Abstract
Objective
To investigate the X-ray-specific sensitive genes and potential signaling pathways involved in the latent period of radiation-induced lung injury (RILI) in mouse models.
Method
Mice were randomized into groups for whole thoracic irradiation with a single fraction of 20 Gy X-ray or 12.5 Gy carbon heavy ion. Lungs were harvested 3 weeks after the irradiation, whole RNA was extracted and detected with the genome-wide transcriptional microarrays. Differentially expressed genes (DEGs) were calculated for each group and the X-ray-specific sensitive genes were determined, followed by the gene enrichment analysis of those DEGs exploring the potentially relevant signaling pathways and biological processes in latent RILI.
Results
Three weeks after irradiation, gene expression levels varied between groups. 76 up-regulated DEGs were determined with mice in the X-ray group and gene ontology enrichment analysis for biological process (GO-BP) obtained several processes which were associated with radiation reaction, mitotic, immune cell chemotaxis or metastasis, immune factors, p53 apoptosis, and tissue remodeling. KEGG signaling pathway enrichment analysis showed that those 76 up-regulated DEGs were enriched in p53, IL-17, FoXO, melanoma, and non-small-cell lung cancer signaling pathways. By comparing the DEGs in X-ray and heavy ion groups, X-ray-specific sensitive genes were determined, the top 10 genes were Adamts9, Aacs, Col6a2, Fdps, Mdk, Mcam, Stbd1, Lbh, Ak3, and Emid1. The expression level of the top 10 genes was found to be significantly higher in the X-ray group than in the control and heavy ion groups.
Conclusion
Our research determined the X-ray-specific sensitive gene set in mice lungs after exposure to radiation. The gene set could be used as a genetic marker to suggest the latency of RILI. The enrichment analysis results suggested that the relevant signaling pathways were potentially involved in the development of RILI. Further validation of those genes and signaling pathways is needed to confirm these findings.
Keywords
Introduction
Radiation pneumonitis is a subacute inflammatory reaction caused by lung tissue damage after radiotherapy for lung cancer, breast cancer, esophageal cancer, malignant lymphoma, and other chest malignant tumors.1-4 It usually occurs during radiotherapy or within 1–6 months after the end of radiotherapy.5,6 In the early stage of radiation-induced lung injury (RILI), the main manifestation is pneumonia (usually 1–3 months after radiotherapy), which is lymphocytic alveolitis that generates an immune response due to the activation of a large number of T lymphocytes. 7 In the late stage of RILI, it may develop into interstitial pulmonary fibrosis. 8 The incidence of radiation pneumonitis was 5%–15%. 6 Clinical symptoms may include persistent dry cough, breath shortness, mild fever, or occasionally high fever which may be secondary to radiation pulmonary fibrosis, and in some cases, even the direct cause of death. 1 In addition to individual sensitivities, the occurrence of radiation pneumonitis is usually related to the dose of radiation, the volume of lung irradiated, and the interval time.9,10 Therefore, radiation injury is one of the key factors limiting the radiation dose of chest tumors, which can affect the local tumor control rate and the quality of life of patients post-radiotherapy.1,11
The clinical symptoms, imaging changes (Chest X-ray, CT), pulmonary function examination, etc., of radiation pneumonitis often show positive evidence 1 month after radiotherapy, 6 or 2.5 to 3 months after the first radiotherapy, the diagnosis can only be confirmed with auxiliary examinations. There was a hiding period before radiation pneumonitis, which lasted nearly 2 months. If we can diagnose or predict the pulmonary inflammatory reaction at this stage, effective intervention measures can be applied as early as possible, such as using low-dose steroids, anti-inflammatory drugs, etc. Early inhibition of the occurrence and development of inflammatory reactions can greatly alleviate or even reverse interstitial pneumonia, 12 which plays a key role in relieving pneumonia symptoms and reducing the morbidity of interstitial pneumonia and fibrosis.
At present, chest radiotherapy mainly adopts three-dimensional conformal radiotherapy, intensity-modulated radiotherapy, or spiral tomography radiotherapy, which mainly relies on linear energy transfer rays such as X-ray and electron rays. X-ray irradiation has been shown to have significant effects on gene expression. X-rays can cause DNA damage and oxidative stress, leading to changes in the regulation of gene expression. These changes can result in either the upregulation or downregulation of specific genes, depending on the dose and duration of exposure.13-17 In this study, low LET radiation represented by X-ray was used to irradiate the lung field of mice to establish lung injury models, and carbon ion ray modeling was used as the positive control of high LET ray to further optimize and select highly specific regulatory genes of X-ray. In the third week after irradiation (early inflammation stage), the genome-wide microarray analysis was used to detect mouse lung tissue to explore the differentially expressed genes related to the hidden progression stage of RILI.
