4pbidc2− ligand chelate to one Cd(II) ion, the O8 atom also bridges the neighbouring Cd(II) ion. Similarly, the coordinated water molecule O9 also links two neighbouring Cd(II) ions. So the adjacent Cd(II) ions are alternately joined through two carboxylate oxygen atoms (Cd1–Cd1#1: 4.0067(1) Å) and two bridging water molecules (Cd1–Cd1#2: 3.9556(1) Å) to form a 1D chain structure running along the crystallographic a axis. Intramolecular O–H···O hydrogen bonds between the carboxyl and carboxylate groups stabilizes the 1D chain. As shown in Figure 3 and Table 3, there are O9–H9B···O4 hydrogen bonds between the coordinated water molecules and carboxylate groups, and N2–H2A···O7 hydrogen bonds between the imidazole groups and carboxylate groups. Therefore, adjacent 1D chains are connected through the above two kinds of hydrogen bonds, leading to an infinite two-dimensional layered structure parallel to the ac plane. Adjacent layers are further linked by O9–H9A···O5 hydrogen bonds between the coordinated water molecules and carboxylate groups, and N4–H4···O2 hydrogen bonds between the imidazole groups and carboxylate groups, forming a three-dimensional supramolecular architecture in the solid state (Figure 4).

A view of the one-dimensional structure of complex

The two-dimensional structure of complex
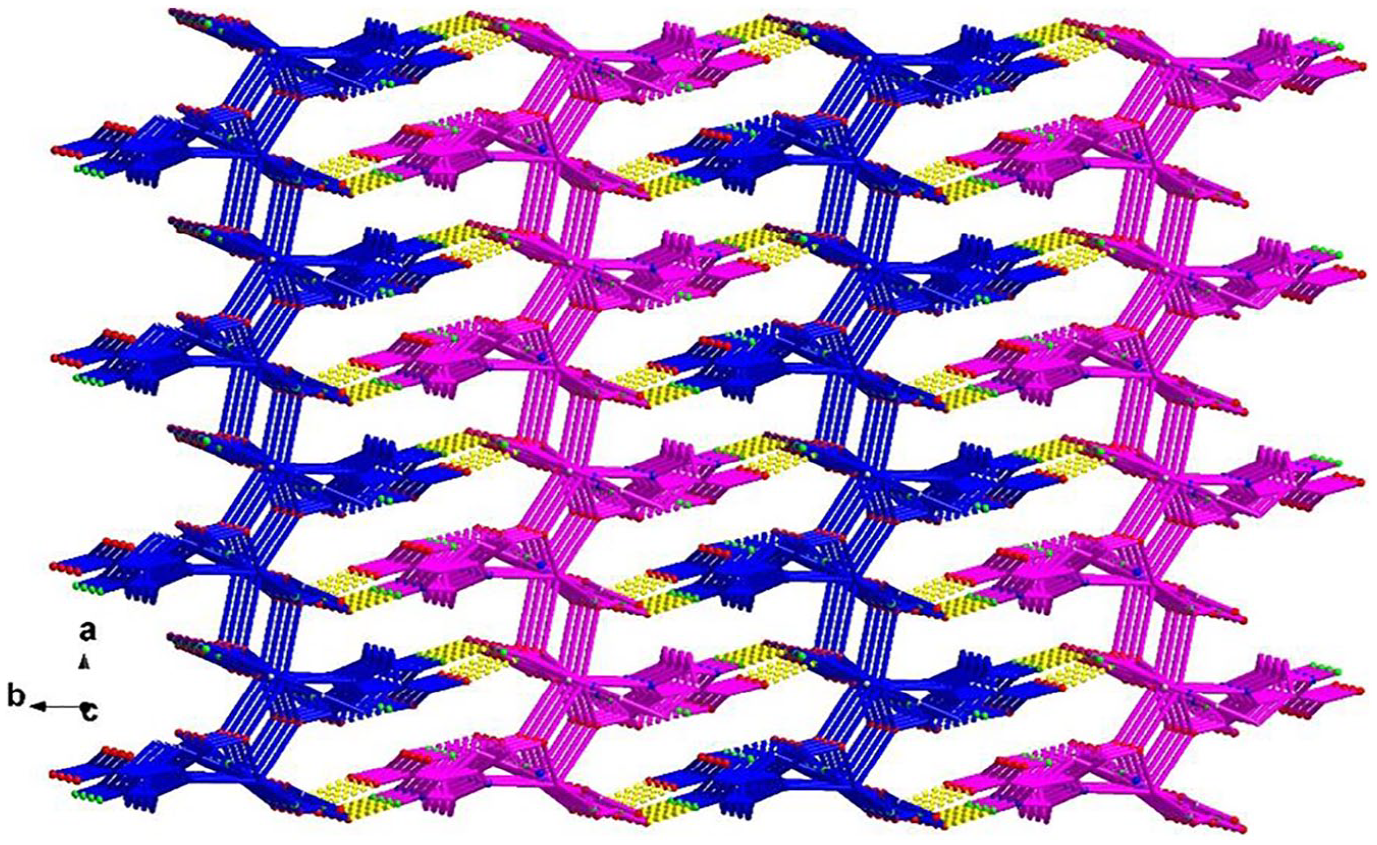
The three-dimensional structure of complex
Through investigation of the literature, we found that four complexes involving H6pbidc ligand, including [Mn(H4pbidc)(H2O)2],
18
[Ni(H4pbidc)(H2O)2]·2H2O,
19
[Ca(H4pbidc)(H2O)3]·4H2O
20
, and [Co(H4pbidc)(pyridine)]·4,4′-bipyridine,
21
have been reported. In this work, a new complex [Cd(H4pbidc)(H2O)]
n
(

