Abstract
Migraine is a neurological disease characterized by severe headache attacks. Combinations of different genetic variations such as copy number variation (CNV) in a gene and microRNA (miRNA) expression can provide a holistic approach to the disease as a pathophysiological, diagnostic, and therapeutic target. CNVs, the Cholinergic Receptor Nicotinic Alpha 7 Subunit (CHRNA7) gene, and expression of gene-targeting miRNAs (hsa-miR-548e-5p and hsa-miR-3158-5p) in migraineurs (n = 102; with aura, n = 43; without aura, n = 59) and non-migraines (n = 120) aged 15–60 years, comparative, case–control study was conducted. Genetic markers were compared with biochemical parameters (BMI, WBC, Urea, GFR, ESR, CRP, HBG). All analyzes were performed by quantitative Real-Time PCR (q-PCR) and fold change was calculated with the 2−ΔΔCT method. The diagnostic power of the CHRNA7 gene, CNV, and miRNAs were analyzed with the receiver operating curve (ROC). CHRNA7 gene and hsa-miR-3158-5p are down-regulated in migraineurs and the gene is controlled by this miRNA via CNVs (p < .05). Both deletion and duplication were detected in patients with migraine for CVN numbers (p = .05). The number of CNV deletions was higher than duplications. When CHRNA7-CNV-hsa-miR-3158-5p was modeled together in the ROC analysis, the area under the curve (AUC) was 0.805, and the diagnostic power was “good”. In migraineurs, the CHRNA7 gene can be controlled by hsa-miR-3158-5p via CNVs to modulate the mechanism of pain. These three genetic markers have diagnostic potential and may be used in antimigraine treatments.
Introduction
Migraine is a complex and neurological disease characterized by recurrent headache attacks. 1 The disease reduces the quality of life of the individual, causes loss of workforce and economic burden, and is a compelling disease that affects all segments of society. 2 Migraine is divided into two major groups (with and without aura). 3 It is unclear whether these two groups are separate disorders or share the same history. The accepted view is that the aura precedes migraine pain. The findings suggest that the inheritance of migraine with aura, changes in brain structure, and treatment responses are also different. 4 Current treatments for migraine are effective in a proportion of patients and have off-target effects. Molecular tools offer a sensitive biomarker-focused approach to develop a more effective targeting rather than a systemic therapy. Many researchers are investigating microRNAs (miRNAs) with epigenetic markers and therapeutic potential and are trying to identify miRNA-based options that may overlap with the disease’s clinical course. However, since the suggested biomarkers are not specific to migraine, there is no approved migraine-specific marker. Biomarker research aims to obtain resources with high specificity, early diagnosis, and explanatory potential for different migraine stages. The biomarker potential of non-coding RNAs and their relationship with migraine have been demonstrated. Despite its dynamic nature, the role of miRNAs in migraine is important. miRNAs are involved in the neuronal and immune system, participate in pain signaling, and can modulate the pain process. Circulating miRNA levels can be regulated by analgesics. This shows that miRNAs can change drug effects and be effective in preventing pain. 5 In addition, the search for gene transcripts such as the CHRNA7 gene (https://omim.org/entry/118511), which encodes proteins effective in the immune system, and the detection of targeting miRNAs will also help to reveal the pathogenesis due to miRNA-mediated control of gene expression. Modulation of the cholinergic system reduces the systemic inflammatory response, and activation of this pathway in migraine treatment may regulate neurotransmitter expression through α7nAChR channels. 6 The possible role of CHRNA7 in this metabolism by the regulation of miRNA suggests that it has an effect on the migraine process. The fact that CHRNA7 contains the 15q13.3 hotspot copy number variations (CNV), which we have shown to be related in migraine patients, 7 and the detection of miRNAs targeting these regions will add a new dimension to the diagnosis and treatment approaches of migraine. Among the universal mechanisms, errors in mitosis and meiosis cause deletions and duplications at the chromosomal level. CNVs occur in many places in the human genome, whether or not they are associated with the disease. Variants that are intermediate between major chromosomal aberrations, minor insertions, and deletions, ie deletions, and duplications, are termed CNV. 8 The number of different base pairs due to CNV between two random individuals is 100 times higher than that of SNPs. 9 CNVs can regulate gene expression and alter gene dosage with insertions and deletions that can be the source of genetic diseases. Deletions may result in the manifestation of a recessive allele, or their interaction with gene regulatory elements may directly result in altered gene expression. CVNs can cause more complex diseases in combination with environmental factors.10,11 In particular, CNVs on CHRNA7 gene may be the target and cause of pain modulation. The regulation of these CNVs by miRNAs may indicate their role in the pathogenesis of migraine.
