Abstract
Introduction:
Hepatocellular carcinoma is a lethal disease worldwide and therefore the establishment of novel diagnostic biomarkers is imperative. In this study, it was hypothesized that an abnormal expression of the lysosomal-associated protein transmembrane 4 beta (LAPTM4B) gene is crucial in the pathogenesis of hepatitis C virus-mediated hepatocellular carcinoma; hence we investigated the expression profile of LAPTM4B in hepatitis C virus-induced hepatocellular carcinoma.
Methods:
A group of 189 consecutive patients (hepatitis C virus-related hepatocellular carcinoma as tumor cases; n=93, hepatitis C virus-related cirrhotics as disease controls; n=96) opting for living donor liver transplantation as a therapeutic surgical regimen were recruited with informed consent. Additionally, paired adjacent non-tumorous tissues (n=93) obtained from cases were also included. Serum LAPTM4B protein concentrations were assessed by third-generation enzyme-linked immunosorbent assay and LAPTM4B mRNA, and protein expressions at tissue level were determined by quantitative real time reverse transcription-polymerase chain reaction (qRT-PCR) and immunohistochemistry techniques, respectively.
Results:
LAPTM4B protein concentrations in sera of patients were higher (p<0.001) in tumor cases (1.25±0.25 ng/ml) compared to disease controls (0.53±0.28 ng/ml). Our study also depicts positive clinicopathological correlations between alpha-fetoprotein titers (b=0.65; p<0.001), quantitative hepatitis C virus RNA copies (b=0.33; p<0.001), and LAPTM4B protein concentrations, all in sera of patients. In addition, qRT-PCR and immunohistochemistry analyses revealed a significantly higher (p<0.05) tissue LAPTM4B mRNA and protein expression, respectively, in tumor cases rather than in non-tumorous tissues and disease controls.
Conclusions:
Together, our results illustrate the LAPTM4B gene as a diagnostic biomarker in patients with hepatocellular carcinoma having documented evidence of chronic hepatitis C virus infection.
Keywords
Introduction
Hepatocellular carcinoma (HCC) is a global health hazard. It is the sixth most common cancer (with ~850,000 new cases each year) worldwide, the second leading cause of cancer-related deaths (with ~800,000 deaths per year) accounting for 85–90% of all primary liver cancers.1–4 Among various risk factors being implicated in the cause of HCC, infection with chronic hepatitis C virus (HCV) has been depicted to be lethal, and the rate of development of HCC ranges between 1% and 3% after 30 years. The highest HCC incidence rates have been reported in recipients of HCV-contaminated blood or blood products (1/1000 person-years for HCC) and in hemophiliacs (0.7/1000 person-years), with the lowest rates being documented in women who received a one-time therapy with contaminated anti-D immune globulin. 5 Research has revealed that the risk of HCC increases 17-fold in HCV-infected patients compared to HCV-negative controls. 6 Until now, the lack of an effective and established HCV vaccine further inhibits its prevention; therefore a complete understanding of the role of various host factors associated in HCV-related HCC is imperative for developing newer treatment avenues for the disease.
In the above context, lysosomal-associated transmembrane protein 4 beta (LAPTM4B) is a recently discovered gene that has been mapped to chromosome 8q22.1; it contains seven exons and six introns, spans ~50 kb, and is an essential factor in maintaining cellular homeostasis.7–8 LAPTM4B-35, an isoform of LAPTM4B has been reported to be upregulated in several cancers including hepatocellular carcinoma, breast carcinoma, pancreatic carcinoma, ovarian carcinoma, colon carcinoma, extrahepatic cholangiocarcinoma, cervical carcinoma, endometrial carcinoma, gallbladder carcinoma, and gastric carcinoma. 9 Studies have illustrated that LAPTM4B promotes tumorigenesis by inhibiting apoptosis, upregulating autophagy, and rendering resistance to chemotherapy. 10 In addition, LAPTM4B has been considered to be a putative novel oncogene with critical roles in increased cell growth, proliferation, and cancer progression towards highly invasive and metastatic stages.11–13
Studies have depicted lower LAPTM4B mRNA levels in normal adult livers. However, much higher LAPTM4B concentrations have been reported in the majority of HCC tissues, which has significantly correlated with the pathological grades of the disease.7,14 Studies have demonstrated a significant upregulation of LAPTM4B mRNA expression in HCC tissues compared to adjacent liver tissues and normal liver tissues. 15 In addition to the expression profile of LAPTM4B in HCC, several lines of evidence have also shown that the LAPTM4B*2 allele is a potential risk factor that may influence an individual’s susceptibility to the development of primary liver cancer. It is associated with poor prognosis in patients opting for surgical resection in HCC.16–18
An in-depth profile of LAPTM4B expression in patients presenting with HCV-mediated HCC and opting for living donor liver transplantation (LDLT) is yet to be reported. Published data have revealed LAPTM4B-35 isoform overexpression to be a critical risk factor for tumor recurrence and poor prognosis in HCC. 19 However, a diagnostic role of LAPTM4B is yet to be documented in HCV-related HCC patients in the setting of LDLT. Therefore, in the present study, we hypothesized that an abnormal expression of LAPTM4B is crucial in the pathogenesis of HCV-related HCC and hence investigated the interrelationships between the expression of LAPTM4B and clinicopathological endpoints of the disease in patients presenting with HCV-associated HCC in an LDLT program. Conclusively, our study is the first report that suggests LAPTM4B as a diagnostic biomarker in patients with HCC having documented evidence of chronic HCV infection.
