Abstract
Background:
In clinical trial development, it is a critical step to submit applications, amendments, supplements, and reports on medicinal products to regulatory agencies. The electronic common technical document is the standard format to enable worldwide regulatory submission. There is a growing trend of using R for clinical trial analysis and reporting as part of regulatory submissions, where R functions, analysis scripts, analysis results, and all proprietary code dependencies are required to be included. One unmet and significant gap is the lack of tools, guidance, and publicly available examples to prepare submission R programs following the electronic common technical document specification.
Methods:
We introduce a simple and sufficient R package, pkglite, to convert analysis scripts and associated proprietary dependency R packages into a compact, text-based file, which makes the submission document self-contained, easy to restore, transfer, review, and submit following the electronic common technical document specification and regulatory guidelines (e.g. the study data technical conformance guide from the US Food and Drug Administration). The pkglite R package is published on Comprehensive R Archive Network and developed on GitHub.
Results:
As a tool, pkglite can pack and unpack multiple R packages with their dependencies to facilitate the reproduction and make it an off-the-shelf tool for both sponsors and reviewers. As a grammar, pkglite provides an explicit trace of the packing scope using the concept of file specifications. As a standard, pkglite offers an open file format to represent and exchange R packages as a text file. We use a mock-up example to demonstrate the workflow of using pkglite to prepare submission programs following the electronic common technical document specification.
Conclusion:
pkglite and the proposed workflow enable the sponsor to submit well-organized R scripts following the electronic common technical document specification. The workflow has been used in the first publicly available R-based submission to the US Food and Drug Administration by the R Consortium R submission working group (https://www.r-consortium.org/blog/2022/03/16/update-successful-r-based-test-package-submitted-to-fda).
Introduction
In clinical trial development, it is critical to submit applications, amendments, supplements, and reports on medicinal products to regulatory agencies. Electronic Common Technical Document (eCTD) is the standard format to enable worldwide regulatory submission. For example, the United States Food and Drug Administration (US FDA) requires that new drug applications and biologics license applications be submitted using the eCTD format.
Within the eCTD submission, the regulatory agencies are increasingly interested in requiring sponsors to submit software programs used to create analysis data sets and analysis results. For example, the US FDA indicated in Section 4.1.2.10 of the study data technical conformance guide that “sponsors should provide the software programs used to create all ADaM datasets and generate tables and figures associated with primary and secondary efficacy analyses.” 1
There is a growing trend of using R for clinical trial analysis and reporting that can be part of the regulatory submission. For example, gsDesign
2
was used in sample size calculation for Moderna’s COVID-19 vaccine trial.
3
DoseFinding
4
was used to evaluate the multiple comparison procedure—modeling (MCP-MOD) approach by the FDA reviewers.
5
However, to the best of our knowledge, we have not observed any public examples of eCTD submission using R, although the FDA has provided a statistical software clarifying statement in 2015:
6
FDA does not require the use of any specific software for statistical analyses, and statistical software is not explicitly discussed in Title 21 of the Code of Federal Regulations [e.g., in 21CFR part 11]. However, the software package(s) used for statistical analyses should be fully documented in the submission, including version and build identification.
One unmet and important gap in creating such submissions is the lack of tools, guidance, and publicly available examples to prepare submission R programs following the eCTD specification. 7 For example, the eCTD specification requires lower case letters in all files and directory names of the submission package that is different from the file name conventions of R packages. 8 This article introduces a simple and sufficient R package, pkglite, to convert analysis scripts and associated proprietary dependency R packages into a compact, text-based file, which makes the submission document self-contained, easy to restore, transfer, review, and submit following the eCTD specification and regulatory guidelines (e.g. the study data technical conformance guide from FDA). The pkglite R package is published on Comprehensive R Archive Network (CRAN) and developed on GitHub. The contribution of pkglite for clinical trial submission is to (1) enable sponsors to transfer internally developed R packages within the eCTD package without using external code repositories (e.g. CRAN, GitHub), and (2) simplify and standardize the workflow in preparing and reviewing R program submissions between sponsors and reviewers with a compliant and reproducible approach.