Materials and Methods
Animals and Groups
C57BL6 female mice (8 weeks old) were used in our experiment and mice were purchased from Guangdong Medical Experimental Animal Center (License: SCXK [Guangdong] 2013-0002), feeding in the SPF barrier environment, temperature 22–24°C, relative humidity 50%–60%, pressure difference ≤10 Pa, 12 h/12 h alternating light and dark, free to eat and drink. The experimental mice were then divided into 3 groups, with 15 mice in each group. Mice in each group were anesthetized by inhalation of 2% isoflurane and by intraperitoneal injection of 80 mg/kg ketamine and properly fixed on the chest irradiation devices. The method was the same as described before. 18 In the X-ray irradiation group, a 6 MV medical linear accelerator was used for whole lung field irradiation, with a single dose of 20 Gy and a dose rate of 3 Gy/min. For mice in the heavy ion irradiation group, the lungs of mice were placed in the Prague peak area, with a peak width of 30 mm, and a single dose of 12.5 Gy (ie, the biologically equivalent dose of 20 GyE) for the whole lung field. The method description is the same as above19,20; mice in the control group were anesthetized but did not receive radiation.
Lung Tissue Acquisition and Total RNA Extraction
All mice were killed by the cervical dislocation method 3 weeks after irradiation, and fresh lung tissue was taken. After TRIzol treatment, total RNA in the lung was extracted with RNeasy Mini Kit (Qiagen), purified RNA with DNase I (Qiagen), and purified RNA was washed with nucleic acid-free water. The purified RNA can be stored in a refrigerator at −80°C for use.
Histopathological Observation
Fresh lung tissue was taken before X-ray irradiation and at the 3rd and 16th weeks after 20 Gy irradiation, fixed with 4% formalin, embedded with paraffin (5 μm thick), and then stained with hematoxylin-eosin (HE) to observe the pathological changes of alveolar structure, alveolar septal thickness, inflammatory cell infiltration, etc. at different time points.
Micro-CT Imaging
Micro-CT scanning imaging was performed before X-ray irradiation and 3 and 16 weeks after 20 Gy irradiation to evaluate the status of pneumonia in mice. CT scanning parameters were as follows: tube voltage 40 kV, current 0.4 mA, the acquisition time of each projection 1s, 360-degree rotation, 240 projections in total. The image is reconstructed to 256 pixels with a size of .13 mm × 256 × 512 Matrix.
Genome-Wide Microarray Detection and Differential Gene Analysis
2100 Bioanalyzer (Agilent) was used to determine the RNA concentration and integrity, and the gene expression level was detected by genome-wide expression microarray (Illumina, Mouse Sentrix-6 V2 Whole Genome BeadChip, BD-201-0202, USA). Adapt to the built-in software processing chip signals, generate original expression files and perform gene annotation and log2 conversion. R V3.4.2 was used for gene expression level analysis, Limma package was used for original data background correction, data standardization, and generation of expression matrix. Set that the fold change (FC) of gene expression level between the 2 groups is greater than 1.2 times, and P < .05 is the differential expression gene. The pheatmap package 21 was used to standardize the relative gene expression data and make a heat map. Cluster analysis was used to identify the expression level of related genes, so as to analyze the difference in gene expression levels in different groups of mice.
KEGG Signaling Pathways Enrichment Analysis
Carry out signaling pathways enrichment analysis on differential genes up-regulated in the X-ray group, use the clusterProfiler package 22 in R/Bioconductor to carry out KEGG pathway 23 gene set enrichment analysis and gene ontology biological process (GO-BP) enrichment analysis. 24 Use P < .01 and Q <.01 as screening conditions, and use GraphPad Prism 7 and R software for data analysis.
Results
Histological Evaluation of Radiation Pneumonitis Progression
In order to determine the inflammation of lung tissues at different time points before and after irradiation, lungs were taken before X-ray irradiation, 3 weeks after 20 Gy irradiation, and 16 weeks after irradiation for HE staining. Compared with lungs harvested before irradiation, there was no significant change in lungs that were harvested 3 weeks after irradiation. At the time point 16 weeks after irradiation, a large number of inflammatory cells infiltrated into the alveoli and widened alveolar septa, and structural disorders were observed with the microscope (Figure 1). It is suggested that the time point 3 weeks after irradiation is still in the early stage of pneumonia, and the pathological changes are insidious and histologically. HE stainings (×20 magnification) for mice lung sections, prior to irradiation, 3 weeks, and 16 weeks post-irradiation. No significant changes were observed in 3 weeks post-irradiation.
