Abstract
Matrix metalloproteinases (MMPs) are a family of zinc-dependent endopeptidases, which play critical roles in cancer progression and metastasis. In recent years, many researchers have been studying the relationship between MMP9 and breast cancer. However, it still remains indecisive. Therefore, we conducted a meta-analysis to draw more accurate conclusions. A total of 21 relevant documents were retrieved, including 25 case-control studies. We quantitatively analyzed the data obtained. To clarify the relationship between MMP9 polymorphism and breast cancer susceptibility under different conditions, we also made a further subgroup analysis for each locus. In summary, we discovered that MMP9 rs3918242 rendered an increased risk for breast cancer, especially among Iranians and Indians. MMP9 rs3787268 could be a protective factor. MMP9 rs17576 and MMP9 rs2250889 have no association with breast cancer risk.
Introduction
Breast cancer is the most common form of cancer and the leading form of cancer-related deaths among females worldwide (22.9% of total cases). 1 Around the globe, it is estimated that the number of diagnosed breast cancer cases was 2,088,849, out of which 626,679 estimated cases resulted in death. 2
Compared to the general population, women who are first-degree relatives of those diagnosed with breast cancer are more prone to this disease. Approximately 5–10% of all cases are due to inherited predisposition, caused by mutations in susceptible genes. 3 Although genetic testing for BRCA1 and BRCA2 mutations has become an integral part of clinical practice for women with family histories of breast cancer, other genetic mutations or variations could be potentially useful in clinical practice.
Matrix metalloproteinases (MMPs) are a family of zinc-dependent endopeptidases that play critical roles in cancer progression and metastasis. 4 The MMP family consists of more than 20 different enzymes and is involved in the degradation of the extracellular matrix (ECM). 5 Among the MMP family, MMP9 (gelatinase B) is demonstrated to have functional roles in tumor angiogenesis, invasion, and metastasis. 6 MMP9 has proteolytic activity against type IV collagen, a major component of the basement membrane. The expression of MMP9 is upregulated in different types of cancer, including breast cancer. 7 The previous meta-analysis has shown that the TT genotype of rs3918242 polymorphism might be a risk factor for breast cancer. 8 However, more studies investigating MMP9 polymorphisms and breast cancer susceptibly have been published since the last meta-analysis. In addition, we included up-to-date literature and more subgroup studies in research to ensure that we obtained more accurate results. Therefore, we updated this meta-analysis to reevaluate the relationship between MMP9 polymorphism and breast cancer susceptibility with recently published studies and more comprehensive analysis.
Materials and methods
Publication search
We have carried out a comprehensive literature search through various databases, such as PubMed, Google scholar, Embase, Web of Science, CNKI and Wanfang database. The search terms are: ((matrix metalloproteinase 9) OR (MMP-9) OR (MMP9) OR (gelatinase B)) AND (cancer OR neoplasms OR tumor OR carcinoma OR malignancy) AND (polymorphism OR SNP OR mutation OR variant OR genotype) AND (breast OR mammary). The last retrieval time of the information and the literature was October 2020. We excluded articles unrelated to this study by evaluating titles and abstracts. We selected the relevant information and literature after careful full-text reading. This study was conducted in accordance with the requirements of Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA).
Inclusion criteria
We have established the following inclusion criteria for retrieved articles. (a) A cohort or case-control study on MMP9 polymorphism and breast cancer susceptibility. (b) Genotype frequency data of case group and control group can be obtained by direct extraction or indirect calculation, and odds ratios (ORs) and 95% confidence intervals (CIs) can be calculated.
Exclusion criteria
Correspondingly, the following conditions should be excluded from the study: (a) repeated publications, case reports, letters, systematic reviews or meta-analyses; (b) publications conducted on cell lines or animal models; (c) studies with missing genotype frequency data; and (d) studies unrelated to breast cancer.
Data extraction
Two investigators (C.Y. and C.S.) independently searched the database. C.Y. was responsible for summarizing all the search results. Then, C.Y. and C.S. independently screened the articles that conformed to the exclusion criteria and inclusion criteria. D.L. summarized the articles screened by the above two researchers again and checked the screening results according to the inclusion and exclusion criteria. Finally, three researchers collectively confirmed that all eligible articles have been included. For disputed issues, the differences were resolved by re-examining and consulting relevant information. Finally, the following data were extracted from the qualified studies: the name of the first author, publication year, country, location, source of control groups, genotyping method, sample size, and allele and genotype frequencies in the cancer cases and controls.
