Abstract
Background:
The epidermal growth factor receptor (EGFR) represents a molecular target for tyrosine kinase inhibitors for non-small cell lung cancer (NSCLC) patients with a mutation in the EGFR gene. Mutations of the EGFR gene that occur at a single position in NSCLC tissue are found as single, whereas two or more mutations on the same allele are poorly detected and investigated.
Patient and methods:
We investigated the presence of the EGFR gene mutations in tumor tissue by Sanger sequencing and ion torrent sequencing in an NSCLC patient at Stage IV of disease.
Results:
We found the presence of three coexisting mutations on the EGFR gene—two of which on exon 21 are present on the same allele, and the third, on exon 20, was analyzed by Sanger sequencing of the peripheral blood lymphocytes. The patient staged as cT4N0M1c (Stage IV) and started afatinib 40 mg daily 8 months ago, showing a clinical benefit.
Conclusion:
In this report we describe the case of an NSCLC patient harboring three coexisting mutations on the EGFR gene, two of which are present on the same allele. This mutation pattern may represent, for patient progeny, a genetic risk of cancer development. Therefore it should be possible to obtain screening guidelines to improve the risk calculation for lung cancer susceptibility in the future.
Introduction
Non-small cell lung cancer (NSCLC) represents the major cause of cancer-related mortality in the world 1 . For lung cancer, and in particular for NSCLC patients, the epidermal growth factor receptor (EGFR) represents a molecular target for tyrosine kinase inhibitors (TKIs) 2 . In clinical practice, the EGFR TKIs confer a better outcome in patients with EGFR activating mutations than those without mutation. To date, approximately 5–15% of NSCLC patients harbor EGFR mutations—most of which occur in exons 19 and 21, each of them accounting for approximately 45% of EGFR gene alterations. The most frequent exon 19 mutations, namely in-frame deletions of amino acids 746–750, are clustered around the catalytic domain and flank the ATP-binding site. While promoting the activation of EGFR TK signaling, the structural changes introduced by these mutations enhance TKI drug binding, allowing a complete blockade of EGFR signaling at relatively low TKI doses. The Leu858Arg substitution is the most frequent mutation occurring in exon 21 within the TK activation loop 3 . The remaining 10% of EGFR TK mutations occur in exons 18 and 20; they do not usually increase the sensitivity to EGFR TKIs, and in some cases they are associated with resistance. Indeed, the Thr790Met (exon 20) often occurs as a secondary mutation in tumors treated with EGFR TKIs and is thought to account for about half of all cases of acquired TKI resistance 4 .
Most often, mutations of the EGFR gene occur at a single position, in NSCLC tissue, and the potential association of two or more mutations on the same allele are poorly detected and investigated.
In this report we describe the case of an NSCLC patient harboring three coexisting mutations on the EGFR gene, two of which are present on the same allele.
Material and methods
DNA extraction
DNA was extracted from formalin-fixed and paraffin-embedded (FFPE) tumor tissue obtained by NSCLC tumor biopsy and from circulating leucocytes; written informed consent was obtained from the patient prior to participation in the study.
Xylene was added once and ethanol was added twice to remove all paraffin from the tissue sample. DNA from FFPE tissue was extracted using QIAamp DNA FFPE Tissue kit (Qiagen GmbH, Hilden, Germany) while DNA from peripheral leucocytes was extracted using QIAamp Blood Kit (Qiagen GmbH, Hilden, Germany) according to the manufacturer’s instructions. Eluted DNA was quantified with Qubit 2.0 Fluorometer (Thermofisher).
IT-PGM sequencing and variant calling
Approximately 10 ng of DNA was required to construct the barcoded and adaptor-ligated library using the Ion AmpliSeq Library Kit 2.0 (Thermofisher). The sample was analyzed using the Ion AmpliSeq Colon and Lung Cancer Research Panel V2 (CLV2) containing a single primer pool to amplify hotspots and targeted regions of 22 cancer genes frequently mutated in colorectal cancers and NSCLCs (https://www.ampliseq.com) 5 . Templated spheres were prepared using 100 pM of each library using the Ion One Touch 2.0 machine. Template-positive spheres were loaded into Ion chip 314 and sequenced by the IT-PGM machine (Thermofisher). Sequencing data were analyzed with the Ion Torrent Suite. Variants with a quality <30 were filtered out. Polymorphism Phenotyping v2 (PolyPhen-2), Sorts Intolerant From Tolerant substitutions (SIFT), and PROtein Variation Effect Analyzer (PROVEAN) computational tools were used to predict the possible impact of amino acid substitutions on the structure and function of protein mutated using straightforward physical and comparative considerations.
