Abstract
Introduction:
5-flourouracil (5-FU) is one of the standard chemotherapeutic drugs used today in the treatment of colorectal cancer patients. Disruption of 5-FU metabolic pathway may contribute to altered effectiveness towards 5-FU-based therapy. Hence, the study of 5-FU metabolizing enzymes might have the potential efficacy to predict survival and response to treatment in colorectal cancer patients.
Materials and methods:
Immunohistochemical localization of 5-FU metabolic enzymes (TS, MTHFR, DPYD, and TP) was evaluated in 143 untreated patients with colorectal cancer; their prognostic and predictive values were also evaluated.
Results:
Immuno-positivity for TS, MTHFR, DPYD, and TP was observed in 77%, 75%, 88%, and 96% of colorectal cancer patients, respectively. Univariate survival analysis in total patients showed that low DPYD expression significantly predicted adverse overall survival (P=0.042). Moreover, subgroup of colon cancer patients with low TS expression was associated with unfavorable prognosis. TP expression also emerged as a prognosticator in the subgroup of early and advanced stage patients. Additionally, when effect of co-expression of 5-FU metabolic enzymes was evaluated in total patients, low coexpression of all four proteins was predictive of poor overall survival than for individuals expressing high coexpression of these proteins (P=0.045). In contrast, none of the 5-FU metabolic enzymes—either singly or on coexpression—emerged as a useful biomarker of potential therapeutic value when evaluated in the subgroup of patients treated with 5-FU alone or 5-FU plus oxaliplatin.
Conclusion:
The above findings suggest that coexpression of 5-FU metabolic enzymes possess significant prognostic value and could be useful biomarkers in colorectal cancer patients.
Introduction
5-Fluorouracil (5-FU) is a commonly used chemotherapeutic agent for the treatment of various tumors and is a key drug in colorectal cancer (CRC) treatment. Despite significant progress in understanding the 5-FU mechanisms, providing of individual 5-FU-based treatment with low toxicity and significant benefit is still an unresolved problem.
Research has indicated that the major mechanism of 5-FU action is primarily mediated through inhibition of thymidylate synthase (TS), for which the pathways have not been fully interpreted (1-3). Briefly, when 5-FU enters the cell, it is catalyzed by thymidine phosphorylase (TP) to fluorodeoxyuridine (FUDR), which is then phosphorylated to its active metabolite fluorodeoxyuridine monophosphate (FdUMP) by thymidine kinase (TK). FdUMP then binds to the nucleotide-binding site of TS, and forms an extremely stable ternary complex among FdUMP, TS, and the cofactor 5,10-MTHF (CH2THF). The level of 5,10-MTHF is regulated by methylene tetrahydrofolate reductase (MTHFR), which catalyzes the irreversible conversion of 5,10-MTHF to 5-methyletetrahydrofolate (5-MTHF). Normally, TS catalyzes reductive methylation of deoxyuridin-5′-monophosphate (dUMP) into deoxythymidine-5′-monophosphate (dTMP). However, the formation of ternary complex (TS, FdUMP and 5,10-MTHF) prevents the binding of the normal substrate dUMP to TS, and thus inhibiting the methylation of dUMP into dTMP catalyzed by TS (4, 5). Thus, the inhibition of TS activity rapidly shuts off DNA synthesis and triggers apoptosis and other cell death processes (6, 7). Moreover, it results in deoxynucleotide (dNTP) pool imbalances and increased levels of deoxyuridine triphosphate (dUTP), leading to DNA damage (4). On the other side, dihydropyrimidine dehydrogenase (DPYD)—a key enzyme involved in the catabolism of uracil and thymine into β-alanine—mediates the inactivation of 5-FU in the liver (5). Thus, TS, MTHFR, DPYD, and TP are the important enzymes of 5-FU metabolic pathway.
Studies so far reveal that all patients with equivalent pathological tumor stage do not respond similarly, and ultimately develop drug resistance. Hence, the most challenging task is to understand the mechanisms underlying chemo-resistance and to identify patient subpopulations that may benefit from therapy. Therefore, identifying molecular markers that can predict clinical outcomes is important to optimize individual therapy for CRC patients (8). Several molecular markers involved in the action mechanisms of 5-FU have been identified during the last decade, but controversial results in different studies complicate their application in clinical use. Hence, it is necessary to elucidate the complex genetic network of 5-FU pathway and the role of 5-FU metabolic enzymes to better understand the antitumor effects of 5-FU. It is evident that these metabolic enzymes, when singly considered, are unable to accurately predict the prognosis in CRC patients. Therefore, the present study aimed to evaluate the efficacy of protein expression of 5-FU metabolic enzymes (TS, MTHFR, DPYD, and TP) singly and in combination to predict the prognosis and treatment response in primary CRC patients.
Materials and methods
Patients
Tumor tissues of 143 untreated patients registered with histologically confirmed CRC were investigated in this study. All patients had undergone surgery between 2007 and 2014 at the Department of Surgical Oncology, The Gujarat Cancer and Research Institute, Ahmedabad, India. Written consent of all the patients was obtained prior to primary tumor tissue collection. Pathologic staging was performed according to the tumor node metastasis (TNM) classification with the World Health Organization (WHO) Grading System. The study was approved by the institution’s review and ethical committees. To study the prognostic value of the studied parameters, out of 143 patients, 114 that could be followed for a minimum period of 36 months or death within that period were included for overall survival (OS) analysis. Out of 114 patients, 13 died due to persistent disease; hence 101 patients were evaluated for relapse-free survival (RFS) analysis. Survival analysis was also performed in the subgroups of patients with early stage and advanced stage disease according to tumor stage, and in colon cancer and rectal cancer patients according to the tumor site. Further, in relation to adjuvant treatment, 113/143 patients were treated with chemotherapeutic regimens. The main chemotherapeutic treatment included were 5-FU/leucovorin, oral capecitabine, or in combination with oxaliplatin. Patients were subgrouped into those treated with 5-FU alone and those treated with 5-FU plus oxaliplatin (5-FU+OX)-based therapy irrespective of radiotherapy (RT). According to adjuvant treatment, out of 101 patients, 83 were included for RFS analysis; and out of 114 patients, 94 were included for OS analysis. Table I depicts the patient and tumor characteristics.