Study quality assessment
The quality of each study was assessed using the Newcastle–Ottawa Scale (NOS) and evaluated based on the following criteria: study ratings of seven to nine stars indicated a high quality; five to six stars indicated a moderate quality; and four stars or fewer indicated a low quality. 9 The assessment was done by three researchers (C.Y., C.S., and D.L.) independently and any discrepancies were discussed after the results were confirmed to ensure that accurate information was being provided.
Statistical analysis
All statistical calculations in this study were performed using Stata 12.0 software (STATA Corp, College Station, TX). The relationship between MMP-9 polymorphism and susceptibility to breast cancer was assessed with ORs and 95% CIs. Heterogeneity within the studies was assessed by the I2 statistics. The random effects model was selected to pool the data. If not, the fixed effects model was adopted. The Z test was used to determine statistical significance. In order to assess the stability of the included studies, sensitivity analyses were also performed by omitting each included study one by one, and switching the fixed effects model to the random effects model or vice versa. Publication bias was assessed by visually inspecting the symmetry of funnel plots, as well as the Begg's test and the Egger's test.
Results
Literature search
We retrieved a total of 1204 related documents from various databases. After checking the inclusion and exclusion criteria, the full text of the literature was evaluated. For studies that could possibly contain data we needed, we also contacted the authors for the original data by Email. Finally, 21 relevant documents10–31 were retrieved, including 25 case-control studies, in which there were two documents containing three case-control studies. A total of 10,336 cases and 11,000 controls were included. The screening flow chart is shown in Figure 1. Among these 25 studies, there are 1510–21 on rs3918242; 4 studies21–24 on rs17576; 3 studies22,23,25 on rs2250889; and 3 studies24,26,27 on rs3787268. The characteristics of the included studies are shown in Table 1.
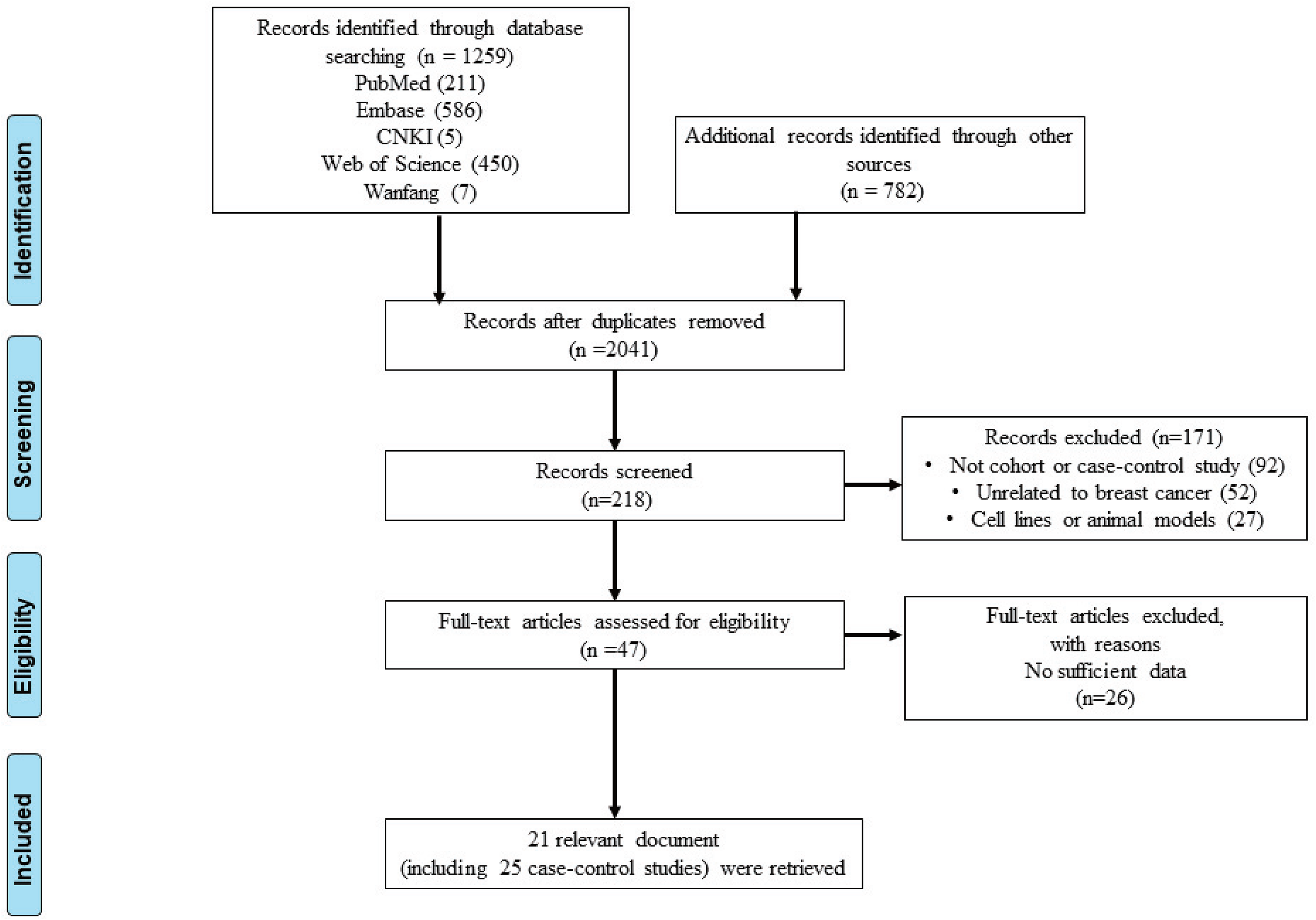
Flow diagram of article selection in this meta-analysis.