Imaging Evaluation of Radiation Pneumonitis Progress
CT and chest X-ray film are common means for clinical diagnosis of pneumonia. The lung micro-CT examination showed that the mouse lung CT had no significant difference before irradiation, 3 weeks after 20 Gy irradiation (latent progression stage), and 16 weeks after 20 Gy irradiation (histological positive stage), which could not effectively indicate the progress of radiation pneumonia (Figure 2). Axial view of mice lungs, prior to irradiation, 3 weeks, and 16 weeks post-irradiations by micro-CT imaging. No significant changes were observed between the 3 time points.
Genome-Wide Transcriptional Profiling in the Latent Stage of Radiation Pneumonitis
Three weeks after a single dose of 20 Gy X-ray or 12.5 Gy heavy ion (20 GyE) whole lung field irradiation, mouse lung tissues were taken for genome-wide microarray sequencing, and the differentially expressed genes were analyzed. Compared with the blank control group (0 Gy), there were 135 differentially expressed genes in the X-ray irradiation group, including 76 up-regulated genes and 59 down-regulated genes (differential expression change fold change (FC) > 1.2, P < .05) (Figure 3A). The heavy ion irradiation group was used as the positive control to further optimize the X-ray-sensitive genes. Compared with the blank group, 303 differentially expressed genes were determined in the heavy ion irradiation group, including 253 up-regulated genes and 50 down-regulated genes (differential expression change FC >1.2, P < .05) (Figure 3B). Volcano plots of sensitive genes in mouse lungs 3 weeks after exposure to X-ray or carbon ions. (A): Gene expression in lungs of mice in X-ray irradiation group, orange dots indicate differential genes. (B): Gene expression in lungs of mice in carbon ions irradiation group, blue dots indicate differential genes.
X-ray-Specific Up-Regulated Gene Enrichment Analysis
Gene ontology biological process (GO-BP) enrichment analysis was conducted on the 76 genes up-regulated in the X-ray irradiation group.
24
The above gene enrichment indicates that it is closely related to multiple radiation reactions, cell mitosis, immunocyte chemotaxis, transfer, immune factors, p53 apoptosis, and biological processes related to cell and tissue remodeling (Figure 4A). The KEGG signaling pathways enrichment analysis showed that the 76 genes were significantly related to p53, IL-17, FoXO, melanoma, and non-small-cell lung cancer (Figure 4B). Enrichment analysis for up-regulated genes in mice irradiated with X-ray 20 Gy. (A): GO-BP enrichment analysis results. (B): KEGG enrichment analysis results.