Patient and tumor characteristics
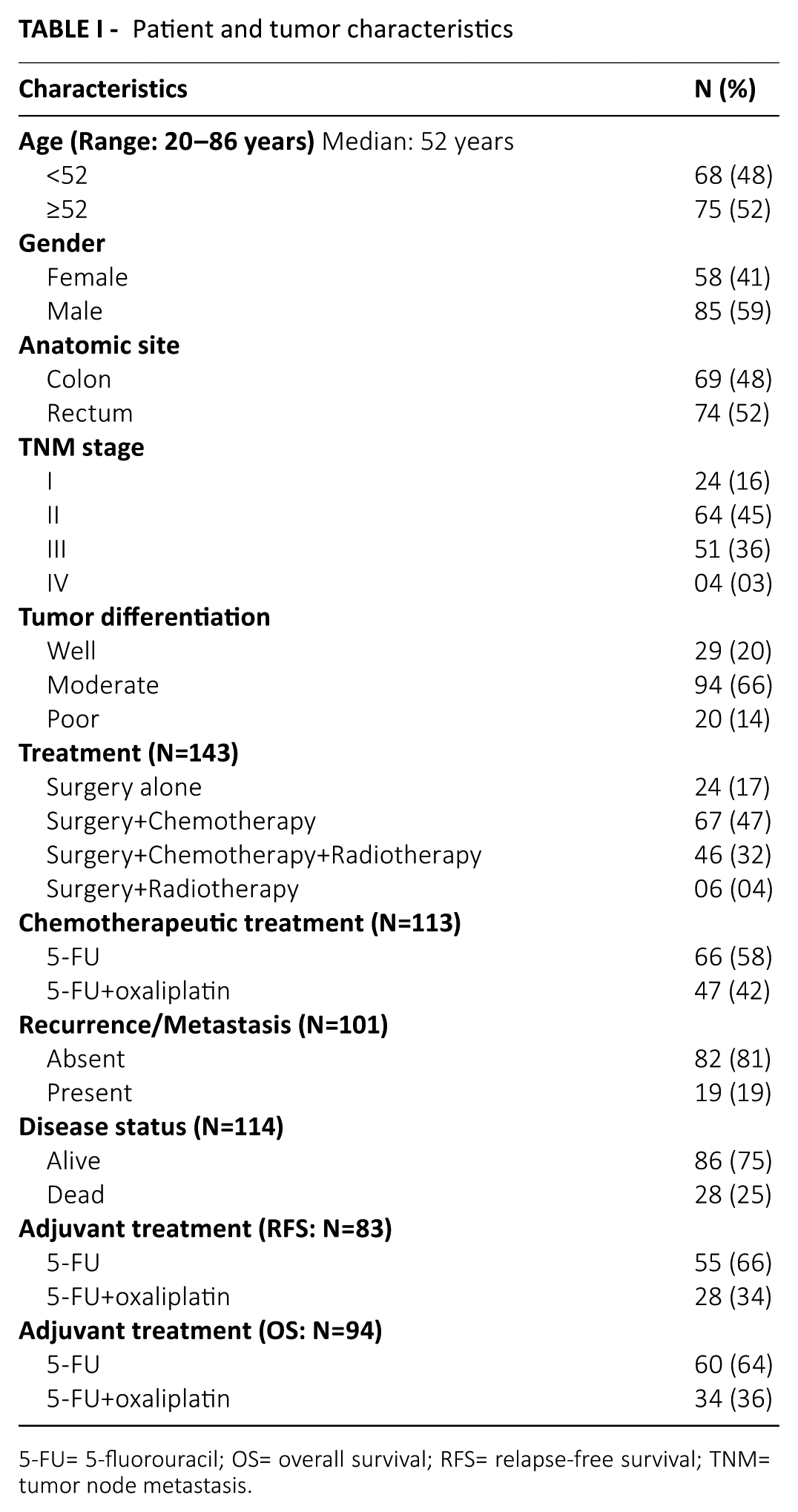
5-FU= 5-fluorouracil; OS= overall survival; RFS= relapse-free survival; TNM= tumor node metastasis.
Sample collection and study of protein expression of 5-FU metabolic enzymes by immunohistochemistry
For immunohistochemistry, paraffin embedded tumor tissue blocks were collected from the pathology department of the institute and were cut into 4 μm thick sections. Commercially available primary antibodies were procured for immunohistochemical staining of TS (Mouse monoclonal TS 106: sc-33679, Biotechnology Inc. Santa Cruz, CA, USA), MTHFR (Mouse monoclonal MTHFR 5D3: GTX60435, GeneTex, Inc., Irvine, CA, USA) DPYD (Rabbit monoclonal DPYD EPR8811: ab134922, Abcam, Cambridge, UK), and TP (Mouse monoclonal TP P-GF.44C: GTX23151, GeneTex, Inc.). Prior to the application of the primary antibody, antigen retrieval was performed for 20 minutes in a pressure cooker. Immunohistochemical localization of 5-FU metabolic enzymes was carried out using Mouse- and Rabbit-specific HRP/DAB (ABC) Detection IHC kit (Abcam). All sections were scored in a blinded fashion by two independent observers familiar with immunohistopathology, unaware of the clinical outcome of the patient. Scoring was done by the modified histoscore (H-score) method (9). Briefly, the staining intensities and the percentage of positive cells were separately assessed for all primary tumor tissues. More specifically, the staining intensity was assessed with a four-point scale from negative: 0; weak: 1; moderate: 2; and strong intensity: 3. The extent of the staining was expressed as a percentage of positive cells (0%–100%) by 10% intervals. The histoscores for TS, MTHFR, DPYD, and TP were counted by multiplying the intensity level by a percentage of positive cells resulting in a value between 0 and 300.
Statistical analysis
The data was statistically analyzed using the Statistical Package for Social Sciences (SPSS) software version 17 (SPSS Inc., Chicago, IL, USA). Two tailed Chi-square (χ2) test was used to assess the correlation of protein expressions with tumor differentiation. RFS and OS were calculated using Kaplan–Meier estimates and the difference in survival curve was calculated using the Log rank test. Correlation between two parameters was calculated using Spearman’s correlation coefficient (r) method. P value ≤0.05 was considered to be significant.