Main characteristics of MMP9 studies.
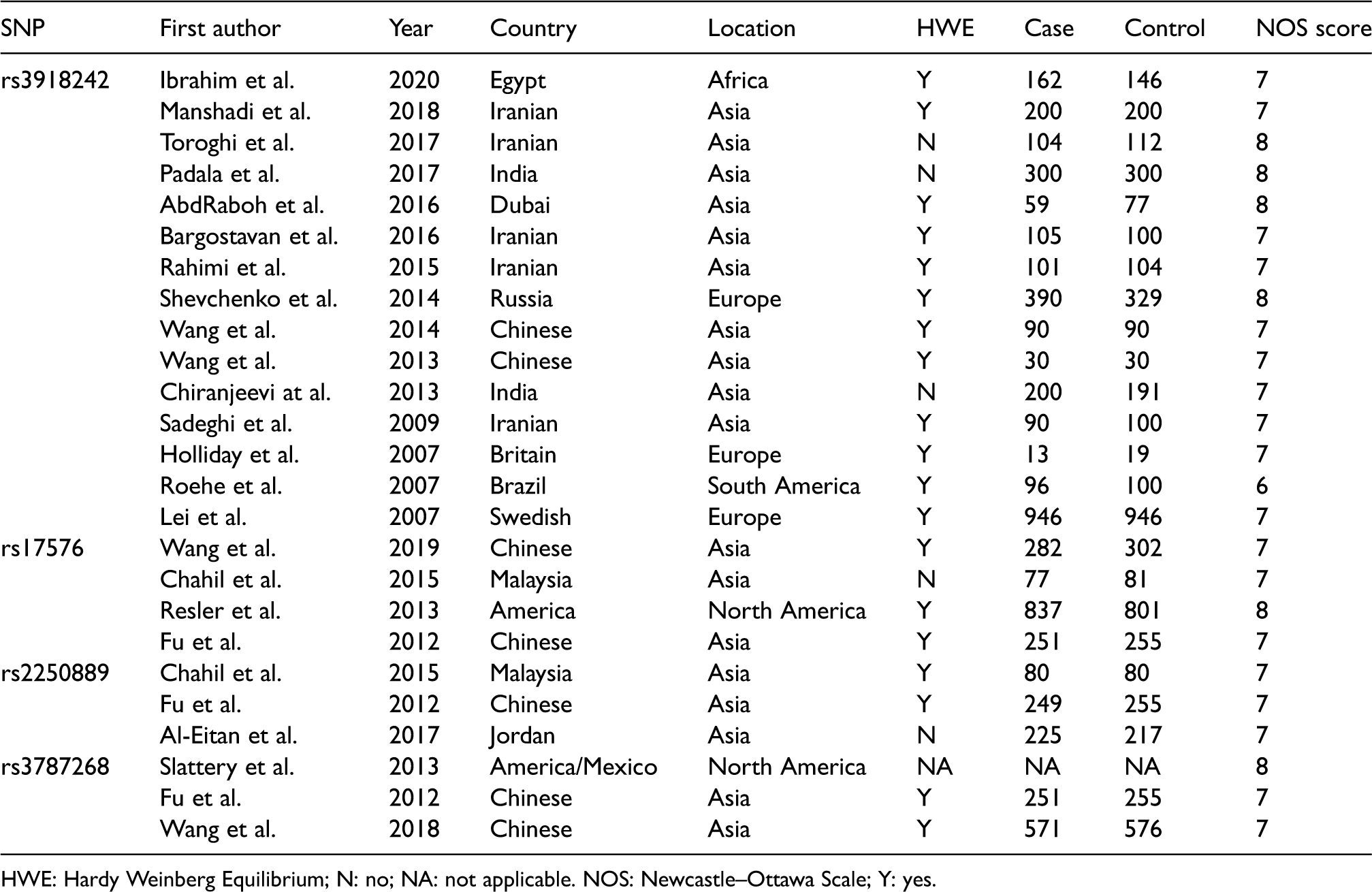
HWE: Hardy Weinberg Equilibrium; N: no; NA: not applicable. NOS: Newcastle–Ottawa Scale; Y: yes.
Quantitative analysis
The results of MMP9 polymorphisms and breast cancer risk analyses are listed in Table 2. From the overall results, for rs3918242, most the overall genetic models showed increased risk (T vs. C: OR = 1.426, 95% CI = 1.137–1.788, P = 0.002, Figure 2(a); CT vs. CC: OR = 1.434, 95% CI = 1.143–1.799, P = 0.002; TT vs. CT + CC: OR = 1.493, 95% CI = 1.168–1.908, P = 0.001). For rs3787268, allelic comparison, homozygote comparison, and heterozygote comparison showed reduced risk (A vs. G: OR = 0.824, 95% CI = 0.716–0.949, P = 0.007; AA vs. GG: OR = 0.701, 95% CI = 0.517–0.951, P = 0.022; AG vs. GG: OR = 0.776, 95% CI = 0.629–0.958, P = 0.018) in the overall populations. As to rs17576 and rs2250889, no significant effects on breast cancer risk were found among the overall populations. To clarify the relationship between MMP9 polymorphism and breast cancer susceptibility under different conditions, we also made further subgroup analysis for each locus.