Analysis of X-ray-Specific Sensitive Genes
Venn diagram analysis was carried out on 2 up-regulation differential gene subsets in the lungs of mice in the X-ray irradiation group and heavy ion irradiation group, and 26 X-ray specific up-regulation genes were obtained (Figure 5A). This gene subset was significantly up-regulated 3 weeks after X-ray irradiation, while there was no significant change in the heavy ion irradiation group (Figure 5B). These results suggest that the above genes have high X-ray-specific transcriptional levels in the early stage of radiation pneumonitis, which has potential differential diagnostic value. The Venn analysis and heatmap view of X-ray specific genes. (A): Analysis of up-regulated gene Venn map in X-ray and heavy ion groups. (B): Heatmap of the expression of X-ray specific sensitive genes in the control group, X-ray group and heavy ion group.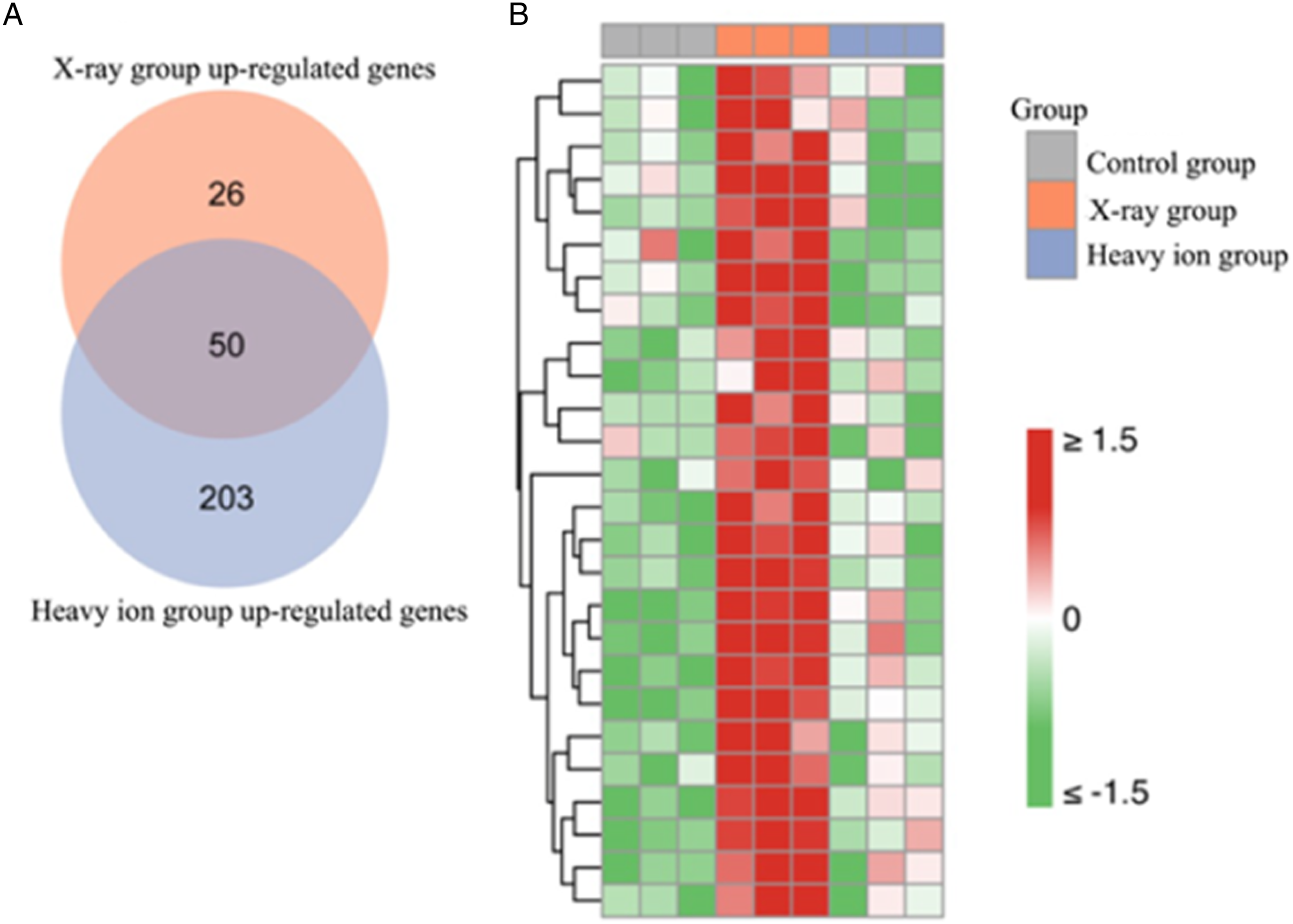
X-ray-Specific Sensitive Gene Marker Combination
Ten most significant differentially expressed genes were selected from the above 26 X-ray sensitive genes as the panel for the latent stage of RILI, including Adamts9 (A disintegrin and metalloproteinase with thrombospondin motifs 9), Aacs (acetoacetyl-CoA synthetase), Col6a2 (collagen type VI alpha 2 chain), Fdps (farnesyl diphosphate synthase), Mdk (Midkine), Mcam (melanoma cell adhesion molecule), Stbd1(Starch Binding Domain 1), Lbh (limb bud and heart development), Ak3 (Adenylate Kinase 3), Emid1 (emilin and multimerin domain-containing protein 1). The expression level of these genes was significantly higher than that in the blank control group and heavy ion irradiation group (Figure 6A). The expression level of a single gene shows an increasing trend (Figure 6B). In the X-ray irradiation group, the overall expression level of the 10 genes was found to be significantly higher than that in the control group or the heavy ion group (P < .0001, P < .0001), which could significantly distinguish the blank control from the high LET radiation (Figure S1). The expression levels of the top 10 X-ray specific sensitive genes in terms of 3 different groups. (A): Heatmap of the expression values of the top 10 X-ray specific sensitive genes in the control group, X-ray group and heavy ion group. (B): The expression value change multiple of the above 10 genes in the X-ray group relative to the control group.