Results
Incidence of TS, MTHFR, DPYD, and TP protein expression
Cytoplasmic immunostaining positivity for TS, MTHFR, and DPYD was observed in 77%, 75%, and 88% of CRC patients, respectively. However, TP showed cytoplasmic and/or nuclear expression in tumor cells and also stromal expression. Total TP protein expression was observed in 96% of patients (Fig. 1). For statistical evaluation, the median H-score value of each parameter was used as a cut-off for grouping the patients in a low expression group (< median H-score) and a high expression group (≥ median H-score).
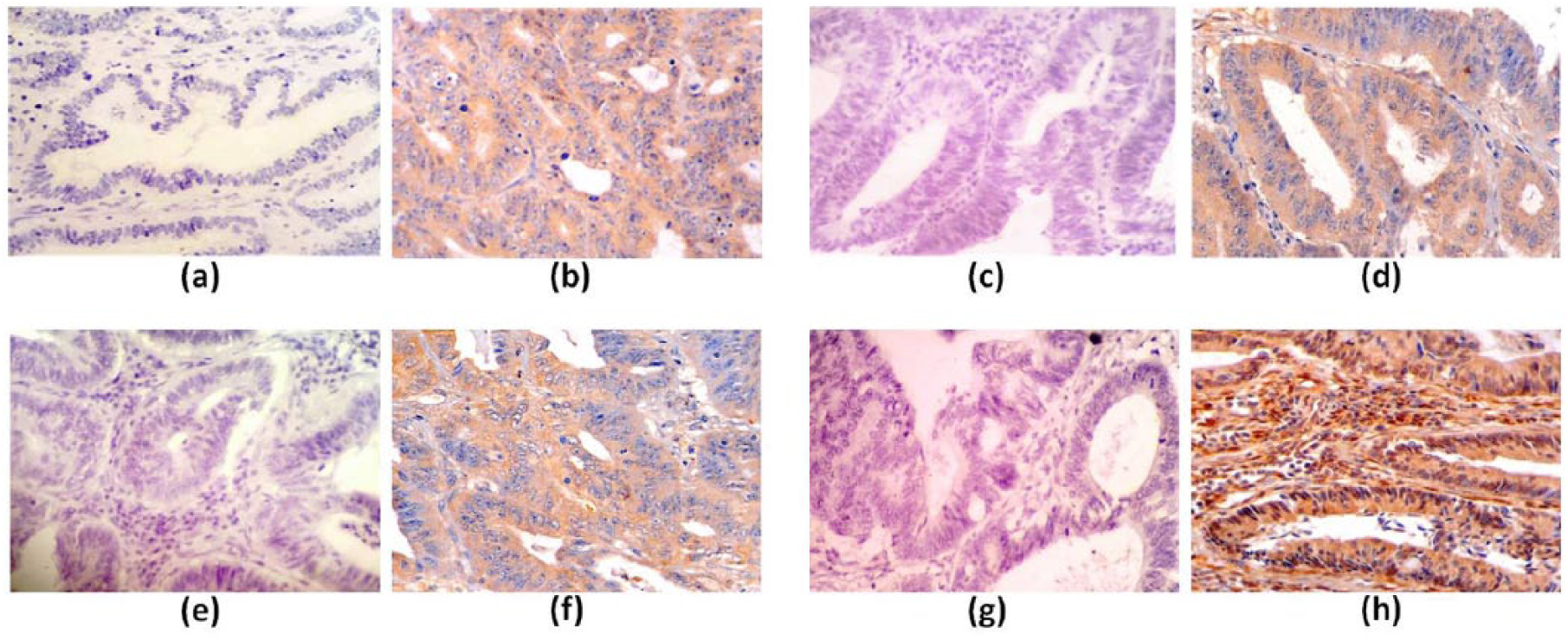
Immunohistochemical expression in primary colon and rectal tumor tissues. Negative expression of a) TS, c) MTHFR, e) DPYD, and g) TP. Cytoplasmic expression of b) TS, d) MTHFR, and f) DPYD. h) Cytoplasmic/nuclear/stromal expression of TP. Magnification: 40X.
Correlation of TS, MTHFR, DPYD, and TP (individual expression and coexpression) with tumor differentiation
When TS, MTHFR, DPYD and TP protein expression were correlated independently, only MTHFR protein expression showed significant correlation with tumor differentiation. The incidence of MTHFR expression was found to be higher in patients having moderately differentiated tumors (63%) as compared to those having well differentiated tumors (21%, P<0.001) and those having poorly differentiated tumors (35%, P=0.022). Moreover, in patients having moderately differentiated tumors, a trend of higher TS (56%) and DPYD expression (62%) was observed as compared to poorly differentiated tumors (TS: 35%, P=0.083 and DPYD: 40%, P=0.075) (Table II). However, TP protein expression showed no significant correlation with tumor differentiation (χ2=0.868, r=–0.049, P=0.564). Additionally, when coexpression of all four proteins correlated with tumor differentiation, no significant association was observed (Table III).
