Abstract
Introduction:
Conducting systematic reviews of clinical trials is time-consuming and resource-intensive. One potential solution is to design databases that are continuously and automatically populated with clinical trial data from harmonised and structured datasets. This scoping review aimed to identify and map publicly available, continuously updated, topic-specific databases of clinical trials.
Methods:
We systematically searched PubMed, Embase, the preprint servers medRxiv, arXiv, Open Science Framework, and Google. We characterised each database using seven predefined features (access model, database type, data input sources, retrieval methods, data-extraction methods, trial presentation, and export options) and narratively summarised the results.
Results:
We identified 14 continuously updated databases of clinical trials, seven related to COVID-19 (initiated in 2020) and seven non-COVID-19 databases (initiated as early as in 2009). All databases, except one, were publicly funded and accessible without restrictions. Most relied on traditional methods used in static article-based systematic reviews sourcing data from journal publications and trial registries. The COVID-19 databases and some non-COVID-19 databases implemented semi-automated features of data import, which combined automated and manual data curation, whereas the non-COVID-19 databases mainly relied on manual workflows. Most reported information was metadata, such as author names, years of publication, and link to publication or trial registry. Only two databases included trial appraisal information (such as risk of bias assessments). Six databases reported aggregate group-level results, but only one database provided individual participant data on request.
Discussion:
Continuously updated topic-specific databases of clinical trials remain limited in number, and existing initiatives mainly employ traditional static systematic review methodologies. A key barrier to developing truly living platforms is the lack of accessible, machine-readable, and standardised clinical trial data.
Keywords
Introduction
The number of published clinical trials and systematic reviews increased tremendously over the past decades.1,2 It is difficult, if not impossible, for any clinician or researcher to stay up to date. Traditional systematic reviews are time-consuming and costly due to the efforts required to manually search databases, screen titles, abstracts, and full-text articles and data extraction. 3 As a result, systematic reviews are often outdated and potentially even misleading by the time of publication.4–7
To address the limitations of traditional systematic reviews, the concept of ‘living systematic reviews’ was popularised. There is no universally accepted definition but living reviews are generally understood to be continuously updated (systematic) reviews.8–11 Living systematic reviews should mitigate delays and reduce the risk of missing important evidence. The COVID-19 pandemic accelerated the development of such ‘living’ projects reflecting the rapidly evolving evidence landscape. These include the COVID-19 trial tracker, 12 the NMA-COVID Project, 13 and WHO’s Living Guideline. 14
Text mining and machine-learning assisted tools have been developed to automate parts of the review process and reduce resource demands15–18 focusing mainly on literature screening, retrieving records, and extracting data from publications.19,20 Whether existing living projects and topic-specific trial databases fully leverage advanced technologies to ensure seamless and – ideally fully automated – flow of clinical trial data into structured databases remains unclear.
We aimed to systematically identify publicly available, continuously updated, topic-specific clinical trial databases of clinical trials to characterise their basic functionalities, infrastructure, and user interface. This scoping review also served as a primer for designing a continuously updated database focused on cancer immunotherapies. 21
Methods
We conducted a scoping review and report it according to the Preferred Reporting Items for Systematic reviews and Meta-Analyses statement for Scoping Reviews (PRISMA-ScR) checklist. 22 Due to the project’s exploratory nature, we did not initially register a protocol for this review.
Eligibility criteria
We included any publicly available continuously updated database retrieving clinical trials (participants allocated to two or more groups); irrespective of retrieval method (e.g. manually, automated, or a combination), and input source (e.g. databases of published literature, trial registries, or both). We did not define a minimum update frequency threshold for the included databases. We considered topic-specific databases only, that is, exclusive to certain conditions, interventions, or indication, such as a COVID-19. Databases including other research designs in addition to clinical trials, such as observational studies, were considered eligible. We did not include broad, topic-unspecific trial databases, such as clinical trial registries. We applied no restrictions on language, release date, intended use (i.e. for research or clinical decision-making), or database status (i.e. live or archived).
Information sources, search strategy, and selection of sources of evidence
We employed a two-step systematic search strategy. We did ‘cold searches’ of various resources to identify key words and references to subsequently build systematic database search strategies.
Step 1: Cold search – One author (KB) searched PubMed (including using the ‘similar articles’ function), preprint servers medRxiv and ArXiv, Open Science Framework, and Google (first five pages for each search) informed by key search terms and databases known to us prior to this scoping review (S1 and S2, Appendix in the Supplementary Material).
Step 2: Systematic search – The PubMed and Embase/Elsevier databases were systematically searched by one author (JH). Two authors (KB, JH) designed a three-component search strategy containing keywords for ‘living’ (e.g. live, updated, and interactive), ‘library’ (e.g. database, platform, and repository), and study design (e.g. trial, and clinical study). See full search strategies and search dates in the Appendix, S3 and S4. The retrieved records were deduplicated using Citavi 23 and imported into the screening tool Rayyan. 24 One author (KB) screened titles, abstracts, and full texts and decided on inclusion in one step. Uncertainties were discussed with another author (PJ or JH) to reach a final decision.
