Abstract
Introduction
Enterococci are generally considered normal bowel commensals, but they are also known to cause urinary tract infections, as well as intraabdominal, pelvic, wound, and soft-tissue infections. Due to their ubiquitous nature they can appear as contaminants in various clinical specimens. When they are isolated from blood cultures, these organisms are taken as the main suspected factors that cause of invasive infections, namely sepsis and endocarditis, occurring in 18% 1 and 5–20%, 2 respectively. One of the most serious clinical manifestations of enterococcal infection is endocarditis, with mortality rates ranging from 15 to 28.9%. 3 Enterococci are, in fact, the third most common bacteria causing infective endocarditis after streptococci and staphylococci. 4 Approximately 90% of cases of the disease are caused by Enterococcus faecalis, and less than 5% by Enterococcus faecium. 4 Infection due to Enterococcus durans is extremely rare: only a few number of cases of endocardial infection have been reported. 5
It has been generally accepted that biofilms are involved in the pathogenesis of endocarditis. Biofilm is a population of cells attached irreversibly on various biotic and abiotic surfaces and encased in a hydrated matrix of exopolymeric substances. Bacteria in those structures are resistant to phagocytosis and high concentrations of antibiotics, which makes them difficult to eradicate.
Little is known about the extent to which various virulence factors contribute to invasive infections due to enterococci. Several factors have been pointed out as potential contributing components to the pathogenicity of enterococci including: adhesins: enterococcal surface protein (which contributes to the colonization of urinary tract epithelial cells); 6 aggregation substance, (contributes to bacterial clumping, survival in neutrophils, and adherence to host tissues); 7 and collagen binding adhesin; 8 invasins: gelatinase (a zinc-metalloprotease); 9 and hyaluronidase; in addition, enterococcal hemolysin is a toxin capable of destroying erythrocytes in several different mammalian species, especially in rabbits, horses, cows, as well as and humans; it is also a cytolysin. 10
In the present study, we have looked into enterococcal isolates which were proven to cause invasive infections in humans, and their ability to form biofilms, the pathogenicity factor genes contained, and antimicrobial susceptibility. Their characteristics were then compared to the same traits of colonizing enterococci from stools.
Materials and methods
Samples and patient data
The study encompassed isolates of enterococci obtained from blood cultures of patients from three clinics of the Clinical Center of Serbia in Belgrade, Serbia, over a three-year period: January 2017-December 2019. During the study period, blood cultures from 2489 patients at the Clinic for Infectious and Tropical Diseases, 920 patients at the Clinic for Hematology and 241 patients at the Clinic for Surgery were examined, resulting in total of 3650 patients. All of them were carried out by the Bacteriology laboratory at the Clinic for Infectious and Tropical Diseases. The strains considered were only those that grew in pure culture and had been isolated more than once. Since they were normally colected from sterile sites, they were registered as “invasive”. 11 Another inclusion criterium for the study was the age of 18 and above, while a single isolate of enterococci isolated from mixed cultures per patient was established as the criterion for exclusion.
Definitions
To make sure that the isolated strains were true pathogens, each case of the disease was discussed with clinicians who provided the diagnosis of sepsis or endocarditis. Sepsis was classified using previously proposed definitions, which introduced the term “SIRS” and infection. The “SIRS” is a systemic inflammatory response syndrome characterized by several clinical symptoms such as elevated body temperature or heart rate, tachypnea, and changes in white blood cell count, among others. 12 The cases of endocarditis were defined according to Duke Criteria, based on pathological criteria, major clinical and minor clinical criteria. 13
The control group of colonizing enterococci consisted of isolates from healthy medical students’ stools. The Ethics Committee of Clinical Center approved the study after all participants had provided their written consent.