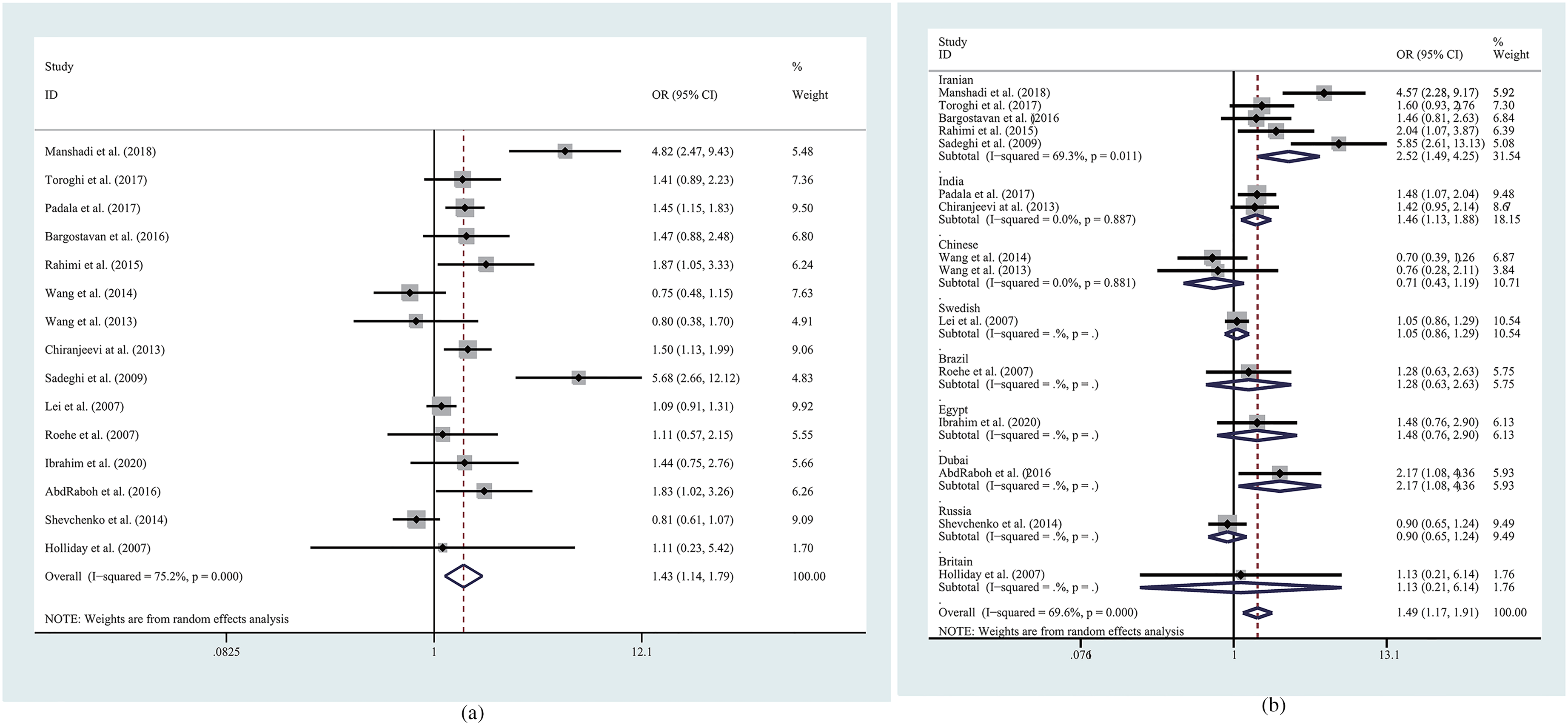
Forest plots of the association between MMP9 rs3918242 polymorphism and breast cancer risk.
Results of meta-analysis for polymorphisms in MMP9 and breast cancer susceptibility.
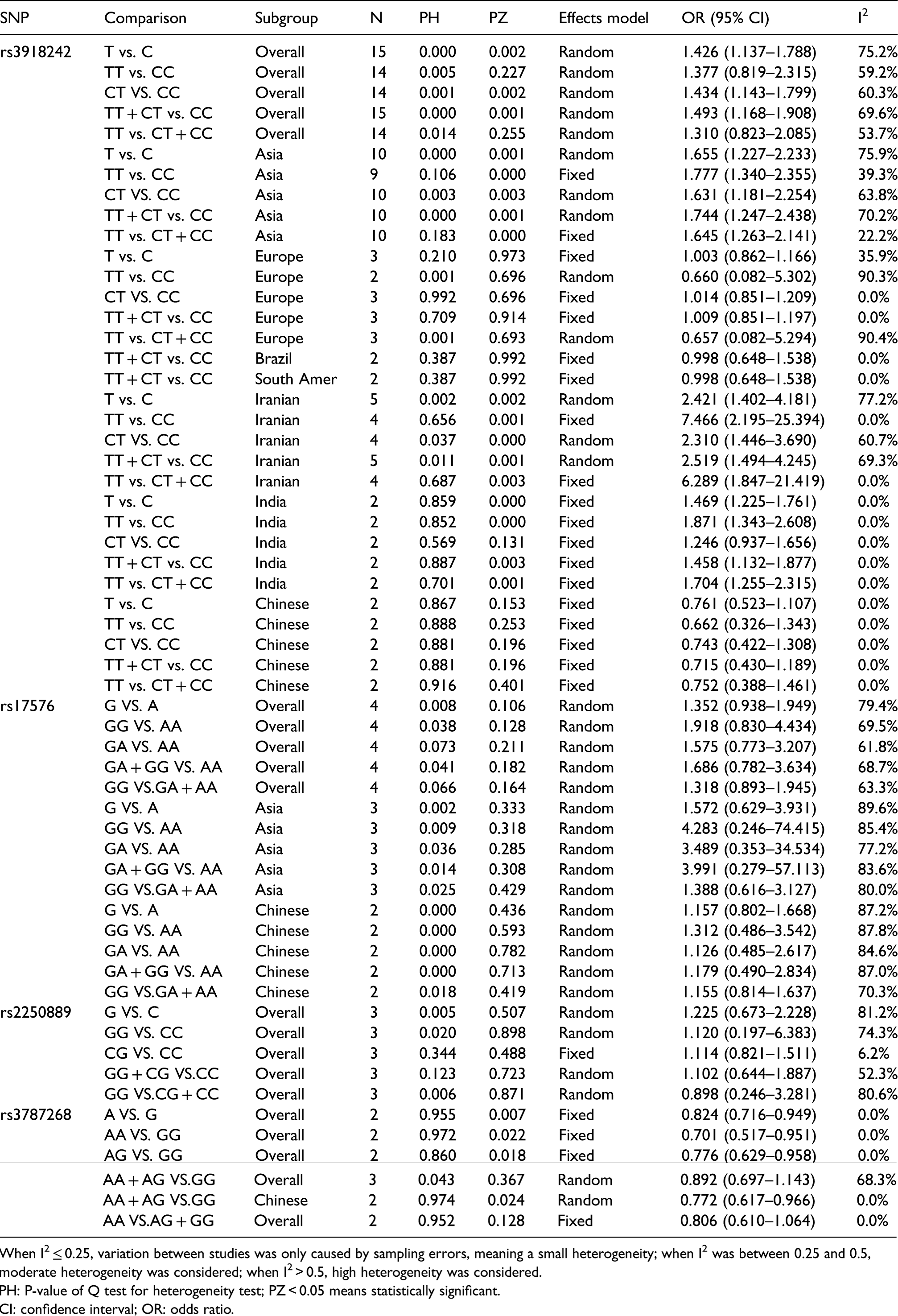
When I2 ≤ 0.25, variation between studies was only caused by sampling errors, meaning a small heterogeneity; when I2 was between 0.25 and 0.5, moderate heterogeneity was considered; when I2 > 0.5, high heterogeneity was considered.
PH: P-value of Q test for heterogeneity test; PZ < 0.05 means statistically significant.
CI: confidence interval; OR: odds ratio.
Stratification analysis by location and country
Subgroup analyses of the susceptibility of MMP9 polymorphisms to breast cancer in different locations showed that rs3918242 was associated with an increased risk of breast cancer for populations in Asia (T vs. C: OR = 1.655, 95% CI = 1.227–2.233, P = 0.001; TT vs. CC: OR = 1.777, 95% CI = 1.340–2.355, P < 0.001; CT vs. CC: OR = 1.631, 95% CI = 1.181–2.254, P = 0.003; TT vs. CT + CC: OR = 1.744, 95% CI = 1.247–2.438, P = 0.001; TT + CT vs. CC: OR = 1.645, 95% CI = 