Data extraction and narrative summary
We narratively summarised the following seven database characteristics based on information available from the database websites and corresponding journal publications (Figure 1). One reviewer (KB) extracted information, and one reviewer (JH) double-checked and confirmed the extractions. Data was tabulated directly in Microsoft Word and screenshots to document the user interface of eligible databases.
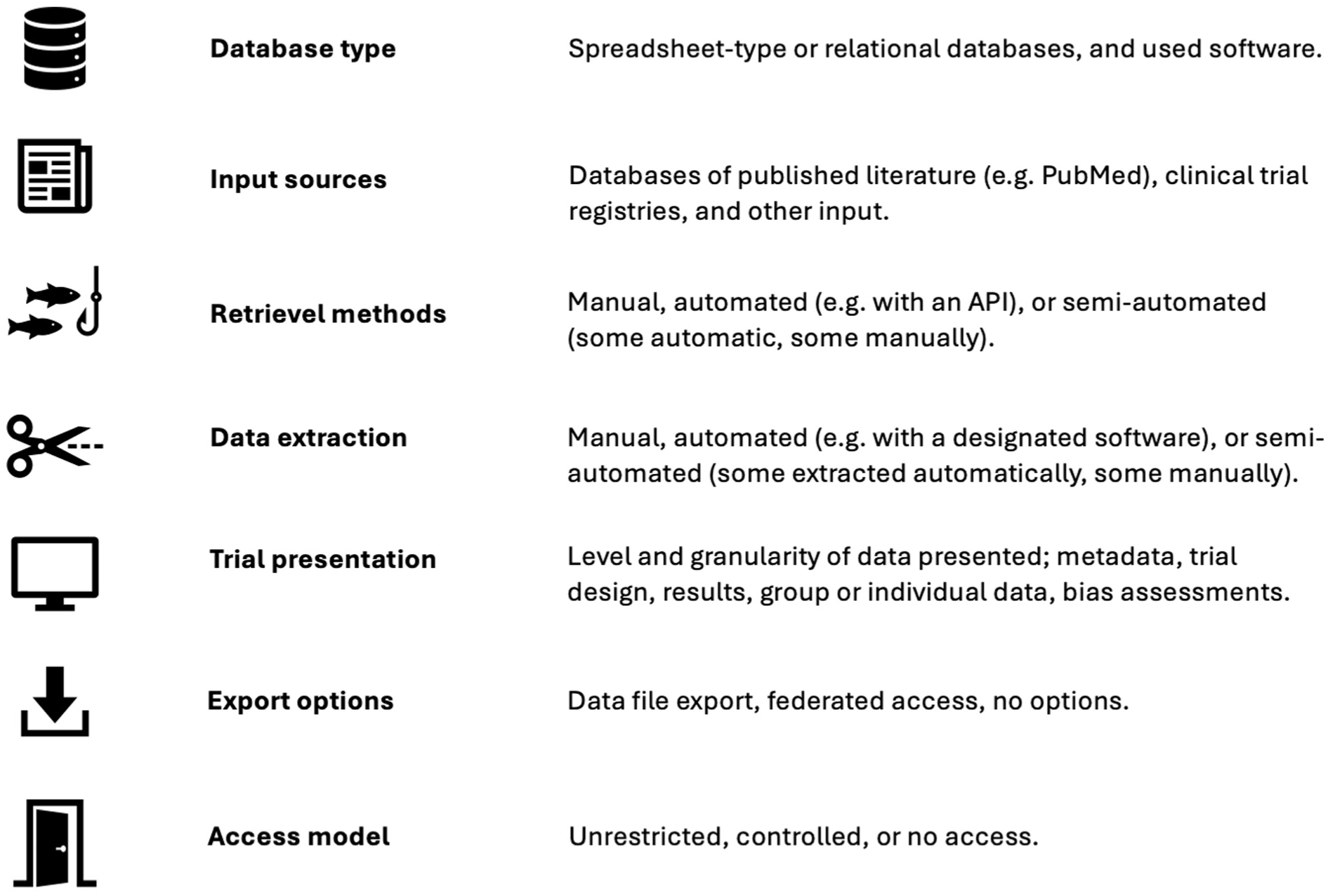
Data-extraction variables on characteristics of continuously updated clinical trial databases.
Database characteristics
Database type: What was the underlying database type and database-management software (IBM Ref)? Options include simple Excel spreadsheets or conventional relational databases, 25 which may be managed with various database-management systems, like RedCap. 26
Input sources: Which data input sources were searched? Databases of published literature like PubMed or Embase, trial registries, for example, ClinicalTrials.gov, or other sources.