Identification and antimicrobial susceptibility testing
Blood samples in the volume of 10 mL were incubated in the semi-automatic blood culture system BACTEC 9240 (Becton Dickinson Diagnostic Instrument Systems, Sparks, Md.). The bottles were incubated for 5 days at the temperature of 35°C, as recommended by the manufacturer of the system. Samples declared positive by the system were plated on blood agar, MacConkey and Chapman agar (Torlak, Belgrade, Serbia) for further examination. Stool samples were collected upon defecation and stored at 4°C for up to 4 h until their cultivation on blood agar. They were obtained by scraping with cotton swab and diluted in 10 mL sterile saline solution. A loopful of suspension, 10 μL, was set up on blood agar. Enterococci, either blood or stool isolates, were recovered from blood agar and presumptively diagnosed by appearance on the medium, Gram stain, catalase test, esculin hydrolisis and mannitol fermentation test. All of them were finally identified by MALDI-TOF. Enterococcal isolates both from blood and stool cultures were preserved on tryptic soy broth supplemented with 20% glycerol. Antibiotic susceptibility tests were performed by agar dilution tests on Mueller-Hinton II agar plates (Torlak, Belgrade, Serbia).
The antimicrobial agents tested were: ampicillin, vancomycin, ciprofloxacin, and aminoglycosides (streptomycin and gentamicin). High level resistance to gentamicin was defined as an Minimal Inhibitory Concentration (MIC) of ≥128 mg/L, and high level resistance to streptomycin as an MIC of ≥512 mg/L. The antibiotics used are those that are commonly prescribed by the Clinical Center of Serbia. Susceptibility testing was interpreted according to EUCAST guidelines, using the recommended breakpoints to define resistance. 14
Pathogenicity factor genes and hemolysin production
Sequence of primers used to detect the virulence genes.

The hemolysin production was determined on Blood Agar base (Torlak, Belgrade, Serbia) supplemented with 5% fresh human blood (group A), but the evaluation was also conducted on agar base that had been supplemented with 5% sheep blood.
Zones of clearing around colonies after 24 h at 37°C indicated capability of β-hemolysis. α hemolysis was characterized by greenish color behind the colonies, while phrase γ hemolysis indicated the situation when no hemolysis occurred.
Biofilm formation assays
Biofilm formation was evaluated with a quantitative adherence assay as described previously, 17 on the wells of a sterile 96-well flat-bottomed polystyrene microplate (Spektar, Čačak, Serbia; prepared by the manufacturer for tissue culture work). The negative control wells contained broth only. The optical density (O.D.) of each well was measured at 550 nm using microplate reader (Diagnostics Pasteur, Strasburg-Schiltigheim, France). Isolates were categorized based on the Stepanović’s approach 17 as strong, medium, weak or nonbiofilm formers while the cut-off O.D. (O.D.c) was defined as three standard deviations above the mean O.D. of the negative control. Each assay was performed in triplicate and the results were averaged.
Statistical analysis
All data were analyzed applying the statistical tool Statistical Package for the Social Sciences (SPSS) Version 20 (IBM Corp., Armonk, NY, USA). The Mann Whitney U test, non-parametric test, was used to determine whether there was a link between the amount of enterococcal biofilm production and pathogenicity factor genes presence in the group of invasive and colonizing isolates. Other statistical calculations were expressed by two-by-two contingency tables (Fisher’s exact test or chi-square test). The p-value ≤0.05 was considered as “statistically significant.”
Results
Patient data
Invasive isolates

E. fs: Enterococcus faecalis; E. fm: Enterococcus faecium; m: male; f: female.
Invasive isolates
The collection of invasive isolates consisted of 14 strains of E. faecalis, 2 E. faecium, and 1 E. durans. All of them were vancomycin susceptible. None of E. faecalis from the study was ampicillin-resistant, while 1 out of 2 E. faecium was resistant to that antibiotic. Susceptibility to high aminoglycoside levels (gentamicin and streptomycin) was both 35.3%.
Pathogenicity factor genes content and β-hemolytic properties of E. durans, E. faecium and E. faecalis.