Correlation of TS, MTHFR, and DPYD with tumor differentiation
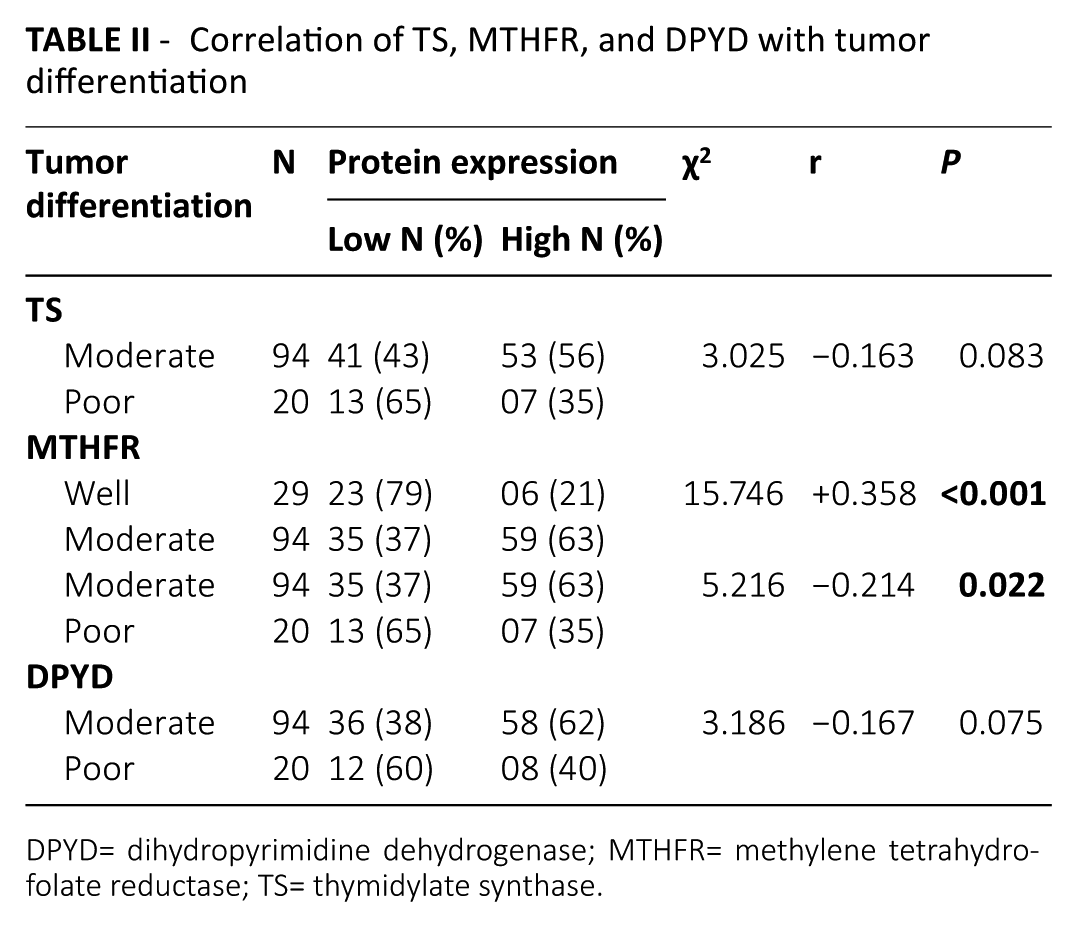
DPYD= dihydropyrimidine dehydrogenase; MTHFR= methylene tetrahydrofolate reductase; TS= thymidylate synthase.
Correlation between coexpression of TS, MTHFR, DPYD, and TP, and tumor differentiation
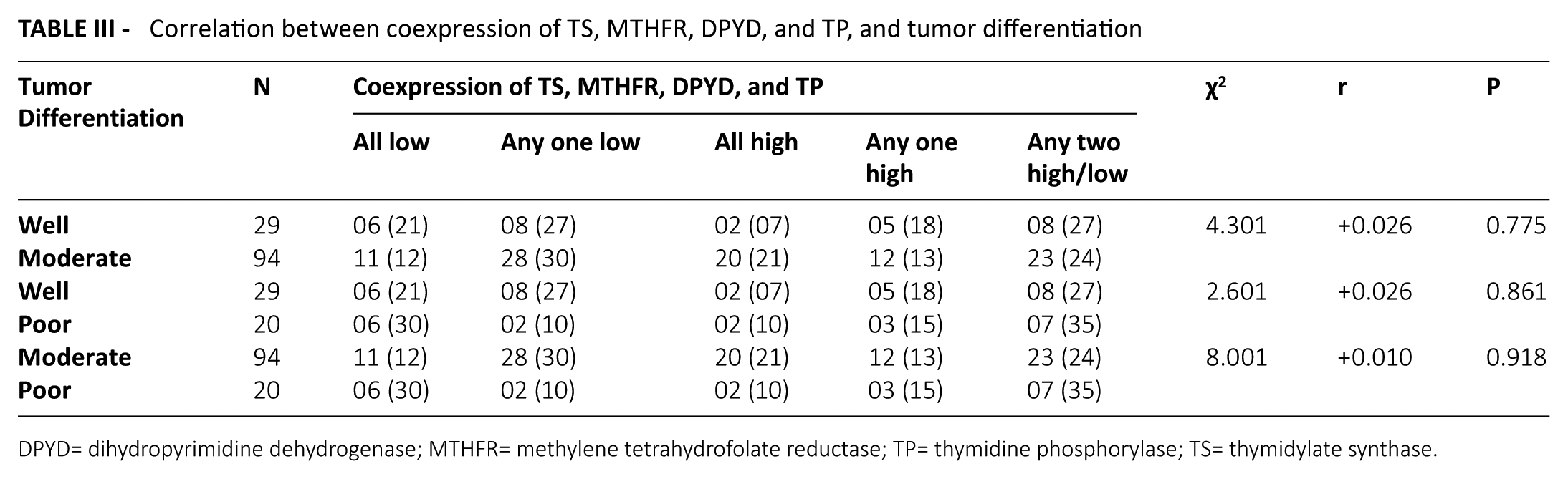
DPYD= dihydropyrimidine dehydrogenase; MTHFR= methylene tetrahydrofolate reductase; TP= thymidine phosphorylase; TS= thymidylate synthase.
Correlation of individual TS, MTHFR, DPYD, and TP protein expression with prognosis
In total patients, Kaplan–Meier univariate survival analysis demonstrated no significant correlation of TS, MTHFR, and TP protein expression with RFS or OS. However, a significantly reduced OS was observed in patients with low DPYD expression compared to those with high DPYD expression (P=0.042; Fig. 2a).
