Abstract
Acute pain that is associated with herpes zoster (HZ) can become long-lasting neuropathic pain, known as chronic post-herpetic neuralgia (PHN), especially in the elderly. HZ is caused by the reactivation of latent varicella-zoster virus (VZV), whereas PHN is not attributed to ongoing viral replication. Although VZV infection reportedly induces neuronal cell fusion in humans, the pathogenesis of PHN is not fully understood. A genome-wide association study (GWAS) revealed significant associations between PHN and the rs12596324 single-nucleotide polymorphism (SNP) of the heparan sulfate 3-
Keywords
Introduction
Post-herpetic neuralgia (PHN) is long-lasting neuropathic pain that is caused by the reactivation of latent varicella-zoster virus (VZV; i.e., herpes zoster (HZ)). Among HZ-afflicted patients, 27–73% develop PHN, which is age-dependent. 1 PHN causes severe pain that lasts for months or even more than 1 year. New ways need to be discovered to prevent PHN and further elucidate the pathogenesis of PHN.
In a previous study, PHN was significantly associated with the intronic rs12596324 single-nucleotide polymorphism (SNP) of the
VZV is an enveloped double-stranded DNA virus that belongs to the varicellovirus genus in the alphaherpesvirus subfamily of the herpesviridae family. Typically, VZV begins initial replication in the mucosal epithelium of the upper respiratory tract. The virus then enters the blood circulation and localizes to the skin, leading to the formation of characteristic blisters. 1 Upon primary infection, the virus causes varicella (chickenpox). Subsequently, sensory neurons that project to the skin become infected, leading to latent infection with VZV that persists for an individual’s lifetime. Immunocompromised hosts and the elderly have a higher susceptibility to the reactivation of VZV, causing HZ.
The DNA genome of VZV encodes over 70 open reading frames (ORFs), and 11 ORFs encode glycoproteins.
1
Glycoproteins on the virion membrane, which are glycosylated through
Heparan sulfate (HS) is a linear polysaccharide that covalently attaches to core proteins, referred to as HS proteoglycans. These molecules exist in the extracellular matrix and at the cell surface.
7
HS on HS proteoglycans is modified by HS-modifying enzymes, resulting in the functional modification of HS proteoglycans. Heparan sulfate glucosamine 3-
The present study was based on our previous findings that a SNP of the
Materials and methods
Human genetic association analysis of genome-wide genotyping data
Subjects with chronic pain and healthy subjects: The human GWAS was performed in our previous study. 2 We reanalyzed the genotyping data in the present study. Briefly, enrolled in the study were 194 adult patients who suffered from chronic pain, including PHN, and had a history of HZ and attended JR Tokyo General Hospital (Tokyo, Japan), Juntendo University Hospital (Tokyo, Japan), or Nihon University Itabashi Hospital (Tokyo, Japan) for the treatment of chronic pain. The patients were apparently Japanese. The detailed demographic data of the subjects and their statistics were provided in previous reports. 2
Enrolled in the study as controls were 282 adult healthy volunteers without any particular diseases or chronic pain who lived in or near the Kanto area in Japan. Detailed demographic data of the subjects and their statistics were provided in previous reports.8,9
The study protocol was approved by the Institutional Review Board of JR Tokyo General Hospital (Tokyo, Japan), Institutional Review Board of Juntendo University Hospital (Tokyo, Japan), Institutional Review Board of Nihon University Itabashi Hospital (Tokyo, Japan), and Institutional Review Board of Tokyo Metropolitan Institute of Medical Science (Tokyo, Japan). Written informed consent was obtained from all of the patients and healthy volunteers.
