Abstract
Background:
Lymph node metastasis is associated with a poorer prognosis in endometrial cancer.
Objective:
The objective was to synthesize and critically appraise existing predictive models for lymph node metastasis risk stratification in endometrial cancer.
Design:
This study is a systematic review.
Data Sources and Methods:
We searched the Web of Science for articles reporting models predicting lymph node metastasis in endometrial cancer, with a systematic review and bibliometric analysis conducted based upon which. Risk of bias was assessed by the Prediction model Risk Of BiAS assessment Tool (PROBAST).
Results:
A total of 64 articles were included in the systematic review, published between 2010 and 2023. The most common articles were “development only.” Traditional clinicopathological parameters remained the mainstream in models, for example, serum tumor marker, myometrial invasion and tumor grade. Also, models based upon gene-signatures, radiomics and digital histopathological images exhibited an acceptable self-reported performance. The most frequently validated models were the Mayo criteria, which reached a negative predictive value of 97.1%–98.2%. Substantial variability and inconsistency were observed through PROBAST, indicating significant between-study heterogeneity. A further bibliometric analysis revealed a relatively weak link between authors and organizations on models predicting lymph node metastasis in endometrial cancer.
Conclusion:
A number of predictive models for lymph node metastasis in endometrial cancer have been developed. Although some exhibited promising performance as they demonstrated adequate to good discrimination, few models can currently be recommended for clinical practice due to lack of independent validation, high risk of bias and low consistency in measured predictors. Collaborations between authors, organizations and countries were weak. Model updating, external validation and collaborative research are urgently needed.
Registration:
None.
Plain language summary
Lymph node metastasis of endometrial cancer is associated with a poor prognosis. There are currently many predictive models. We summarized and evaluated them in this article.
Keywords
Introduction
In 2022, there were expected to be 84,520 new cases and 17,543 new deaths of uterine corpus cancer in China, with endometrial cancer (EC) constituting the majority. 1 EC with lymph node metastasis (LNM) signifies at least the International Federation of Gynecology and Obstetrics (FIGO) 2023 stage IIIC1/2 2 and suggests a poorer prognosis. 3 Even micrometastasis predicts an inferior outcome, 4 which has already been integrated into FIGO 2023 staging system (IIIC1i and IIIC2i). 2
Therefore, accurate prediction of LNM informs decisions on lymphadenectomy extent while minimizing surgical complications without affecting staging. Recent data indicated LNM promotes distant spread through tumor-immune tolerance, 5 partially explained the association between LNM and poor prognosis, emphasizing the need of a predictive model for LNM. Plenty of researches have exhibited the association between LNM and traditional/novel parameters, for example, tumor diameter, histological subtype, myometrial invasion (MI), carbohydrate antigen 125 (CA125), microcystic, elongated and fragmented (MELF) pattern and molecular subtype.3,6,7 In recent years, dozens of predictive models for LNM in EC with diverse performance were built and published,8,9 with some studies evaluating existing models as well.10,11 However, despite abundant models published, there is still no universal predictive model for LNM in EC. Here, we systematically summarized predictive models retrieved from the Web of Science (WOS), evaluating risk of bias, with bibliographic analysis conducted as well, to present advances and inform clinical application of LNM prediction in EC.
Materials and methods
Literature retrieval and data extraction
We searched the WOS for articles reporting models predicting LNM in EC with entries in Supplementary Table 1. The initial search was performed on 18 September 2022 with weekly alerts for newly published models through 1 December 2023. Inclusion criterion was articles building or validating LNM predictive models for EC. Exclusion criteria were as follows: (1) articles discussing LNM risk factors only without predictive modeling; (2) not original researches, for example, meta-analysis and review; and (3) full texts unavailable, for example, conference abstracts or patents. Initial screening for eligibility was independently performed by two authors (H.L. and J.W.) through a systematic review of titles and abstracts based on predefined inclusion and exclusion criteria. Full-text articles passing initial screening underwent further assessment for eligibility by both authors, with any discrepant decisions reconciled through collaborative discussion mediated by a third author (Z.W.). In addition, we have conducted similar searches on other platforms, for example, PubMed and Scopus, to include articles meeting the criteria as possible. Custom data abstraction forms listing variables of interest were created a priori (Supplementary Table 2) and used by the two independent authors (H.L. and J.W.), with any variability in extracted information between reviewers again resolved via consensus discussion. The systematic review adhered to the Preferred Reporting Items for Systematic reviews and Meta-Analyses (PRISMA) statement, with a completed checklist in Supplementary Table 3.