Retrieval methods: How was the database populated? Manually, automated, for example, by application programming interfaces (APIs), or via community-based data submissions where researchers submit data to the database, like ClinicalTrials.gov or the nucleotide sequencing database GenBank. 27
Data extraction: How was the trial information extracted and curated? Manually, fully automated with designated software tools,15–17 or a combination of machine-assisted manual curation, usually referred to as ‘semi-automated’. 28
Trial presentation: What trial information were reported on the website? Meta data (e.g. title, author names, publication or trial registry information), detailed trial information (e.g. study design, sample size, treatment descriptions, funding), results (outcomes and effect estimates), type of results (aggregate group level or individual participant data), and trial appraisal (e.g. risk of bias assessments, limitations in the trial design).
Export options: What were the options for downloading and reusing data? Free download of data (e.g. as CSV or Excel files), federated access 29 (e.g. users can access the data remotely, usually in browser-based applications, but without options for downloading it to their own personal computers, such as Vivli.com), 30 or no option for exporting data.
Access model: How was the database content accessed? Unrestricted access (i.e. data is freely available without any constraints), controlled access (e.g. users must submit requests and the provider grants or denies access), or a hybrid between the two.
Results
Search results
Our systematic database search returned 1707 hits (after deduplication); 15 were assessed in full-text (see reasons for exclusion in S6, Appendix in the Supplementary Material) and one database (TrialsResultsCenter31,32) was included. One database (Evidence Finder33–35) was found through other sources, nine databases (Cochrane COVID-19 Study Register, 36 COVID-evidence, 37 COVID TrialsTracker, 12 COVID-NMA initiative, 13 EPPI Centre Covid-19 Living Map of the Evidence, 38 MetaEvidence breast cancer, 39 MetaEvidence COVID, 40 MetalO, 41 and MetaPreg 42 ) were known to us before our systematic search. Our cold searches yielded three eligible databases (Infectious Disease Data Observatory (IDDO), 43 Worldwide Antimalarial Resistance Network (WWARN),44–46 and EvidenceMap47–49 (S5, Appendix in the Supplementary Material).
In total, we included 14 databases (Table 1), seven non-COVID-19 databases on youth mental health (Evidence Finder), pancreatic surgery (Evidence Map), cardiology and oncology trials (TrialResultsCenter), cancer (MetaEvidence breast cancer and MetalO), pregnancy (MetaPreg), and malaria (WWARN) and seven COVID-19 databases (Cochrane COVID-19 Study Register, COVID-evidence, COVID TrialsTracker, COVID-NMA initiative, EPPI Centre Covid-19 Living Map of the Evidence, IDDO, and MetaEvidence COVID).
Included continuously updated field-specific clinical trials databases.
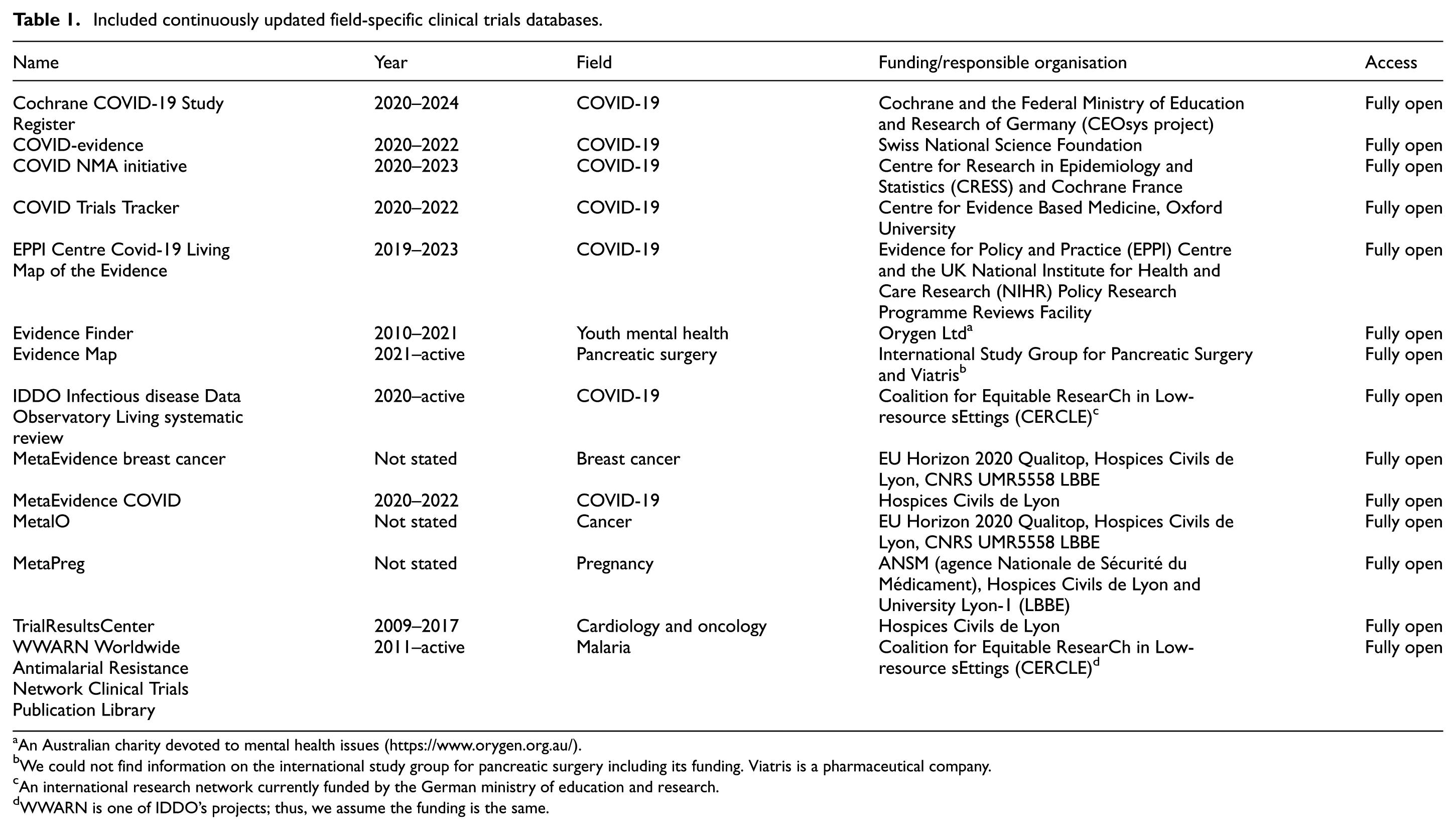
An Australian charity devoted to mental health issues (https://www.orygen.org.au/).