Methods
Patient recruitment
A group of 233 consecutive patients who presented with HCV-related HCC with underlying liver cirrhosis and HCV-related cirrhosis without any evidence of HCC, and who opted for LDLT between January 2013 and January 2016, were prospectively recruited from the Centre for Liver and Biliary Surgery, Indraprastha Apollo Hospital, New Delhi, with written informed consent. The study protocol conformed to the provisions of the 1975 Declaration of Helsinki (as revised in Seoul, Korea, October 2008) and was approved and acquitted by the Institutional Ethical Clearance Committee (IECC) of Jamia Millia Islamia.
Sample size estimation and power analysis
In this study, we hypothesized that an abnormal expression of LAPTM4B may play a crucial role in the pathogenesis of HCV-related HCC and hence calculated the sample size considering the statistical parameters of 80% power and 95% confidence level.
Inclusion criteria
In the context of our study, HCC patients with underlying cirrhosis and documented evidence of chronic HCV infection (positive anti-HCV antibody and quantitative HCV RNA titers for ≥6 months) were included in the study as “tumor cases” (n=93). The diagnostic criteria for tumor cases were followed as described elsewhere. 20 Cirrhotic patients with documented evidence of chronic HCV infection (positive anti-HCV antibody and quantitative HCV RNA titers for ≥6 months) without any evidence of HCC were included as “disease controls” (n=96). The diagnostic criteria for disease controls were followed according to standard clinical practices. The disease control group was age- and sex-matched with the tumor cases. In addition, adjacent non-tumorous tissues (NTs) taken at least ≥5 cm distant from the primary tumor from another liver segment that did not contain the primary tumor was defined as a separate “NT” group (n=93), which was derived from the tumor cases. The sampling of NTs from a liver segment other than the primary tumor ruled out the possibility of the presence of satellite nodules of the primary tumor, which maintained the integrity of NT samples. Furthermore, NT specimens collected were blinded to at least two additional surgical pathologists who performed histopathological analyses to determine the tissue integrity and the absence of malignant cells. Their results were 100% concordant.
Exclusion criteria
Patients excluded from the study were those with: (a) HBV, HBV/HCV co-infections, and HCV/HIV co-infections (n=13); (b) other forms of cirrhosis, and/or HCC without documented evidence of chronic HCV infection (n=23); and (c) a history of any immunosuppressive drug regimen, alcohol consumption, and psychiatric illnesses resulting in an inability to provide informed consent (n=8).
Patient records
Maintenance of records for each patient has been detailed in the Supplementary Material.
Biochemical and serological profile of the patients
All biochemical and serological data pertaining to each patient were collected from the hospital records. All biochemical and serological data are pre-operative.
Assessment of LAPTM4B serum protein concentrations
The activated human LAPTM4B protein concentrations in the sera of tumor cases and disease controls were quantitated using commercially available third-generation human lysosomal-associated transmembrane protein 4B, enzyme immunoassay (ELISA) kit (MyBioSource, Cat. No# MBS9332479, San Diego, CA, USA) according to the manufacturer’s instructions. Sera of patients were collected by venipuncture in sterile vacutainers (red-capped plain vials) prior to LDLT; they were bar-coded, aliquoted, and stored at −80°C before LAPTM4B serum protein analysis. Data recorded below or beyond the reference range of the kit were discarded.
Evaluation of LAPTM4B protein expression in tissues
The evaluation of the localization and expression of tissue LAPTM4B protein was achieved through immunohistochemistry (IHC) following standard pathological procedures. Briefly, positive control tissues used in the IHC experiments were known HCC tissues that stained positive for Glypican-3 (GPC3) protein. The negative control tissues were incubated with 1X PBS (pH 7.4) instead of the primary antibody. The simultaneous use of both negative and positive controls ensured the specificity and reliability of the staining procedure. Light microscopy was utilized to decipher the immunostained sections to facilitate the assessment of the prevalence of positive cases and the localization of immunostaining within tissues. Positivity was described as tumor cells with unequivocal staining of the granular cytoplasm. At least 20 microscopic fields were captured using different magnifications in each case, and subsequently the best fields were represented. At least two independent staining experiments confirmed all negative and positive cases. In addition, staining were reviewed and documented by at least two independent people; their results were 100% concordant. The primary and secondary antibodies used were anti-LAPTM4B (1:100 dilutions, ABCAM, Cat. No# ab82810, Cambridge, MA, USA) and DakoRealTM EnvisionTM, horse-radish peroxidase (HRP) rabbit/mouse conjugate (1:200 dilutions, Santa Cruz Biotechnology Inc., Dallas, Texas, USA), respectively. The chromogenic substrate used was DAB (3, 3′-diaminobenzidine tetrahydrochloride; Sigma Fast DAB, Sigma-Aldrich, St. Louis, MO, USA). The grading of LAPTM4B protein expression on the tissues was scored and classified according to an immunoreactive score (IRS) as previously reported 21 (Supplementary Table 1).
Analysis of LAPTM4B mRNA expression in HCV-mediated HCC
The expression of LAPTM4B mRNA was quantified using quantitative real-time reverse transcription-polymerase chain reaction (qRT-PCR) technology on liver tissue samples of disease controls as well as tumor tissues and adjacent NTs obtained from patients in accordance with an earlier study. 15 LAPTM4B qRT-PCR was conducted on a Rotorgene Q, 5-Plex HRM Thermal Cycler (Qiagen GmbH, Hilden, Germany). Catalogues of total RNA extraction, cycling conditions for LAPTM4B qRT-PCR, and the interpretation of LAPTM4B mRNA expression have been discussed in Supplementary Materials and Methods.
Reporting of diagnostic accuracy
A standard STARD (Standards for Reporting of Diagnostic Accuracy Studies) flow diagram (http://www.equator-network.org/reporting-guidelines/stard) representing the completeness and transparency of reports relevant to our study is shown in the Supplementary Figure.