Coordination modes of H4pbidc2− anion found in the reported M-H6pbidc complexes (a) and the new coordination mode in complex
Fluorescence quenching spectra and quenching mechanism
Fluorescence measurements were carried out to investigate whether complex
The effects of complex
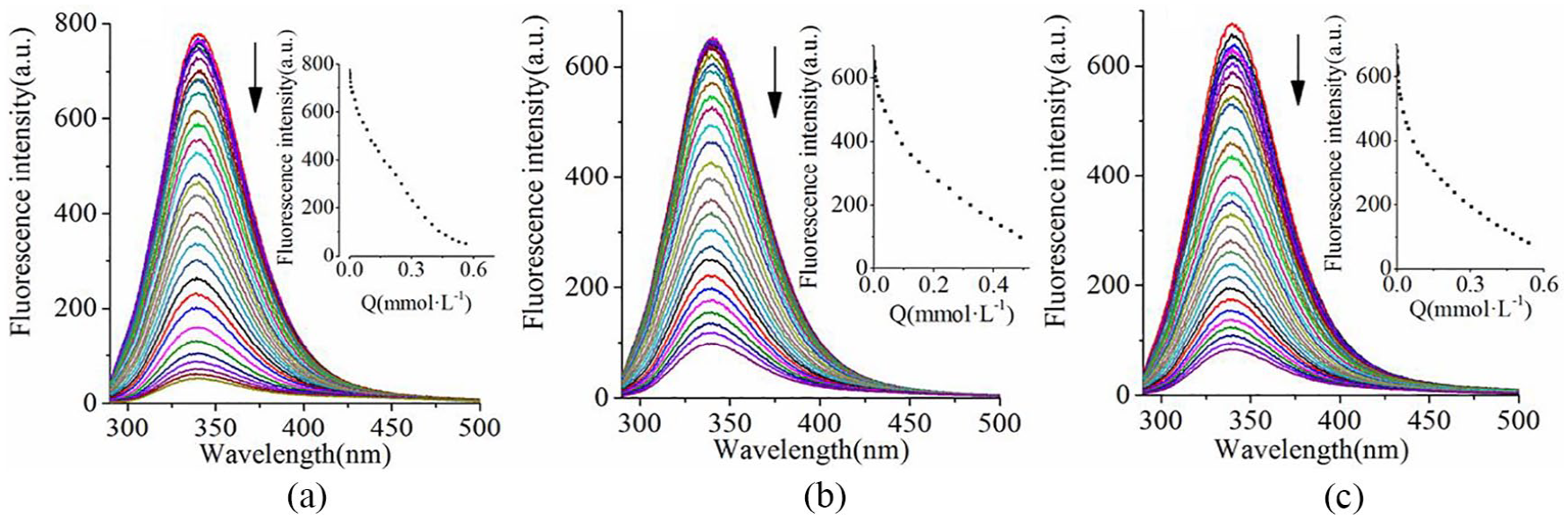
The quenching effect of complex
Quenching can be classified as either dynamic or static quenching by different mechanisms. Dynamic quenching results from collision between fluorophore and quencher, and static quenching is due to the formation of ground-state complex between fluorophore and quencher. The quenching mechanism of the drugs with BSA was probed using the Stern–Volmer equation 26

