Abstract
Background:
Hodgkin lymphoma, a highly curable malignancy is currently treated with an adriamycin, bleomycin (BLM), vinblastine, and dacarbazine (ABVD) regimen. BLM-induced pulmonary toxicity (BPT) is one of the dose-limiting toxicities. Previous reports have revealed that genetic variants rs1050565, rs11077, and rs1800562 are involved in the development of BPT. These results cannot be extrapolated to the South Indian population because of their ethnic difference. This study aimed to determine the frequency of rs1050565, rs11077, and rs1800562 variants in South Indian healthy individuals and Hodgkin lymphoma cases. These frequencies were compared with 1000 genome populations’ data. We also assessed if these variants modified the risk to Hodgkin lymphoma.
Material and methods:
A total of 200 healthy individuals and 101 cases with Hodgkin lymphoma were recruited for this case-control study after ethical approval. Blood samples were collected from the study participants and DNA was extracted. Genotyping of rs1050565, rs11077, and rs1800562 variants was done using real-time polymerase chain reaction. A chi-square test was used to assess the differences in genotype frequency data between cases and controls.
Results:
The minor allele frequencies of rs1050565 and rs11077 were 4.3% and 39%, respectively, whereas all the individuals were wild-type for rs1800562 mutation. The frequencies significantly differed from 1000 genome data. The variants did not alter the risk for Hodgkin lymphoma.
Conclusions:
We determined the frequencies of rs1050565, rs11077, and rs1800562 variants in South Indian healthy individuals, and the frequencies differed significantly from 1000 genome populations. We also found that the studied polymorphisms are not associated with Hodgkin lymphoma risk in the South Indian population.
Keywords
Introduction
Hodgkin lymphoma (HL), a B-cell neoplasm that affects the lymphatic system is highly curable. Hodgkin and Reed-Sternberg (HRS) are the malignant cells involved in HL. 1 In 2015, 11,200 new cases were diagnosed with HL in India, and in 2016, an estimated 8500 new cases were diagnosed in the USA.2,3 Currently, the standard therapy for HL is a chemotherapy regimen that consists of adriamycin, bleomycin (BLM), vinblastine and dacarbazine (ABVD). This regimen shows a high cure rate of about 80% in the early stage of the disease, which reduces to 60%–70% in the advanced stage. In addition, this regimen causes long-term toxicities in the form of BLM-induced pulmonary toxicity (BPT), adriamycin-induced cardiomyopathy, and vinblastine-induced sensory neuropathy. 1 Hence, identifying prognostic factors that will optimize the treatment of each patient by increasing efficiency and lowering toxicity is important to improve the treatment outcome in HL. Many enzymes are involved in detoxification and transport of these chemotherapeutic drugs, and polymorphisms in these enzymes can lead to higher toxicity. 4 In this study, we focused on polymorphisms that are involved in BLM metabolism.
Bleomycin hydrolase (BLMH) is one such enzyme that inactivates BLM. BLMH is a neutral cysteine protease and is absent in skin and lungs, which makes them vulnerable to BLM toxicity. In contrast, increased activity of BLMH is associated with BLM resistance. The gene coding BLMH is thought to play an important role in regulating the enzymatic activity of BLMH. The gene consists of 12 exons and a single nucleotide polymorphism (SNP) rs1050565 in BLMH gene results in Ile443Val amino acid substitution. 5 So far, only two studies have been conducted to assess the role of rs1050565 variant in the development of BPT, and the results of both studies are contradictory. One study conducted in 2005 reported that SNP in BLMH was not associated with the occurrence of BPT, while the other study conducted in 2016 found that bleomycin-treated patients with the A/A genotype had favorable pulmonary function test results.5,6 In 2008, de Haas et al. 7 found that G/G genotype of BLMH was associated with reduced survival in germ cell tumor patients.