Statistical analysis
All statistical computations were performed considering 80% power and 95% confidence interval (CI) using IBM SPSS, Ver20 (http://www-01.ibm.com/software/analytics/spss/) statistical software. For the purpose of this study, data were normalized using log10 transformation wherever required. For all computations, a two-tailed probability level of p<0.05 was considered statistically significant.
Results
Clinical, biochemical, and serological characteristics of the study population
The clinical, biochemical and serological attributes of the study population are illustrated in Table 1 and Table 2.
Clinical characteristics of the study population.
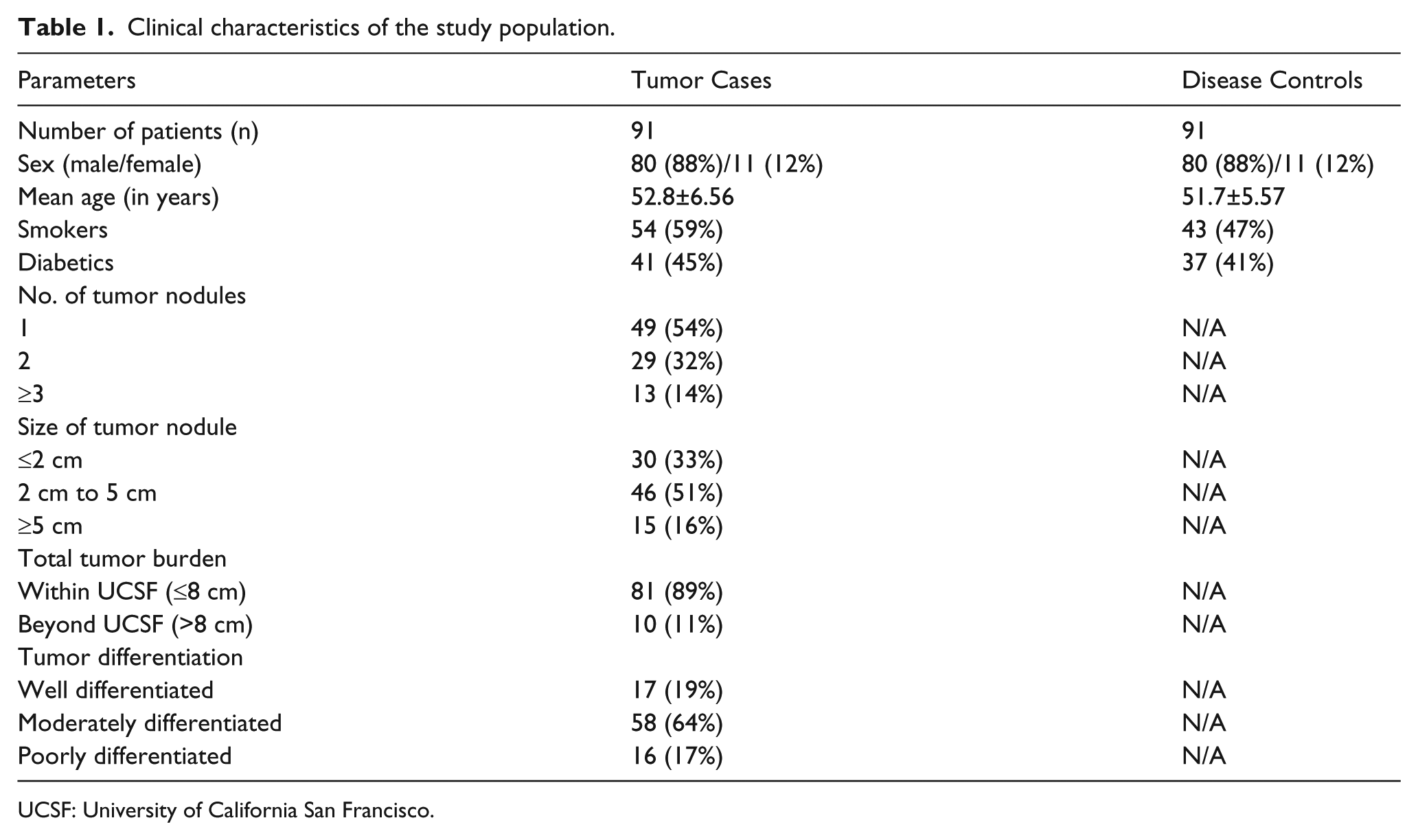
UCSF: University of California San Francisco.
Biochemical and serological attributes of the study population*.