We could not find information on the international study group for pancreatic surgery including its funding. Viatris is a pharmaceutical company.
An international research network currently funded by the German ministry of education and research.
WWARN is one of IDDO’s projects; thus, we assume the funding is the same.
Key findings
There were important trends and differences between the 14 identified databases indicating that the COVID-19 databases leveraged methods and technology to automatise several of the time-consuming tasks, whereas the non-COVID-19 databases largely replicated methods from traditional systematic reviews without integrating methods to streamline the workflow. While the COVID-19 databases were launched during the pandemic in 2020 (and all but one is no longer maintained), the oldest non-COVID-19 databases date back to 2009–2011. The database types were unclearly reported for most databases, and the input sources were generally similar. The most important differences between COVID-19 and non-COVID-19 databases pertain to the retrieval and data-extraction methods. The non-COVID-19 databases largely used manual retrieval methods (four manual, one semi-automated, two unreported) and data extraction (five manual, two unreported); meanwhile the COVID-19 databases were more advanced, and five databases used semi-automated retrieval and data-extraction methods and only two used manual methods. All databases reported basic meta-data information (including title, authors, and links to publications/registry entries), but only few databases reported more granular results and bias assessments.
Database characteristics – non-COVID-19 databases
The TrialResultsCenter was active from 2009 to 2017 and is no longer updated. EvidenceFinder was released in 2010 with the last update in July 2021, though its current maintenance status is uncertain. WWARN was released in 2011, but its maintenance status is also unclear; the last trial in their Data Inventory 44 dates from 2018, but their trial summary 43 says it is current until 2022. EvidenceMap was launched in 2021 with the most recent update on July 26, 2023. The release dates were not specified for three databases (MetaEvidence breast cancer, MetalO, and MetaPreg). Six of the seven non-COVID-19 databases were either publicly funded or funded by non-profit organisations while one (Evidence Map) was funded by a pharmaceutical company. All database websites were fully accessible with no restrictions.
The databases primarily searched common sources, such as PubMed/MEDLINE, Embase, and Web of Science, employing methods similar to those used in systematic reviews, including manual database searches, data extraction, and curation. EvidenceMap was built on a relational Microsoft SQL server database, and WWARN used a proprietary data management tool (RedCap). The database type for the remaining five was not specified.
Regarding data availability, five databases (EvidenceMap, MetaEvidence breast cancer, MetalO, MetaPreg, and WWARN) made their data readily available for download. One database (TrialResultsCenter) provided data upon request, and another (EvidenceFinder) did not specify download options (see Table 2).
Database characteristics (non-COVID 19).