Statistical analysis: A GWAS was conducted for a subgroup of the 96 patient subjects with a history of HZ. A total of 282 control subjects were used in the analyses. To explore associations between the SNP and a history of HZ, Pearson’s
Plasmid construction
The human HS3ST4-expressing plasmid was provided by Dr Jian Liu (University of North Carolina, Chapel Hill). 10 Plasmids that encoded the VZV glycoproteins gB (ORF31), gH (ORF37), and gL (ORF60) with a pFN21A or pFC14A HaloTag CMV Flexi vector (Promega) backbone were constructed. Fragments of gB, gH, and gL were amplified from VZV clinical isolate 1710 strain genome DNA using KOD DNA polymerase (KOD FX Neo, Toyobo) with the following primer sets: gB (forward, cagagcgataacgcgatgtccccttgtggctatta; reverse, agcccgaattcgtttttacacccccgttacattct), gH (forward, actatagggctagcgatgtttgcgctagttttagc; reverse, gttggctcgagagctctcacggctccacagaga), and gL (forward, actatagggctagcgatggcatcacataaatggtt; reverse, gttggctcgagagctggcgcgggggcggacgggcc). The gB fragment was cloned into a pFN21A-HaloTag vector, which adds a HaloTag at the amino terminus, and gH and gL were cloned into the pFC14A-HaloTag vector, which adds a HaloTag at the carboxyl terminus using the In-Fusion HD Cloning Kit (Takara Bio).
Cells
The human skin MeWo melanoma cell line was cultured in high-glucose Dulbecco’s modified Eagle medium (DMEM; Fujifilm Wako Pure Chemical Corporation) supplemented with 10% fetal bovine serum (FBS; Biological Industries) and a penicillin-streptomycin solution (Thermo Fisher Scientific). MeWo cells were seeded in 24-well culture dishes (Corning) 1 day before transfection. The plasmid that encoded HS3ST4-cDNA with a pcDNA3 backbone or the vector plasmid pcDNA3 (Thermo Fisher Scientific) was transfected in MeWo cells using FuGENE 6 Transfection Reagent (Promega). After incubation for 5 min at room temperature, the mixture was added to the cells and incubated in a 5% CO2 incubator at 37°C for 24 h, followed by medium exchange. Two days later, the medium was changed to DMEM that contained 0.3 mg/mL G418 (Thermo Fisher Scientific). Selection continued for 14 days. Before the end of selection, all of the control MeWo cells without plasmids were dead. MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) clones were then obtained by limiting dilution. We selected one clone for use in the experiments. We repeated the experiments using another clone and did not find significant differences between clones.
Virus
The VZV wildtype strain 1710, which was isolated in our laboratory from the blister contents of a 9-year-old girl with HZ, was used in this study. MeWo cells that were infected with VZV were collected and frozen using CELLBANKER1 (Takara Bio) until the experiments were conducted. For the infection experiments, the cells were infected with the frozen infected cells after thawing. The titers of infected cells were determined using a plaque assay of frozen infected cells after thawing on MeWo cells.
Western blot
To quantify protein expression, MeWo, MeWo-HS3ST4 (+), and MeWo-HS3ST4 (−) cells at passage eight (P8) or P17 in 10 cm dishes were washed and suspended in electrophoresis sample buffer (Santa Cruz Biotechnology), followed by sonication. The samples were loaded onto e-PAGEL(R) 5–20% precast gels (ATTO). The protein bands were transferred to a polyvinyl difluoride membrane (Trans-Blot Turbo Midi 0.2 μm PVDF Transfer Packs, Bio-Rad) using a Trans-Blot Turbo Transfer System (Bio-Rad), followed by blocking with Blocking One (Nakalai) and blotting with anti-HS3ST4 sheep antibodies (R&D) and anti-glyceraldehyde-3-phosphate dehydrogenase (GAPDH) 1E6D9 mouse antibodies (Proteintech) as primary antibodies and peroxidase donkey anti-sheep IgG (H+L) and peroxidase donkey anti-mouse IgG (H+L) (Jackson Immunoresearch) as secondary antibodies. The membrane was treated with horseradish peroxidase substrate (WBKLS0100 Immobilon Western Chemiluminescent HRP Substrate, Merck). Chemiluminescence was quantified using a Lumino Image Analyzer (ImageQuant LAS-4000mini, Cytiva).
Plaque assay
MeWo cells were seeded in 12-well culture dishes (Corning) 1 day before infection with VZV. Frozen infected cells were thawed immediately and transferred to DMEM, followed by centrifugation. The pellet was resuspended in fresh DMEM. The serially diluted infected cells were added to naive cells in the well, and then the cells were incubated in a 5% CO2 incubator at 37°C until fixation and staining. Six days after infection, the cells were fixed in crystal violet dyes with HCHO for 2 h at room temperature, followed by washing and drying and then observing and counting plaques.