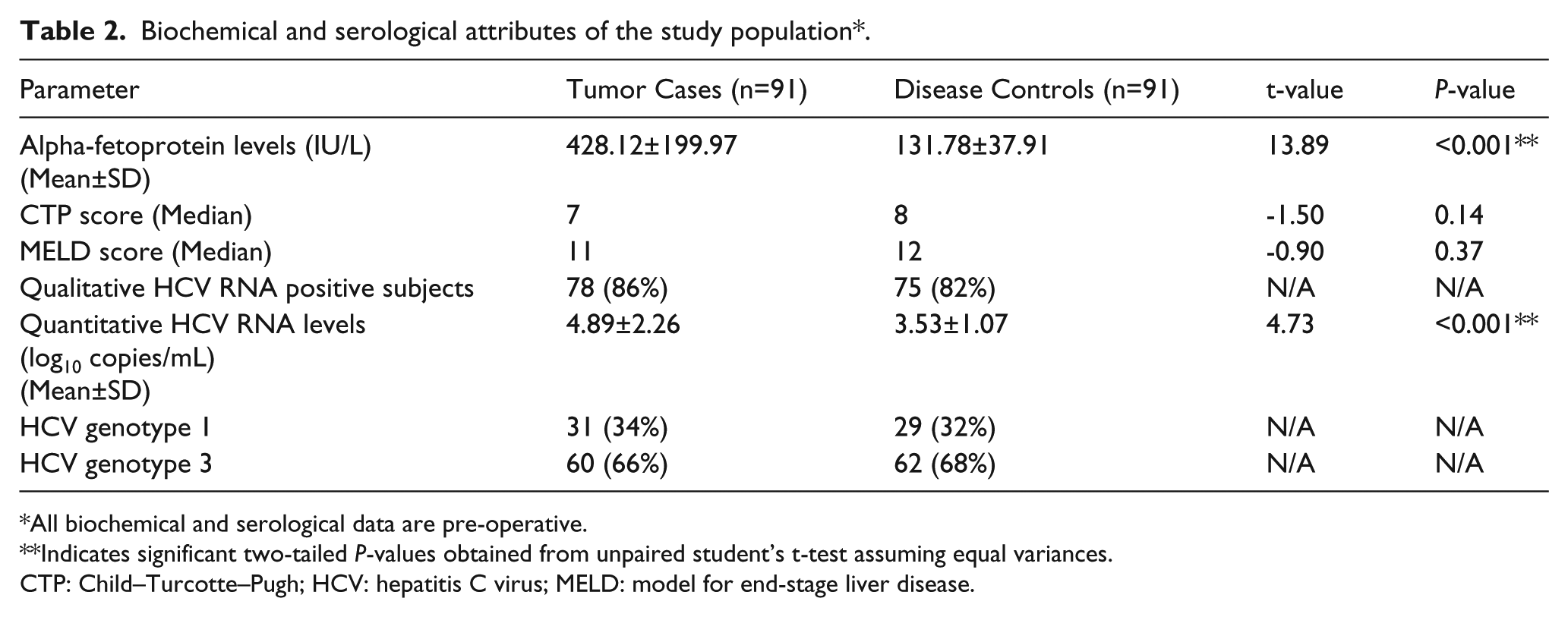
All biochemical and serological data are pre-operative.
Indicates significant two-tailed P-values obtained from unpaired student’s t-test assuming equal variances.
CTP: Child–Turcotte–Pugh; HCV: hepatitis C virus; MELD: model for end-stage liver disease.
Serum LAPTM4B protein expression
A significant difference (p<0.001) in serum LAPTM4B protein concentrations (in ng/mL) between tumor cases (n=91; 1.25±0.25) and disease controls (n=91; 0.53±0.28) was observed (Figure 1(a)).
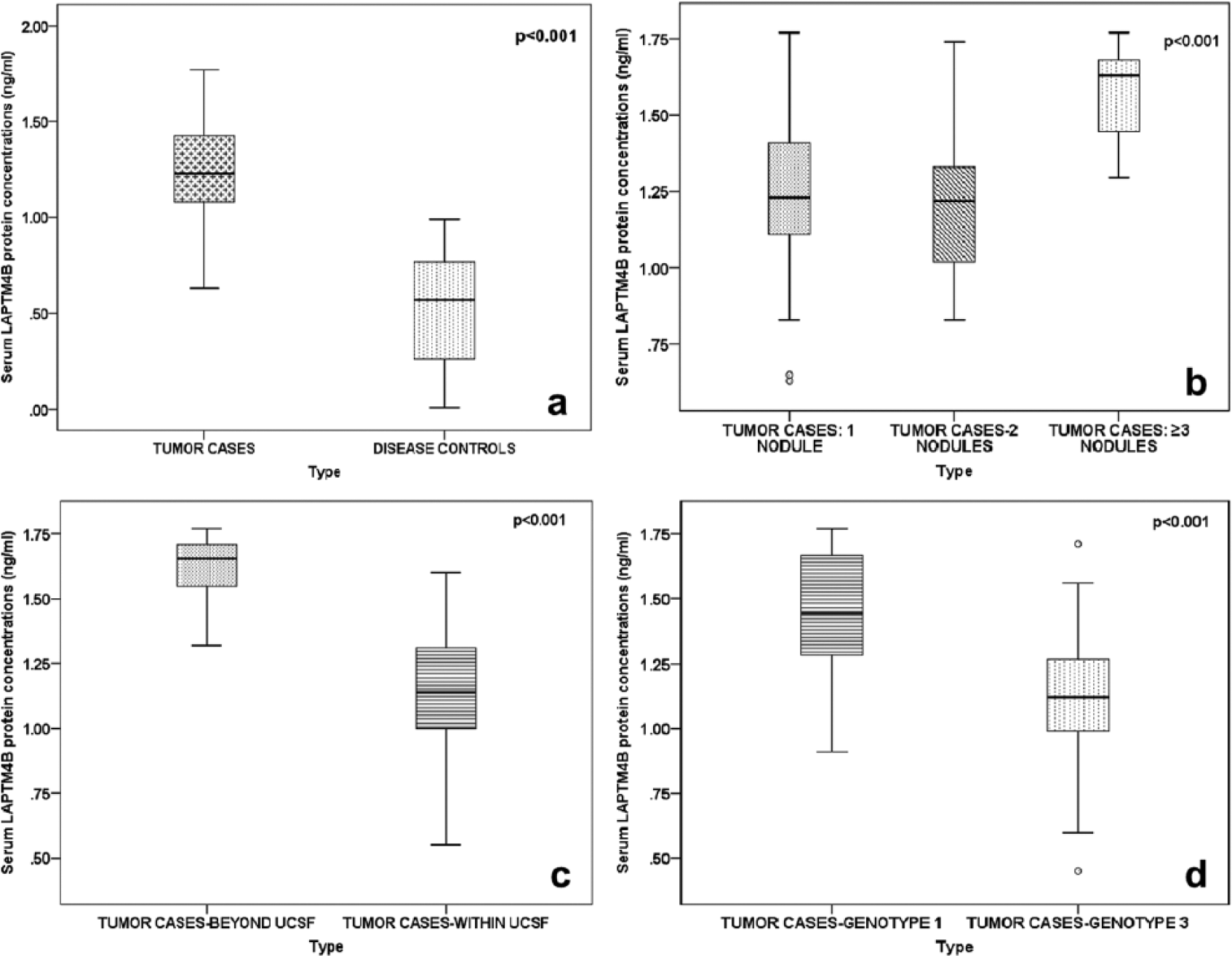
A difference in serum LAPTM4B protein expression patterns: (a) Between tumor cases and disease controls. (b) Between different numbers of tumor nodules in tumor cases. (c) Between tumor cases who opted for living donor liver transplantation within (≤8 cm) and beyond (>8 cm) the UCSF criteria. (d) Between different HCV genotypes in tumor cases.