The Gene Ontology Biological Process Enrichment Analysis
This analysis was carried out for those 26 X-ray-specific sensitive genes. The results indicated that the pathway was mainly related to the biological process of cell growth regulation, such as cell cycle DNA replication maintenance of fidelity, cell response to growth hormone stimulation, etc. (Figure S2, (A)). Further analysis of the protein-to-protein interaction network for X-ray-specific genes determined 4 genes (Cenpf, Rad51, Plk1, and Cdk1) with high node values, including 2 X-ray-specific genes (Cenpf, Rad51). The results suggest that Cenpf and Rad51 are the main binding points of protein interaction (Figure S2, (B)). The biological pathway enrichment analysis shows that X-ray-specific sensitive genes (Cenpf and Rad51) played a major role in the PPI network, as well as in the pathway of cell growth regulation (Figure S2, (C)).
Discussion
RILI is a common complication of radiotherapy for chest tumors. The early symptoms are always not obvious and examinations have a certain lag, it is difficult to detect RILI in advance. Early diagnosis and intervention are important in limiting the development of pneumonia and relieving clinical symptoms. 25 Therefore, performing early detection of radiation pneumonia has practical value. 26 At present, there is still a lack of specific biomarkers for detecting pneumonia clinically. Cell biology studies suggest that when external stimuli act on cells, cells will change the level of gene expression, and regulate the expression level of downstream proteins, leading to status changes in signaling pathways. 26 This regulation mechanism also exists in the cell stress response process to radiation. 15 This provides a theoretical basis for us to search for reliable genetic markers of early RILI. We can also reveal the occurrence and development of radiation pneumonitis by studying the evolution of the transcription level of gene signatures. In this research, an average dose of 20 Gy X-ray delivered to mouse whole lung is sufficient to induce symptomatic pneumonia and inflammation-related histological and pulmonary function changes, which consists of literature reports. 27 Heavy ion radiation is used as a positive control, and the biological dose of 12.5 Gy (ie 20 GyE) is also used for modeling. Three weeks after irradiation, the genome-wide transcriptional profiling was detected, and the highly specific and sensitive genes of X-ray-specific were screened. The sensitive gene panel can provide a prediction for the occurrence of RILI. Meanwhile, the enrichment analysis of GO-BP and KEGG of differentially expressed genes showed that many signaling pathways were involved in biological process changes, indicating the signaling pathways were closely related to the early changes of RILI. Post-exposure to X-ray radiation, a series of cascade reactions of cellular factors and chemokines occurred in cells, changing the permeability of capillaries and regulating the migration, infiltration, and differentiation of immune cells. In this process, pulmonary cells undergo remodeling changes, which may activate the endogenous apoptosis mechanism through the p53 signaling pathway, potentially suggesting a similar biology process in non-small-cell lung cancer (NSCLC) while elevated p53 levels trigger pyroptosis to produce anti- NSCLC effects. 28 The negative regulation of G1/S phase transformation of cell mitosis and proliferation of vascular smooth muscle cells (such as vascular remodeling to form a loop) was also involved in this pathological change. Evidence currently available indicates that checkpoint kinases can stimulate p53-dependent G1 arrest via regulation of transcription, which is a manifestation that occurs after DNA damage. 29
Our study proposed potential gene signatures and investigated the molecular mechanism and changes in signaling pathways during the asymptomatic stage of pneumonia caused by medical X-ray exposure in mice. This provides a basis for further screening and identification of reliable biomarkers for radiation-induced lung injury (RILI) and for identifying populations susceptible to radiation-induced pneumonia. Further verification of this gene signature panel is required through PCR, protein level analysis, and clinical sample analysis.
Supplemental Material
Supplemental Material - Gene Signatures for Latent Radiation-Induced Lung Injury Post X-ray Exposure in Mouse
Supplemental Material for Gene Signatures for Latent Radiation-Induced Lung Injury Post X-ray Exposure in Mouse by Tongtong Zhang, Zhaoming Zhou, Lei Wen, Changguo Shan, Mingyao Lai, Jing Liao, Xin Zeng, Gang Yan, Linbo Cai, Meijuan Zhou, and Minghua Wang in Dose-Response
Footnotes
Acknowledgments
Thanks to all the staff who contributed to this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Approval
The animal experiments were carried out in accordance with ethical procedures and the protocols were approved by the Ethics Committee for Laboratory Animals of the Southern Medical University. (L2019074).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