In this study, we evaluated the expression of the CHRNA7 gene in patients with migraine, comparative quantitative measures of CNVs on the gene, and the expressions of two miRNAs targeting these CNVs. We analyzed the contribution of the gene-CNV-miRNA axis to the pathogenesis of migraine and their potential as a biomarker.
Materials and methods
The study protocol and blood sampling
The study group consisted of 220 individuals, 102 patients with migraineurs (43 with aura and 59 without aura), and 120 nonmigraineurs, between the ages of 19–60. The number of subjects in the study group was determined by power analysis. The study was approved by Gaziantep University Clinical Research Ethics Committee with the number 2020/275. Patient and healthy volunteers were recruited from individuals who applied to Gaziantep University Neurology Clinic and were followed up. The patients were diagnosed according to The International Classification of Headache Disorders 3rd Edition (ICHD-3) diagnostic criteria. Healthy individuals did not have a family history of migraine or any neurological disorder. Covariates in migraine patients; Biochemical parameters (BMI; body mass index, WBC; white blood cell, Urea, GFR; glomerular filtration, ESR; erythrocyte sedimentation rate, CRP; C-reactive protein, HBG; hemoglobin), age, sex, child number, income status, education, alcohol, and cigarette use, medical history, family history for any disease, drug use for headache, dominant hand, types of headache, duration of headache attacks, pain side, pain attack origin, pain intensity, autonomic findings, phonophobia, photophobia, migraine frequency, mood changes, age of onset and Migraine Disability Assessment Test (MIDAS) score. 12
CNV specific miRNA selection on CHRNA7 gene
15q13.3 microdeletion syndrome is associated with many neurological diseases such as epilepsy, schizophrenia, autism, attention deficit hyperactivity disorder. This region is genomically unstable and prone to recombination. CHRNA7, one of the seven genes (KLF, TRPM1, FAN1, MTMR10, MIR211, OTUD7A) covering this region, is likely responsible for the disease. The fact that neuroinflammation has an important role in the pathogenesis of chronic migraine suggests that deletion or additions in this unstable region may be associated with the disease. 13 If instability has the power to distinguish patients and healthy people, it will gain a new biomarker as a diagnosis and treatment target. Demonstrating the effect of miRNAs targeting CNVs, as a continuation of our study in which we demonstrated this instability, brought to mind the question of how the CNV-miRNA couple and the control of the CHRNA7 gene contribute to migraine. 7 CHRNA7 (α7nAChR) has been investigated in chronic migraine by in vitro and in vivo studies. 6 However, migraine types and miRNA-CNV linkage were not shown. miRNAs are being investigated as potential biomarkers and therapeutic targets. These dynamically structured epigenetic molecules have a wide range of effects on their own, and the variations of the regions they target play an important role in determining their behavior (such as SNP, mutation, methylation, and CNVs in their targets). Therefore, in addition to the resources, we obtained through bioinformatics analyzes in our previous study, 7 we searched for miRNAs targeting CNVs using various databases and algorithms.
Bioinformatic analyzes for miRNAs targeting CNVs
In the first step, the mRNA sequence of the CHRNA7 gene was detected in the NCBI database (https://www.ncbi.nlm.nih.gov/gene/) (Reference Sequence: NM_000746.6) for miRNAs targeting CNVs. In the second step, miRNAs targeting the CHRNA7 gene were detected using the mirDB database (https://mirdb.org)
14
and validated in the Target Scan database (https://www.targetscan.org/vert_72/) (Release 7.2: March 2008) (Accession Date: 02 May 2021).
15
Afterward, the 15q13.3 region and the region covered by the CHRNA7 gene were determined in the DECIPHER V11.4 database (https://www.deciphergenomics.org/) (Accession Date: 02 May 2021).
16
The target sequences of miRNAs were detected using the NCBI Blast menu (https://blast.ncbi.nlm.nih.gov/Blast.cgi) for specific matching. On the first identified CHRNA7 mRNA sequence, the last obtained matches were screened to identify miRNAs targeting CNVs in the gene. In this determination, among the miRNAs obtained in miRDB, those with high target scores were chosen (Figure 1). The workflow of CNV-miRNA-CHRNA7 discovery analysis.
Total RNA and DNA preparation
Fresh DNA and RNA were obtained from all blood samples of patients and control individuals. DNA extraction was performed according to the following manufacturer's protocol (PureLink™ Genomic DNA Mini Kit, Thermo Fisher Scientific, Waltham, USA) from 1 mL of whole blood in EDTA tubes. Total RNA extraction was performed using Trizol and modified 17 for 500 μL whole blood. The DNA and RNA (in RNA later solution) were aliquoted and stored at −80°C for further analysis. The quantity and purity of DNA and RNA were traditionally measured by spectrophotometer (Multiskan GO) (Thermo Scientific, San Diego, USA).