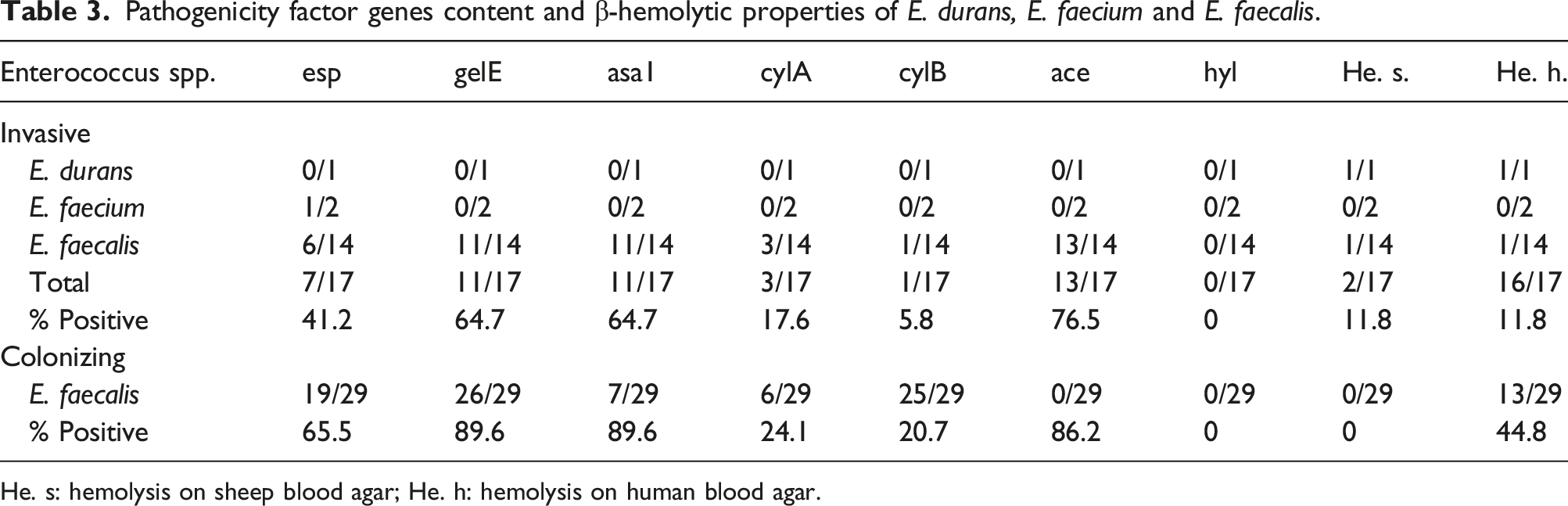
He. s: hemolysis on sheep blood agar; He. h: hemolysis on human blood agar.
Antimicrobial susceptibility and biofilm formation of E. durans, E. faecium and E. faecalis.

Biofilm formation: 0: non biofilm former; +: weak; ++: medium; +++: strong biofilm former; Amp: ampicillin sensitive; Van: vancomycin sensitive; Cip: ciprofloxacin sensitive; Gen: gentamicin sensitive; Strep: streptomycin sensitive.
Colonizing isolates, and virulence factor genes comparisons in two groups
Since all invasive isolates were vancomycin susceptible and E. faecalis was the predominant species (82.3%), only strains of E. faecalis (n = 29) susceptible to that antibiotic were chosen to be members of the control, or colonizing group (Table 4). Antibiotic susceptibility rates were higher in that group, with susceptibility to gentamicin and streptomycin at 62.1% and 51.7% respectively. The susceptibility to ampicillin and vancomycin was 100% for both of them. However, differences between the two groups were only significant in regards to ciprofloxacin susceptibility (p = 0.014). Genes esp, gelE, asa1, ace, cylA, and cylB were present in 19 (65.5%), 26 (89.7%), 26 (89.7%), 25 (86.2%), 7 (24.1%) and 6 (20.7%) isolates respectively (Table 3). Colonizing isolates possessed more virulence factor genes than invasive ones (2–6 per strain), however the differences were not statistically significant.