Included articles were categorized per Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis (TRIPOD) statement: Type 1a: Development only; Type 1b: Development and validation using resampling; Type 2a: Random split-sample development and validation; Type 2b: Nonrandom split-sample development and validation; Type 3: Development and validation using separate data; and Type 4: Validation only. 12 Specially, for each article, the LNM rate was calculated from the main database used to develop/validate the model (TRIPOD Types 1, 2 and 4). If the article introduced a new database for validation (TRIPOD Type 3), we extracted the LNM rate of the data sets developing the models. “Recruitment period” and “number of patients” also followed this standard. As for the effectiveness of the model, only those from the data set developing the model were recorded (except for TRIPOD Type 4, with only the validating data set available). Besides, we evaluated whether authors provided accessible clinical applications, for example, nomograms, scoring systems or website-based calculators.
Evaluation of model performance
The Prediction model Risk Of BiAS Tool (PROBAST) was used to assess the risk of bias and concern for applicability per model, using signaling questions in four different domains: participants, predictors, outcomes and analysis. Overall risk was judged as “low” if all domains indicated low risk. “High” overall risk resulted from at least one domain with high risk. “Unclear” overall risk was assigned when at least one domain had unclear risk, with all others having low risk. This scoring system was also applied to the assessment of concerns of study applicability, indicating the extent to which the participants, predictors and outcomes in the study matched the review question. C-statistics were adopted to measure discrimination, reflecting a model’s ability to distinguish patients with versus without LNM.
Statistical analysis