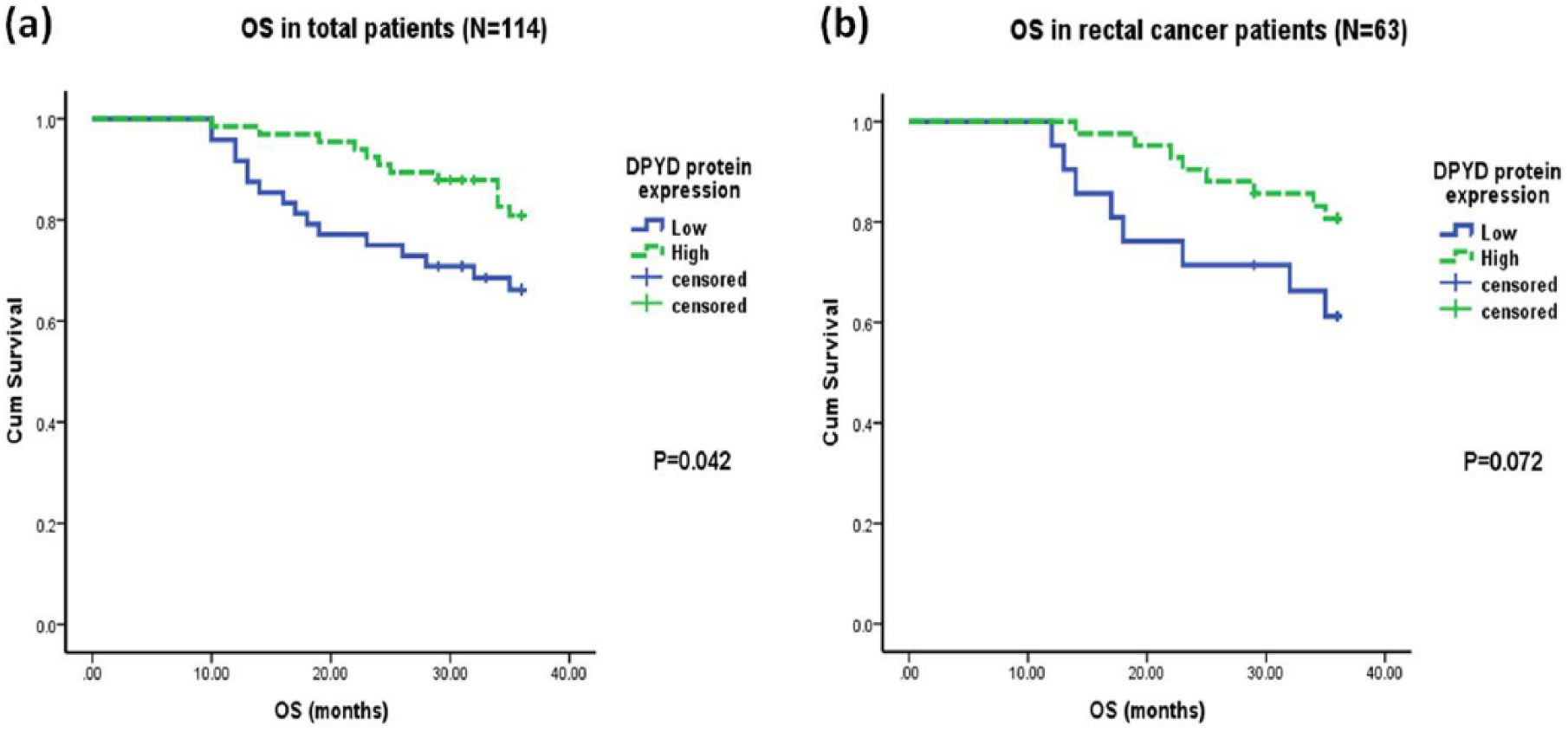
Kaplan–Meier overall survival curves with respect to high and low DPYD protein expression: a) total CRC patients; b) rectal cancer patients.
According to tumor stage, in early stage patients, a trend of reduced RFS was observed in those with low TP expression compared to high TP expression (P=0.071; Fig. 3a). Conversely, in advanced stage patients, high TP expression was significantly associated with reduced RFS compared to low TP expression (P=0.046; Fig. 3b). Further, the rest of the 5-FU metabolic enzymes did not correlate with the prognosis according to tumor stage.
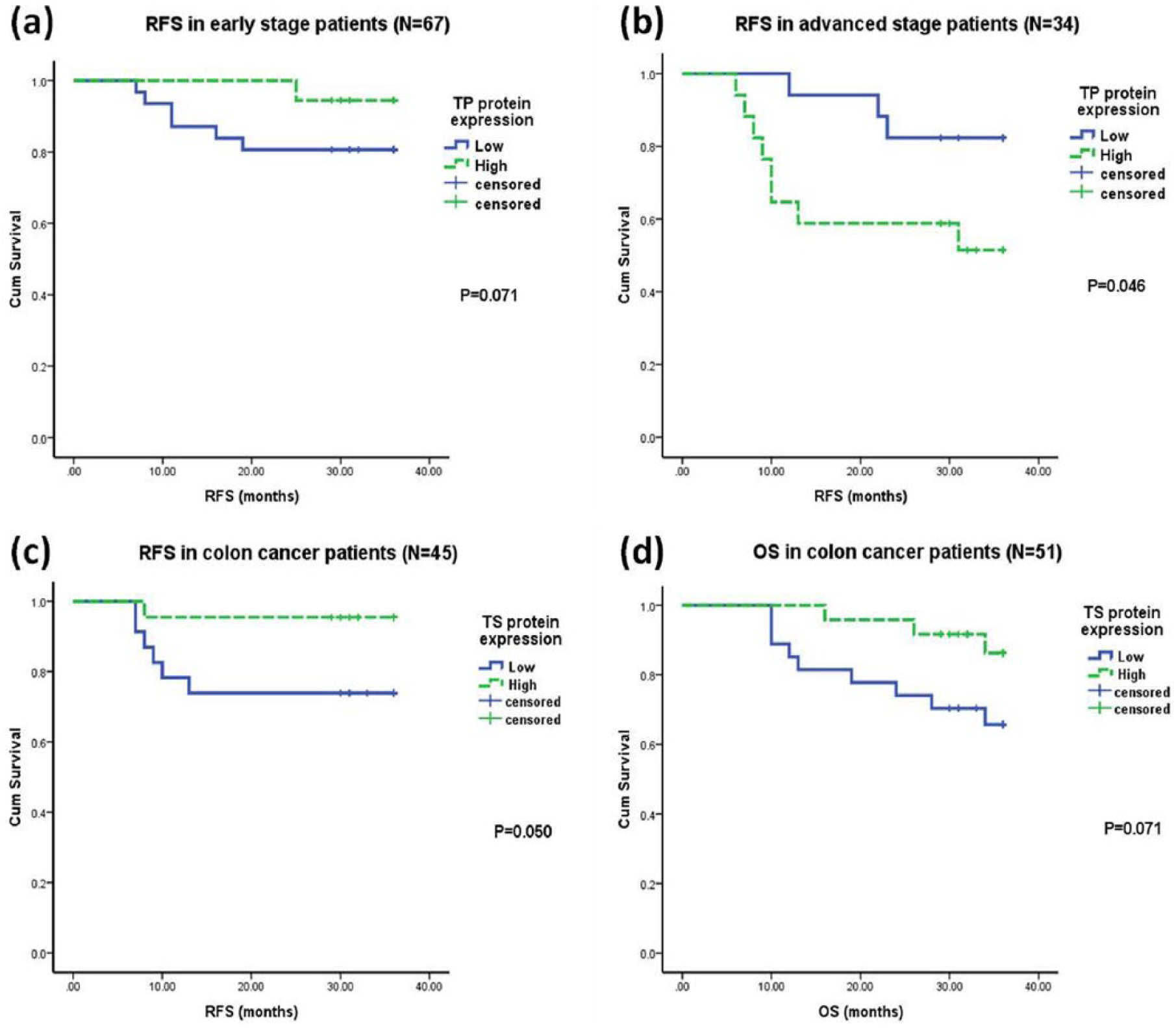
Kaplan–Meier survival curves according to disease stage and tumor site. a) Reduced RFS with low TP expression in early stage patients. b) Reduced RFS with high TP expression in advanced stage patients. c) Reduced RFS with low TS expression in colon cancer patients. d) Reduced OS with low TS expression in colon cancer patients.