Reverse transcriptase PCR (RT-PCR) for complementary DNA (cDNA) analysis
Complementary DNA were obtained using the Qiagen RT kit (QuantiTect®, ReverseTranscription Kit, Germantown, MD, USA) according to the manufacturer’s protocol. A 12.5 μL reaction mix was prepared to contain 10 ng total RNA, 5X miScript HiSpec Buffer, 10X miScript Nucleics Mix, miScript Reverse Transcriptase Mix, and RNAse free water. Thermal cycling conditions were performed on an automatic Thermal Cycler (Veriti, Applied Biosystems, USA) for 30 min at 18°C, 30 min at 40°C, and 5 min at 85°C. cDNAs were immediately placed on ice and stored at 20°C until expression.
Quantitative-comparative real-time PCR (qPCR) analysis
Analysis for gene expressions was performed on Rotor-Gene-Q (Qiagen, Germantown, MD, USA) and SYBR Green technology. ACTB gene (Hs_ACTB_1_SG, [NM_001101], Qiagen) was used for standardization of CHRNA7 gene (Hs_CHRNA7_1_SG QuantiTect Primer Assay (QT00074732)), and reference human RNA was used for RNA control (Life Technologies; AM7155). 20 μL PCR reaction was prepared. The reaction mix contained 5 μL of RT-PCR sample, 2X SYBR Green Universal PCR Master Mix (Qiagen), and 900 nM of each specific primer (Qiagen). Thermal cycler condition; pre-incubation at 96°C, incubation for 10 min at 95°C for 1 cycle, 40 cycles at 90°C for 20 s, at 60°C for 25 s, and 72°C for 35 s. All reactions were performed in triplicate, and ΔΔCT data were used for analyzes, and fold change values were calculated according to the 2−ΔΔCT method. 18
Comparative-quantitative CNVstandardization
CHRNA7 gene (Thermo Fisher Scientific, USA) CNV analyzes were standardized according to the reference TERT gene (4403316, Applied Biosystem, Life Technologies, Pleasanton, CA) and TaqMan kit protocol (Applied Biosystem, Life Technologies, Pleasanton, CA) using by Rotor-Gene Q (Qiagen). A total of 10 μL of the reaction mixture for each sample; 2X master mix, 20X TaqMan copy number Assay, 20X TaqMan Copy Number Reference Assay, and RNAse-free water. For each sample, 10 μL reaction mixture was pipetted into 0.2 mL (4-strip) PCR tubes. 8 μL of diluted DNA samples were added to the tubes. Thermal cycling conditions; 1 cycle of 10 min at 90°C, 25 s at 95°C, and 35 cycles at 72°C for 50 s (Rotor-Gene Q, Qiagen). All samples were run in triplicate. Relative quantitation analysis was performed between an unknown sample (CHRNA7-FAM probe) and a calibrator (TERT-VIC probe) to determine the copy number according to manufacturer kit protocol. Two assays were combined in a single reaction and analyzed by Real-Time PCR. Relative quantitation calculated with ∆∆Ct method. 18 Several calculations were performed. The measured Ct values were subtracted for the FAM-labeled test sample and the VIC-labeled reference assay (RNase P) (CtTest − CtReferance assay). The same calculation was used for the calibrator sample. The ΔCt value obtained for the calibrator was subtracted from the ΔCt obtained for the test and the ΔΔCt were calculated (ΔCtTest-ΔCTCalibrator = ΔΔCt for each sample). Finally, copy numbers were determined with 2−ΔΔCt x 2 (copy number of calibrator). 19 Copy number was evaluated as <2; deletion, 2; neutral, and >2; duplication.
Biostatistical analyses
The number of subjects in the study was determined with G-Power 3.1. In order to determine the difference of 0.6 units between groups using retrospective data, the required minimum sample size was calculated as 100 individuals in each group under 5% Type I error and 80% power conditions. The normality of the data and sex differences were evaluated with the Shapiro-Wilk test. The data were analyzed using the independent t-test when the distribution was normal and the Mann–Whitney U test when the distribution was not. Kruskal-Wallis and Dunn's multiple comparison test was used to compare more than two groups. Differences in miRNA and CHRNA7 expression, CNV variation were identified using a one-way ANOVA with Bonferroni correction for multiple comparisons at a cut-off α = 0.05. Diagnostic values of the CHRNA7 gene, CNV, and miRNAs were evaluated by receiver operating characteristics (ROC) analysis. The diagnostic Powers of miRNAs were evaluated according to the rating of Hosmer and et al. 20 Correlation analyzes were performed to examine the effects of CHRNA7 gene expression, CNV and miRNA expressions on migraine, and comparative contributions of the CNV-miRNA-CHRNA7. Logistic regression analysis was performed to determine the diagnostic values of these parameters together. The heat mapper was used to calculate the heatmap. 21 The significance value was p < .05. The SPSS software version 22.0 was used for the analyses (IBM Corp. Released 2013. IBM SPSS Statistics for Windows, Version 22.0. Armonk, NY: IBM Corp).