1.263–2.141, P < 0.001). An increased risk of breast cancer for rs3918242 was also found among Iranian populations (T vs. C: OR = 2.421, 95% CI = 1.402–4.181, P = 0.002; TT vs. CC: OR = 7.466, 95% CI = 2.195–25.394, P = 0.001; CT vs. CC: OR = 2.310, 95% CI = 1.446–3.690, P < 0.001; TT vs. CT + CC: OR = 6.289, 95% CI = 1.847–21.419, P = 0.003; TT + CT vs. CC: OR = 2.519, 95% CI = 1.494–4.245, P = 0.004, Figure 2(b)). Similarly, there were statistically significant increased breast cancer risks for rs3918242in the allelic comparison, homozygote comparison, dominant model, and recessive model of the Indian subgroup (T vs. C: OR = 1.469, 95% CI = 1.225–1.761, P < 0.001; TT vs. CC: OR = 1.871, 95% CI = 1.343–2.608, P < 0.001; TT vs. CT + CC: OR = 1.704, 95% CI = 1.255–2.315, P = 0.001; TT + CT vs. CC: OR = 1.458, 95% CI = 1.132–1.877, P = 0.003). However, there was no increased risk of MMP9 rs3918242 on breast cancer susceptibility in Europe, South America, or among the Chinese population. In the subgroup analysis of MMP9 rs3787268, dominant model reduces breast cancer risk (AA + AG vs. GG: OR = 0.772, 95% CI = 0.617–0.966, P = 0.024).