Serum LAPTM4B protein expression among different numbers of tumor nodules in tumor cases
Patients presenting with ≥3 tumor nodules in HCV-related HCC, had the highest serum protein concentrations (1.58±0.16) of LAPTM4B (in ng/mL) and significantly differed (p<0.001) among patients with two tumor nodules (1.22±0.24) and subjects with a solitary nodule (1.25±0.26) (Figure 1(b); Supplementary Table 2).
Serum LAPTM4B protein expression between tumor cases within and beyond the UCSF criteria
Individuals in tumor cases who had a total tumor burden beyond the UCSF criteria (>8 cm) had significantly higher (p<0.001) concentrations (1.62±0.14 ng/ml) of serum LAPTM4B protein than those (1.12±0.24 ng/ml) within the criteria (≤8 cm) (Figure 1(c); Supplementary Table 2).
Serum LAPTM4B protein expression between different genotypes of HCV in tumor cases
HCV-mediated HCC patients with HCV genotype 1 had significantly higher (p<0.001) concentrations (1.46±0.25) of serum LAPTM4B protein (in ng/mL) than patients with HCV genotype 3 (1.11±0.24) (Figure 1(d); Supplementary Table 2).
Expression profile of LAPTM4B in tissues
Group-wise comparisons illustrated a significant difference in tissue LAPTM4B protein expression between tumor cases and disease controls (p<0.0001). Our results display no significant difference in tissue LAPTM4B protein expression between tumor tissues and adjacent NTs obtained from tumor cases (p=0.17). However, tissue LAPTM4B protein expression was significantly different between adjacent NTs and disease controls (p=0.02) (Figure 2(d) to (f); Supplementary Table 3).
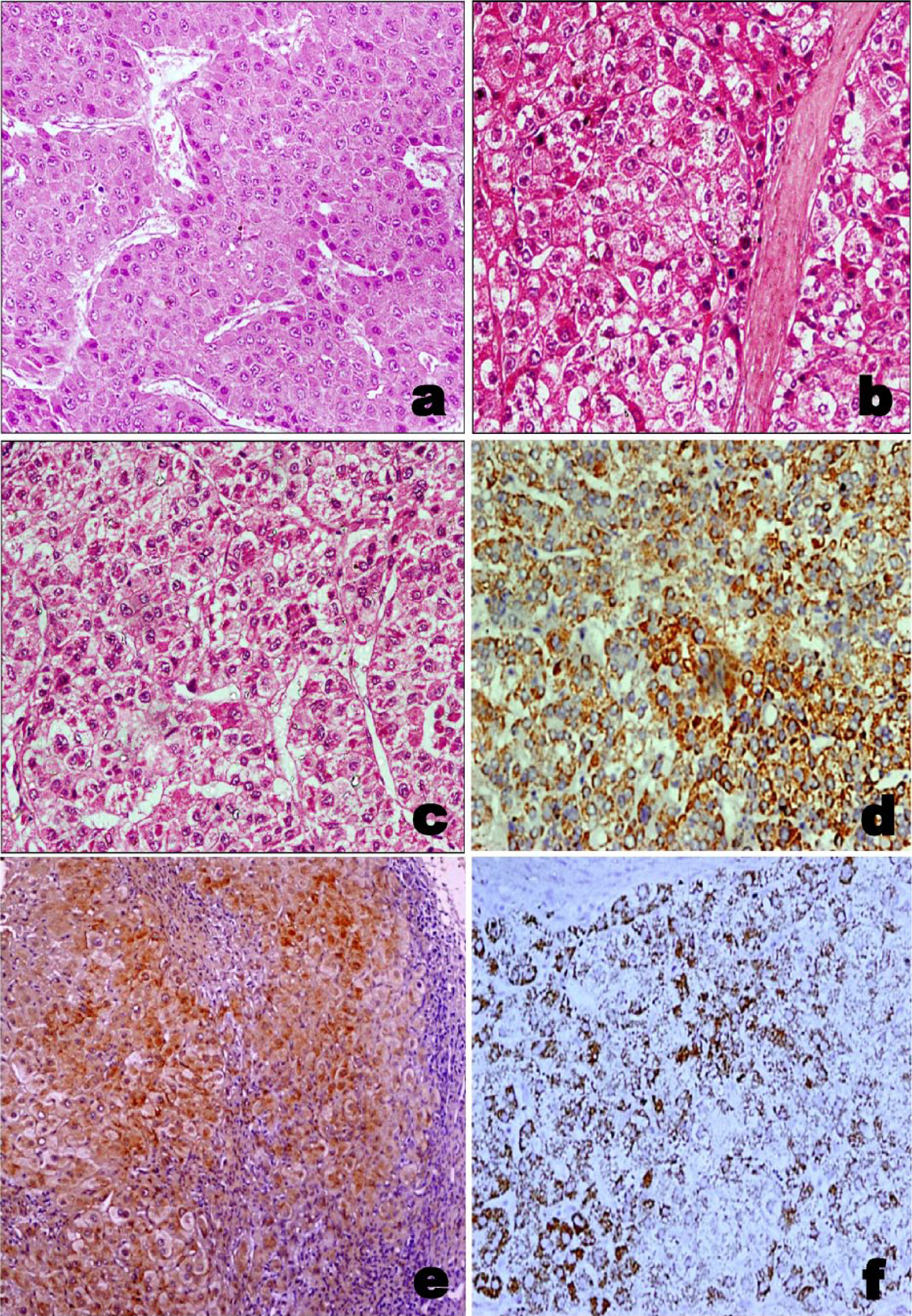
Representative photomicrographs of: (a) Hematoxylin and eosin staining patterns in HCV-related HCC tissues (magnification: 20X). (b) Hematoxylin and eosin staining patterns in paired adjacent NTs obtained from HCV-related HCC patients (magnification: 20X). (c) Hematoxylin and eosin staining patterns in HCV-related cirrhotic tissues without HCC (magnification: 20X). (d) Tissue LAPTM4B protein expression representing the localization of LAPTM4B protein in HCV-related HCC (magnification: 20X). (e) Tissue LAPTM4B protein expression representing the localization of LAPTM4B protein in paired adjacent NTs obtained from HCV-related HCC patients (magnification: 10X). (f) Tissue LAPTM4B protein expression representing the localization of LAPTM4B protein in HCV-related cirrhotic tissues without HCC (magnification: 20X).