C-statistics were presented using forest plots. Articles were collected in structured forms by IBM SPSS Statistics 26 (SPSS Inc., Chicago, IL, USA). 13 Statistics were conducted by R 4.3.1. 14 Bibliographic analysis in this research was conducted by VOSviewer 1.6.18. 15
Results
Overview of the models
The systematic literature search yielded 64 included articles by 1 December 2023 (Supplementary Table 2), with the selection process shown in Figure 1.
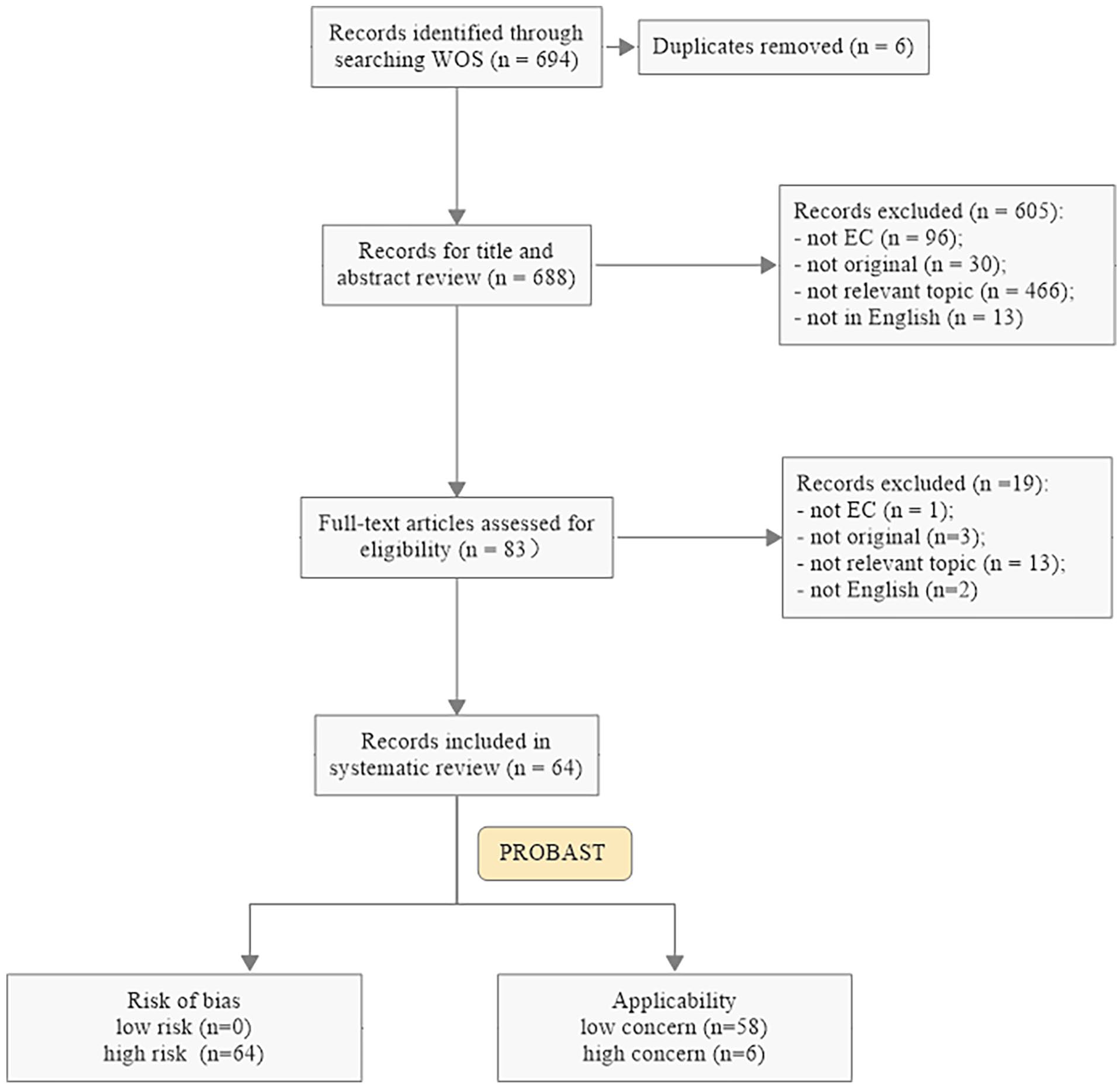
Flow chart of the study selection procedure. EC: endometrial cancer; PROBAST: Prediction model Risk Of BiAS Tool; WOS: Web of Science.
With a recruit period between 1977 and 2022, all the articles were published between 2010 and 2023, with an increase of publications after 2020 (31/64, 48.4%). The most common articles were “development only” (TRIPOD Type 1a, 21/64), followed by Type 4 (13/64) and 3 (12/64). Terminology describing predictive approaches primarily included “model” (37/64) and “nomogram” (6/64). The median number of patients of the articles was 333 (interquartile range (IQR) = 175–524.5), with an obviously skewed distribution observed (skewness = 5.897), resulting mainly from two studies based upon large nation-scaled databases.16,17 The median LNM rate reported was 12.5% (IQR = 10.0%–17.3%), which was absent in two articles.10,18 The minimum (1.0%) was reported by Convery et al. 19 in 2011, whose study included low-grade endometrioid EC (EEC), and the maximum (40.8%) was reported by Asami et al. 20 in 2022, whose study only included EC patients having 10 or more LNs removed (Figure 2).

Summary of characteristics of included articles. (a) Numbers of documents published per year by TRIPOD classification. (b) Distribution of TRIPOD types. (c) Model name terminology usage. (d) Distribution of reported LNM rates. LNM; lymph node metastasis; MRI: magnetic resonance imaging; TRIPOD: Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis.