Sanger sequencing
In order to investigate the presence of the germline mutation on EGFR, exons 18, 19, 20, and 21 were sequenced. DNA was amplified by polymerase chain reaction (primers are listed in Table 1). Sequencing was performed using the BigDye Terminator v3.1 Cycle Sequencing Kit and a 3130XL Genetic Analyzer (Applied Biosystems). Sequences were finally analyzed with the SeqScape v2.5 software (Applied Biosystems) 6 .
Primer sequences.
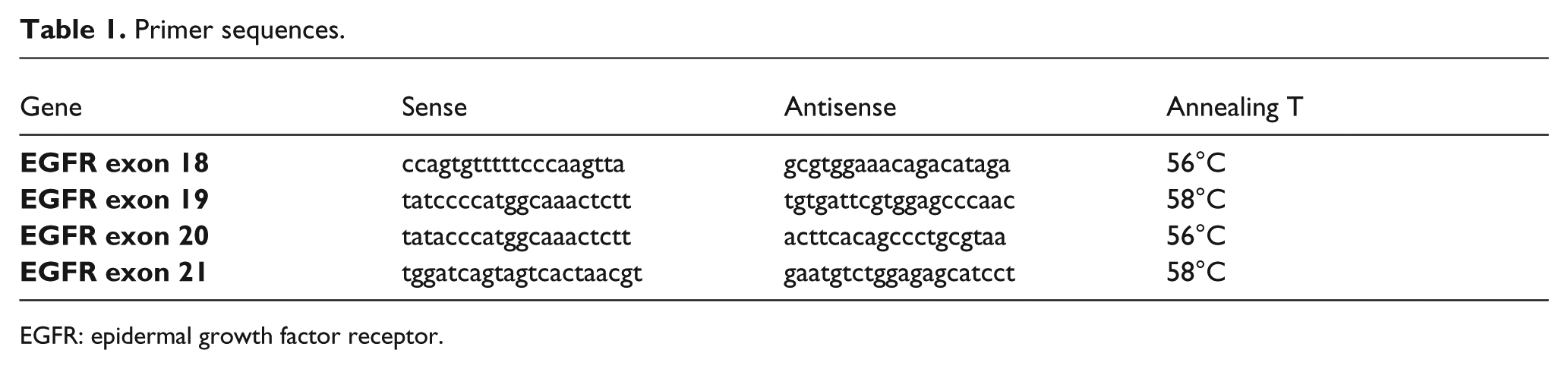
EGFR: epidermal growth factor receptor.
Case description
In June 2017, a 54-year-old Caucasian female, a smoker (30 pack/years), was admitted to the Emergency Department of Policlinico Umberto I Rome with headache, dizziness, nausea, and nystagmus. The cytological exam from a lumbar puncture showed the presence of neoplastic cells. The patient underwent a total body computed tomography (CT) scan with evidence of a mass in the left lower lobe (16×14mm), multiple bilateral lung metastases and several bone lesions. The histological examination, obtained by IR-guided primary lung lesion biopsy, revealed an NSCLC, CK7+ and TTF1+ adenocarcinoma. EGFR mutation analysis was performed, showing Leu858Arg (exon 21), Leu861Arg (exon 21), and Arg776His (exon 20) mutations. The neurological symptoms were linked to leptomeningeal carcinomatosis even if the brain magnetic resonance imaging (MRI) did not show any sign of brain metastases or leptomeningeal involvement. She was staged as cT4N0M1c (Stage IV) according to the AJCC Cancer Staging Manual (8th edition). In August 2017 the patient started afatinib, 40 mg daily, which showed a clinical benefit with a partial regression of neurological symptoms since the first few months of treatment. The patient experienced TKI-related cutaneous toxicities: skin xerosis G2, pruritus G1, and peripheral edema. The total body CT scan performed in November 2017 revealed a partial response on the lung mass and nodules, and stable disease on the bone lesions (Figure 1). The TKI treatment is currently ongoing with a progression-free survival of 8 months.