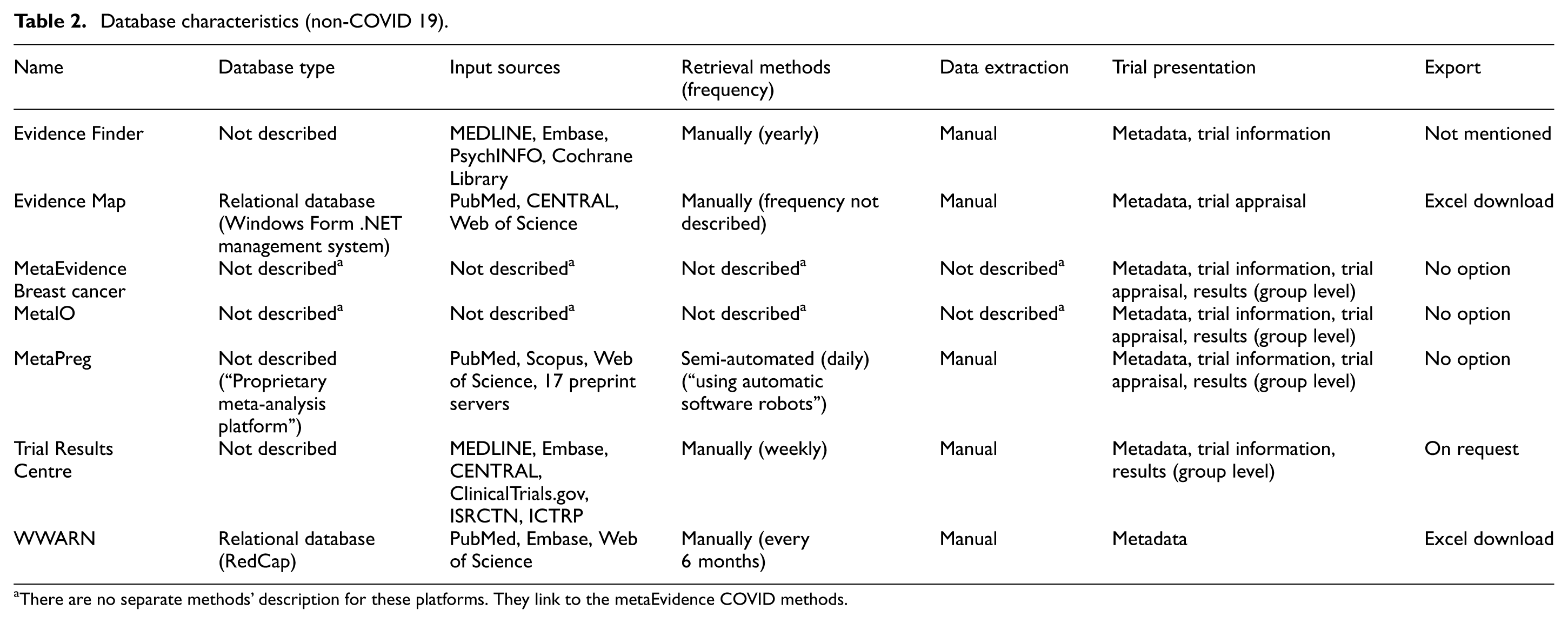
There are no separate methods’ description for these platforms. They link to the metaEvidence COVID methods.
All databases included basic metadata information, such as title, author names, and links to publication or trial registries. Three databases (MetaEvidence breast cancer, MetalO, and MetaPreg) reported aggregated group-level results and risk of bias assessments. One (TrialsResultsCenter) reported aggregated group-level results while two databases (Evidence Finder and TrialsResultsCenter) reported some trial information. EvidenceMap uniquely reported trial appraisal as a risk of bias assessment (Table 2; S7, Appendix in the Supplementary Material).
Database characteristics – COVID-19 databases
The seven COVID-19 databases were initiated shortly after the SARS-CoV-2 outbreak in early 2020. By January 2024, the last of these databases, the Cochrane COVID-19 Study Register had ceased updates. All databases were publicly funded or supported by non-profit institutions and were openly accessible to the public without restrictions.
Three of the seven COVID-19 databases (Cochrane COVID-19 Study Register, EPPI Centre Covid-19 Living Map of the Evidence, and MetaEvidence COVID) primarily searched common sources, including databases of published reports such as PubMed/MEDLINE and Embase. In addition, six of the seven databases also searched one or more trial registries, broadening the scope of evidence retrieval.
Five databases (Cochrane COVID-19 Study Register, COVID-evidence, COVID TrialsTracker, EPPI Centre Covid-19 Living Map of the Evidence, and Meta-evidence) used semi-automated methods for study retrieval and data extraction, particularly for trial registry records.
In contrast, two databases (COVID NMA Initiative and IDDO) relied solely on manual data retrieval and extraction.
Two databases (IDDO and COVID-evidence) used proprietary database-management tools (RedCap and Directus). The database-management systems for the other five were not specified.
In terms of data availability, six databases (Cochrane COVID-19 Study Register, COVID-evidence, COVID TrialsTracker, EPPI Centre Covid-19 Living Map of the Evidence, MetaEvidence COVID, and IDDO) provided data for direct download, while one database (COVID NMA initiative) made data available upon request (Table 3).
Database characteristics (COVID-19 specific).