Results
Population data
The mean ages of migraineurs and nonmigraineurs were 34.39 ± 10.03 and 34.52 ± 7.79, respectively (p = .703). There was statistical significance between the groups depending on the difference in sex numbers (χ2 = 21.120, df = 1, 95% CI = 1.871–5.924, OR = 4.830, p < .0001). Migraineurs consisted of 90 females (88.24%), 12 males (11.76%), and nonmigraineurs 73 females (60.83%) and 47 males (39.17%). WBC, ESR, and HBG were significant between groups. Wbc and HBG were found to be low (P WBC = 0.025, P HBG = 0.027) and ESR to be high in migraineurs (p = .007). 7 The high ESR was associated with the high number of females in migraine patients.
Detection of miRNAs targeting to CNVs on CHRNA7 gene
After the mRNA sequence of the CHRNA7 gene was determined, the miRNAs targeting this gene were scanned with algorithms, and 85 miRNAs with target scores ranging from 50-92% were detected. Then, the 15q13.3 hotspot (15:31,923,438–32,173,018) and the CHRNA7 gene were determined (15:30,618,103–32,153,206). Target screening was performed for 85 miRNAs on this sequence and two mature miRNAs with high target scores were selected (hsa-miR-548e-5p (Score = 92%) (Gene ID: 100,313,921) and hsa-miR-3158-5p (Score = 91%) (Gene ID: 100,422,900)). Using target specificity algorithms of miRNAs, it was determined that hsa-miR-548e-5p is specific for two different positions on CHRNA7 (374-380 and 1712–1718), while hsa -miR-3158-5p interacts with one position (791–798) (Figure 2). In silico analyzes also indicate that these miRNAs can be used in gene regulation by targeting CNVs on the gene and in the pathogenesis and diagnosis of migraine disease. In silico analysis for study design. 1-CHRNA7 mRNA sequence detection for target mature miRNA sequence search. 2- Detection of miRNAs for target CHRNA7 gene. 3- CNV search on 15q13.3 hotspot. 4- Sequences of Mature miRNAs searched in BLAST database. 5- Match between the CHRNA7 mRNA sequence and two selected mature miRNAs.
Evaluation of CNV, CHRNA7, and miRNA expressions
Expression analysis of CHRNA7 gene, CNV, and two mature miRNAs for groups.

*Statistically significant p < .05. CHRNA7 gene, CNV numbers, and mature hsa-miR-3158-5p are significant according to the independent sample t-test.
SD; standard deviation, SE; standard error.
The Mann–Whitney U test was used to reveal differences in CHRNA7, CNVs, and miRNAs for demographic, clinical, and biochemical parameters between groups. As a result, a statistically significant difference was found between the two groups for parameters (Figure 3). miRNA and CHRNA7 expressions, CNV, and sex effect between the groups. CHRNA7, hsa-miR-3158-5p, CNV, and sex differences were statistically significant in migraineurs according to the nonmigraineurs. The significance level is p < .05. hsa-miR-548e-5p were not associated with migraine disease (p > .05).
The relationship between hsa-miR-3158-5p, which was statistically significant, and biochemical parameters (BMI, WBC, Urea, GFR, ESR, CRP, HBG) was evaluated by Spearman's Rank Correlation analysis (Figure 4). Correlations were considered significant at the p < .05 and p < .01 levels. Accordingly, although hsa-miR-3158-5p showed a positive correlation with WBC, urea, GFR, ESR, and CRP, and negatively with BMI, it was not statistically significant (p > .05). However, a statistically significant negative correlation was detected between hsa-miR-3158-5p expression levels and HBG ( p = .001). In addition, no correlation was found between CHRNA7 gene expression levels, CNVs and biochemical parameters (p > .05). WBC (p = .025), ESR (p = .007), and HBG (p = .027) are statistically significant in migraineurs compared to nonmigraineurs.