Microscopy imaging
MeWo cells were seeded in 12-well culture dishes (Corning) 1 day before infection with VZV. Frozen infected cells were thawed immediately and transferred to DMEM, followed by centrifugation as described in the plaque assay section. The pellet was resuspended in fresh DMEM. The diluted infected cells were added to 4 × 105 naive cells in the well at a multiplicity of infection (MOI) of 0.005, and then the cells were incubated in a 5% CO2 incubator at 37°C. Live-cell images that were acquired with an inverted microscope (Axiovert 200, Carl Zeiss, Digital Sight DS-L1, Nikon Solutions) 2 days after infection in Figure 3.
Cytotoxicity assay
As a cytotoxicity assay, a lactate dehydrogenase (LDH) release was quantified using the CytoTox 96 Non-Radioactive Cytotoxicity Assay (Promega) according to the manufacturer’s protocol. 11 MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells were seeded in 96-well culture dishes (Corning) 1 day before infection with VZV. The infected cells were added to the cells in 96-well culture dishes at an MOI of 0.05. After incubation for 2 h in a 5% CO2 incubator at 37°C, the medium was exchanged with 100 μL of fresh DMEM with FBS. Zero hours after infection (h.a.i.) was set at this point in the experiment. The dispensed supernatant (50 μL) and 100 μL of the exchanged fresh medium with the cells were frozen at −80°C at 0, 6, 24, 48, 72, and 96 h.a.i. The LDH release ratio of 50 μL of the supernatant with or without cells after the freeze-thaw cycle was examined.
Quantitative real-time polymerase chain reaction assay
MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells were seeded in 24-well culture dishes (Corning) 1 day before infection with VZV. The infected cells were added to cells in 24-well culture dishes at an MOI of 0.005. After incubation in a 5% CO2 incubator at 37°C for 4 h, the medium was exchanged.
Total DNA was extracted from the cells with and without infection in 24-well culture dishes (Corning) using the QIAamp DNA Mini Kit (Qiagen). VZV DNA and ribonuclease (RNase) P as a copy number reference were quantitatively examined using TaqMan Real-Time Polymerase Chain Reaction (PCR) Master Mix (Thermo Fisher) and the 7500 Fast Real-Time PCR System (Applied Biosystems). The probe (final concentration of 0.25 μM) and primers (final concentration of 0.9 μM) for the VZV genome were the following: ORF28 probe (TCC AGG TTT TAG TTG ATA CCA FAM-TAMRA), sense primer (CGA ACA CGT TCC CCA TCA A), and antisense primer (CCC GGC TTT GTT AGT TTT GG). 12 For RNase P, the TaqMan Copy Number Reference Assay, human, RNase P (VIC-TAMRA, Thermo Fisher) was used for detection as a copy number reference. The PCR conditions for VZV and RNase P were the following: one cycle of 50°C for 2 min and 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 1 min. The copy numbers of the VZV genome were corrected relative to the copy numbers of RNase P, which represented the total number of nuclei. To determine cell proliferation rates, quantitative real-time PCR with RNase P detection was performed. Rates were similar among the cell lines.
Electroporation
MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells were treated with the Amaxa 4D Nucleofector Kit SF (Lonza) according to the manufacturer’s protocol and transferred to a 16-well cuvette. 2.5 × 106 cells per sample were electroporated with 0.4 μg of each of the plasmids that encoded the VZV glycoproteins gB, gH, and gL and 0.2 μg of the control plasmid pmaxGFP (Lonza) by the Amaxa 4D Nucleofector X unit (Lonza). After 10 min of incubation at room temperature, cells with fresh DMEM and FBS were transferred to 4 wells of a 96-well culture dish per sample. The medium in the 96-well culture dishes was exchanged with 100 μL of fresh medium 4–6 h after electroporation. As a control for electroporation efficiency, the control plasmid pmaxGFP was co-electroporated. Efficiency was monitored 24 hours after electroporation by observing fluorescence (Axiovert 200, Carl Zeiss, Digital Sight DS-L1, Nikon Solutions), which was sufficiently high in every well (∼95–98%). Glycoprotein expression was monitored by observing the fluorescence of glycoproteins that were tagged with HaloTag after the addition of HaloTag TMRDirect Ligand (Promega). Twenty-four hours after electroporation, 50 μL of the dispensed supernatant and 100 μL of the exchanged fresh medium with cells were frozen. The cytotoxicity of 50 μL of the supernatant with and without cells after the freeze-thaw cycle was examined using the CytoTox 96 Non-Radioactive Cytotoxicity Assay (Promega), similar to the cytotoxicity assay section.