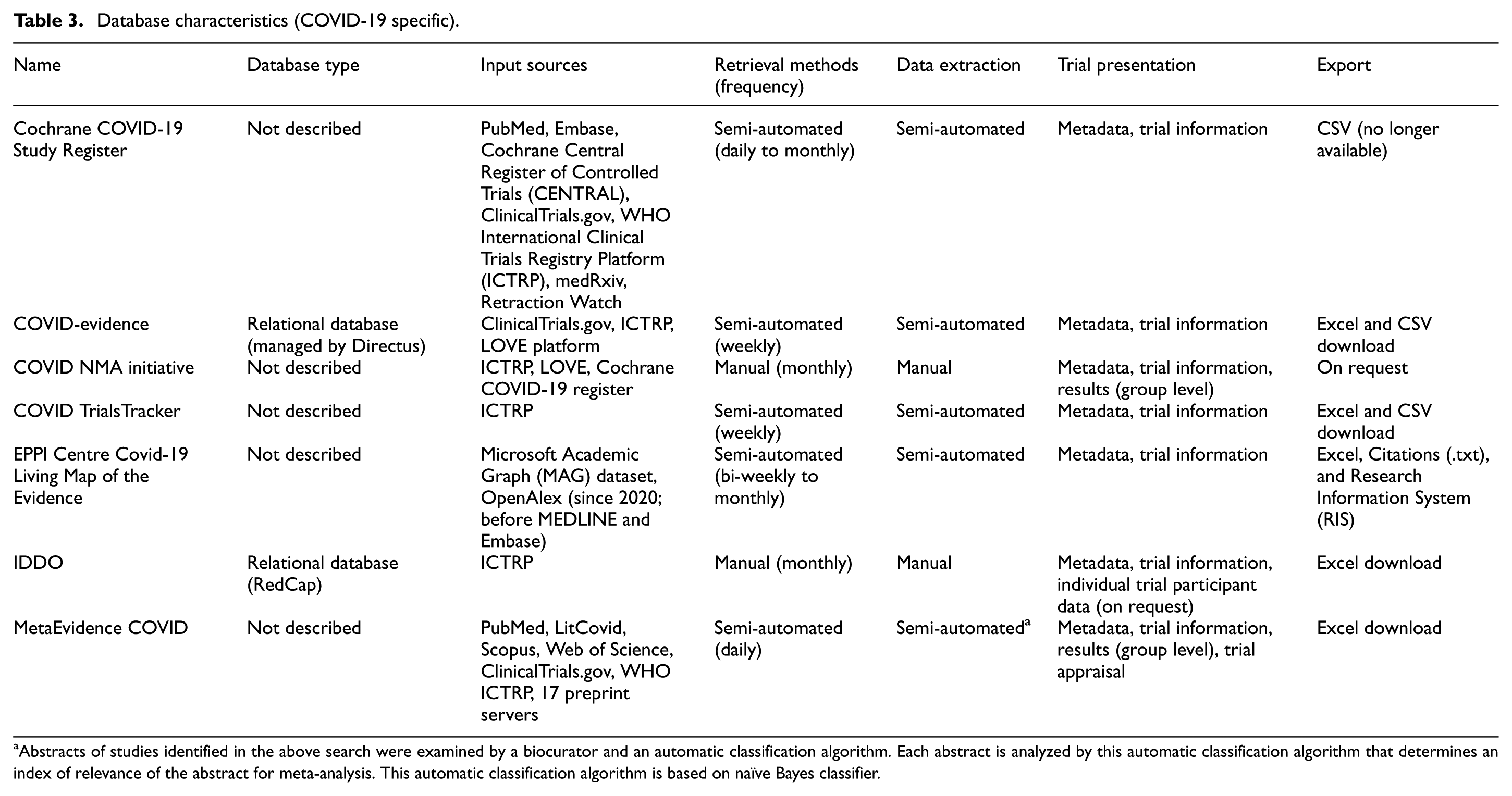
Abstracts of studies identified in the above search were examined by a biocurator and an automatic classification algorithm. Each abstract is analyzed by this automatic classification algorithm that determines an index of relevance of the abstract for meta-analysis. This automatic classification algorithm is based on naïve Bayes classifier.
All databases reported basic metadata, including trial information such as study type or design, sample size or expected starting date (extracted from ClinicalTrials.gov). Two (Covid NMA Initiative and MetaEvidence COVID) also reported aggregate group-level results, and while MetaEvidence COVID uniquely included trial appraisal through risk of bias assessments (Table 3; S7, Appendix in the Supplementary Material).
Discussion
Summary of evidence
We identified 14 clinical trial databases, of which seven were related to COVID-19. The databases differed significantly in their data retrieval methods and maintenance approaches. COVID-19 databases predominantly used semi-automated methods to retrieve and extract data, primarily from clinical trial registries. In contrast, non-COVID databases mainly relied on manual methods inspired by systematic reviews, which are typically time-consuming and difficult to maintain. While all databases mainly provided basic metadata, the availability of curated trial information, results, or trial appraisal was sparse. Only one database (IDDO) explicitly stated to share individual participant data (on request). This highlights a gap in data sharing and transparency for most databases.
Findings in context
Our results highlight a significant gap in the availability of continuously updated, topic-specific clinical trial databases. Existing databases largely borrow methods and technology from static article-based systematic reviews requiring manual retrieval, data extraction, and curation from unstructured journal articles. In addition, data captured or exported from clinical trial registries are often not easily integrated into structured databases, further compounding the issue.