Sensitivity analyses and publication bias
In order to assess the impact of different models, we conducted a sensitivity analysis by deleting individual studies and the replacement of effect models, whereas the overall statistical significance did not change, indicating that the results are stable (Figure 3(a)). We used the Begg's and Egger's tests for the detection of publication bias; the results showed that there was no obvious publication bias (Figure 3(b)).
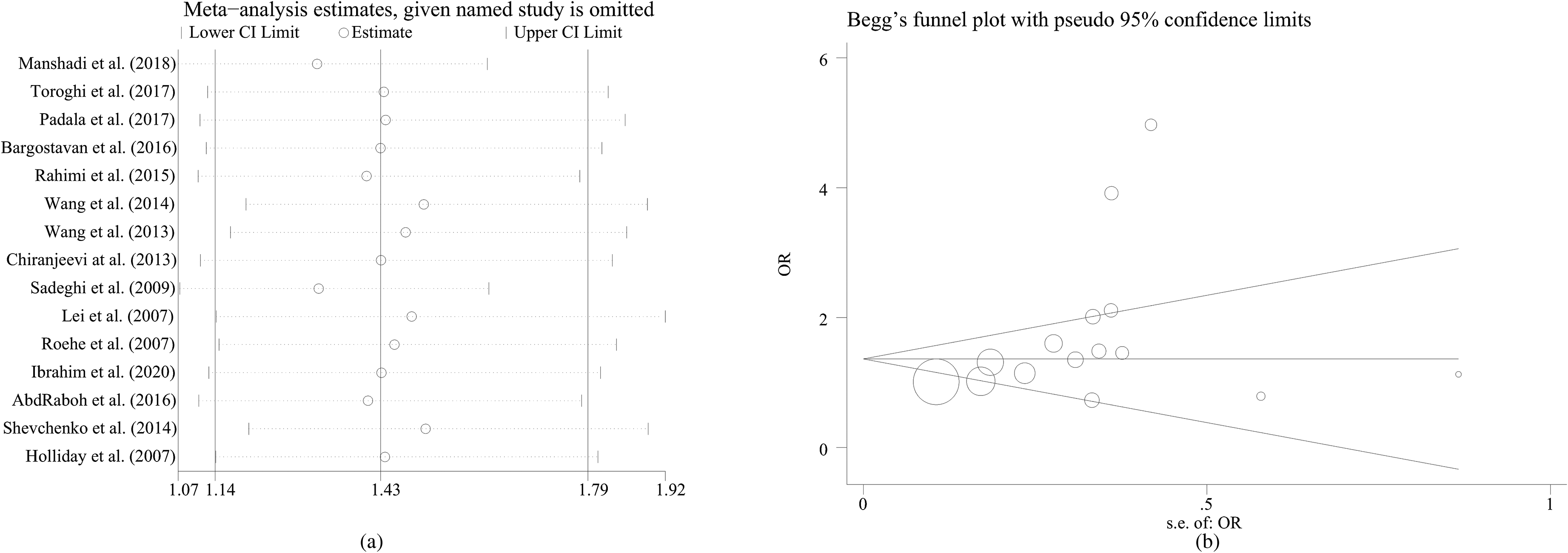
Sensitivity analysis and Begg's funnel plot for publication bias under MMP9 rs3918242 polymorphism.