Statistical analysis of assay data
Significant differences between two groups were evaluated using Student’s
Results
Herpes zoster was significantly associated with an HS3ST4 SNP
Demographic data of patients and healthy control subjects, stratified by rs12596324 genotype.

The data are expressed as numbers and the mean ± SD (range).
Association between HZ and the rs12596324 SNP of

*
Virus-mediated cell fusion occurred robustly in cells with HS3ST4 infected with VZV
To clarify whether HS3ST4 contributes to VZV infection, we first produced cells that expressed HS3ST4. MeWo cells were introduced with a plasmid that encoded HS3ST4 cDNA (MeWo-HS3ST4 (+)) or a vector plasmid (MeWo-HS3ST4 (−)). To examine whether MeWo-HS3ST4 (+) cells stably express HS3ST4, the expression of HS3ST4 in MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells at passages eight and 17 were analyzed by Western blot (Figure 1). MeWo-HS3ST4 (+) cells showed a distinct band around 60 kD, detected by anti-human HS3ST4 antibodies, at passages eight and 17, whereas MeWo-HS3ST4 (−) cells and the original MeWo cells showed no band around 60 kD. These results suggested that MeWo-HS3ST4 (+) cells stably expressed HS3ST4 protein. HS3ST4 is expressed in MeWo-HS3ST4 (+) cells. HS3ST4 was analyzed in MeWo cells, MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells at passage eight (P8) or P17 by Western blot. Bands were detected around 60 kD by anti-HS3ST4 antibodies. GAPDH was detected as an internal control. Black triangles indicate the predicted positions of the antigens.
To determine whether HS3ST4 contributes to VZV infection, we performed a plaque assay and observed plaque phenotypes of cells with and without HS3ST4 infected with VZV. MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells that were infected with VZV in 12-well plates were fixed 6 days after infection. As shown in Figure 2, plaques on MeWo-HS3ST4 (+) cells had distinct borders (Figure 2(a)), whereas plaques on MeWo-HS3ST4 (−) cells had unclear borders (Figure 2(b)). These results suggested that infected MeWo-HS3ST4 (+) cells peeled off with engulfed