There were 14.1% (9/64) articles not providing accessibility for clinical application; most (8/9) from model development studies (TRIPOD Types 1–3). The nomogram was the most common approach provided (15/64). Model complexity ranged from simple cutoffs of human epididymis protein 4 (HE4) or CA125 levels 21 to unsupervised machine learning algorithms. 22
Articles developing models: TRIPOD Types 1–3
Majority of included articles (51/64) focused on model development (TRIPOD Types 1–3) (Figure 2(b)). Most common development approaches were “risk stratification” (17/51), “nomogram” (12/51) and “risk score” models (11/51). And traditional clinicopathological parameters remained the mainstream, including serum tumor marker in 43.1% (22/51) articles, MI and grade in 35.3% (18/51), histological subtype and lymphovascular space invasion (LVSI) in 21.6% (11/51) (Supplementary Table 2).
Six articles developed gene signature–based models.18,22 –26. Most (4/6) sequenced tumor samples from their own centers,18,22,23,25 with 4/6 incorporating machine learning.18,22,24,25 In 2017, Ahsen et al. developed a sparse classification algorithm based upon support vector machine (SVM), predicting LNM in EC from an 18-miRNA signature, achieving 100% accuracy in the training cohort, 90% sensitivity and 80% specificity in the testing cohort. 18 In 2018, Kang et al. 23 combined multivariable logistic regression and principle component analysis to predict LNM from genechip in low-risk EEC, reaching an area under curve (AUC) of 0.72, a sensitivity of 100% and a specificity of 42%.
Four studies built models upon radiomics, two utilizing machine learning.27 –30 Yang et al. 30 built a decision tree by machine learning based on RadScore, tumor grade and size, with RadScore calculated from four parameters of diffusion-weighted imaging (DWI) in magnetic resonance imaging (MRI), including short-axis diameter of LN. Performance yielded 93.8% sensitivity, 79.9% specificity and 99.2% negative predictive value (NPV). Analogously, Soydal et al. 28 applied machine learning to predict LNM based on nine texture features from 18F-FDG Positron Emission Tomography/Computed Tomography (PET/CT), attaining 80% diagnostic accuracy.
It is noteworthy that a pioneer research by Feng et al. first combined digital histopathological images with deep learning via multiple instance-learning (MIL) framework (AUC = 0.938; 83.0% sensitivity, 91.1% specificity, 87.2% NPV). Moreover, the histological explanation of the model was identified: enhanced texture, voluminous extracellular mucin, cleft-like structures and elongated angular glandular lumen were associated with LNM in EC, shedding light on further clinical translation. 31
Specially, two articles reported predictive models for para-aortic LNM (PALNM) in EC. Karube et al. 32 defined high-risk patients for PALNM as those with obturator/common iliac LNM ± G3 ± MI ⩾ 1/2 (100.0% sensitivity, 100.0% NPV). Similarly, Turan et al. 33 stratified high risk as non-endometrioid histology, LVSI, serosal spread, adnexal metastasis or pelvic LNM (PLNM)—attaining 94% sensitivity, 53.7% specificity and 98.6% NPV. However, both the models incorporated postoperative elements like LVSI, limiting preoperative utility.
Articles specially validating models: TRIPOD Type 4
Thirteen articles were designed to validate existing models only (TRIPOD Type 4), with a total of 14 models validated (Supplementary Table 4). The most commonly validated models were the Mayo criteria and a nomogram by Bendifallah et al. 34 in 2012, with four and three independent validations, respectively.
Being the most widely applied model for LNM in EC, different variants of Mayo criteria were assessed. Both adopting the conventional Mayo criteria (histology, MI and tumor diameter), Convery et al. 19 only included EEC patients with grade 1/2 and MI ⩽ 50%, using tumor diameter ⩽ 2cm for stratification (NPV 98.2%), while Kang et al. 35 included all histologically confirmed EC (NPV 97.1%). Tuomi et al. 36 adopted more accurate Mayo criteria, with (1) MI stratified by 50% and 66% and (2) adnexal metastasis included, dividing patients into four risk groups for LNM, with only stage I–III EEC patients included in the study, reaching an AUC of 0.781. Among postoperative models validated by Koskas et al., 11 a “Mayo clinical nomogram” was composed of grade, MI, cervical invasion, LVSI and tumor diameter, reaching an AUC of 0.75. However, no identified Mayo variants integrated molecular characteristics, for example, the Cancer Genome Atlas (TCGA) molecular classification.