According to the tumor site, in colon cancer patients low TS expression was associated with reduced RFS (P=0.050; Fig. 3c) and OS (P=0.071; Fig. 3d). Additionally, in rectal cancer patients, a trend of unfavorable OS was observed with low DPYD expression compared to high DPYD expression (P=0.072; Fig. 2b).
However, according to adjuvant treatment, protein expression of any of the 5-FU enzymes was not associated with the prognosis in the subgroup of patients whether treated with single agent 5-FU or with combined 5-FU+OX-based therapy (data not shown).
Effect of coexpression of TS, MTHFR, DPYD, and TP on prognosis
In total CRC patients, coexpression of all four proteins was analyzed and a significantly worse OS was observed in patients with low coexpression of all four proteins (TS, MTHFR, DPYD, and TP) compared to high coexpression of these proteins (P=0.045; Fig. 4). Similarly, in total patients, when the effect of coexpression of three proteins (TS, MTHFR, and TP) was considered, no significant difference in RFS (Log rank=0.717; df=3; P=0.869) and OS (Log rank=1.794; df=3; P=0.616) was observed (Tab. IV). Further, coexpression of two proteins (TS and MTHFR, TS and DPYD, TS and TP, DPYD and TP, MTHFR and TP, MTHFR and DPYD) was correlated with survival. However, no significant association was observed (data not shown). Similarly, no such association was observed in the subgroups of patients treated with 5-FU alone or 5-FU+OX-based therapy.
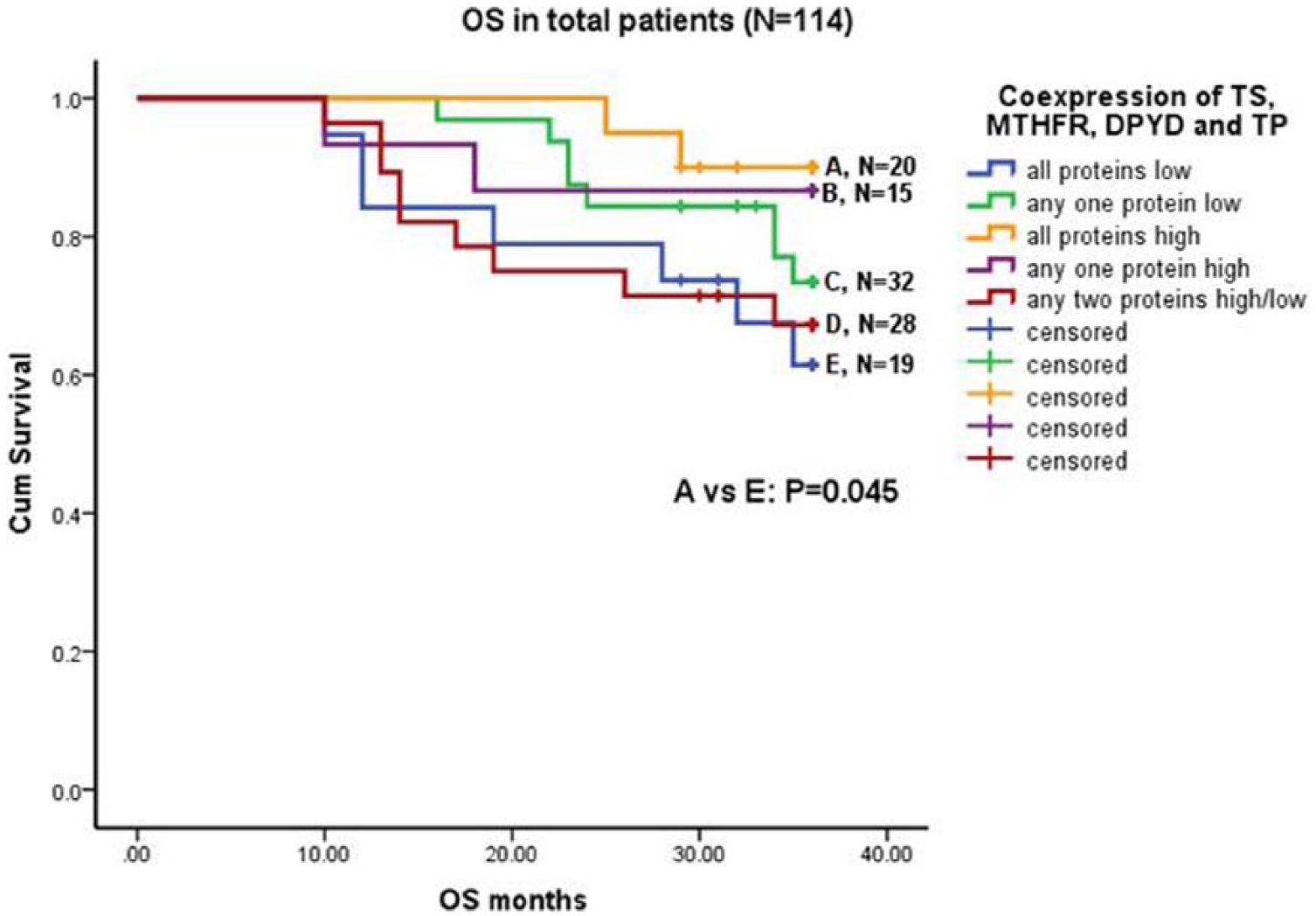
Overall survival curve with coexpression of all four proteins in total CRC patients. Low coexpression of TS, MTHFR, DPYD, and TP had significantly reduced OS compared to high coexpression of these proteins.