Despite the development of various databases, none of the 14 platforms effectively addressed the fundamental problem of automatically integrating structured, machine-readable datasets from diverse sources. Currently, there are no publicly available standardised datasets that could seamlessly feed into such databases.
The first requirement to enable such continuously updated databases of clinical trial data is to have all trial data available in machine-readable and harmonised formats. 50 The idea of such machine-readable repository of RCTs dates back over 25 years back to the Global Trial Bank which was envisioned to go beyond trial registration and provide trial results in a computable format.51–54 Although the Global Trial Bank never materialised, the 2004 policy by the International Committee for Journal Medical Editors requiring trial registration as a condition for publication marked an important step. 55 This was followed by requirements to post summary results on ClinicalTrials.gov (from 2007) and the EU Clinical Trials Register (from 2012). 56
Some researchers have argued that clinical trial data should be made available in a structured dataset, such as the Common Technical Document format used for commercial drug applications. 57 Others have proposed that evidence synthesis, like systematic reviews, should rely primarily on clinical trial registries as the main data source, rather the traditional publication-based sources, to enable more timely and comprehensive synthesis. 58 The concept of the Global Trial Bank and of continuously updated, clinical trial databases lies at the intersection of these two ideas.
Future directions or emerging solutions
To sum up, the challenges to create living, continuously updated, topic-specific, trial databases can be addressed on several levels taking into consideration the life cycle of an individual clinical trial; (1) data may be captured and integrated directly from a clinical trial into a live database (green path); (2) data may be shared to clinical trial registries and reused in a uniform standardised format (yellow path); (3) data may be accessed and reused from traditional journal publications (red path) (Figure 2).
Capturing and reusing data directly from the clinical trials in standardised have been tested in initiatives, such as FDA Critical Path Initiative, 59 which has developed databases on Alzheimer’s,60,61 Multiple Sclerosis, 62 Parkinson’s, 63 and tuberculosis 64 clinical trial data. However, these resources remain private or public-private and are not universally accessible. The prevalence and accessibility of similar data platforms are not well documented, indicating a need for more transparent and accessible clinical trial data infrastructure.
There are several reasons why clinical trial registries in their current shape are challenging as data sources for creating topic-specific trial databases. These limitations include a lack of data harmonisation, data quality issues, and data export options (see Table 4 for details).
One potential tool to automatise data extraction from journal publications is TrialStreamer, a mega-collection of PubMed-indexed clinical trials (n = 852,723 as of December 2023), which includes annotated PICO information (population, intervention, comparator, outcome). 65 Trial Streamer utilises a blend of machine-learning and crowd-sourced manual extraction to gather trial data. While useful for meta-epidemiological studies, it lacks the granularity required for clinical applications or guideline development.
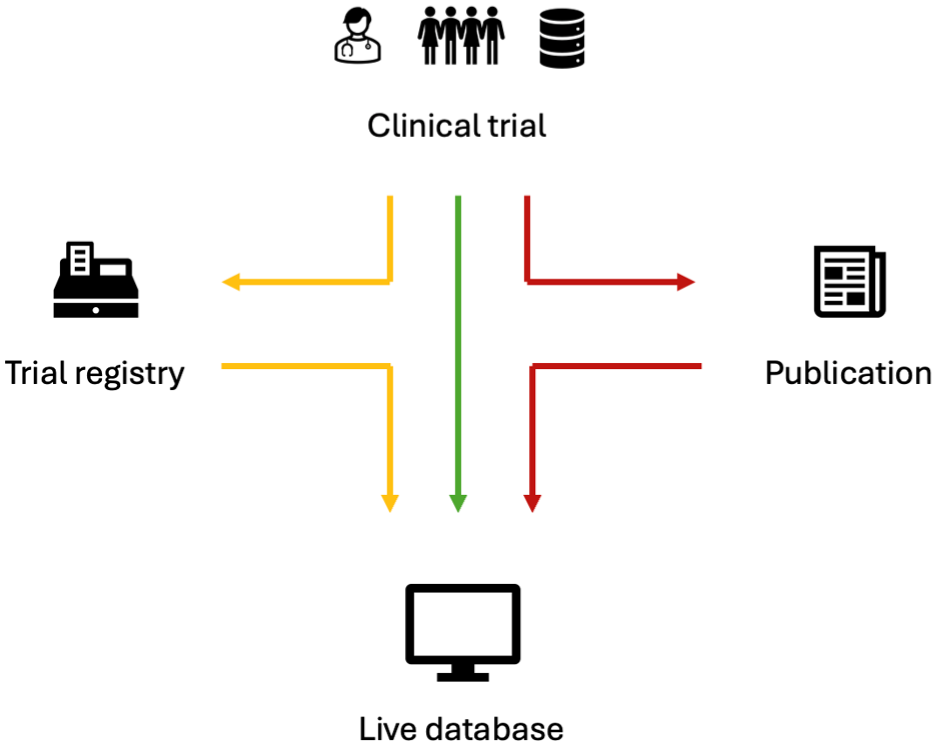
Pathways from clinical trial data into live topic-specific databases.