7
No correlation was detected between CHRNA7, CNV, miRNA measurements, and covariates associated with migraine (p > .05). There was no correlation between CNVs and CHRNA7 gene expression, MIDAS score, and migraine types (with or without aura) (p > .05). Likewise, no correlation was detected between MIDAS scores, migraine with and without aura, and hsa-miR-3158-5p expressions (p > .05). Correlation analysis for hsa-miR-3158-5p and biochemical parameters. A Spearman's rank-order correlation was run to determine the relationship between hsa-miR-3158-5p expression levels and biochemical parameters in 102 (N) migraineurs. There was a strong, positive, or negative correlation that was statistically significant between miRNA expressions and some parameters. A statistically significant negative correlation was found between hsa-miR-3158-5p expression level and HBG. BMI; body mass index, WBC; white blood cell, GFR; glomerular filtration rate, ESR; erythrocyte sedimentation rate, CRP; C-reactive protein, HBG; hemoglobin, N; the number of individuals.
Diagnostic values of CHNRA7, CNV, and hsa-miR-3158-5p in migraine disease
To explore the possibility that the CHRNA7 gene, CNV, and miRNAs could serve as novel and potential biomarkers for migraine disease, ROC analysis was performed in two recruited cohorts. CHRNA7 and CNVs have the power to discriminate migraineurs from nonmigraineurs. Accordingly, the AUC values for CHRNA7 and CNVs were statistically significant and were 0.731 (Specificity = 80.8, Sensitivity = 66.7, Criterion ≤19.79, p < .001) and 0.669 (Specificity = 75 Sensitivity = 70.6, Criterion ≤1, p < .001), respectively. CHRNA7 gene expression and CNV variations have the potential to be a biomarker that can be used to distinguish patient individuals from healthy individuals. Although hsa-miR-3158-5p is statistically significant (p = .044), its diagnostic value is fair (AUC = 0.578).
Multiple logistic regression analyses of miRNA predictor variables for association with migraine disease.
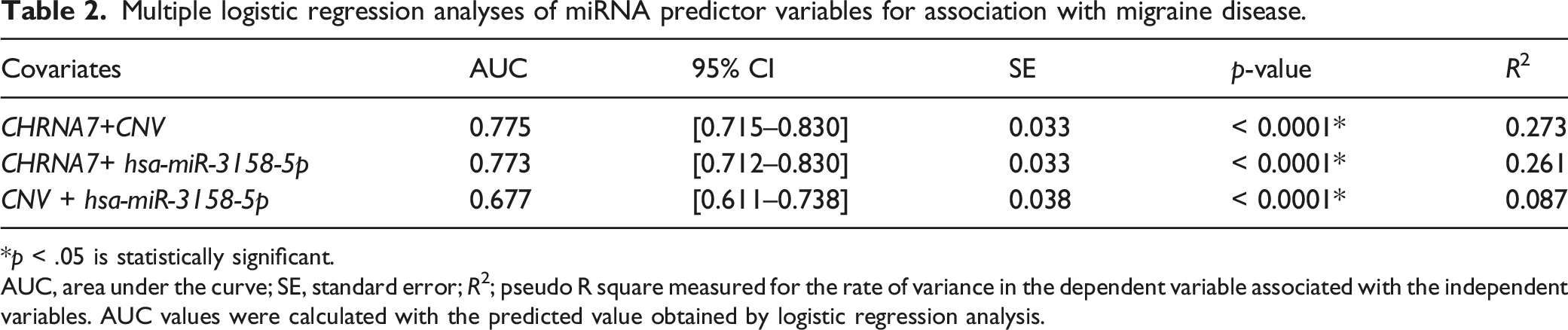
*p < .05 is statistically significant.
AUC, area under the curve; SE, standard error; R2; pseudo R square measured for the rate of variance in the dependent variable associated with the independent variables. AUC values were calculated with the predicted value obtained by logistic regression analysis.