where F0 and F are the fluorescence intensity of BSA in the absence and presence of quenching agent (complex
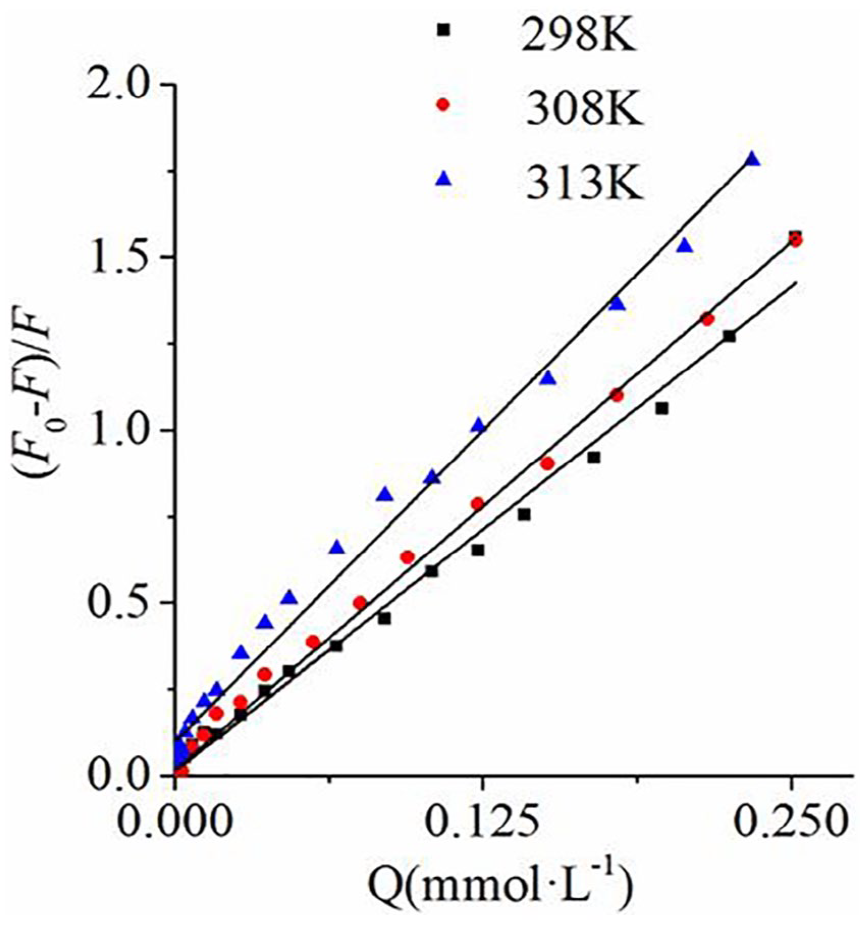
The Stern–Volmer curves for quenching of complex
The quenching constants between complex

Binding parameter and binding mode
For static quenching, the relationship between fluorescence quenching intensity and the concentration of quenchers can be described by the modified form of the Stern–Volmer equation 28

where Kb is the binding constant and n is the number of binding sites.
Plots of Log((F0 − F)/F) versus log[Q] yields logKb as the intercept and n as the slope (see Figure 7). As can be seen from Table 5, the binding constants Kb decreased with the increasing of temperature, which may indicate forming an unstable compound. The unstable compound would be partly decomposed with the rising temperature; therefore, the values of Kb decreased. The values of n at the experimental temperatures were approximately equal to 1, which indicated the existence of a single binding site in BSA for complex

