Abstract
The carcinogenicity of radon has been convincingly documented through epidemiological studies of underground miners. However, there is a lack of early warning indicators for radon radiation damage. In this study, mixed serum samples of 3 groups were collected from 27 underground uranium miners and seven aboveground miners according to the radiation exposure dose. The differentially expressed proteins in the serum were identified using the isobaric tags for the relative and absolute quantitation (iTRAQ)-based method. Some differentially expressed proteins were validated by enzyme-linked immunosorbent assay (ELISA) in 84 underground and 32 aboveground miners. A total of 25 co-differentially expressed proteins in 2 underground miner groups were screened, of which 9 were downregulated and 13 were upregulated. Biological process analysis of these proteins using Metascape showed that 5 GO terms were enriched, such as negative regulation of very-low-density lipoprotein particle clearance, endocytosis, and regulated exocytosis. The results of the ELISA for the expression levels of GCN1, CIP2A, and IGHV1-24 in the serum of 116 miners’ serum showed that the levels of GCN1 and CIP2A were consistent with the iTRAQ results. In conclusion, APOC1, APOC2, APOC3, ORM1, ORM2, ANTXR1, GCN1, and CIP2A may be potential early markers of radon radiation damage.
Keywords
Introduction
Estimating the long-term biomedical effects of small doses of ionizing radiation is a complicated issue. It has been reported that more than 60% of the ionizing radiation a person receives every year can be caused by natural sources of radiation. 1 Radon, an imperceptible naturally occurring radioactive noble gas, is the largest single fraction of radiation exposure from natural sources. 2 It is the second most common cause of lung cancer after smoking. A correlation between radon and the frequency of lung cancer has been consistently demonstrated in several studies of indoor exposure in dwellings, as well as for uranium miners. 3 It is important to note that the current models estimating the risk of radiation-related hazards are based on the analysis of data collected from irradiated miners. In addition to using dosimetry methods to evaluate the risk of harmful radiation effects, the severity of such a risk should be evaluated using a biological marker. Biological test systems can evaluate the risks of radiation and provide a thorough assessment of the quality of the environment and its suitability for humans. 1 Due to its good liposolubility, radon and its progeny can easily reach the lung, enter blood circulation, and act on the human body. 4 Therefore, with increases in radon exposure, we investigate whether there are potential biomarkers in the peripheral blood that can reflect the changes corresponding to radon damage.
Proteomics is an emerging discipline that studies the composition and functions of all proteins in a particular type of cell, tissue, or body fluid in a large-scale, high-throughput, and systematic manner. Novel proteomics named isobaric tags for relative and absolute quantification (iTRAQ) combined with mass spectrometry technology (LC-MS/MS) now represents a powerful tool for the identification of cancer biomarkers. The iTRAQ technology has been successfully applied to biomarker screening of multiple tumors and diseases in both tissue and serum samples. 5 The aim of this study was to screen for differentially expressed proteins in serums from former uranium miners exposed to high doses of radon using iTRAQ based method, and to provide basic experimental support for the study of early warning molecular biomarkers of radon damage.
Materials and Methods
Study Population and Sample Preparation
This cohort study involved 116 male former uranium miners aged from 50 to 86 yr old at a uranium mine in Hunan Province, China, who had no acute infectious diseases, malignant tumors, or serious family history of genetic diseases. Miners were divided into 2 groups according to their workplace: 32 were aboveground miners (controls) and 84 were underground miners (the cumulative exposure dose of radon was 2.10–251.70 working level months [WLM]). All serum samples from miners were frozen following venipuncture and centrifugation and stored in liquid nitrogen until analysis.
Isobaric Tags for the Relative and Absolute Quantitation Analysis
Based on the smoking factors of uranium miners, 34 serum samples of uranium miners who were smokers were selected for iTRAQ detection and analysis. Three mixed serum samples were prepared according to the radiation exposure dose. The mixed serum group 1 consisted of serum samples from 11 underground uranium miners with an exposure dose of 21.00–54.57 WLM. The mixed serum group 2 consisted of serum samples from 16 underground uranium miners with an exposure dose of 82.32–217.05 WLM. Group 3 was a mixed serum sample of 7 aboveground miners and was used as the control group.