For invasive strains, the Mann-Whitney U test could not identify a correlation between the presence of each virulence factor gene and the amount of biofilm production. In colonizing strains the test indicated that the absence of ace gene was linked to higher amount of biofilm formation (p = 0.047). Only one colonizing strain was not a biofilm former (3.4%); others produced it in a small (9, or 31.0%), moderate (14, or 48.3%) or large (5, or 17.2%) amount. There was no statistical difference in the abundance of produced biofilm between invasive and colonizing groups of enterococci (p = 0.233).
In six isolates obtained from patients who died of sepsis, three produced biofilm in a small amount, one in a medium amount, and one in a large amount, while one did not produce any (E. faecium), thus leading to the conclusion that the production of biofilm could not be prerequisite for a deadly outcome. However, all endocarditis isolates were biofilm formers.
Hemolysis production and its correlation with cyl genes
Out of 46 isolates, 2 (4.3%) were β-hemolytic on agar supplemented with sheep blood, and 15 (32.6%) were β-hemolytic on agar supplemented with human blood. While 13 colonizing versus 2 invasive isolates produced β-hemolysis on agar supplemeted with human blood (p = 0.021), none of colonizing versus 2 invasive isolates was β-hemolytic on agar with sheep blood. Either cylA or cylB were detected in 3 (17.6%) of invasive and 12 (41.4%) of colonizing isolates (p = 0.1), while both genes were present in one member of each groups (5.9% and 3.4% respectively). The presence of any cyl gene correlated with β-hemolysis on agar supplemented with human blood: cylA (p = 0.008) and cylB (p = 0.003). Based on the study of the invasive and colonizing strains separately, a correlation with β-hemolysis was only noted on agar supplemented with human blood when members of the group of colonizers had the cylA gene (p = 0.019).
Discussion
Enterococci are important pathogens in humans, but they can also be contaminants of patients’ samples and therefore an irrelevant part of the microbiological results. If an isolate is obtained in a single blood culture, there is always a risk that it is not clinically significant. Our analysis, therefore, focused only on cases when repeated isolations of the same and single microorganism in two or more blood cultures were obtained and when clinicians confirmed invasive infection. In Khatlib’s study 18 of what differentiates bacteremia from blood culture contamination yielding enterococci, vancomycin and ampicillin resistance were identified as contamination predictors, along with the presence of commensal organisms and nonurinary/abdominal sources of sepsis. Considering the purity of the culture and sensitivity to vancomycin, the isolates from blood culture in the present study were comparable to Khatlib’s estimation of invasive strains: enterococci were all sensitive to vancomycin and were isolated in pure culture, no matter what species was diagnosed. In addition, according to other authors vancomycin-resistant Enterococcus often exists as a colonizing organism that does not necessarily contribute to infection, which makes it more difficult to determine exactly when and how these patients should be treated. 19 Regarding the source of infection, the majority of underlying diseases in the study group were associated with the abdominal/urinary system (70.6%): liver, gall bladder, pancreas, rectum, and prostate.
The antimicrobial susceptibility testing of invasive strains revealed a 5.9% resistance to ampicillin since only one isolate was resistant. It was not surprising that it was E. faecium since the majority of hospital-associated members of that species are resistant to ampicillin. 20 Susceptibility rates to gentamicin and streptomycin high levels were 35.3% for both of them. The given data are different and better than the results obtained from another study conducted partly at the same institution, which consisted of analysing all blood culture strains from within the entire Clinical Center. The results of this study indicated that 71.4% of E. faecalis strains were resistant to both streptomycin and gentamicin, while resistance rates were 100% and 95% respectively in the case of E. faecium. Furthermore, resistance to glycopeptides was 28.6% in E.faecalis and 95% in E. faecium. 21 This study included primary isolates from all clinics but did not require a careful evaluation of the clinical significance of each isolate of enterococci. As they were not evaluated for true invasiveness, some of them sometimes appearing in just one blood culture, it is highly probable that some of the results could have been skewed by contaminants.
There was no significant difference in antimicrobial susceptibility among the groups of invasive and colonizing strains, except for ciprofloxacin, to which invasive strains were more resistant (p = 0.014).