surrounding cells. Plaque phenotype of MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells infected with VZV. MeWo-HS3ST4 (+) (a) and MeWo-HS3ST4 (−) cells (b) that were infected with VZV in 12-well plates were fixed 6 days after the infection. The images below are twice-enlarged images of pink rectangles in the upper images.
To determine why plaques of infected MeWo-HS3ST4 (+) cells had distinct borders, we observed cells with and without HS3ST4 after infection under an inverted microscope 2 days after infection (Figure 3). MeWo-HS3ST4 (+) cells that were infected with VZV had larger cell fusions (Figure 3(a)) compared with MeWo-HS3ST4 (−) cells that were infected with VZV (Figure 3(b)). Mock-infected cells had no cell fusion (Figures 3(c) and (d)). These results suggested that HS3ST4 accelerated cell fusion after VZV infection. Virus-mediated cell fusion occurred robustly in cells with HS3ST4 infected with VZV. MeWo-HS3ST4 (+) (a and c) and MeWo-HS3ST4 (−) cells (b and d) were infected (a and b) or mock-infected with VZV (c and d) and observed under an inverted microscope 2 days after infection. Scale bar = 100 μm.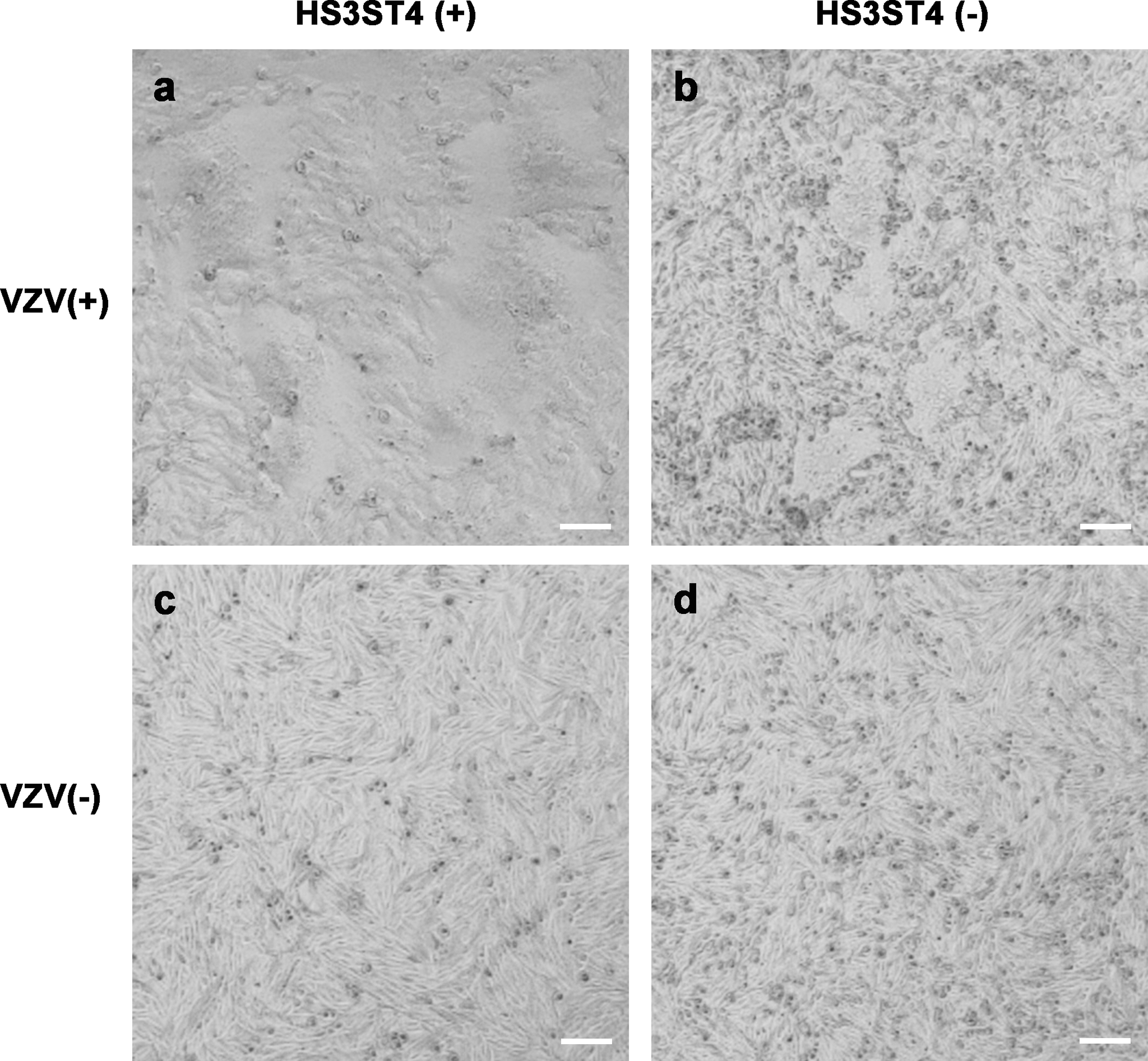
Higher cytotoxicity in cells with HS3ST4 after infection
To quantitatively determine the time course of cell fusion after infection, cells with and without HS3ST4 were infected or mock-infected with VZV, and supernatants were collected on each day after infection and analyzed using the cytotoxicity assay (Supplementary Dataset S2). As shown in Figure 4, infected MeWo-HS3ST4 (+) cells had a higher LDH release ratio, that is, cytotoxicity, 1–3 days after infection compared with infected MeWo-HS3ST4 (−) cells ( Higher cytotoxicity in cells with HS3ST4 after infection with VZV. MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells were infected or mock-infected with VZV, and supernatants were collected on each day after infection and analyzed by the cytotoxicity assay. *
Expression of VZV glycoproteins increased the cytotoxicity in cells with HS3ST4
Glycoproteins gB, gH, and gL of VZV were reported to contribute to cell fusion.
