Abstract
μ-opioid receptor (MOP) plays a critical role in mediating opioid analgesic effects. Genetic variations, particularly those in the MOP gene (Oprm1), significantly influence individual variations in opioid efficacy and side effects across species, highlighting the need for pharmacogenomic research in human and veterinary contexts. This study aimed to identify single-nucleotide variations (SNVs) within Oprm1 in 100 cats of various breeds. Oprm1 spans over 170 kb and consists of five exons that combine to yield three splice variants in the cat Ensembl database. Among these variants, Oprm1-202 is an ortholog of the MOR-1 transcript, which is the most abundant in humans and mice. Oprm1-202 shares 92% and 87% coding sequences (CDS) and 96% and 94% amino acid sequence identity with human and mouse MOR-1, respectively. Phylogenetic trees were constructed from the CDS and amino acid sequences of nine species, including humans, cats, and mice. Both the CDS and amino acid sequences of MOP in cats showed phylogenetic development closer to that of primates than of rodents. Four SNVs were identified in the CDS of Oprm1. One SNV was located in exon 1 and the other three in exon 2 of Oprm1, all of which were synonymous substitutions. Although synonymous mutations generally have a limited functional impact, they may influence splicing and receptor expression. Further research is required to assess the effects of these SNVs on opioid efficacy, receptor expression, and analgesic responses across breeds, considering the potential breed-specific genetic factors in cat species.
Introduction
Opioid receptors, classified into μ, δ, and κ subtypes, are seven-transmembrane G protein-coupled receptors. Among which, the μ-opioid receptor (MOP) is the primary molecular target for clinically utilized opioids, such as morphine and fentanyl. Studies utilizing MOP gene (Oprm1)-knockout mice have shown that the pharmacological effects of opioids are mediated by MOP.1,2 Despite the broad-spectrum application of opioids in pain management, individual differences in the analgesic responses and side effects to opioids have been documented in both humans and animals. These variations are primarily influenced by genetic and environmental factors, highlighting the importance of pharmacogenomics for determining opioid efficacy.3,4
A considerable body of research has focused on the role of single nucleotide variations (SNVs) in the OPRM1 gene, specifically, their impact on analgesic efficacy in humans. 5 For example, the dosage requirements of fentanyl varies according to different OPRM1 genotypes.6,7 Notable SNVs such as A118G and IVS3+A8449G have been associated with the varied dosage requirements of opioids such as morphine, fentanyl, and alfentanil.6–8 More than 700 SNVs have been identified in the human OPRM1 gene and documented in databases such as dbSNP by the National Center for Biotechnology Information and the National Human Genome Research Institute (dbSNP) and a database for gene polymorphisms developed by the National Center for Biotechnology Information and the National Human Genomic Research Institute (www.ncbi.nlm.nih.gov/SNP/). In addition to human studies, genetic variations within Oprm1 have been linked to the functional effects of variants in animal models. For example, in rhesus macaques, Oprm1 C77G has been reported to influence alcohol response and consumption,9,10 pharmacogenetic sensitivity to naloxone,10,11 and stress response and aggression.12,13 In dogs, the Oprm1 C-15A variant is potentially associated with opioid dysphoria. 14 In murine models, Shigeta et al. 4 and Han et al. 15 reported that short tandem repeat polymorphisms and transposable elements in the 5′ flanking and 3′ untranslated regions of Oprm1 in mice correlated with morphine-induced antinociceptive effects among inbred strains, suggesting genetic influence on analgesic variability in response to opioids.
With a growing emphasis on animal welfare, particularly in companion animals, effective perioperative pain management has become a priority. Similar to humans, opioids, such as morphine, buprenorphine, and fentanyl, are used for pain relief in these animals. 16 However, research on the pharmacogenomics of opioid analgesics in veterinary species, despite the evidence of individual variability in antinociceptive responses in cats, is limited. Johnson et al. 17 reported such variability in cats that were administered opioids, suggesting a need to elucidate cat-specific genetic factors influencing opioid responses.
Considering the need to enhance pain management strategies in companion animals, particularly cats, this study aimed to identify SNVs in the feline Oprm1 gene. This investigation sought to advance our understanding of the genetic influence on the behavioral responses to MOP agonists in cats, potentially improving individualized pain management and welfare standards in veterinary practice.
Materials and methods
Animals
A cohort of 100 client-owned cats were examined as part of a preoperative assessment conducted at the Sendai Animal Care and Research Center. The cats were scheduled for elective spaying or neutering procedures. Each animal was determined to be healthy based on a thorough preoperative evaluation, which included a comprehensive physical examination, complete blood cell count, biochemical profile analysis, and screening tests for Feline Leukemia Virus and Feline Immunodeficiency Virus.
The procedures for this study were reviewed and approved by the Institutional Animal Care and Use Committee of the Akita Cerebrospinal and Cardiovascular Center (approval no. 21-07). Written informed consent was obtained from each owner before inclusion of their cats in the study. In addition, the owners provided specific consent for blood sampling and analysis of nucleotide variations in genomic DNA.