Methods
We recommend organizing all analysis scripts of a clinical study following the R package folder structure to enhance reproducibility and ensure consistency.9,10 A minimal R package includes (1) a DESCRIPTION file stating important metadata such as authors, version, and dependencies of other R packages, (2) a NAMESPACE file scoping the exported and imported R functions, (3) an R/ directory of .R files containing R functions, (4) a man/ folder containing documentation files, (5) a vignette/ folder with analysis programs, and (6) a test/folder containing formal automated testing scripts. While not required for use with pkglite, it has significant advantages to use the R package folder structure. First, it is a well-defined format that organizes a self-contained bundle of assets, such as analysis programs, documentation, and tests, in a standard and consistent structure. Second, it can detect many common issues and enhance reproducibility by passing a collection of automated compliance checks using
After a clinical study project has been developed and organized as an R package folder structure, pkglite helps the sponsor prepare submission R programs. Typically, the eCTD package can include R functions, analysis scripts, analysis results, and all proprietary R package dependencies. For this entire scope of assets, pkglite can convert them into a compact, text-based file, which makes it easy to restore, transfer, review, and submit following the eCTD specification and regulatory guidelines. Once reviewers receive the text file in the eCTD package, they can unpack it by following the steps in the Analysis Data Reviewer’s Guide (ADRG). We provide a concrete example of the pack and unpack flow in the Results section.
The complete workflow to apply pkglite in preparing R packages for eCTD submission is shown in Figure 1. First, organize the R-based analysis and reporting code following the R package folder structure, which we refer to as the “project-specific R package” in the remainder of the article. Next, pack the project-specific R package and other proprietary R packages into a text file using pkglite. Then, this text file with submission programs will be placed in module 5 (clinical study reports) of the eCTD submission package. When reviewers receive the eCTD submission package, they can unpack and install the project-specific R package and other proprietary R packages from the text file to reproduce analyses performed using R. In comparison, the canonical workflow to distribute R packages requires two steps that are not compatible with the eCTD submission requirements: (1) using R CMD build to create a source tarball—a binary file that cannot be included in the eCTD submission, and (2) submitting the tarball to a public or access-controlled package repository (e.g. CRAN or GitHub) for distribution, which could be difficult for confidential file transfer between sponsors and reviewers due to security and compliance constraints and will not make the eCTD submission self-contained.
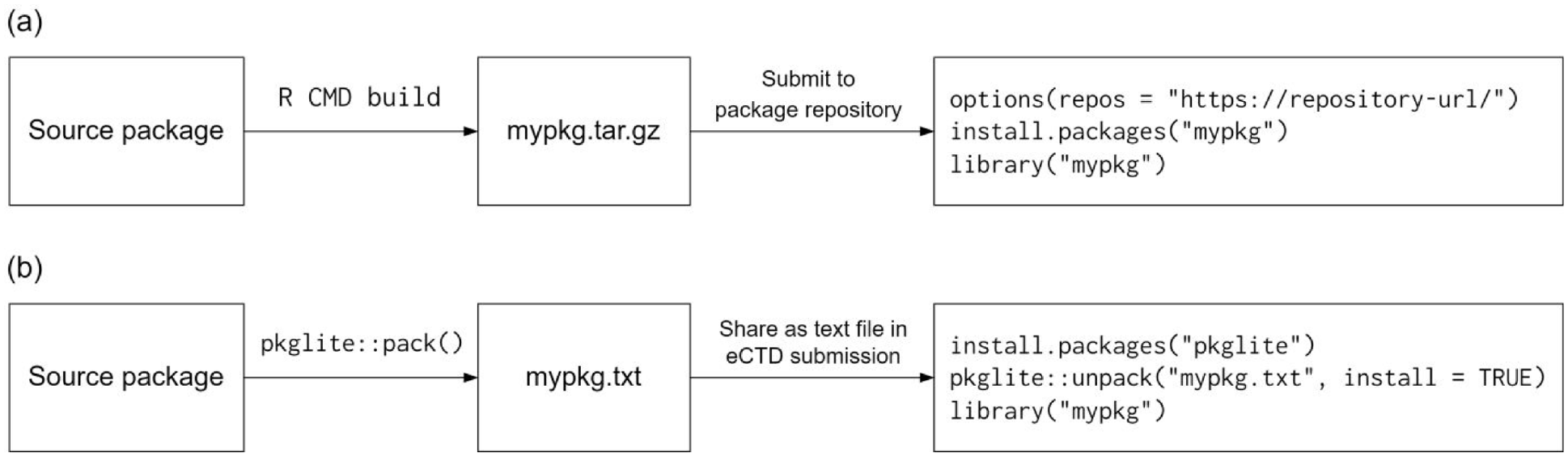
A comparison between the canonical and pkglite workflow to distribute an R package: (a) the canonical workflow for R package distribution and (b) the pkglite workflow for R package distribution.