Isobaric Tags for the Relative and Absolute Quantitation analysis was conducted at Shanghai Applied Protein Technology Co.(Shanghai, China). Total proteins were extracted from samples using SDT lysis (4% SDS, 0.1 M DTT, 100 mM Tris/HCl pH7.6). Serum pools were depleted of the most abundant proteins using an Agilent Human 14 Multiple Affinity Removal System Column (Applied Biosystems, USA) following the manufacturer’s instruction. Ultrafiltration tubes were used for desalination and concentration of low-abundance components. Protein concentrations were quantified using the bicinchoninic acid (BCA) Protein Assay Kit (Bio-Rad, USA). Subsequently, samples were labeled using an 8-plex iTRAQ labeling kit (Applied Biosystems) according to the manufacturer’s protocol. iTRAQ-labeled peptides were fractionated by strong cation exchange (SCX) chromatography using the AKTA purifier system (GE Health care, USA). Each fraction was injected into the nanoscale liquid chromatography coupled to tandem mass spectrometry (nano-LC-MS/MS) system for analysis. High-resolution LC-MS/MS analysis was performed on a Q Exactive mass spectrometer (Thermo Fisher Scientific) operated in positive ion mode coupled to an EASY-nLC liquid chromatograph (Thermo Fisher Scientific). The MS data were acquired in a data-dependent acquisition mode. The raw files were processed using Proteome Discoverer 1.4 (Thermo Fisher Scientific) and searched using the Mascot search engine (version 2.2, Matrix Science, UK) against the UniProt protein human database. The false discovery rate (FDR) for the peptides was set to 1%. Among the identified proteins, proteins with more than 1.2-fold and P< .05 compared to control were defined as up-regulated expression, while proteins with less than 0.83-fold and P< 0.05 were defined as down-regulated expression.
Bioinformatics Analyses
Metascape (http://metascape.org) is a web-based portal designed gene-list tool that provides gene annotation, enrichment analysis, and a protein-protein interaction (PPI) network based on over 40 independent knowledge bases. 6 In our study, Metascape was employed for GO, KEGG, and PPI analysis to further explore the complex mechanism of co-differentially expressed proteins in 2 underground miner groups. We selected term enrichment significance according to P-value < 0.01, minimum count of 3, and enrichment factor > 1.5.
Enzyme-Linked Immunosorbent Assay
The human CIP2A enzyme-linked immunosorbent assay (ELISA) Kit, human GCN1 ELISA kit, and human IgHV1-24 ELISA kit were all purchased from AndyGene Biotechnology Co. (China) and used to detect the levels of CIP2A, GCN1, and IgHV1-24, respectively, according to the manufacturer’s instructions.
Statistical Analysis
Statistical analysis was performed using the Statistical Program for Social Sciences (SPSS, Inc., Chicago, IL, USA). The data are reported as the mean ± standard deviation (SD), and the quantitative variables were analyzed using Student’s t-tests between 2 groups. Statistical significance was set at < 0.05.
Results
Differential Expression Analysis of Serum Proteins in Underground Miners Using iTRAQ
The Differentially Expressed Proteins in Group 1 Compared to Group 3.

The Differentially Expressed Proteins in Group 2 Compared to Group 3.

Co-differentially Expressed Proteins in the 2 Underground Miner Groups (groups 1 and 2).

Functional Enrichment Analysis of Differentially Expressed Proteins
The differentially expressed serum proteins of groups 1 and 2 were considered for gene ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis using Metascape.
6
The results showed that 9 biological process terms were significantly enriched in the 2 groups: regulated exocytosis, hemostasis, protein-lipid complex remodeling, immunoglobulin production, endocytosis, zymogen activation, post-translational protein phosphorylation, cell-substrate adhesion, and contractile actin filament bundle assembly (Figure 1). To further investigate the relationship between the differentially expressed proteins in groups 1 and 2, a PPI network was constructed based on the information from the Metascape online database. The PPI network and molecular complex detection (MCODE) components identified in the gene lists are shown in Figure 2, and the top 4 modules with the highest scores were obtained from the PPI network. Figure 2 shows that the biological functions mainly related to MCODE1 were post-translational protein phosphorylation, regulation of Insulin-like Growth Factor (IGF), transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs), and regulated exocytosis. In MCODE2, these were plasma lipoprotein particle remodeling, protein–lipid complex remodeling, and protein-containing complex remodeling. In MCODE3, these were platelet degranulation, platelet degranulation, and response to elevated platelet cytosolic Ca2+. The biological function related to MCODE4 was leukocyte migration. Heatmap of enriched pathway terms of differentially expressed proteins in groups 1 and 2, colored according to P-values. Protein-protein interaction network and MCODE components identified in the protein list of groups 1 and 2.