Discussion
The gene MMP-9 is located on chromosome region 20q11.2-q13.1. 32 Studies have shown that the expression of MMP9 is controlled by different biological mechanisms. One study demonstrated that the C to T substitution in the promoter region of the MMP9 gene has a higher promoter activity at position −1562, 33 which could lead to higher risk of carcinogenesis due to the angiogenesis and tumor progression by MMP9. 34 However, other studies also found opposite effects of MMP9 on tumorigenesis; for example, one study showed that the absence of MMP9 significantly delayed tumorigenesis in the C3(1)-Tag model of basal-like triple-negative breast cancer. 35 Due to the complexity of mechanisms of MMP9 effects on tumorigenesis, it is of great value to explore the impact of MMP-9 polymorphisms on breast cancer risk.
MMP9 rs3918242 is a functional polymorphism in the promoter region. 35 Our results suggested that T-allele and CT genotype in MMP9 rs3918242 had a significant association with an increased risk of breast cancer in the overall analysis. This is consistent with the findings of previous research, which have shown that MMP9 was highly expressed in the breast cancer cell, 36 and T-allelic promoter at position −1562 had higher promoter activity than the C-allele. 35 In the subgroup analysis of location, the same conclusions could be drawn for Asia. In the subgroup analysis of ethnicity, increased risks were also found among the Indian populations, except for CT versus CC, which could be explained by the T allele in CT still being a relatively strong risk factor. Among the Iranian population, the T allele, CT genotype, and TT genotype were also found to be associated with increased risk. However, among the Chinese and Brazil populations, no significant associations were found, which could be either due to the small number of original studies included, or attributed to the true difference between the Chinese population and other ethnic groups.
Previous studies have found that the rs17676 polymorphism is associated with the risk of multiple cancers. However, in this study, no significant alterations of breast cancer risk were found in the overall analysis, the subgroup analysis of Asia, or the hospital-based study of MMP9 rs17576. Some studies suggest that this is because the site plays different roles in the development of different cancers. 24 However, more studies are needed to confirm this conclusion.
For rs3787268, statistically significant decreases of breast cancer risk were found to be associated with the presence of the A allele in the allelic comparison, homozygote comparison, and heterozygote comparison. In the dominant model, there was no change of risk in the overall group, but a decreased risk was found among the Chinese population. These results suggested that the A allele, AA genotype, and AG genotype might be a protective factor for breast cancer. Our results confirm the conclusions of Wang et al. 27 However, Slattery et al. 26 came to the opposite conclusion. Their study suggests that the rs3787268 polymorphism increases the risk of breast cancer. However, to clarify the associations more studies are needed.
For rs2250889, Al-Eitan et al. 25 argued that there was no significant correlation between the risk of breast cancer; this was also found in our study. However, due to the small number of the original studies focusing on these polymorphisms, further studies are needed to confirm our conclusions.
Results of the sensitivity analysis show that there is some heterogeneity in the study of rs17576, which may be the result of the quality difference. More studies are needed to confirm or exclude the relevant heterogeneity.
However, our research had several limitations. First, is the paucity of data; most results of the subgroups analyses are not very robust because of the limited number of studies. As we do not have enough published data, some loci studies of small samples can only provide a certain reference value. Second, significant heterogeneity was observed in some of the overall comparisons and subgroup analysis. Subgroup analysis showed that ethnicity and different techniques could be one of the sources of heterogeneity. Third, this meta-analysis may suggest the direction for future studies, but gives no support to the speculation of actual differences between ethnic subgroups; more well-designed, larger studies in various populations are needed.
In conclusion, we discovered that MMP9 rs3918242 rendered an increased risk for breast cancer, especially among Iranians and Indians. MMP9 rs3787268 could be a protective factor. Also, MMP9 rs17576, rs2250889 has no association with breast cancer risk. To confirm the associations found in this meta-analysis, more well-designed and larger studies in various population groups are needed.
Footnotes
Author contributions
Cunye Yan, Chenyu Sun, and Dengwei Lu contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Chongqing, China and Medical Science and Technology Innovation Fund of Chongqing General Hospital (grant number cstc2020jcyj-msxmX0668, Y2020MSXM14).
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Provenance and peer review
Not commissioned; externally peer reviewed.
Patient consent for publication
Not required.