Double-log plots of complex
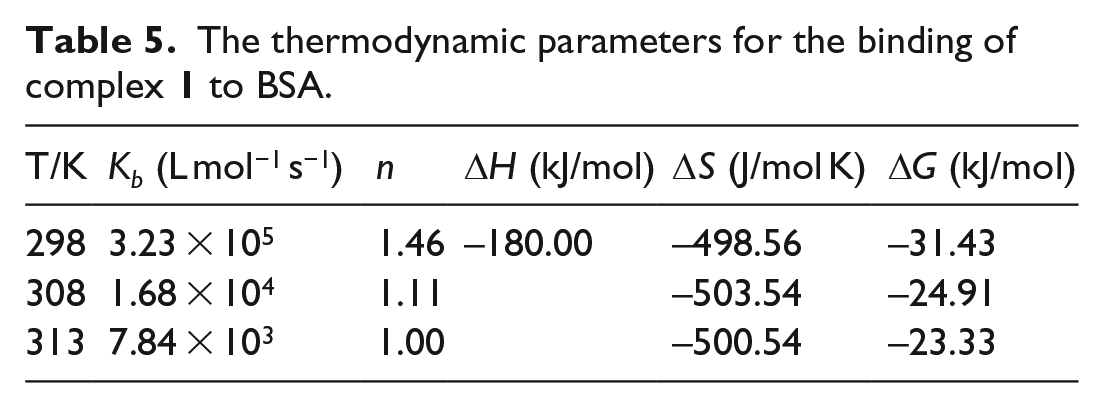
The thermodynamic parameters for the binding of complex
Essentially, there are four types of non-covalent interactions that could play a key role in drug binding to proteins including hydrogen bonds, van der Waals forces, electrostatic and hydrophobic interactions.
29
The thermodynamic parameters, enthalpy change (ΔH) and entropy change (ΔS) for the binding reaction are the main evidence to confirm binding modes. From the thermodynamic standpoint, ΔH > 0 and ΔS > 0 imply a hydrophobic interaction; ΔH < 0 and ΔS < 0 reflect the van der Waals forces and hydrogen bond interactions; and ΔH ≈ 0 and ΔS > 0 suggest an electrostatic force.
26
In order to elucidate the binding mode, the binding constant of complex
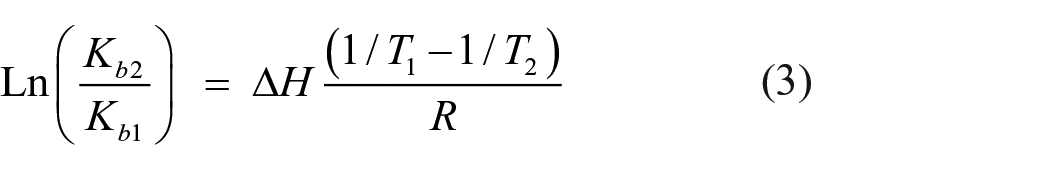


where Kb is the binding constant at the corresponding temperature, and R is the gas constant.
As shown in Table 5, the enthalpy change (ΔH) and entropy change (ΔS) of the combination processes at different temperatures are all negative, which indicated that these bindings are an exdothermic and entropy decreasing processes. The negative value for Gibbs free energy (ΔG) suggests that the binding process for the interaction between complex
The interactions between BSA and the free ligand H6pbidc or free CdCl2 have also been investigated under the same experimental conditions. As shown in Supplemental Figures S1 and S2 (see supporting information), the fluorescence intensity of BSA was little changed with the increase addition of the free ligand H6pbidc or free CdCl2. Furthermore, we have determined the molecular weight of the complex
Conclusion
Through the solvothermal reaction of CdCl2·2.5H2O with H6pbidc, a new complex, [Cd(H4pbidc)(H2O)]
n
(
Supplemental Material
Supporting_Information – Supplemental material for Synthesis, crystal structure and bovine serum albumin–binding studies of a new Cd(II) complex incorporating 2,2′-(propane-1,3-diyl)bis(1H-imidazole-4,5-dicarboxylate)
Supplemental material, Supporting_Information for Synthesis, crystal structure and bovine serum albumin–binding studies of a new Cd(II) complex incorporating 2,2′-(propane-1,3-diyl)bis(1H-imidazole-4,5-dicarboxylate) by Xiao-Fei Li, Li-Gang Ma, Yan-Qiu Yang, Yan-Ju Liu, Xiang-Ru Meng and Huai-Xia Yang in Journal of Chemical Research
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the by programme for the key scientific research project of colleges and universities in Henan Province (grant no. 16A350002) and Henan Province basic and frontier technology research projects (grant no. 152300410214).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