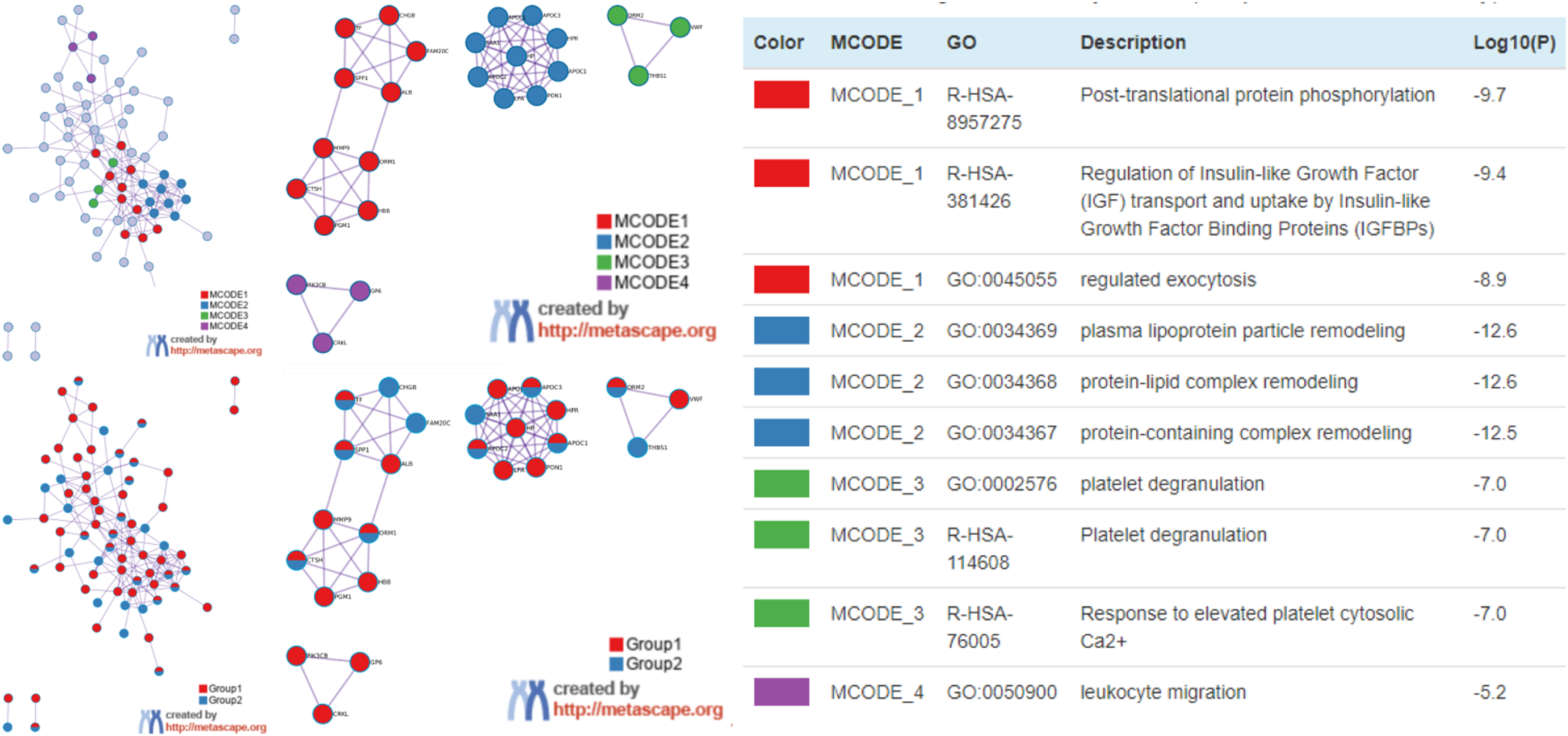
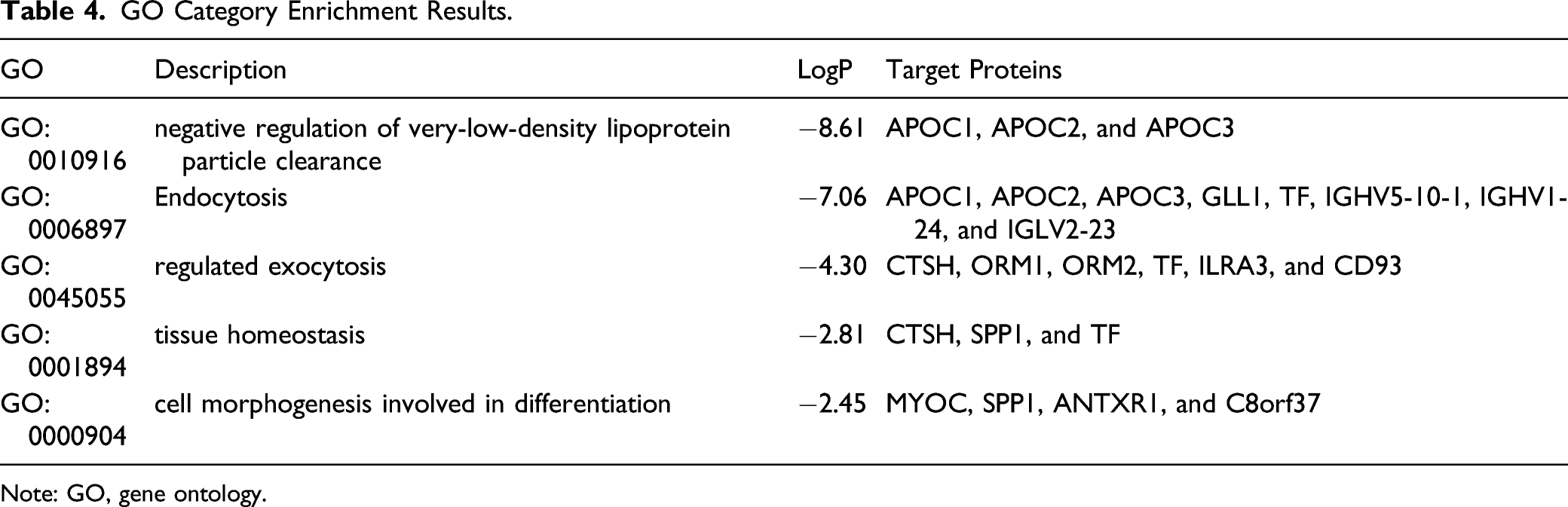
To further understand the mechanism and function of co-differentially expressed proteins, bioinformatic analyses were performed using Metascape. As summarized in Figure 3, 5 GO terms were mainly enriched: negative regulation of very-low-density lipoprotein particle clearance, endocytosis, regulated exocytosis, tissue homeostasis, and cell morphogenesis involved in differentiation (Figure 3A and 3B, and Table 4). Moreover, to investigate the relationship between co-differentially expressed proteins, a PPI network was constructed based on the information from the Metascape online database. The identified PPI network and MCODE1 component are shown in Figure 3C. The biological function related to the PPI network was very-low-density lipoprotein particle clearance, involving proteins such as apolipoprotein C1 (APOC1), C2 (APOC2), and C3 (APOC3). DisGeNET GO analysis in Metascape showed that co-differentially expressed proteins were associated with 30 GO terms, such as hyperlipoproteinemia type III, hyperinsulinism, drug toxicity, etc. The top 20 GO enrichment pathways are shown in Figure 4 and Table 5. Enrichment analysis of co-differentially expressed proteins in 2 underground miner groups in Metascape. (a) Heatmap of enriched terms across input gene lists, colored by P-values; (b) Net graph of enriched terms, Different colors represent different cluster ID. (c) PPI network and MCODE components identified in the protein lists. Note: PPI, protein-protein interaction; MCODE, molecular complex detection. GO Category Enrichment Results. Note: GO, gene ontology. Enrichment analysis of co-differentially expressed proteins related disease in Metascape. Top 20 Enriched Protein Patway-Related Diseases.