Genotyping
Genomic DNA was extracted from whole blood samples using the QIAamp DNA Mini Kit (Qiagen K.K., Tokyo, Japan) according to the manufacturer’s instructions. Nucleotide sequences of the feline Oprm1 gene were identified by direct sequencing with Polymerase Chain Reaction (PCR) products that covered exons 1–4 of Oprm1 of the reference sequences in GenBank and Ensembl (GenBank ACCESSION no. NC_058372.1, REGION: 138328277-138383427; Ensembl Transcript ID no. ENSFCAT00000005362.6). PCR was performed using Pwo SuperYield DNA polymerase (Roche Diagnostics Japan, Tokyo, Japan), and the specific primers for exons 1–4 of the feline Oprm1 gene were as follows: exon 1 (forward, 5′-GAA AAG TCT GGG TGC TCG CTG TT-3′; reverse, 5′-TCC CTT TAC GCC CCT ATT AGC TG-3′), exon 2 (forward, 5′-GCT CAT CTG TAT TTC ATG TTG GTC T-3′; reverse, 5′-CAG CAT GTC GGT CTA TAA ACC CTT-3′), exon 3 (forward, 5′-ATG GCT TTA AGT TAG CTC TGG TCA A-3′; reverse, 5′-TCC CTT AGT TAC CAA TCC ACT TCT A-3′), and exon 4 (forward, 5′-CCC ATA AAT GCC CAC ATA CAG GAA-3′; reverse, 5′-ATA CTT CCA CAG AGA TGG TGC CTT-3′). The cycle sequencing reaction with the BigDye™ Terminator Cycle Sequencing Kit (version 3.1; Applied Biosystems Japan, Tokyo, Japan) was performed according to the manufacturer’s instructions, and the reaction products were purified. Nucleotide sequences of the products were determined using an ABI PRISM 3130xl Genetic Analyzer and Sequencing Analysis software (version 5.4; Applied Biosystems Japan).
Genetic similarity analysis
The coding sequences (CDS) and translated amino acid sequences of the feline Oprm1 were compared with those of humans (Homo sapiens), chimpanzees (Pan troglodytes), rhesus monkeys (Macaca mulatta), squirrel monkeys (Saimiri boliviensis), cattle (Bos taurus), dogs (Canis lupus familiaris), rats (Rattus norvegicus), and mice (Mus musculus). Molecular Evolutionary Genetics Analysis (MEGA) X software (version 10.1.7) was used for evolutionary phylogenetic tree analysis. 18 The CDS and translated amino acid sequences of Oprm1 in these nine species were aligned using ClustalW with the parameters of Gap Open Penalty 15.0 and Gap Extension Penalty 6.66 for Pairwise and Multiple sequence alignments. Phylogenetic trees of the CDS and protein of Oprm1 were constructed using the neighbor-joining method.
Results
The Oprm1 gene spans over 170 kb and consists of five exons that combine to yield three splice variants in the Cat Ensembl database. Among these variants, Oprm1-202 (Ensembl Transcript ID no. ENSFCAT00000005362.6) is an ortholog of the MOR-1 transcript, which is the most abundant transcript in humans and mice. Oprm1-202 shares 92% and 87% CDS and 96% and 94% amino acid sequence identity with human and mouse MOR-1, respectively (Figure 1). Phylogenetic trees were constructed from the CDS and amino acid sequences of nine species, including humans, cats, and mice (Figure 2). The CDS sequences used for phylogenic analysis are shown below: NM_000914.4 (H. sapiens), NM_001033915.1 (P. troglodytes), NM_001032824.1 (M. mulatta), AY665244.1 (S. boliviensis), NM_174408.2 (B. taurus), NM_001314122.1 (C. lupus familiaris), NM_013071.2 (R. norvegicus), and NM_001302793.1 (M. musculus). Both the CDS and amino acid sequences of the feline MOP were closely related to those of companion and domestic animals, such as dogs and cattle, and showed phylogenetic development closer to primates than to rodents.

Homology analysis of CDS sequences in the Oprm1 gene.

Phylogenic analysis of the Oprm1 gene. The phylogenic trees were constructed from (a) CDS and (b) amino acid sequences of the Oprm1 gene in nine species using neighbor-joining method.