MicroRNAs (miRNA) inhibit translation initiation by messenger RNA (mRNA) destabilization; hence they can control many processes in the tumor cell. They can inhibit the translation of either oncogenes or tumor suppressor genes. Thus, miRNAs play a crucial role in tumor development. Navarro et al. 8 found that miRNAs are expressed in HRS cells and are involved in pathophysiology of HL. SNP in miRNA (MiR-SNP) leads to deregulation of miRNA expression that can alter treatment outcome, toxicity profile and risk to develop disease. SNP rs11077 in Exportin 5 (XPO5) miRNA was found to be associated with a higher incidence of BPT and a lower response rate. 1
The HFE gene is highly similar to HLA class 1 gene. Its two most common polymorphisms (rs1799945 and rs1800562) have been found to alter iron metabolism in patients with hereditary hemochromatosis (HH). 9 As bleomycin exerts its toxic effect by generation of reactive oxygen species, accumulation of iron can contribute to toxicity. 10 Another study also observed that two SNPs in HFE did not increase the risk for HL. 11
Inter-ethnic differences exist in the distribution of allele and genotype frequency; therefore the frequency data of other population cannot be extrapolated to the South Indian population. 12 So far, very few studies have reported the prevalence of BLMH rs1050565, MiR-SNP XPO5 rs11077 and HFE rs1800562 gene polymorphisms in India. Hence, we aimed to establish the normative genotype and allele frequencies of these three polymorphisms in the South Indian population. The frequencies were compared with the data of 1000 genome populations. The study polymorphisms have been reported to be associated with BPT but the role of these polymorphisms in HL susceptibility has not been studied. Therefore, before studying the role of concerned polymorphisms in the development of BPT, we also assessed whether these polymorphisms are associated with HL risk.
Materials and methods
Ethical approval was obtained from the JIPMER institutional ethics committee (ECR/342/Inst/PY/2013). Written informed consent was obtained from participants before their enrolment in the study. In the case of a child participant, consent was obtained from their legally accepted guardian, and for children above seven years old their own assent was obtained. In this case-control study, 101 patients identified with HL at the Medical Oncology Department, JIPMER, were enrolled. Controls (N=200) were unrelated healthy individuals, age- and gender-matched from the indigenous population of south India with no prior family history of HL.
Sample collection and processing
From all the study participants, 5 mL samples of venous blood were collected for DNA extraction in tubes containing 10% liquid EDTA. The tubes were centrifuged and the cells were separated and stored at −30°C until DNA extraction. Genomic DNA was extracted for genotyping by the standard phenol–chloroform method from peripheral leucocytes. DNA samples were quantified using bio photometer (Eppendorf BioPhotometer Plus) and were diluted to 50 ng/μL of optimal concentration. Diluted samples were stored at 4°C.
Genotyping
Genotyping of BLMH rs1050565 A/G, XPO5 rs11077 A/C, and HFE rs1800562 A/G variants was done using real-time polymerase chain reaction (PCR) (AB7300; Applied Biosystems). TaqMan SNP assay kits were used for genotyping as per the manufacturer’s instructions. To reconfirm, in 15% of the randomly selected samples, genotyping was carried out in triplicate.
Statistical analysis
GraphPad Instat version 3 (GraphPad Software Inc., San Diego, CA, USA) was used for statistical analysis. A chi-square test was used to verify if genotype frequency data in the control group was consistent with the Hardy–Weinberg equilibrium. To check for the differences in distribution of gender, age, and genotype frequency data between cases and controls, a chi-square test was done. A chi-square test was again used to compare the allele and genotype frequencies of the South Indian healthy population with 1000 genome project data. A two-sided P value <0.05 was considered statistically significant.
Results
A total of 200 healthy controls and 101 HL cases were analyzed in this case-control study. They were genotyped for three SNPs—the details of which are shown in Table 1. The mean age ±SD of patients and healthy controls were 27.8 ±18.5 and 24.5 ±5.6 years, respectively. Among the cases, 76.2% were under age 40 and 23.8% were above age 40 years. There were 114 (57%) males and 86 (43%) females in the control group, and 69 (68.3%) males and 32 (31.7%) females in the case group. Baseline characteristics, such as gender and age, were similar in both groups.
List of studied SNPs with rs IDs and chromosome locations.