6
To determine whether these glycoproteins play essential roles in cell fusion after infection in MeWo-HS3ST4 (+) cells, glycoprotein-encoding or vector plasmids were expressed in cells with and without HS3ST4. Twenty-four hours after electroporation, the cells were observed under a fluorescent microscope (Figure 5). MeWo-HS3ST4 (+) cells with gB, gH, and gL showed large fused cells (Figure 5(a)), whereas MeWo-HS3ST4 (−) cells with gB, gH, and gL showed smaller fused cells (Figure 5(b)). MeWo-HS3ST4 (+) nor MeWo-HS3ST4 (−) cells with vector plasmid showed no fused cells (Figure 5(c) and (d)). These results implied that the expression of HS3ST4 with the glycoproteins accelerated cell fusion. In parallel with fluorescent microscopy, the supernatants were collected and quantitatively analyzed using the cytotoxicity assay (Supplementary Dataset S3). As shown in Figure 6, glycoprotein expression significantly increased a relative value of the LDH release ratio (LDH release ratio in cells that expressed glycoprotein-encoding plasmid per LDH release ratio in cells that expressed the vector plasmid) in MeWo-HS3ST4 (+) cells compared with MeWo-HS3ST4 (−) cells ( Expression of HS3ST4 along with VZV glycoproteins accelerated cell fusion. Twenty-four hours after electroporation, the cells were observed under a fluorescent microscope. (a–d) Fluorescent images of MeWo-HS3ST4 (+) cells with gB, gH, and gL (a), MeWo-HS3ST4 (−) cells with gB, gH, and gL (b), MeWo-HS3ST4 (+) cells with the vector plasmid (c), and MeWo-HS3ST4 (−) cells with the vector plasmid (d). Scale bar = 100 μm. Expression of VZV glycoproteins increased cytotoxicity in MeWo-HS3ST4 (+) cells

VZV genome replication had similar efficiency in the cells with HS3ST4
To investigate the way in which HS3ST4 contributes to viral genome replication, we analyzed copy numbers of the viral genome chronologically. Cells with and without HS3ST4 infected with VZV were collected at the indicated time points. Copy numbers of the VZV genome were compared with 0 hour, based on quantitative real-time PCR (Figure 7, Supplementary Dataset S4). From 0 to 96 h after infection, no significant difference in copy numbers of the VZV genome was found between MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells ( VZV genome replication had similar efficiency in cells, regardless of HS3ST4 expression. MeWo-HS3ST4 (+) and MeWo-HS3ST4 (−) cells that were infected with VZV were collected at the indicated time points. Copy numbers of the VZV genome were analyzed by quantitative real-time PCR. *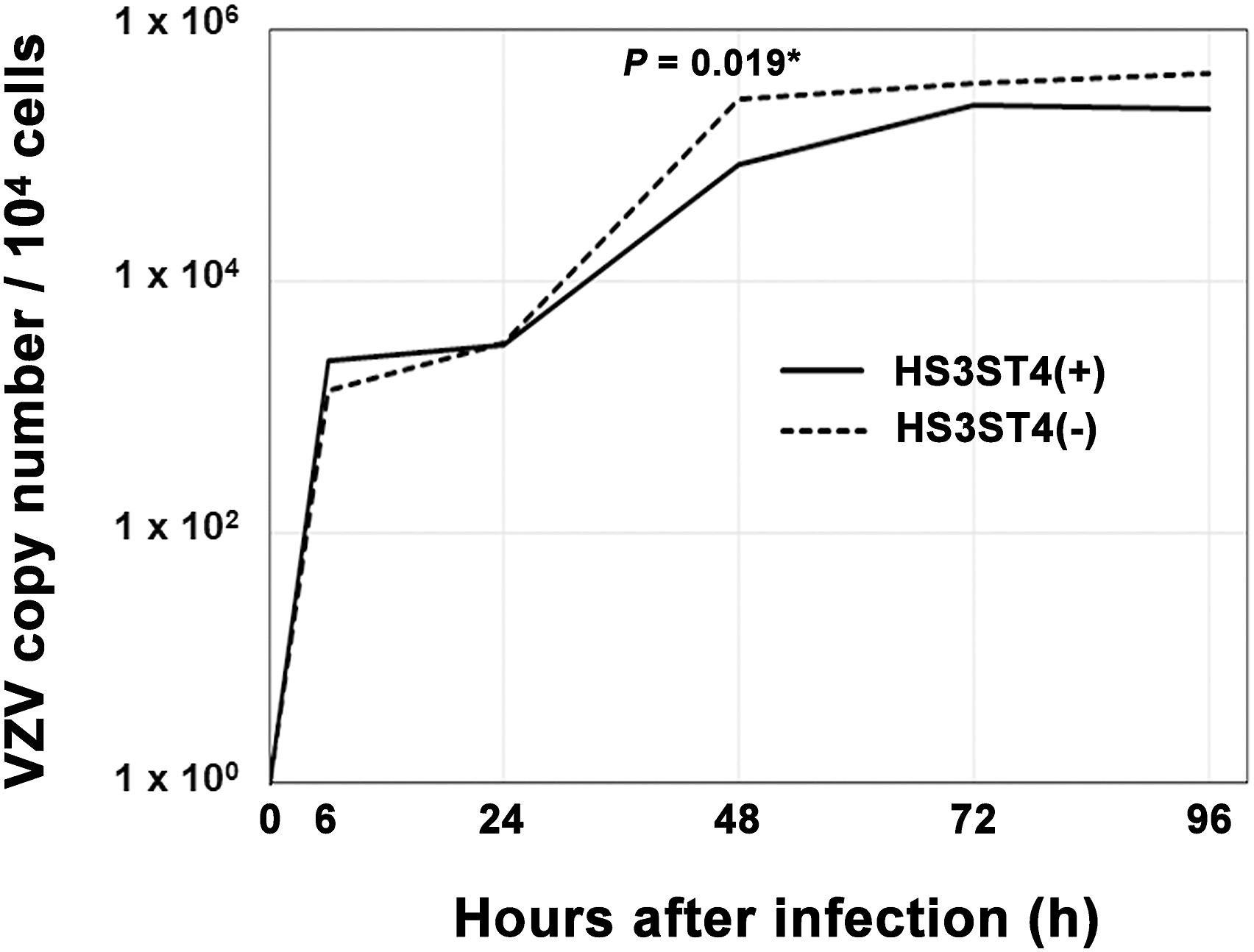