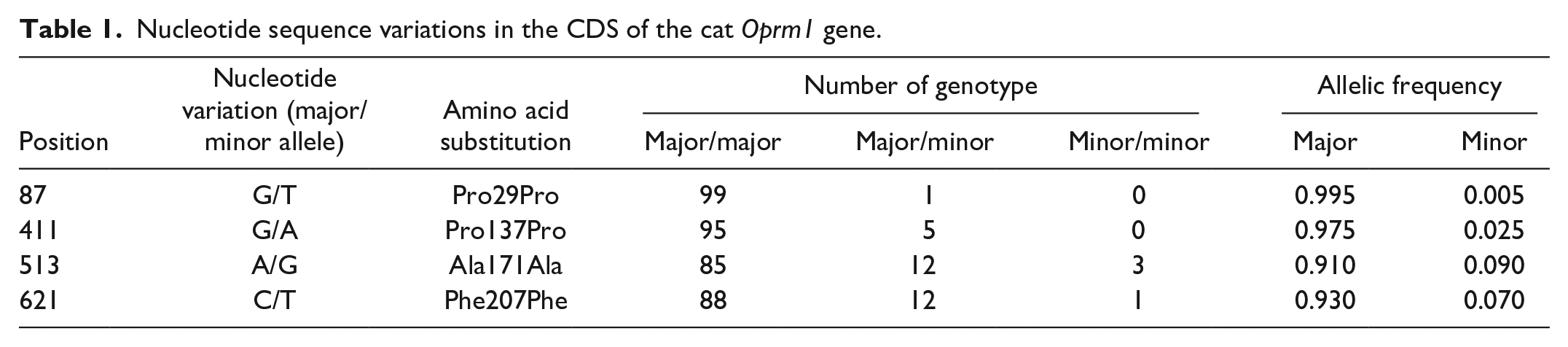
In the cohort of 100 cats analyzed in the present study, four SNVs were identified in the CDS of Oprm1 (Table 1). One SNV was located in exon 1 and other three in exon 2 of Oprm1, all of which were synonymous substitutions.
Nucleotide sequence variations in the CDS of the cat Oprm1 gene.
Discussion
To our knowledge, this is the first study identifying SNVs in cats. The study sample consisted of 100 cats, including Abyssinians, American Shorthairs, Bengals, Chinchillas, Japanese cats, Maine Coons, Munchkins, Russian Blues, Scottish Folds, and various other mixed breeds. Given the extensive diversity of cat lineages, including numerous purebred lines, further analyses of untested and mixed breeds may reveal additional polymorphisms.
The four SNVs identified in this study were synonymous variants with no amino acid substitutions. Synonymous mutations associated with diseases are likely to be much rarer than disease-associated nonsynonymous changes, suggesting that only few synonymous mutations may have a significant impact on splicing. However, there are numerous examples of synonymous mutations that induce diseases by disrupting the splicing process. 19 In companion cats, opioids such as buprenorphine, morphine, and fentanyl are used for perioperative, cancer, and wound pain management. 16 Variations in the nucleotide sequence within CDS regions and SNVs located in untranslated and intronic regions (per the Ensembl database) may cause structural changes in the gene and alter MOP expression, potentially leading to interindividual differences in clinical opioid efficacy. Because this study did not examine the relationship between opioid responsiveness during surgery and the identified single SNVs, it is implausible to conclude from these study findings alone that these SNVs influence gene expression, receptor availability, or individual responses to opioids.
It remains uncertain whether the SNVs identified in this study resulted from the sequence differences between specific purebred lineages or if these variations occurred at some point and subsequently spread across individuals. Statistical analysis revealed no significant differences in allele frequency associated with breeds (data not shown). This lack of significance may be attributed to the small sample size of each breed in this study and the ambiguity in identifying purebred lineages within crossbred felines.
Further investigations are required to explore the relationship between single SNVs and intraoperative opioid responsiveness across various breeds beyond those examined in this study. Understanding breed-specific responses to opioids is critical because genetic diversity among breeds may influence the pharmacokinetics and pharmacodynamics of these drugs. By expanding data collection to include a broader range of breeds, researchers can better assess the generalizability of SNV effects on opioid responsiveness, thereby developing more personalized and effective pain management strategies in cats.
The focus and findings of this study, examining SNVs associated with the efficacy of MOP agonists in pain management in cats, has the potential for advancing translational research with applications in human medicine. In cats, the MOP CDS and protein are more closely aligned with those found in primates than in rodents (Figures 1 and 2). Therefore, given the genetic similarities between humans and cats, information obtained from opioid-based pain management in cats may be beneficial for human opioid pain management, and insights from human opioid protocols may support the refinement of opioid-based pain management in cats. In addition, researchers can gain insight into the genetic determinants of pain perception and opioid responsiveness by investigating how SNVs influence the effectiveness and pharmacodynamics of MOP agonists in cats.
In conclusion, four SNVs were identified in the coding sequences of MOP across 100 cats representing several breeds, all of which were synonymous substitutions. Further investigations are necessary to elucidate the relationship between opioid responsiveness in cats and the SNVs identified in this study. These findings have the potential to advance personalized pain management strategies in both veterinary and human healthcare.
Footnotes
Acknowledgements
We would like to thank Editage for language editing.
Author contributions
Kazumasu Sasaki: Study conception and design, investigation, data interpretation, literature research, and manuscript drafting. Shinya Kasai: Study conception and design, investigation, data analysis, data interpretation, literature research, and manuscript drafting. Junko Hasegawa: Investigation. Tatsuya Ishikawa: Interpretation and literature research. Kazutaka Ikeda: Interpretation and literature review. All authors contributed to and approved the submitted manuscript.
Data availability statement
The original contributions of this study are included in the article. Further inquiries can be directed to the corresponding author.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Grant-in-Aid for Scientific Research from Japan.
Society for the Promotion of Science (19KK0225), intramural research funding from the Akita Cerebrospinal and Cardiovascular Center (21-01), and Sendai Animal Care and Research Center Fund.