Association of TS, MTHFR, and TP coexpression with RFS and OS
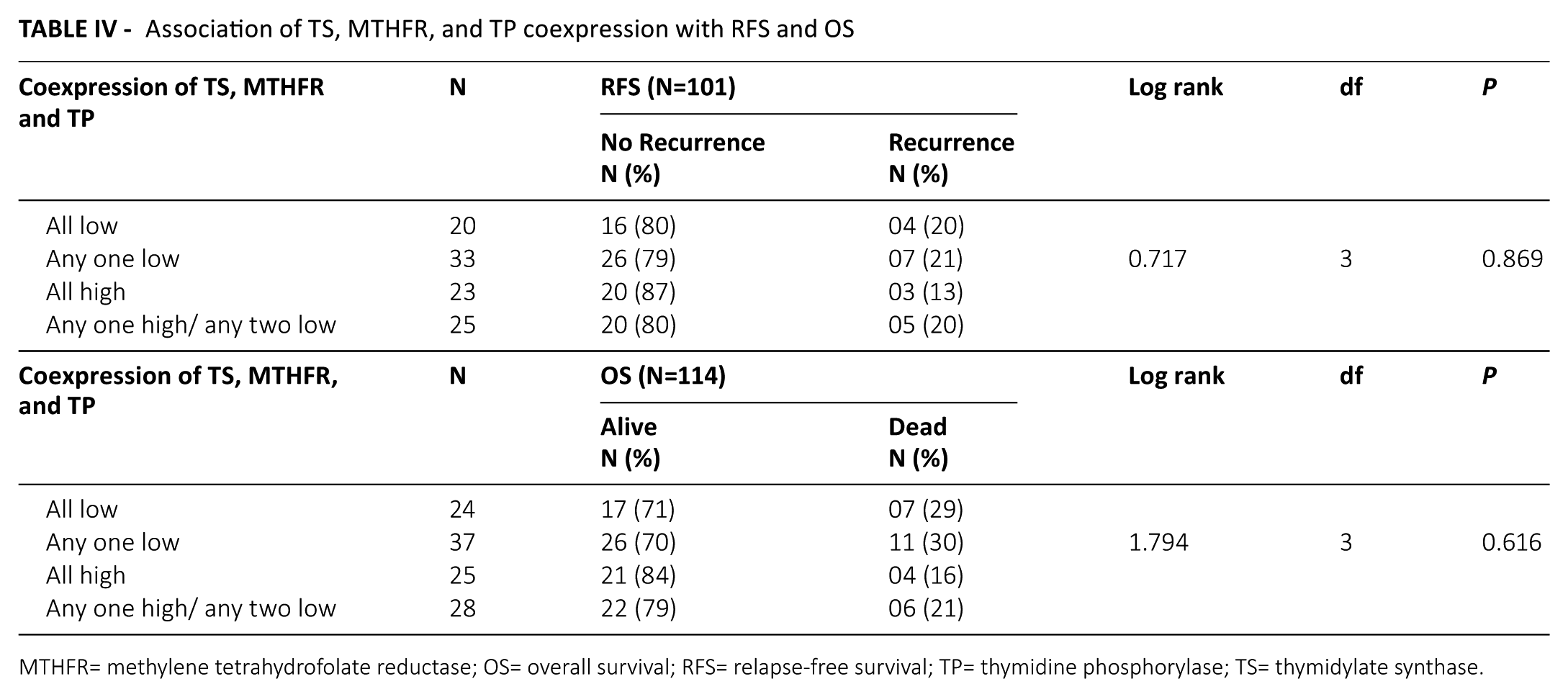
MTHFR= methylene tetrahydrofolate reductase; OS= overall survival; RFS= relapse-free survival; TP= thymidine phosphorylase; TS= thymidylate synthase.
Intercorrelation of TS, MTHFR, DPYD, and TP protein expression levels
The non-parametric Spearman’s correlation revealed that TS protein expression showed a significant positive correlation with MTHFR (r=+0.273; P=0.001) and DPYD expression (r=+0.310; P<0.001), and a trend of inverse correlation with TP expression (r=−0.154, P=0.067). Moreover, a significant positive correlation was observed between MTHFR and DPYD expression (r=+0.260; P=0.002).
Discussion
In CRC, it has been well documented that the molecules related to therapeutic response or failure would identify the patients with risk of recurrence and persistent disease, and also be helpful in selecting the best treatment for them. Thus, with this view point, the current study was embarked on to evaluate the possible role of important molecular enzymes involved in the metabolism of chemotherapeutic drug 5-FU (TS, MTHFR, DPYD, and TP).
5-FU, the standard treatment for CRC, mainly acts by inhibiting the activity of the TS enzyme. Due to the controversial reports regarding the prognostic and predictive role of TS protein expression in CRC, its clinical significance is yet to be elucidated. In the present study, immunohistochemical localization revealed that 77% of CRC patients exhibited heterogenous cytoplasmic TS protein expression. Similarly, Westra et al. (10) demonstrated cytoplasmic TS immunoreactivity in 90% of primary tumor samples in stage III colon cancer patients. Edler et al. (11) reported that TS protein expression was high in 76% of CRC patients. In contrast, a study in non-small cell lung cancer (NSCLC) showed that cytoplasmic TS positive protein expression was found in 57.4% of patients (12). There have been previous observations that it also may be expressed in the nucleus of human cells (13, 14). Additionally, Soong et al. (15) observed cytoplasmic and nuclear localization of TS protein expression in 61% of CRC patients. When the correlation was evaluated between TS protein expression and prognosis, the current study reported no association in total CRC patients. Similarly, several studies failed to show a relationship between TS protein expression and prognosis in CRC patients (16, 17). Allegra et al. (18) also showed no significant correlation of TS protein levels with survival in locally advanced colon cancer patients. In contrast, CRC patients having high levels of TS had poorer survival compared to those with low TS levels (19, 20). Similarly, patients with TS negative tumors had a better outcome than those with TS positive tumors in gastric and gastro-esophageal adenocarcinoma (21). Gao et al. (22) also showed a correlation of high TS expression with unfavorable survival and lower response rates to FU-based chemotherapy in gastric cancer. In contrast, several studies reported that high/positive TS protein expression correlated with better clinical outcome in patients with gastric cancer (23) and NSCLC (12, 24). The current study is consistent with these findings, but only in the subgroup of colon cancer patients. This likely suggests that colon cancer (presumed to be a biologically less aggressive tumor site) patients with high TS protein levels are probably more sensitive to adjuvant therapies, which confers to a better prognosis. This contradicts the theory of high TS and 5-FU resistance (25).