Results
This section provides a real-world example to illustrate the procedure to submit a project-specific R package using pkglite. We assume the project-specific R package is esubdemo (available at https://github.com/elong0527/esubdemo). The goal is to pack and save it under module 5 of an eCTD submission package. In this example, the eCTD package is ectddemo, available at https://github.com/elong0527/ectddemo.
To achieve the goal above, we first use
Here, “esubdemo/” is the assumed folder path for the project-specific R package (unpacked). The
After packing the project-specific R package and other proprietary R packages into a plain text file, we can save the packed text files in module 5 of the eCTD package, specifically under “m5/datasets/ectddemo/analysis/adam/programs/” following the eCTD specification. It is acknowledged that under the adam/ folder for analysis data, there are two sub-folders: programs/ and datasets/. The datasets/ sub-folder consists of clinical trial data following the ADaM implementation guide. The programs/ sub-folder contains the submission program, including the packed project-specific package, proprietary R packages, as well as the R scripts for each analysis (tlf-*.txt files). It is worth noting that R Markdown files can be transferred into R scripts by using
Next, after verifying the correctness of the packed R packages, the sponsor should update the Analysis Data Reviewer’s Guide (ADRG) to provide guidelines for the reviewers to restore the packed assets. An ADRG template demonstrating the impacted sections for the ectddemo example is available at https://github.com/elong0527/ectddemo/blob/master/m5/datasets/ectddemo/analysis/adam/datasets/adrg.pdf. Following the ADRG, reviewers can easily unpack and install the R packages after receiving the submission package by using
After installing the packages, reviewers can reproduce the trial analysis results by running the R scripts for each analysis (tlf-*.txt files), which often rely on the functions from the proprietary packages.
Conclusion
In this article, we demonstrate the usage of pkglite, that is, allowing sponsors to prepare R packages into compact text files. To provide hands-on experience to readers, we present a mockup example, where we introduce the recommended workflow to pack R packages into text files that can be incorporated into the regulatory submission package following the eCTD specification. After regulatory agencies receive the submission package, we provide methods to unpack R packages from the text file that can be used to execute all analysis scripts.
In addition to the usage of pkglite, we also demonstrate the advantages of using pkglite in preparing R-based analysis and reporting programs for eCTD submissions. First, by applying pkglite, project-specific R packages are packed and restored as plain text files as required by the FDA submissions gateway, making it off-the-shelf for both sides of the submission to store, transfer, and review. Second, the packed project-specific R packages are presented in an unambiguous, human-friendly, and machine-readable format. Third, pkglite is pipe-friendly (e.g. using the native pipe operator|
Following similar steps, the R Consortium R submission working group has prepared and submitted an eCTD submission package to the FDA using pkglite. The FDA reviewer is able to reproduce analysis results following instructions in the ADRG. The submission package is publicly available at https://github.com/RConsortium/submissions-pilot1-to-fda. The final response letter from FDA is at https://github.com/RConsortium/submissions-wg/blob/main/Documents/Summary_R_Pilot_Submission2.pdf.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