Three articles by Koskas et al.10,37,38 validated a nomogram built by Bendifallah et al. 34 based on five preoperative factors—age, race, histology, grade, MI and cervical stroma invasion (CSI), achieving an AUC of 0.76–0.83. Besides, the formula and website calculator versions of this model were also accessible, 10 convenient for clinical application.
Performance of the models
Calibration
The C-statistics for the predictive models, as illustrated in Figure 3 (ranging from 0.7 to 0.947), provide a comprehensive overview of their performance. The prediction intervals (PIs) for C-statistics were broad, indicating substantial between-study heterogeneity.

PIs of the C-statistics for different models. PI: prediction intervals (PI).
Risk of bias and applicability
Figure 4 provides a summary of the risk of bias and applicability, revealing an overall high risk of bias across all models.

Risk of bias (a) and applicability (b) assessment of the models based upon PROBAST (weighted with sample size). PROBAST: Prediction model Risk Of BiAS Tool.
For the risk of bias, 73%, 58%, 49% and 99% of the studies were classified as high risk in the fields of “Participants,” “Predictors,” “Outcomes” and “Analysis” (Supplementary Table 5). Notably, “Analysis” received the highest proportion of high-risk classifications, primarily attributed to insufficient events per variable (EPV) or the grading of continuous variables.
Bibliometric analysis
A further bibliometric analysis revealed a relatively weak collaboration network between authors, organizations and countries in models predicting LNM in EC (Figure 5(a)–(c)). Among the 64 articles, the most cited was a stratification system created by Kang et al. 39 (citation count = 11) (Figure 5(d)), composed of clinicopathological parameters, that is, MRI reported MI, extension beyond corpus, enlarged LNs, serum CA125.

Bibliometric analysis of co-authorship of authors (a), organizations (b) and countries (c) and citation of documents (d).
According to the “total link strength” of the 64 articles included, the top five organizations were Institut Curie, Université Pierre et Marie Curie from France, National Cancer Center from Korea, Tenon Hospital and CENTRE HOSPITALIER UNIVERSITAIRE DE REIMS from France. This is consistent with France’s overall high contribution depicted in the country collaboration map (Figure 5(c)). The top five keywords are “endometrial cancer,” “carcinoma,” “lymph node metastasis,” “lymphadenectomy,” and “risk” (Supplementary Figure 1), reflecting the primary research focus in this domain.
Discussion
LNM is associated with a poor prognosis in EC, and lymphadenectomy in the definitive surgery remains essential for evaluating LN status. However, overuse of non-therapeutic lymphadenectomy has led to a significant increase in systemic morbidity, lymphedema and lymphocyst formation. 40 This highlights the need for accurate predictive models to guide personalized decisions on lymphadenectomy in EC.
In many hospitals in China (including ours), the Mayo criteria are still adopted to identify candidates for sparing lymphadenectomy, that is, MI < 1/2, tumor size < 2 cm and G1/2, which are assessable preoperatively by ultrasound/MR and hysteroscopic biopsy. Nevertheless, other parameters, for example, CA125, molecular classification, which are related to LNM3,26 and preoperatively available as well, are promising to provide added value for LNM prediction. Given the outburst of clinical predictive models in recent years, we sought to summarize published models for LNM in EC, aiming to reach an individualized lymphadenectomy strategy with both ideal accuracy and accessibility.
In alignment with trends across predictive model research, articles on predicting LNM in EC are mainly those developing models, that is, TRIPOD Types 1–3, and only a few articles were designed exclusively for validation (Type 4). Specially, in TRIPOD Type 4 articles, those designed to validate models developed by the authors’ teams accounted for a large proportion (Supplementary Table 4), suggesting a possible publication bias. Thus, multicenter prospective studies that validate and directly compare models developed independently are imperative to establish credibility and justify clinical translation.