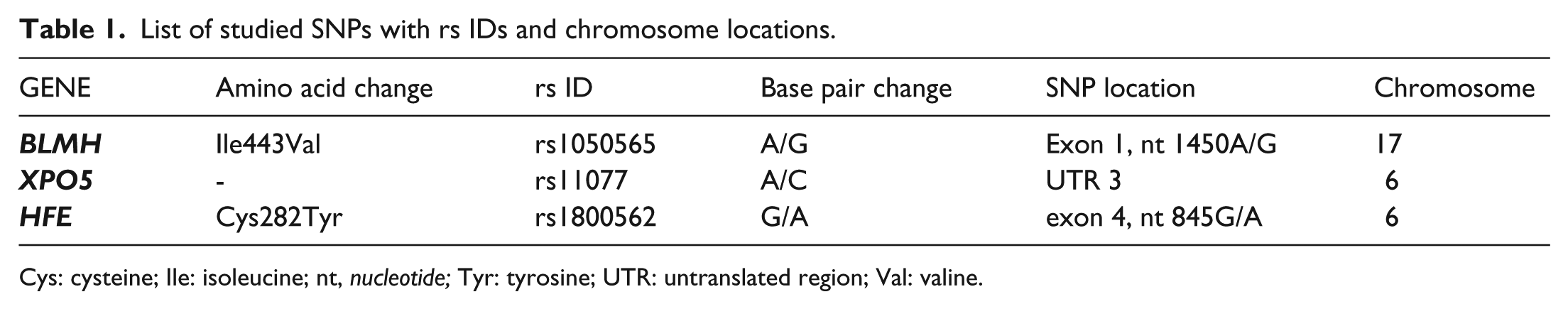

Cys: cysteine; Ile: isoleucine; nt, nucleotide; Tyr: tyrosine; UTR: untranslated region; Val: valine.
Association of SNPs with HL risk
The genotype frequencies of rs1050565 and rs11077 polymorphisms were found to be concordant with the Hardy–Weinberg equilibrium (P>0.05) whereas rs1800562 polymorphism was discordant. The genotypic and allelic frequencies distribution of rs1050565, rs11077, and rs1800562 polymorphisms in patients and controls is shown in Table 2. There was no significant statistical difference in the distribution of genotypic and allelic frequencies of the studied variants between the patients and controls.
Comparison of genotype and allele frequencies of South Indian population with 1000 genome populations
The distribution of allele and genotype frequencies of studied polymorphisms in HL cases and healthy individuals.
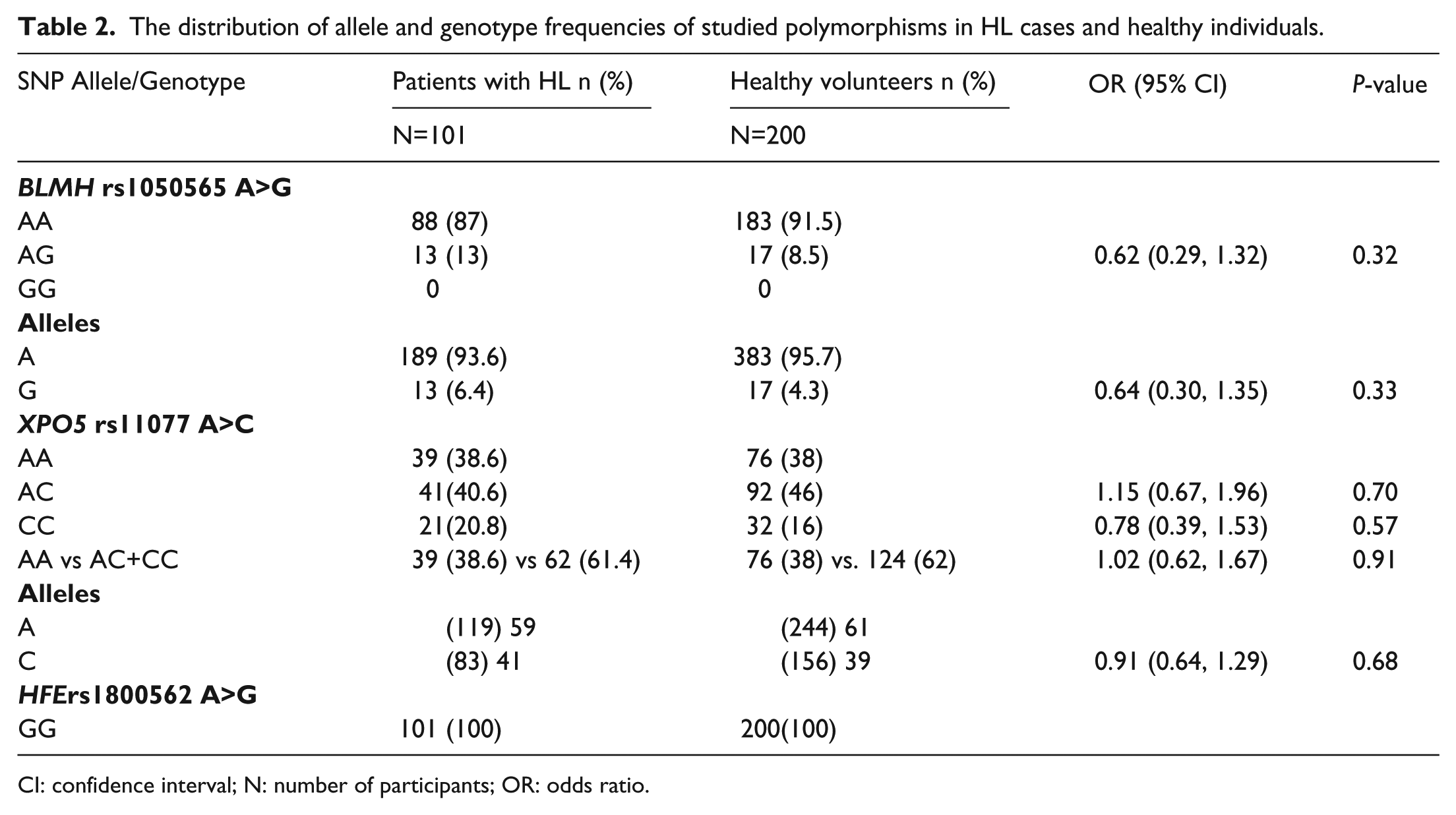
CI: confidence interval; N: number of participants; OR: odds ratio.
The allele frequency of rs1050565 polymorphism was significantly different from all 1000 genome populations. In the case of variant rs11077, the frequency distribution was significantly different from the African and east Asian populations. The allele frequency distribution of rs1800562 deviated significantly from American and European populations (Table 3).