Resistance to ciprofloxacin is related to risk factors like hospital-acquired Urinary Tract Infection (UTI), previous treatment in a urological department, and transfer from a health care center. 22 In a French study with healthy donors none of the participants carried fluoroquinolone-resistant E. faecalis in the beginning. But, after ciprofloxacin treatment in some of them developed a high-level resistance to that group of antibiotics. 23 Given the diagnoses of the recidivant or a chronic character of the diseases or earlier surgical interventions in our patients, it is likely that they had spent some time in hospitals previously, during which they could have acquired the ciprofloxacin resistant enterococci. In the present study susceptibility rates to this antibiotic were low, 41.2% and 51.7% in invasive and colonizing isolates respectively, but in some studies on chronic patients they were considerably lower, from 10.9%-0%.24,25 The results of the current study indicate that ampicillin or vancomycin are still efficient antibiotics for the treatment of invasive enterococcal infections while the combination of ampicillin and aminogycosides has a synergistic effect in about one third of the cases.
Although 3 species of enterococci caused invasive diseases, E. faecalis was the most common. The members of that species carried many more virulence genes (2–5) than E. faecium, in which only esp in one strain was identified. This observation is in keeping with the results of some other studies, which came to similar conclusions.26,27
In an attempt to estimate the importance of the esp gene in enterococci causing endocarditis, the latter was detected in only 2 out of 7 isolates from patients with this condition. Hence, even though all esp positive strains in the study (6 invasive and 19 noninvasive) formed biofilms, the conclusion was that the esp gene was not required for endocarditis pathogenesis or biofilm formation. All endocarditis strains (E. faecalis, E. faecium and E. durans) were able to form biofilm at least in a small amount, while E. durans (isolate B1) and one strain of E. faecalis (isolate B5, causing recidivant infection) produced a moderate amount of biofilm. This is the evidence that demonstrated the critical role biofilms play in endocarditis.
The other virulence factor genes (gelE, asa1, ace, cylA or cylB) could not have influenced biofilm production of enterococci, except ace, whose absence correlated with heavier biofilm formation among colonizing isolates. Other authors agree that none of the genes analyzed can accurately predict pathogenicity or biofilm production in the tested bacteria28–30 explaining that biofilm formation is more dependent on the adhesion properties and environmental factors like medium composition, than on virulence genes. In light of this, we now understand that the negative coupling of the ace gene and the more pronounced biofilm production observed in colonizing isolates is the result of factors other than pathogenicity factor genes.
Virulence genes are characterized by their capability of colonizing and invading of the host’s tissues or affecting the host by producing toxins or causing inflammation. Since their presence enhances the pathogenicity, it is expected that invasive strains carry them in a greater number than the colonizing ones, but this was not the case. They were almost equally present in both groups of enterococci: curiously, the percentages were even larger in colonizing group. Similarly, it was the Bulgarian author Strateva 31 who noted that esp and asa1 gene, well known for their adhesive properties, are more prevalent among colonizing than invasive enterococci.
No single virulence gene or pattern of genes from this study could be ascribed specifically to endocarditis, sepsis or intestinal carriage. This observation is in line with Fahmy’s, study 32 who reported that there was no link between the distribution of virulence markers and the type of infection either for invasive or non-invasive infections. Interestingly, both genes gel E and asa1 were noted exactly in the same ratio in invasive (64.7%) and colonizing group (89.7%). They were the most frequently detected genes in all colonizing strains, followed by ace gene (86.2%), while ace (76.5%) was most prevalent among invasive strains, followed by gel E and asa1. Thus, the 3 genes predominated in both groups of isolates. Hyl gene was not detected in any of isolates, but the lack of its detection, especially in E. faecalis, has been documented previously. 33
Finally, there was no proof that E. durans produced any detectable genes of virulence in Patient 1 when it caused endocarditis. The strain produced strong β-hemolysis on blood agar supplemented not only with human but also with sheep erythrocytes. It is known that sheep erythrocytes are more resistant to hemolysin mediated lysis than erythrocytes of human origin. 10 The reason might lie in the lipid composition of the sheep erythrocyte membrane whose level of lipids is considerably higher than the one found in erythrocytes of other mammalian species, including humans. Membrane lipids, namely phosphatidylcholine and sphingomyelin, inhibit lysis of erythrocytes by hemolytic E. faecalis. Erythrocytes of mammalian species contain phosphatidylcholine in high levels, while sheep erythrocytes do not, and that compound might be the binding component for hemolysin from E. faecalis on the membrane of erythrocites. 34 Consequently, it could be that the two features of human erythrocite membrane (high phosphatidylcholine and low total lipid content) enable some enterococci to produce visible hemolysis on human blood agar.