Discussion
In addition to significant associations that were found between the rs12596324 SNP of
Introns occasionally function to regulate transcription.
13
Although it remains unclear whether the G allele on intronic rs12596324 SNP of the
VZV-infected MeWo cells fuse with the human neuronal somata.
4
Similarly, VZV infection was reported to provoke neural cell fusion
HS3ST4 has minimal effects on VZV replication and markedly enhances fusogenic activity in the presence of VZV glycoproteins. Hyperfusogenic measles virus and other paramyxovirus mutants induced strong cytopathology through cell fusion in infected cells, despite the suppression of viral replication. 11 Accordingly, it is reasonable that HS3ST4 accelerates cytopathology through virus-induced fusogenic activity despite almost no changes in the viral replication rate.
In Figure 6, MeWo-HS3ST4 (−) cells had a relative value > 1, indicating that the glycoproteins induced cytotoxicity even in MeWo-HS3ST4 (−) cells, consistent with Figure 5(b) compared with Figure 5(d). MeWo-HS3ST4 (+) cells had more robust cytotoxicity after expressing these glycoproteins, consistent with Figure 5(a) compared with Figure 5(c). These results suggested that these glycoproteins induced cytotoxicity, including fusogenic activity, in MeWo-HS3ST4 (+) cells. This implies that the glycoproteins gB, gH, and gL are sufficient to accelerate cell fusion through the HS3ST4-related pathway. HS that is modified by HS3ST4 has been shown to assist herpes simplex virus (HSV)-1 entry in Chinese hamster ovarian (CHO-K1) cells, which are not naturally susceptible to HSV-1.15,17 Since HSV-1 is closely related to VZV, HS3ST4 likely plays similar essential roles in VZV entry. In the present study, we assayed cell fusion using human cells that are naturally susceptible to VZV. The assumed receptors for VZV glycoproteins on the cell surface would be modified by HS3ST4 solely in cells with HS3ST4. This modification of HS in receptors on the cell surface would promote cell-cell attachment, followed by cell fusion.