Comparison of genotype and allele frequencies of the study population with 1000 genome populations’ data.
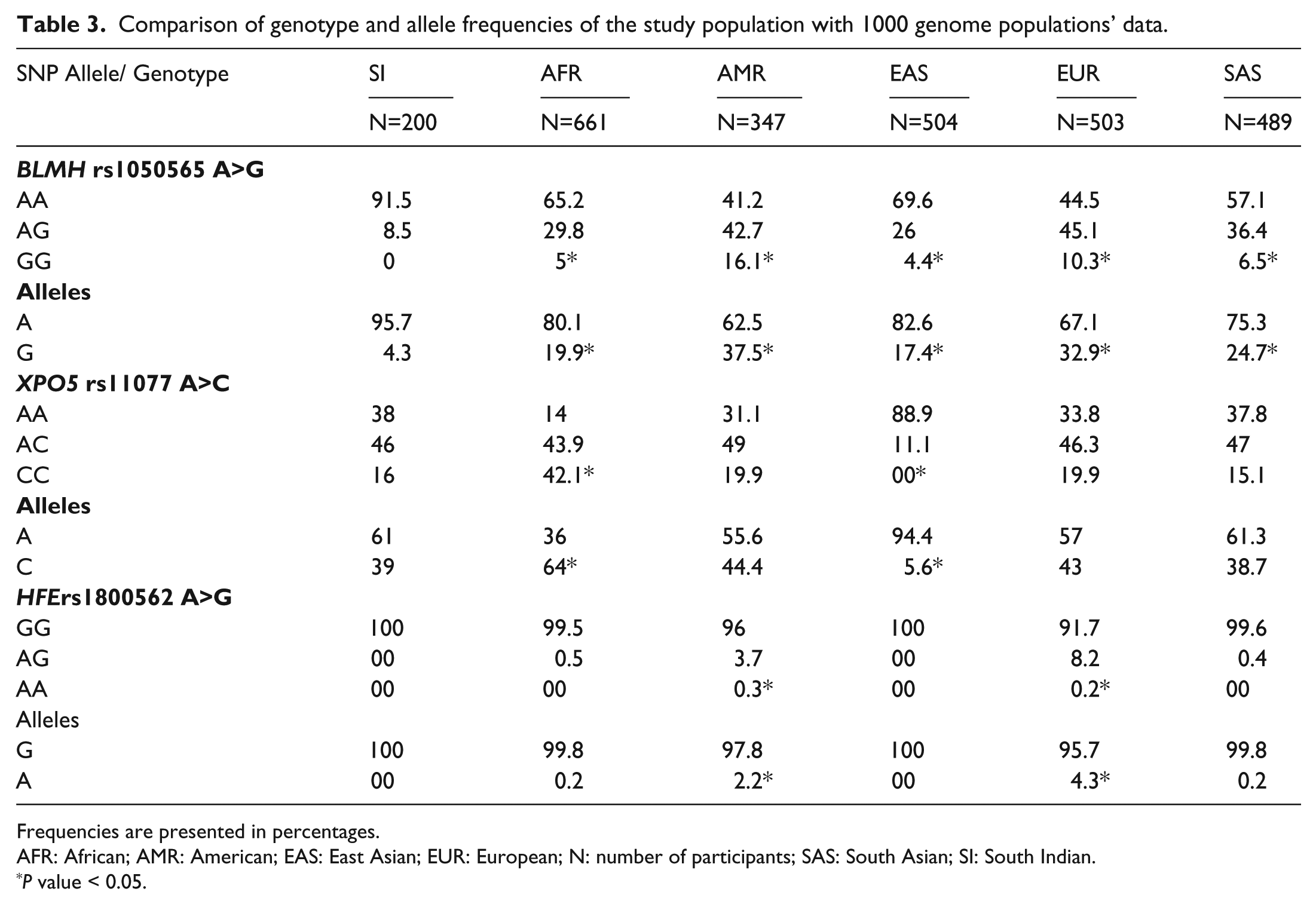
Frequencies are presented in percentages.
AFR: African; AMR: American; EAS: East Asian; EUR: European; N: number of participants; SAS: South Asian; SI: South Indian.
P value < 0.05.
Discussion
Although BLMH rs1050565, MiR-SNP XPO5 rs11077, and HFE rs1800562 genetic polymorphisms have been found to be associated with BPT, their involvement in the development of HL has not been studied. The present study established the allele and genotype frequency of these variants in South Indian healthy individuals and in HL patients.
The role of BLMH rs1050565 in the development of HL has been investigated for the first time in this study. BLMH is a cysteine protease and its exact physiological function is not known. In a meta-analysis, Llorca et al. 13 included six studies that evaluated the role of the BLMH variant in the development of Alzheimer disease (AD). Two studies reported the association of the BLMH variant with AD in the Caucasian population, while three other studies contradicted these results, and one study reported a lack of association in the Japanese population. The G/G genotype of BLMH was not a significant risk factor in the Tunisian and Brazilian populations.14,15
BLMH inactivates bleomycin, and the polymorphism in the BLMH gene alters aminopeptidase activity, which leads to increased BLM sensitivity and may increase the risk of BPT. In 2005, the Nuver et al. 6 study did not find any association between the BLMH gene polymorphism and the development of BPT. The study was conducted in the Netherlands and the frequency of A/A, A/G, and G/G genotypes were 45.9%, 43.5%, and 10.6%, respectively, in germ cell tumor patients. In another study conducted in the Netherlands population, de Haas et al. 7 reported that germ cell tumor patients with the G/G genotype had reduced survival compared to patients who carried heterozygous and wild type genotypes; their frequencies were 10.2%, 43.8%, and 46%, respectively. Jóna et al. 5 examined the role of this SNP in the development of long-term pulmonary adverse drug reactions after treatment with bleomycin. They reported that the A/G+G/G genotype of rs1050565 was associated with bleomycin-induced pulmonary toxicity, and their distribution of A/A, A/G, and G/G genotypes were 55.0%, 33.6%, and 11.5%, respectively.