Most models rely heavily on traditional clinicopathologic factors, with some incorporating common IHC markers like ER and p53, but it is still not enough to reflect advances in molecular biology of EC, especially those in TCGA molecular classification. The broad application of transcriptomics- and radiomics-based models is currently restricted in clinical practice, since there is no general imaging software and genechip for LNM in EC, considering even polymerase epsilon (POLE) sequencing is challenging in many community hospital laboratories. 41 And the increasing application of deep learning in models actually reduces their accessibility to some extent. Thus, balancing predictive performance and accessibility poses an ongoing challenge requiring creative solutions. Moreover, clinical utility should directly inform model development. For instance, incorporating LVSI has questionable real-world value since it is only derivable after surgery. To drive progress, we advocate introducing a preoperative prediction model combining clinical, imaging and essential molecular variables (e.g. TCGA classification) into future guidelines for personalized surgical decisions regarding lymphadenectomy in EC,42,43 consistent with the opinion from the editorial by Jędryka. 44 For instance, ongoing efforts like the ROME study assessing radiomic and molecular profiles hold promise through integrating predictive and prognostic information available prior to definitive treatment. 45
A key priority in developing LNM prediction models is identifying low-risk candidates eligible for lymphadenectomy omission without compromising staging accuracy or forgoing indicated adjuvant therapy. Therefore, minimizing false negative rate is imperative to avoid under-treatment, as Koskas et al.10,11 emphasized. 46 For high-risk cases classified by the model, sentinel lymph node mapping and intraoperative frozen section pathology and/or one-step nucleic acid amplification still enables personalized decisions on nodal evaluation extent (Supplementary Figure 2), as supported by recent opinions.47 –49
According to the theory of tumor metastasis, LNM requires primary tumors to create permissive pre-metastatic niches in draining LNs via secreted growth factors, exosomes, etc.47,50,51 This partially explains that LNM often occurs in cases with MI ⩾ 1/2, tumor diameter ⩾ 2 cm and high grade, where tumors are more likely to have larger volume and more disordered transcriptome, producing above factors more effectively. For instance, during an creative exploration of LNM in EEC by RNA-sequencing, after matching for tumor size, grade and MI, the molecular classification distributions between LNM and non-LNM cases were not statistically different, and a further machine learning method even found no genes were discriminant between the two groups, which seemed counterintuitive and inconsistent with the latest report by Jamieson et al. 3 Eventually, the originally planned hierarchical clustering by the authors was accomplished by introducing a complicated unsupervised clustering analysis. In this work, there seemed to be an over-matching of clinicopathological parameters between LNM and non-LNM cases, factors suggesting intrinsic mechanism of LNM (see above) were regarded as confusing variables and were “matched,” thus bringing new bias. 22 Therefore, future models for advancing personalized surgical decisions will require balancing predictive accuracy, biological consistency and clinical utility with patient safety at the forefront.
Despite the abundance of prediction models that have been published, their clinical applicability seems limited for several concerning reasons. First, nearly all studies had high risk of bias mainly driven by constrained sample sizes (including small EPV). Such sparse cases relative to parameters risk overfitting statistical noise, while underpowering validation efforts can distort generalizability. Furthermore, the models demonstrated only moderate and inconsistent discriminative and calibrated performance overall, indicating significant potential for inaccurate classification and risk stratification applications clinically.
On top of the previous discussion, a large heterogeneity in model performance (as illustrated by wide prediction intervals) was found throughout different studies and populations. Substantial heterogeneity and inconsistency in predictive performance were observed across divergent validation cohorts (wide prediction intervals). Typically, poorer accuracy in external settings prompted developing new models—exponentially multiplying tools lacking standardization. Rather than recurrently constructing models “from scratch,” progressively enhancing earlier proposed tools by integrating new variables or updating statistical approaches upon emerging evidence is advised.