Challenges in using trial registries for topic-specific databases.

The importance of living evidence synthesis has been increasingly recognised, exemplified by the Wellcome Trust’s recent investment of GBP 45 million over 5 years to advance data infrastructure for dynamic evidence projects. This initiative reflects an acknowledgement of the need for more adaptable, continuously updated data ecosystems. 66
Limitations
Our scoping review has several limitations. We did not prespecify our methodology and register a protocol for this work. We believe that the impact is minimal considering the scope of the review and its exploratory nature without any quantitative analyses. We may have missed eligible databases during our searches due to their scarcity and lacking common terminology, and if such databases have not been described in related published articles, they may be invisible. It is likely that several ‘stealth’ 67 commercial solutions exist. An example of a public commercial solution is the LiveSLR programme,68,69 which is a software combining an annotated library of publications (called LiveRef) with a machine-assisted search/screening/ranking workflow, which creates an interactive live platform used in drug applications and health technology dossiers. Unfortunately, there is little transparency when it comes to such commercial solutions. We decided to not make detailed assessments of the included databases’ infrastructure, including granular descriptions of data retrieval and extraction from sources such as ClinicalTrials.gov. We did so mainly because these workflows were not transparently reported on websites or corresponding journal publications. Transparent and exhaustive reporting of the database infrastructure should be a priority because automation is the main methodological challenge. For example, the metaPreg platform reported to use ‘automatic software robots’ to screen databases, with no further specifications. 50 There may be several reasons for vaguely describing the methodology, for example, if setups are updated and adjusted frequently. However, open and exhaustive descriptions of platform infrastructure, including how automation is attempted and achieved, is encouraged.
Conclusions
This review is the first to systematically assess the characteristics and data management of continuously updated clinical trial databases. It provides insights into their design, limitations and highlights key technical and methodological challenges. We found only few topic-specific databases of clinical trials, which mainly relied on manual searches and manual data extraction. Such approaches are inefficient and time-consuming maintenance, compromising data timeliness and utility. Recent COVID-19 trial databases implemented semi-automated workflows, albeit confined to clinical trial registries. However, these workflows remain insufficient for integrating data from journal publications. A major challenge remains in automatically extracting and structuring trial data from publications. It seems timely to reconsider ideas such as the Global Trial Bank to centralise not only trial registration but also trial results reporting and data sharing. Facilitating the shift from article-based publications to a fully ‘digital’ machine-readable system of clinical trial data could enable true living evidence synthesis.
Supplemental Material
sj-pdf-1-ctj-10.1177_17407745251400635 – Supplemental material for Topic-specific living databases of clinical trials: A scoping review of public databases
Supplemental material, sj-pdf-1-ctj-10.1177_17407745251400635 for Topic-specific living databases of clinical trials: A scoping review of public databases by Kim Boesen, Lars G Hemkens, Perrine Janiaud and Julian Hirt in Clinical Trials
Supplemental Material
sj-pdf-2-ctj-10.1177_17407745251400635 – Supplemental material for Topic-specific living databases of clinical trials: A scoping review of public databases
Supplemental material, sj-pdf-2-ctj-10.1177_17407745251400635 for Topic-specific living databases of clinical trials: A scoping review of public databases by Kim Boesen, Lars G Hemkens, Perrine Janiaud and Julian Hirt in Clinical Trials
Footnotes
Author contributions
KB, PJ, and JH made substantial contributions to the conception and design of the work; KB have drafted the work, and all authors substantively revised it; all authors have approved the submitted version; all authors have agreed both to be personally accountable for the author’s own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature.
Kim Boesen: Conceptualisation, Methodology, Validation, Formal analysis, Investigation, Data Curation, Writing–Original Draft, Writing–Review & Editing, Visualisation, Supervision, Project administration
Lars G. Hemkens: Writing–Review & Editing, Funding acquisition
Perrine Janiaud: Conceptualisation, Methodology, Validation, Writing–Review & Editing, Supervision, Funding acquisition
Julian Hirt: Conceptualisation, Methodology, Validation, Formal analysis, Investigation, Data Curation, Writing–Review & Editing, Supervision, Project administration
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: LGH, PJ, and JH designed and worked on the COVID-evidence and PragMeta database. All authors worked on the continuously updated trial database, Cancer Immunotherapy Evidence Living (CIEL). All authors declare no financial conflicts of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The CIEL Project was funded by the Basel Cancer League (KLbB-5577-02-2022). RC2NB (Research Centre for Clinical Neuroimmunology and Neuroscience Basel) is supported by Foundation Clinical Neuroimmunology and Neuroscience Basel.
Data sharing statement
All data related to this review are available in the paper and the Appendix.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