In this study, conducted with southern Indians, the frequency of A/A was 91.5%; 8.5% had the heterozygous (A/G) genotype and none of the patients carried the homozygous mutant (G/G) genotype. The frequency distribution in southern Indians is significantly different from all the above-mentioned populations as well as from all 1000 genome populations including the South Asian population. This shows that genotype frequency differs from population to population. While the frequency distribution of this variant was similar in the cases and the control groups, it was not associated with HL risk. The G/G genotype, which is implicated in the development of BPT, is absent in the study population. Further studies are needed to investigate the role of this variant in the development of BPT in the South Indian population.
Exportin 5 (XPO5) helps in the transportation of pre-miRNAs into cytoplasm from the nucleus. After transportation into the cytoplasm, pre-miRNAs undergo a further maturation process and converts into functional miRNAs that regulate the transcription of mRNAs. Knockout of XPO5 was found to inhibit the transport of pre-miRNAs into cytoplasm, which led to decreased miRNA expression, whereas overexpression of XPO5 triggered increased miRNA expression. As XPO5 plays an essential role in the occurrence and progression of cancers, Mir-SNP XPO5 rs11077 was considered an important biomarker in the prediction of cancer prognosis and treatment-related toxicities. 16 Mir-SNP XPO5 rs11077 has been found to be associated with many cancers, such as non-small cell lung cancer, laryngeal cancer, and thyroid cancer in previous studies.16-18 This variant in our study did not seem to increase the risk for HL. In Navarro et al., study, incidence of BPT was higher in patients who carried A/A or C/C genotype compared to patients with A/C genotype in Spanish population. A/A or C/C genotype was also associated with lower response rate, disease-free and overall survival in HL patients. The frequency of A/A, A/C, and C/C genotypes in the Navarro et al. 1 study was 19.7%, 52.8%, and 27.6%, respectively, which was significantly different from our population (38%, 46%, and 16%, respectively). The frequency distribution was also different compared to the African and East Asian populations. Thus, further studies are required to evaluate the involvement of this polymorphism in the onset of BPT, the response rate, the disease-free and overall survival in southern Indian HL patients.
Two SNPs (rs1799945 and rs1800562) in the HFE gene have been found to cause HH in the European population. 9 van der Schoot et al. 10 carried out a study to investigate the role of these polymorphism in the development of BPT as the HFE gene is known to regulate iron metabolism, and the SNPs in these genes may contribute to toxicity. They reported that rs1799945 was associated with a higher incidence of BPT, but they could not analyze the role of rs1800562 as the frequency of the mutant genotype was very low. 10 Here, we have determined the frequency of rs1800562 only, and we found that all the individuals were wild type (G/G) for the mutation. This finding was consistent with other studies conducted in both the southern and northern Indian populations.19,20 An allele frequency was found to be 8.33%, 4.4%, 7.6%, and 8.5% in the Brazilian, German, French, and British populations, respectively. 9 When compared to the 1000 genome project data, the frequencies in the study population were significantly different from the American and European populations.
To conclude, we have determined the allele and genotype frequencies of BLMH rs1050565, MiR-SNP XPO5 rs11077, and HFE rs1800562 variants in the southern Indian healthy population and the HL patients. We found that the distribution of these polymorphic frequencies in the study population differed significantly from other world populations. The genotype and allele frequencies of the mentioned polymorphisms were similar in both the cases and the controls, and were not associated with HL risk.
Footnotes
Acknowledgements
We acknowledge JIPMER for providing intramural grant to carry out this study. We also acknowledge late Dr. Steven A. Dkhar for his guidance in conducting this study.
Authors’ note
The manuscript data was presented in European society for medical Oncology Asia-2017 (ESMO) congress conducted in Singapore on 17–19 November 2017 and the abstract was published in annals of oncology supplements (Vol. 28, Suppl. 10, November 2017)
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