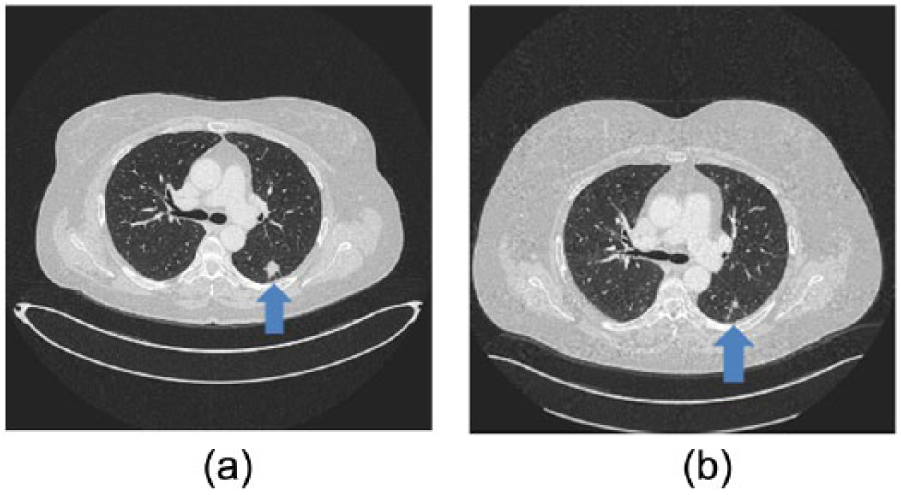
(a) CT scan before treatment with afatinib. (b) CT scan 3 months of treatment with afatinib.
Our next-generation sequencing (NGS) revealed the presence of three previously reported missense mutations on the EGFR gene: a Leu858Arg and Leu861Arg (exon 21) occurring with an extremely similar allele frequency (40.3% and 41.3%, respectively) and lying on the same allele (Figure 2(a)); the third mutation was an Arg776His (exon 20) occurring with an allele frequency of 67.5%. In order to detect the possible germline origin of such mutations we performed Sanger sequencing on DNA extracted from lymphocytes. The results obtained revealed that only Arg776His was germline (Figure 2(b)).
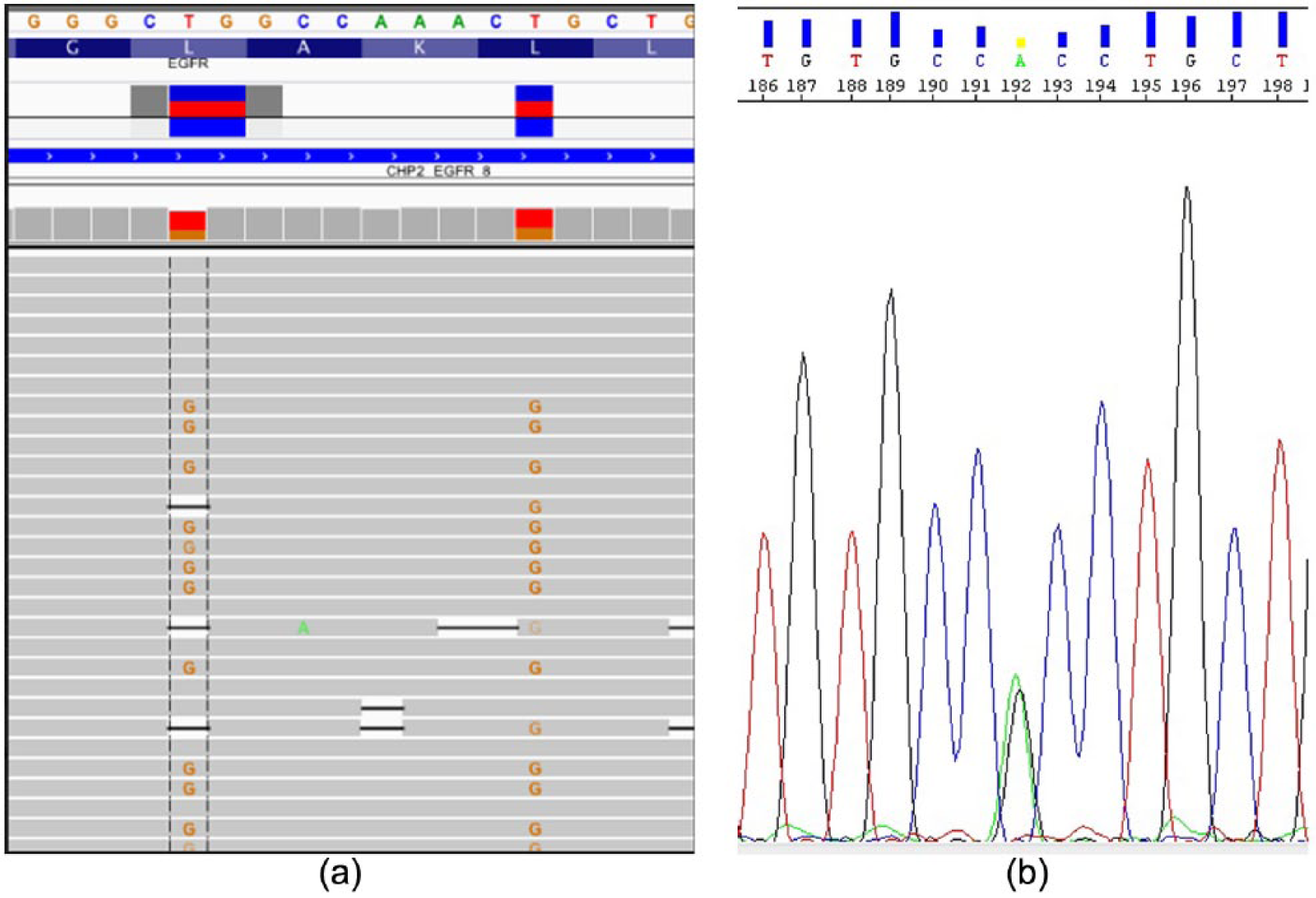
(a) Visualization of Leu858Arg and Leu861Arg mutations of EGFR gene (exon 21); the figure shows the coexistence of two mutations on the same allele. (b) Electropherogram of germline mutation Arg776His.