To increase the diagnostic value of the three covariates, we performed ROC curve analyzes with their combinations. CHRNA7 (Coefficient = −0.066, SE = 0.011, OR = 0.935, 95% CI = 0.914–0.960, p < .0001), CNV (Coefficient = −0.742, SE = 0.222, OR = 0.480, 95% CI = 0.310–0.740, p = .0009), and hsa-miR-3158-5p (Coefficient = −0.050, SE = 0.018, OR = 0.951, 95% CI = 0.920–0.985, p = .0049) were statistically significant in the overall fit model when tested by logistic regression analysis (p < .0001, R2 = 0.293, cut-of value p = 0.5, AUC = 0.805, SE = 0.031, 95% CI = 0.746–0.855) (Figure 5). When CHRNA7, CNV, and hsa-miR-3158-5p were modeled together, the diagnostic value was found to be “good”. These results indicate that the combination signature has good potential diagnostic value in distinguishing migraineurs from nonmigraineurs. It is possible to use these 3 predictive variables to predict disease according to ROC curve analysis. Diagnostic value of CHRNA7, CNV, and hsa-miR-3158-5p in migraine disease. AUC values indicate 2−ΔΔCT. The diagnostic value of three covariates was determined as good (AUC = 0.805) according to logistic regression analyzes.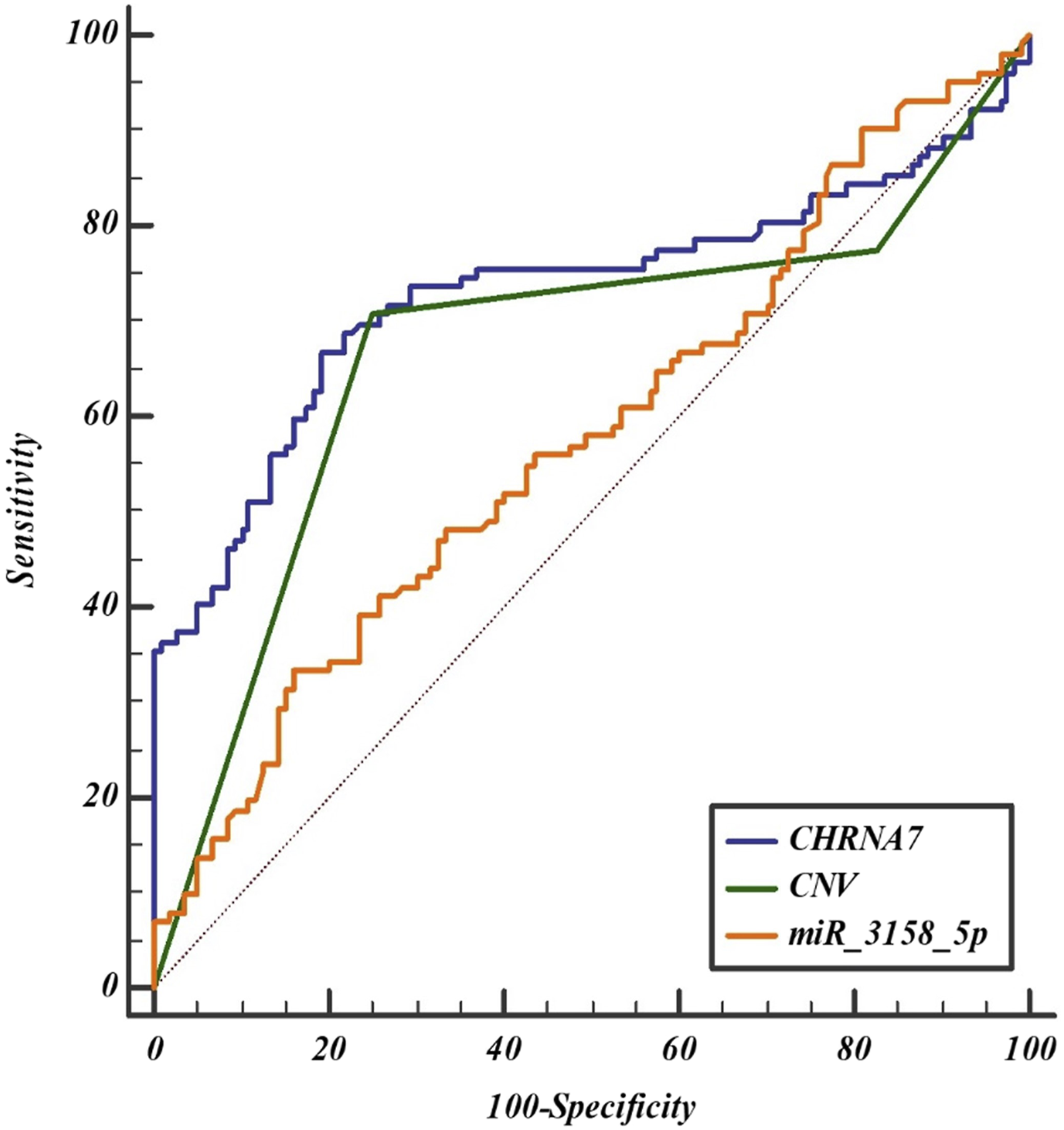
Discussion
Migraine is a common neurological disorder. The diagnosis of migraine is made by the clinical reflections of the symptoms accompanying the headache attacks. Migraine accompanied by sensory and motor disturbances manifests itself with and without aura. Since migraine is a multifactorial disease and causes increasing disability and loss in quality of life, it is important to define the mechanisms and factors associated with the disease.