Our study represents the first systematic review and bibliometric analysis specifically evaluating LNM prediction model credibility and accessibility for EC, highlighting predictor distribution, accessibility, performance and risk of bias. However, there exist some limitations in our study. First, due to the inherent diversity of predictive model articles, parameters such as the number of patients, LNM rate and effectiveness of the model had great heterogeneity, confining the value of systematic review.19,20 Large-scale external validation is thus needed to delineate generalizability of existing models. Second, because of the small number of articles included in the systematic review, the bibliometric analysis based upon which was constricted, unable to reveal the evolution of models predicting LNM in EC over time. However, our bibliometric analysis did visually exhibit a relatively isolated collaborative network between authors, organizations and countries in this area (Figure 5(a)–(c)), partly explaining the absence of a universal predictive model for LNM in the era of precision medicine. This re-emphasized the necessity of multicenter/international collaboration.
Conclusion
A number of predictive models for LNM in EC have been developed. Although some exhibited promising performance as they demonstrated adequate to good discrimination, few models can currently be recommended for clinical practice due to lack of independent validation, high risk of bias and low consistency in measured predictors. Collaborations between authors, organizations and countries were weak. Model updating, external validation and collaborative research are urgently needed.
Supplemental Material
sj-csv-7-whe-10.1177_17455057241248398 – Supplemental material for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis
Supplemental material, sj-csv-7-whe-10.1177_17455057241248398 for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis by He Li, Junzhu Wang, Guo Zhang, Liwei Li, Zhihui Shen, Zhuoyu Zhai, Zhiqi Wang and Jianliu Wang in Women’s Health
Supplemental Material
sj-docx-5-whe-10.1177_17455057241248398 – Supplemental material for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis
Supplemental material, sj-docx-5-whe-10.1177_17455057241248398 for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis by He Li, Junzhu Wang, Guo Zhang, Liwei Li, Zhihui Shen, Zhuoyu Zhai, Zhiqi Wang and Jianliu Wang in Women’s Health
Supplemental Material
sj-pdf-1-whe-10.1177_17455057241248398 – Supplemental material for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis
Supplemental material, sj-pdf-1-whe-10.1177_17455057241248398 for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis by He Li, Junzhu Wang, Guo Zhang, Liwei Li, Zhihui Shen, Zhuoyu Zhai, Zhiqi Wang and Jianliu Wang in Women’s Health
Supplemental Material
sj-pdf-2-whe-10.1177_17455057241248398 – Supplemental material for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis
Supplemental material, sj-pdf-2-whe-10.1177_17455057241248398 for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis by He Li, Junzhu Wang, Guo Zhang, Liwei Li, Zhihui Shen, Zhuoyu Zhai, Zhiqi Wang and Jianliu Wang in Women’s Health
Supplemental Material
sj-xlsx-3-whe-10.1177_17455057241248398 – Supplemental material for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis
Supplemental material, sj-xlsx-3-whe-10.1177_17455057241248398 for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis by He Li, Junzhu Wang, Guo Zhang, Liwei Li, Zhihui Shen, Zhuoyu Zhai, Zhiqi Wang and Jianliu Wang in Women’s Health
Supplemental Material
sj-xlsx-4-whe-10.1177_17455057241248398 – Supplemental material for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis
Supplemental material, sj-xlsx-4-whe-10.1177_17455057241248398 for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis by He Li, Junzhu Wang, Guo Zhang, Liwei Li, Zhihui Shen, Zhuoyu Zhai, Zhiqi Wang and Jianliu Wang in Women’s Health
Supplemental Material
sj-xlsx-6-whe-10.1177_17455057241248398 – Supplemental material for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis
Supplemental material, sj-xlsx-6-whe-10.1177_17455057241248398 for Predictive models for lymph node metastasis in endometrial cancer: A systematic review and bibliometric analysis by He Li, Junzhu Wang, Guo Zhang, Liwei Li, Zhihui Shen, Zhuoyu Zhai, Zhiqi Wang and Jianliu Wang in Women’s Health
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