Correlation between alpha-fetoprotein levels and serum LAPTM4B protein concentrations
A significant (p<0.001), direct and positive correlation (r=0.65) between alpha-fetoprotein (AFP) (in IU/mL) levels and serum LAPTM4B protein concentrations (in ng/mL) was observed in the study population (Figure 3).
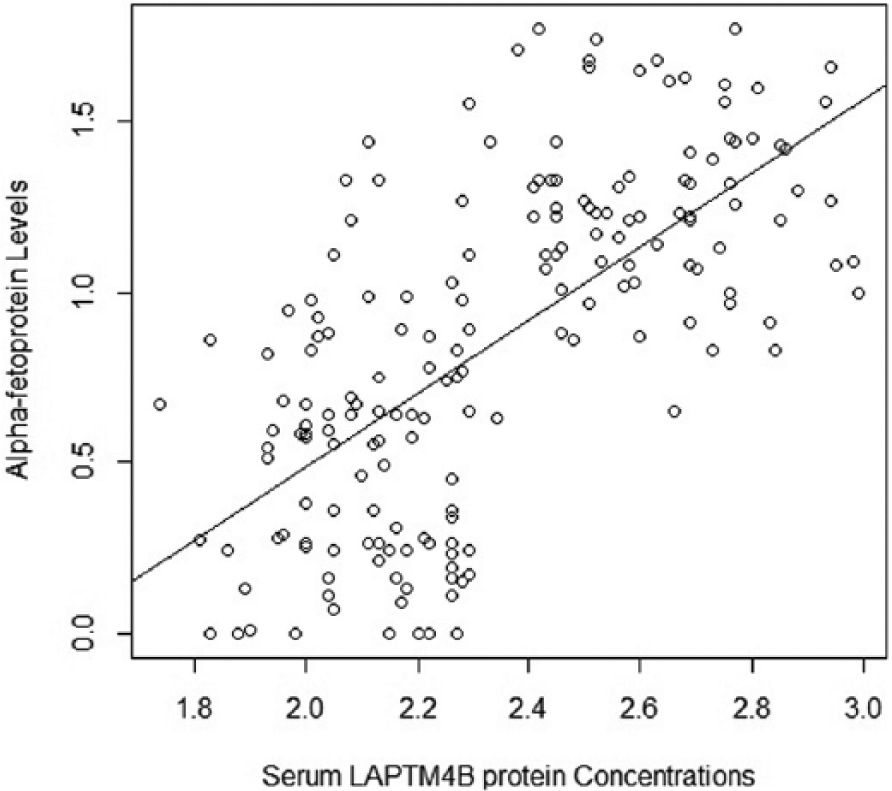
Scatter plot to show correlation between alpha-fetoprotein levels (in IU/mL) and serum LAPTM4B protein concentrations (in ng/mL) in the study population.
LAPTM4B mRNA expression in liver tissues
Analysis of LAPTM4B mRNA expression in liver tissues of patients revealed a significant increase (p<0.05) in expression in tumor tissues compared to disease controls and adjacent NTs (Figure 4).
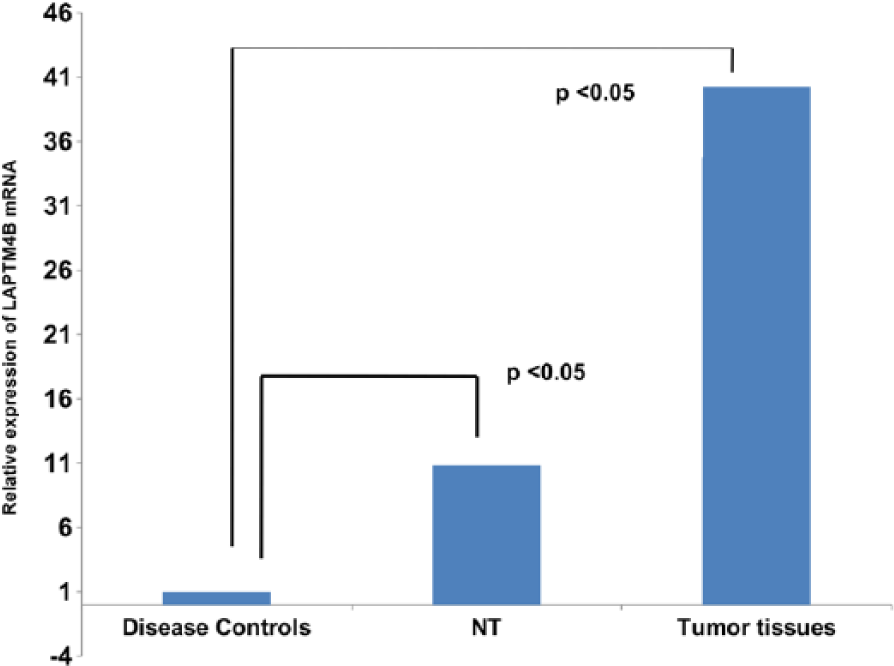
Quantitative reverse transcription-polymerase chain reaction (qRT-PCR) analysis of LAPTM4B mRNA levels in the liver tissues of disease controls, adjacent NTs, and tumor tissues.