The multifactorial nature of migraine challenges researchers when investigating both its pathogenesis and therapeutic targets. The absence of a primary culprit and the effect of covariates can create uncertainty in the modulation of the genetic basis of the disease. Many studies demonstrate the role of genetic factors in the pathogenesis of migraine. 22 The effect of epigenetics in neurological diseases is known and partially responsible for the chronification of migraine. 23 Genome-wide association analyzes (GWAS), the most common of these genetic studies, identified 13 migraine-associated loci. 24 The detection of these loci provides an enormous resource for miRNA research. The availability of known gene targets facilitates miRNA screenings. Thus, the detection of miRNA-target gene expressions helps to explain the pathogenesis of the disease and reveals their potential for use as biomarker studies and therapeutic targets.
Apart from known genetic rules, DNA-independent epigenetic approaches also have potential as a diagnostic and therapeutic target in many diseases including migraine. As an epigenetic tool, miRNAs are small non-coding RNAs (∼22 nt) and although mature miRNA (∼7–8 nt) sequences are short, they have great implications as they specifically regulate gene expression. 25 These epigenetic tools can be informative when they overlap with the clinical course of the disease. Therefore, covariants in the migraine clinic can modulate the expression of DNA and their association with miRNAs may provide a better understanding of the pathogenesis of migraine. In addition to the effects of miRNAs in the pathogenesis of the disease as a whole, it may help to elucidate their significance in migraine with and without aura and whether these types occur by the same or different mechanisms. Indeed, studies show the importance of miRNAs in migraine. In a study examining the severity of the disease episodic and chronic, miRNA-mediated biomarkers were investigated in the transition between these two stages in the effect of drugs. 26 In another study, a high level of calcitonin gene-related peptide (CGRP), whose therapeutic target and biomarker potential was recently investigated, was associated with downregulation of miR-382-5p after detoxification in chronic migraine patients using medication overdose. Another research group showed that hsa-miR-34a-5p and hsa-miR-375 in saliva can be used as potential biomarkers in migraine pain. 27 In a study showing the status of dynamic miRNAs during attacks, it was found that 32 of 372 miRNAs were expressed differently in migraine patients and 4 of them were associated with migraine without aura. 5
There is evidence to suggest that activation of the a7nAChR may reduce the neuroinflammatory response. 28 Neuroinflammation has an important role in the pathogenesis of chronic migraine. However, the effect of α7nAChR in chronic migraine is not clear. α7nAChR is one of the ligand-gated ion channels and participates in neurotransmitter modulation and is thought to have a neuroprotective effect. 29 Studies are showing that antagonists of this receptor are effective in neuropathic pain. 30 In a study investigating the role of α7nAChR in rats with a chronic migraine model, it was found that the mechanical threshold was increased and the pain was relieved by activating the ligand. 6 α7nAChR is encoded by CHRNA7 gene. In a study involving the CHRNA7 gene, the pain-relieving properties of a medicinal formula called Fufang Danshen (FFDS) used in Chinese medicine were investigated. 31 In this bioinformatics study, 45 therapeutic targets for 10 pain disorders were investigated, and all of the targets were directly or indirectly associated with pain disease proteins. CHRNA7 is one of these proteins. In our study, we suggest that the CHRNA7 gene may be effective in modulating pain in patients with migraine. However, it should be remembered that many factors can affect gene expression simultaneously or separately. 15q13.3 hot spot CNVs, whose effects we have investigated in migraine patients, are also known as the cause of microdeletion syndrome in which the CHRNA7 gene is involved. Therefore, they may have a functional role in the expression of the gene. In the relative analysis, CNV gains and losses were determined. Regarding the copy number of the TRESK K2P potassium channel (KCNK18) gene in patients with migraine, in the closest study to our study, no alteration was found in patients and controls. 32 In conclusion, we predict that CNV alterations are not effective through the CHRNA7 gene in migraine patients. The researchers reported that loss and gains in the CHRNA7 gene decreased α7 nAChR-associated calcium flux. The source of the deletion is the change in gene dosage (haploinsufficiency) caused by the loss of a functional allele, while the gains show increased ER stress due to increased expression of CHRNA7 mRNA and increased expression of nAChR-specific ER chaperones. The clinical reflection of this situation is neuropsychiatric diseases, cognitive deficits, 33 motor delays, hypotonia, schizophrenia, and specific phenotypes that vary from person to person.34,35 No other copy variation was detected in CHRNA7. Therefore, it can be thought that the gains and losses in the gene are pathogenic and the penetration is variable. 36 In conclusion, although CHRNA7 is not under CNV control, it is low expressed in migraineurs. If so, could miRNAs regulate the expression of this gene via CNVs? The database we searched for CNVs reports possible pathogenicity and association with pain sensitivity of duplicate CNVs in this gene (DECIPHER). Our results suggest that miR-3158-5p may have a function in regulating CHRNA7 gene expression in migraineurs. The findings suggest that hsa-miR-3158-5p may have a function in regulating CHRNA7 gene expression in migraineurs. The findings suggest that hsa-miR-3158-5p may have a function in regulating CHRNA7 gene expression in migraineurs. CNV-based genotyping can be performed to elucidate the effect of duplications associated with gene dosage on decreased CHRNA7 gene expression and miRNA expressions.