Genes that encode the glycoproteins gB, gH, and gL are classified into late genes for VZV. 18 In the human TG, these glycoproteins are expressed only at low levels, if at all, during latent infection. 19 After robust reactivation of the virus, these glycoproteins are sufficiently expressed on the cell surface to enhance fusogenic activity, possibly in cells with higher HS3ST4 expression.
Supplemental Material
sj-xls-1-mpx-10.1177_17448069211052171 – Supplemental Material for Heparan sulfate 3-O -sulfotransferase 4 is genetically associated with herpes zoster and enhances varicella-zoster virus–mediated fusogenic activity
Supplemental Material, sj-xls-1-mpx-10.1177_17448069211052171 for Heparan sulfate 3-
Supplemental Material
sj-xlsx-2-mpx-10.1177_17448069211052171 – Supplemental Material for Heparan sulfate 3-O -sulfotransferase 4 is genetically associated with herpes zoster and enhances varicella-zoster virus–mediated fusogenic activity
Supplemental Material, sj-xlsx-2-mpx-10.1177_17448069211052171 for Heparan sulfate 3-
Supplemental Material
sj-xlsx-3-mpx-10.1177_17448069211052171 – Supplemental Material for Heparan sulfate 3-O -sulfotransferase 4 is genetically associated with herpes zoster and enhances varicella-zoster virus–mediated fusogenic activity
Supplemental Material, sj-xlsx-3-mpx-10.1177_17448069211052171 for Heparan sulfate 3-
Supplemental Material
sj-xlsx-4-mpx-10.1177_17448069211052171 – Supplemental Material for Heparan sulfate 3-O -sulfotransferase 4 is genetically associated with herpes zoster and enhances varicella-zoster virus–mediated fusogenic activity
Supplemental Material, sj-xlsx-4-mpx-10.1177_17448069211052171 for Heparan sulfate 3-
Footnotes
Acknowledgments
We thank Mr Michael Arends for editing the manuscript. We are grateful to the volunteers for their participation in the study and anesthesiologists and surgeons for collecting the clinical data.
Author contributions
Conceptualization: Seii Ohka, Kazutaka Ikeda.Data curation: Seii Ohka, Daisuke Nishizawa, Hideko Arita, Kazuo Hanaoka, Masako Iseki, Jitsu Kato, Setsuro Ogawa, Ayako Hiranuma, Shinya Kasai, Junko Hasegawa, Masakazu Hayashida, Kazutaka Ikeda. Formal analysis: Seii Ohka. Funding acquisition: Seii Ohka, Daisuke Nishizawa, Masayuki Saijo, Kazutaka Ikeda. Investigation: Seii Ohka, Kazutaka Ikeda. Methodology: Seii Ohka, Souichi Yamada, Daisuke Nishizawa, Shuetsu Fukushi, Masayuki Saijo, Kazutaka Ikeda. Project administration: Seii Ohka, Kazutaka Ikeda. Resources: Seii Ohka, Souichi Yamada, Daisuke Nishizawa, Yoshiko Fukui, Hideko Arita, Kazuo Hanaoka, Masako Iseki, Jitsu Kato, Setsuro Ogawa, Ayako Hiranuma, Shinya Kasai, Junko Hasegawa, Masakazu Hayashida, Shuetsu Fukushi, Masayuki Saijo, Kazutaka Ikeda. Supervision: Shuetsu Fukushi, Masayuki Saijo, Kazutaka Ikeda. Validation: Seii Ohka. Visualization: Seii Ohka. Writing-original draft preparation: Seii Ohka. Writing-review & editing: Seii Ohka, Souichi Yamada, Daisuke Nishizawa, Yoshiko Fukui, Hideko Arita, Kazuo Hanaoka, Masako Iseki, Jitsu Kato, Setsuro Ogawa, Ayako Hiranuma, Shinya Kasai, Junko Hasegawa, Masakazu Hayashida, Shuetsu Fukushi, Masayuki Saijo, Kazutaka Ikeda.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Japan Society for the Promotion of Science (JSPS) KAKENHI [JP16H06276 (AdAMS) KI, 20K07774 SO, 17K09052 SO, 20K09259 DN, 17K08970 DN, 17H04324 KI, 18K07894 MS]; and Japan Agency for Medical Research and Development (AMED) [JP19ek0610011 KI].
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