Associative relationships among quantitative HCV RNA levels, LAPTM4B tissue protein expression levels, and the degree of carcinogenesis
To study the associative relationships among quantitative HCV RNA levels, LAPTM4B tissue protein expression, and the degree of carcinogenesis in the tissues, odds ratios (OR) were calculated. In this context, our results displayed that the presence of detectable HCV RNA copies in the serum to be poorly associated (OR<1) with LAPTM4B expression in the tissues of tumor cases, adjacent non-tumors, and disease controls (Supplementary Table 4). However, the expression of LAPTM4B was found to be strongly associated (OR >1) with the degree of carcinogenesis in the tissues (Supplementary Table 5). Needless to say, it was reasonable to consider adjacent tumor tissues as less carcinogenic than tumor tissues—the tissues of disease controls being the least carcinogenic.
Discussion
Our study illustrates an aberrant expression profile of the LAPTM4B gene in patients presenting with HCV-induced HCC and opting for LDLT. In this context, both serum and tissue LAPTM4B protein expressions exhibited several intriguing aspects that suggested the LAPTM4B gene as a diagnostic biomarker in HCV-mediated HCC.
The demographic profile of the study population illustrated a male preponderance (88%) over females (12%). The majority of the tumor cases (54%) presented with a solitary tumor and only a handful of them (11%) presenting with HCV-related HCC opted for LDLT beyond the UCSF criteria (>8 cm). Evaluations in post-transplant surgical pathology revealed the least percentage of poorly differentiated tumors (17%), which have the worst prognosis.
Serum AFP levels (in IU/L) significantly differed (p<0.001) between tumor cases (428.12±199.97) and disease controls (131.78±37.91), and HCV genotype 3 were more represented in the study population than HCV genotype 1. Our data are congruent with previous reports from India, which have illustrated HCV genotype 1 to be significantly associated with a higher viral load, greater liver disease severity, faster disease progression, and more non-responders to interferon compared to HCV genotypes 3 and 2.22-25
Several lines of evidence have suggested that LAPTM4B is a potential proto-oncogene, whose overexpression is crucial in the carcinogenesis and progression of HCC. 7 Reports indicate that potencies of recurrence, metastasis, and progression of HCC are associated with protein expression of the LAPTM4B-35 isoform. This is indicative of its application as a novel biomarker for recurrence prediction and metastasis potency of HCC, which may facilitate the improvement of diagnosis, prognosis, and treatment of HCC in future. 19 Elevated expression of LAPTM4B-35 has been assessed to be an independent prognostic factor for overall survival and disease-free survival in patients with HCC.26,27 Likewise, data also show an overexpression of LAPTM4B-35 in combination with a positive argininosuccinate synthetase expression to be a negative prognostic marker for HCC. 28 In contrast, studies have also opined that LAPTM4B-35 protein is a weak tumor-associated antigen candidate even though it is associated with cell proliferation. 29
In our study, serum LAPTM4B protein concentrations significantly differed between tumor cases and disease controls. In this study, we found that patients presenting with ≥3 tumor nodules in HCV-related HCC bore the highest serum LAPTM4B protein concentrations compared to patients with two tumor nodules or a solitary nodule. Interestingly, individuals in tumor cases with a single tumor nodule had slightly higher serum LAPTM4B protein concentrations than those with two tumor nodules. This might be attributable to the fact that the initial process of carcinogenesis may involve an increased production of LAPTM4B, and subsequently active forms of the protein might rapidly incorporate into extracellular fluid, thereby resulting in a quick increase in blood concentration.
HCV-related HCC patients with total tumor burden beyond the UCSF criteria (>8 cm) had significantly higher concentrations of serum LAPTM4B protein than those within the criteria (≤8 cm). Additionally, tumor cases with HCV genotype 1 had significantly higher serum LAPTM4B protein concentrations than tumor cases with HCV genotype 3, which justifies the report of a significant association of HCV genotype 1 with a greater liver disease severity. 22
In this study, levels of AFP tightly correlated with LAPTM4B protein concentrations as evaluated in the sera of patients. The interpretation was that an increase in AFP levels, an established serum biomarker of HCC, may result in a corresponding increase in serum LAPTM4B protein concentrations in patients.