The association of miRNAs with biochemical parameters may be reflected in the pathology of the disease. This result may be an advantage in miRNA-mediated diagnosis. Studies have shown that changes in hemoglobin levels are associated with migraine attacks. 37 The low HBG level we experience in migraineurs may be a result of iron deficiency. On the other hand, there are also studies in which HBG controls miRNA levels. 38 Neuroinflammation increases after autophagic reactions triggered by intracerebral hemorrhage. In the study conducted on the role of miRNAs in this microglia-mediated mechanism, a relationship was found between the expression of miR-144, the mTOR target, and activation of autophagy and inflammation of microglia. The researchers suggested that mTOR exerted an effect on the reduction of microglia inflammation as a result of promoting the expression of miR-144 in microglia treated with HBG. 38 There is limited evidence for the role of microglial involvement in migraine. It is thought that neuroinflammation can be reduced by activating the a7nAChR receptor. 39 However, the relationship between the receptor and neuroinflammation is not clear. Downregulation of the CHRNA7 gene reduces the activation of the pathway. It is known that neuroinflammation plays an important role in chronic migraine. It is possible that CNV variation on the gene and decrease in miRNA expression may increase neuroinflammation by affecting the control of the gene. As an important result, chronic migraine pathology can be therapeutically intervened using the Gene-CNV-miRNA relationship. It is thought that miR-3158-5p is regulated by hemoglobin modulation in the mechanism maintained under the responsibility of the CHRNA7 gene. In studies, ESR is presented as an inflammation marker in migraine. In the findings supporting this idea, ESR, platelet count and RDW were found to be higher in migraine compared to the control group. 40 In studies, ESR is presented as an inflammation marker in migraine. In the findings, ESR, platelet count and RDW are higher in migraine compared to the control group. Another study with similar results had high ESR levels. ESR and CRP are neuroinflammatory markers that are frequently used clinically. 41 Migraine burden differs according to gender. Migraine, which has a higher incidence in women than in men, is more severe in women in terms of attacks. Although the mechanisms causing this difference are not clear, it is likely that the balance of estrogen and progesterone is responsible. 42 Although the mechanisms causing this difference are not clear, the modulation of vascular tone caused by the balance of estrogen and progesterone has been implicated as responsible. 43 In addition, polymorphisms in the ESR gene were found to be significant in female migraine patients, supporting these findings. 44 Genetic susceptibility to migraine is accepted, but there is no definitive framework. Among the molecules currently under investigation, miRNAs have significant potential. 45 Migraine-specific miRNAs have been discovered in miRNA analyzes of peripheral blood monocytes of migraine patients. In our study, miR-3158-5p has potential as one of the migraine diagnosis candidates. Supporting it with ESR gene expression will show the effect of this miRNA more clearly.
The diagnostic power of miRNAs has been suggested in previous migraine studies. It has been determined that miR-382-5p is not only upregulated but also present during attacks. 46 It is difficult to consider miRNA expressions as migraine-specific markers because they vary in painful and non-painful states. 47 However, our results showed that hsa-miR-3158-5p together with CHRNA7-CNV has strong diagnostic value in patients with migraine. The abnormal expression of the CHRNA7 gene and hsa-miR-3158-5p in the whole blood of migraine patients identified in this study and the change in CNV numbers may be the basis for establishing minimally invasive biomarkers for the detection and monitoring of migraineurs. Not only are miRNAs useful as biomarkers, antimigraine treatments can also affect miRNA expressions.
Footnotes
Acknowledgements
We would like to thank Gaziantep University Scientific Research Project Unit for supporting our study (Project Number: TF.UT.19.28 and SBF.14.01). This study makes use of data generated by the DECIPHER community. A full list of centers that contributed to the generation of the data is available from ![]() and via email from
and via email from
Author contributions
All authors contributed equally to every stage of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was financially supported by the Scientific Research Project Unit of Gaziantep University with project number TF.UT.19.28 and SBF.14.01).
Ethical approval
The work described in this article has been carried out by The Code of Ethics of the World Medical Association (Declaration of Helsinki) for experiments involving humans; Uniform Requirements for manuscripts submitted to biomedical journals.
Informed consent to participate
All blood donors participating in this study gave written informed consent.
Informed consent for publication
Written informed consent was obtained from all authors.
Data availability statement
Data are available upon request from the authors. Those who performed the original analysis and collection of the data are not responsible for any further analysis or interpretation of the data for DECIPHER.