MTHFR, a vital enzyme involved in folate metabolism, also plays a role in 5-FU metabolism. A single in vitro study in colon cancer cell lines by Sekhon et al. (26) investigated the MTHFR protein expression by Western blot analysis, and reported that MTHFR may be required for tumor cell survival, and its inhibition should be considered for anti-tumor therapy. To the best of our knowledge, ours is the first study that investigated MTHFR protein expression by immunohistochemistry (IHC) and evaluated its prognostic role in CRC patients. MTHFR is a cytoplasmic enzyme (27) and in present study, cytoplasmic immunoreactivity was observed for MTHFR in 75% of CRC patients. The current study failed to observe any association of MTHFR protein expression and prognosis. Interestingly, MTHFR expression could discriminate patients according to tumor differentiation. These data should be treated as preliminary due to the small number of patients in well/poorly differentiated tumor groups.
DPYD is a rate-limiting enzyme of 5-FU catabolism and the present study reported 88% cytoplasmic DPYD immunoreactivity. These results were consistent with the view of Bai et al. (28), who documented cytoplasmic staining pattern in 67.7% of patients with CRC. Soong et al. (15) also observed low (34%) but predominant cytoplasmic staining of DPYD expression along with nuclear staining in a minority (<5%) of CRC patients. The reported data regarding role of DPYD expression and prognosis are conflicting. High/positive DPYD expression was associated with shorter survival rates in CRC patients receiving adjuvant 5-FU (29-31). On the other hand, DPYD protein expression was not associated with survival in stage III colon cancer patients treated with adjuvant 5-FU (10). Besides CRC, other malignancies, such as head and neck cancer (32) and OSCC (33), also showed an association of low DPYD levels with high response rates to 5-FU. In contrast to the above studies, the negative DPYD expression was significantly associated with the enhanced tumor cell proliferation and poorer prognosis in patients with intrahepatic cholangiocarcinoma (34). In accordance with this, the present results indicated low DPYD as a poor prognosticator to predict OS in total CRC patients and in the subgroup of rectal cancer patients. One possible explanation for this suggests that DPYD expression is related to tolerance and response to 5-FU therapy. In particular, low levels or absence of DPYD was found to be associated with the accumulation of 5-FU, which frequently leads to significant toxicities and adverse events, even death (35).
Studies have suggested that aberrant TP expression is a promoter of tumor growth affecting prognosis and treatment response. Hence, the current study explored the clinical utility of TP protein expression and documented no significant association with the prognosis in all the patients. However, in early stage patients, high TP expression was associated with better RFS; whereas in advanced stage patients, high TP expression correlated with adverse prognosis. The probable explanation for such variation could be due to the dual function of TP in predicting the clinical outcome. In early stage patients, because of its role in activation of 5-FU metabolic pathway, high TP expression in tumor tissue may lead to an increase in the intratumoral concentration of 5-FU (36), which inhibits DNA synthesis. Thus, it may improve the curative effect of 5-FU, which is important in the treatment of CRC and hence confers better survival (37). However, in advanced stage patients, the correlation of high TP expression with adverse prognosis may be due to the angiogenic role of TP as the disease advances. Thus, TP expression could be useful as a prognosticator in CRC patients according to the tumor stage. Similarly, several studies have suggested the prognostic role of TP in CRC (28, 38-40).
Since colorectal cancer is a heterogeneous disease with diverse growth rates and metastatic potential, it manifests different clinical outcome in patients with similar stages. Interestingly, when coexpression of the 5-FU metabolic enzymes was evaluated, it was observed that only the group of patients with low coexpression of all the four 5-FU metabolic enzymes (TS, MTHFR, DPYD, and TP) portends a significant reduced three-year overall survival. This indicates that the downregulation of these 5-FU enzymes’ coexpression might have a negative influence on survival.
Conclusion
Concomitant low expression of TS, MTHFR, DPYD, and TP identified patients with unfavorable prognosis and thus could be a useful prognostic indicator in CRC. Hence, combined use of this panel of four 5-FU metabolizing enzymes might emerge as a valuable biomarker in identifying the high-risk group of CRC patients with poor prognosis. However, for a definite conclusion a larger patient series is warranted.
Footnotes
Disclosures
Financial support: This study was financially supported by The Gujarat Cancer & Research Institute/ Gujarat Cancer Society.
Conflict of interest: None of the authors has financial interest related to this study to disclose.
List of abbreviations
5-FU 5-Fluorouracil
5-FU+OX 5-FU plus oxaliplatin
5-FUDR 5-fluoro-2′-deoxyuridine
5-MTHF 5-Methyletetrahydrofolate
CH2THF 5,10- Methylene tetrahydrofolate
CRC Colorectal cancer
dNTP Deoxynucleotide
DPYD Dihydropyrimidine dehydrogenase
dTMP Deoxythymidine-5′-monophosphate
dUMP Deoxyuridin-5′-monophosphate
dUTP Deoxyuridine triphosphate
FdUMP Fluorodeoxyuridine monophosphate
FUDR Fluorodeoxyuridine
H-score Modified histoscore
MTHFR Methylene tetrahydrofolate reductase
OS Overall survival
RFS Relapse-free survival
TK Thymidine kinase
TP Thymidine phosphorylase
TS Thymidylate synthase