Our data display a significant increase in tissue LAPTM4B mRNA expression in tumors compared to disease controls and adjacent NTs. This is consistent with a previous report correlating the overexpression of the LAPTM4B gene with the malignant phenotypes of HCC. 15 However, our results indicate a weak association of LAPTM4B tissue protein expression with the presence of detectable HCV RNA in sera. This may be explained thus: once the expression of LAPTM4B is triggered by HCV RNA, it maintains its elevated expression even when HCV RNA is undetectable in the serum (the molecular mechanism is yet to be explained). HCV has been reported to induce autophagy in order to facilitate its replication by counteracting the apoptotic response of infected cells. Therefore, in this context, the induction of LAPTM4B by HCV is a plausible justification as it is known to induce autophagy and suppress apoptosis. 30 Our observation is further supported by a report on the induction of autophagy promoting lysosomal-associated membrane protein 2A (LAMP2A) in an HCV cell cultural model. 31
Our study also demonstrates a strong association of LAPTM4B tissue protein expression with the degree of carcinogenesis in the tissues, which is again in agreement with previous reports that LAPTM4B is overexpressed in HCC and adjacent NTs compared to the tissues of a healthy liver. 7
LAPTM4B protein has been reported to be overexpressed in HCCs compared with paired adjacent non-tumorous liver or normal liver tissues. Significantly higher expression has been found in poorly differentiated tumors when compared with moderately or well-differentiated HCCs, which strongly suggests that LAPTM4B is an important regulator of cell growth, proliferation, and differentiation. 7 In our results, LAPTM4B protein expression in tissues illustrates an overexpression of the protein in HCV-related HCC patients compared to a weak expression in HCV-related cirrhotics. A possible explanation for this phenomenon is an upregulation of LAPTM4B transcription factors in HCC. Promoter analyses have described potential transcription factor binding sites within the 5′-flanking sequence of LAPTM4B, which includes c-Ets-1, LYF-1, E2F, c-JUN, and c-Myb. Studies on human HCCs have shown that the c-Ets-1 oncogene was upregulated and involved in the overexpression matrix metalloproteinase-7. 32 However, whether an increase of c-Ets-1 is directly responsible for an overexpression of LAPTM4B in HCC remains to be investigated.
In addition, an intermediate expression of adjacent NTs—as found in our study—and the lack of a significant difference in LAPTM4B protein expression in tissues between tumorous tissues and NTs obtained from the tumor cases may be interpreted as plausible pre-cancerous transformations. These might have started to take effect in the adjacent NTs, resulting in such aberrant expression of the LAPTM4B protein, which corroborates the observation in a setting of HCV un-induced HCC. 10 This indicates that the overexpression of LAPTM4B is associated with the initiation of malignancy in HCV-induced HCC. In this study, adjacent NTs in patients with HCV-related HCC was studied in order to obtain insights into the pre-cancerous stages, which may open up possible avenues of preventive intervention.
Taken together, to the best of our knowledge, this is the first report on the role of the LAPTM4B gene as a diagnostic biomarker in patients presenting with HCV-related HCC in the setting of LDLT. In this regard, several molecular biomarkers, such as LAMA4, CCT, and TGF-alpha, have been identified with the prognosis in HCC.10,33,34 However, it is unclear whether such markers are effective replacements for clinical application, or are in addition to the prognostic factors already in use. Therefore, it remains to be investigated whether the LAPTM4B gene is competent enough singly, or in combination with other biomarkers as a valuable tool in enhancing the effectiveness of the early diagnosis and prognosis of HCC.
Conclusions
Studies have suggested that LAPTM4B inactivation could be a novel therapeutic target for cancer. 35 Recent reports are also indicative of the fact that LAPTM4B can facilitate pro-survival functions of epidermal growth factor receptor (EGFR) in cancer cells and this may be regarded as a therapeutic target for EGFR-positive cancers or a combined target for anti-EGFR therapies. 36 In the same vein, a high homology (46%) of LAPTM4B to LAPTM4A suggests that LAPTM4B might play a critical role in multidrug resistance;37-38 therefore it is reasonable to speculate that targeted inhibition of LAPTM4B protein levels might be a new vista for targeted HCC therapy. Infection with HCV alone does not explain the development of primary liver cancer and therefore it is imperative that real-time investigations on other hepatocarcinogenic patient profiles (e.g., HBV-induced HCC, non-HBV, non-HCV-mediated HCC, cryptogenic HCC, HBV, and HCV co-infection in HCC, and most importantly in HBV associated with HCC without any evidence of underlying cirrhosis (as evidenced in <10% of HCC cases), 39 especially in LDLT) be undertaken. In this context, a larger cohort of patients involving multiple centers needs to be included with adept clinical algorithms that will eventually prevent patients’ drop-out. Different host–viral and host–host interactions, such as TGF-β1 and LAPTM4B, 40 require special consideration for better functional insights into the molecular mechanisms of disease pathogenesis in HCV-related HCC. Finally, it is no less important to translate laboratory findings into real improvements in clinical practice for the benefit of patients presenting with HCV-related HCC.
Key Points
HCV-related HCC is a global health problem and early diagnostic biomarkers are imperative.
This study reports the expression profile of the LAPTM4B gene.
A transcriptional and translational expression signature of LAPTM4B in liver tissues of patients is depicted.
Data also illustrate positive clinicopathological correlations with LAPTM4B protein concentrations.
Conclusively, the results elucidate the LAPTM4B gene as a diagnostic biomarker for HCV-related HCC.
Supplementary Material
Supplementary Material, Supplementary_Material – Expression signature of lysosomal-associated transmembrane protein 4B in hepatitis C virus-induced hepatocellular carcinoma
Supplementary Material, Supplementary_Material for Expression signature of lysosomal-associated transmembrane protein 4B in hepatitis C virus-induced hepatocellular carcinoma by Gaurav Roy, Papai Roy, Atanu Bhattacharjee, Mudassar Shahid, Mohammad Misbah, Subash Gupta and Mohammad Husain in The International Journal of Biological Markers
Footnotes
Acknowledgements
The authors would like to thank Mr. Soumya Chatterjee, proprietor and creative head, Skylark Studio, Paikpara, Kolkata, West Bengal, India, for his invaluable input and suggestions on the figures in this manuscript. In addition, this study was aptly supported by the Department of Biotechnology, Jamia Millia Islamia, New Delhi, India, in terms of infrastructure.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