While it has been documented that cytolysin activity requires the complete cyl operon (genes cyl L L L S ABM),15,35 E. durans did not have the complete operon or at least not cylA and cylB genes, which indicates that they were not essential for producing β-hemolysis in that species. It was evident that other additional determinants in E. durans control biofilm formation. Interestingly, despite the observations that vegetations due to cytolytic enterococci exhibited a significant increase in lethality compared to those caused by isogenic strains specifically defective in cytolysin production, 36 both of the patients whose isolates exhibited β-hemolysis on blood agar supplemented with sheep erythrocytes made a full recovery and were in a good condition on the follow up. Generally, colonizing isolates more often produced β-hemolysis on agar supplemented with human blood compared to invasive ones (p = 0.021). The occurrence of β-hemolysis on agar supplemented with human blood appears to be a trait of colonizing enterococci from stool samples and, according to the present data, cylA gene seems to play an important role in that process (p = 0.019).
All patients diagnosed with endocarditis survived, while 6 of those suffering from sepsis did not. There was no correlation between the amount of biofilm produced by strains recovered from cases of enterococci-related deaths and the mortality rate as they produced different amounts of biofilm or none at all. It is highly probable that the fatal cases may not be solely caused by enterococcal infections, but rather as a result of serious underlying diseases that have contributed to the unfavorable outcome.
The weakness of the study is the small number of strains tested, especially species other than E. faecalis. The second, the sample size calculation was not done. The third, a limited number of virulence genes were examined, and the results, in conjunction with some others, could shed new light on the conclusions reached.
Conclusion
In conclusion, the content of virulence genes between the colonizing and invasive groups of isolates was not significantly different. A blood agar medium supplemented with human blood served as a suitable medium for detecting cyl genes in colonizing enterococci. A small number of isolates included in the present study indicates that further research is required in that direction.
Supplemental Material
Supplemental Material - Comparative study of virulence factor genes, β-hemolysis and biofilm production in invasive and colonizing enterococci
Supplemental Material for Comparative study of virulence factor genes, β-hemolysis and biofilm production in invasive and colonizing enterococci by Milica Jovanović, Branko Velebit, Tanja Tošić, Gina Maki, Sladjana Pavić, Snežana Jovanović, Rajica Stošović and Marcus J Zervos in European Journal of Inflammation.
Footnotes
Acknowledgements
We would like to express our gratitude to the doctors at the Clinic for Infectious and Tropical Diseases, the Clinic for Hematology, and the Clinic for Surgery, for providing us with the information about patients' diagnoses and for the discussion on their illnesses.
Authors contributions
MJ and MZ designed the trial. TT and SJ have conducted the work and were involved in data collection. GM, MJ and BV conducted experiments. SP and RS participated in interpretation of results. All authors supported forarticle writing and final approval of the article.
Declaration of conflicting interests
Marcus Zervos, M.D. discloses conflicts of interests for grants: Cempra, Paratek and Genentech consulting medicines company. The other authors have no conflict of interest.
Funding
The authors received financial support for the research from grants: Cempra, Paratek and Genentech consulting medicines company.
Ethics approval
Ethical approval to report this case was obtained from The Ethic Board of Clinical Center of Serbia (Approval number 447/5). Written informed consent was obtained from all study paarticipants.
Informed consent
Written informed consent was obtained from all subjects before the study.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