Conclusions
Ninety percent of the somatic mutations in the EGFR gene are clustered around the activation loop of the ATP binding pocket of the TK domain causing increased EGFR signaling. In particular, the Leu858Arg mutant EGFR, found in our patient, typically shows an increased catalytic activity due to the disruption of the auto-inhibitory mechanism, but it is also characterized by a 20-fold higher affinity to gefitinib compared to wild type EGFR 3 . Similar to patients with a Leu858Arg mutation, lung cancer patients with an L861R EGFR mutation demonstrated a favorable response to gefitinib 7 .
In contrast, patients with double EGFR mutations were previously shown to be characterized by a low objective response rate to EGFR TKI compared with those carrying a single EGFR mutation in either exon 19 or exon 21 8 . Wei et al. 8 described the co-occurrence of exon 19 and 21 mutations, but did not address whether the two mutations occurred on the same allele. Thus, it is possible that the low response rate of patients harboring these two mutations may be due to their presence on different alleles or by the presence of different tumor clones.
In contrast, in our case, the two mutations (Leu858Arg and Leu861Arg) coexist on the same EGFR allele (exon 21) and with a very similar allele frequency (40.3% and 41.3%, respectively), which suggests they also co-occur in the same cancer cells. Upon treatment with the TKI afatinib, in this case we obtained a partial response on the lung mass and nodules, a stable disease on bone lesions, and a progression-free survival of 8 months. Therefore, in our patient, the association of two different EGFR mutations seems to be associated with a favorable response to TKI therapy. While a single case report does not allow us to draw a major conclusion, it is tempting to speculate that the particular mutation pattern we found in this patient might confer increased response to TKI therapy, or in particular, to Afatinib, which needs to be further tested experimentally.
The reported low frequency of detection Arg776His somatic mutation among the NSCLC patients, its reported association with a second EGFR mutation 9 , and the previous finding of a germline Arg776Gly mutation co-occurring with an Leu858Arg and Gly719Ser, prompted us to consider the possibility that it occurred on the germline. Therefore, we searched whether the Arg776His also could be detected in the germline. To this purpose, we Sanger sequenced DNA extracted from the patient’s peripheral blood lymphocytes, which confirmed that the Arg776His mutation is constitutive in the patient. According to recent literature, this mutation could have contributed to the development of the lung cancer in our patient 10 . Although in vitro data seem to suggest that the presence of double mutations confer resistance to EGFR TKI, the association of the Arg776His with other EGFR mutations was not reported to confer resistance to the erlotinib or gefitinib treatment 10 ; rather, Kobayashi et al. 11 reported a case of partial response in a woman harboring Leu858Arg+Arg776His.
In the “biomarkers era,” the presence of detectable mutations in tumor tissue confer, at least in some instances, the possibility to predict a response to specific molecular targeted therapy. Due to the presence of a germline mutation, in this particular case it also allowed us to understand that this patient, and potentially her progeny, might be exposed to a genetic risk of cancer development. Therefore it should be possible to obtain screening guidelines to improve the risk calculation for lung cancer susceptibility in the future.
Footnotes
Author contributions
Francesca Belardinilli and Angela Gradilone contributed equally to this work.
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Associazione Italiana per la Ricerca sul Cancro (IG17734), Ministry of University and Research, PRIN projects and Istituto Pasteur-Fondazione Cenci Bolognetti to G. Giannini.