Validation of Differentially Expressed Proteins in Serum Using ELISA
To validate the iTRAQ results, ELISA was used to measure 3 candidate proteins: GCN1, IGHV1-24, and CIP2A in the serum of 116 former uranium miners. Figure 5 illustrates that GCN1 and CIP2A levels were significantly increased in underground miners (U-M) compared to aboveground miners (A-M). There were no significant differences in serum IGHV1-24 levels between underground and aboveground miners. Thus, GCN1 and CIP2A levels were consistent with the iTRAQ results. Expression of GCN1, IgHV1-24, and CIP2A in serum of miners.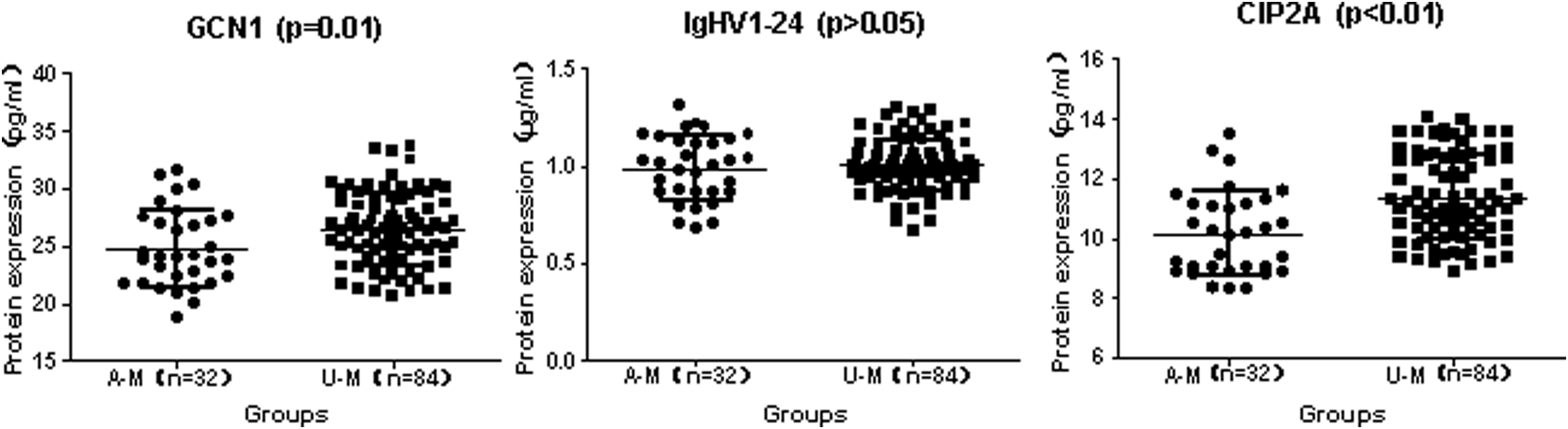
Discussion
Uranium minerals are associated with radioactive elements such as radium and radon in the ore which arises from the radioactive decay. There are mainly 3 parts of total effective dose of uranium miners which contains radon exposure, gamma radiation, and long-lived alpha emitters. Uranium underground miners are primarily exposed to internal ionizing radiation from radon decay products especially α-emitters. Radon is a naturally occurring radioactive noble gas that contributes the largest single fraction of radiation exposure from natural sources. Underground uranium miners are primarily exposed to ionizing radiation internally through inhalation of radon decay products, especially α-emitters such as radon-222 (222Rn), polonium-218 (218Po), plumbum-214 (214Pb), bismuth-214(214Bi), and polonium-214 (214Po), along with dust with high contents of crystalline silica.7,8 Upon inhalation, radon progeny in a conjugated state are mainly deposited in the lung, whereas most of its unconjugated forms are deposited in the nasal pharynx and trachea bronchus. Due to good liposolubility, radon and its progeny easily reach the lung and are absorbed into the blood. Thus, lung and blood cells become the main targets of irradiation from radon exposure. 9 Animal and cell studies, clinical trials, and epidemiological investigations have confirmed that long-term exposure to high levels of radon and its progeny can initiate biological effects at the molecular and cellular levels and pose potential health risks to the exposed populations. 10 Radon has already been defined as a lung carcinogen by the International Agency for Research on Cancer (IARC) based on acohort of radon-exposed miners from 1988. 11 Primary prevention is fundamental in lung cancer, which is characterized by a long latency period. Therefore, it is necessary to explore early warning molecular biomarkers for radon-induced lung cancer.
Serum is an attractive sample for identifying potential protein biomarkers because of its easy accessibility. Moreover, it is an archive of information on physiological and pathological conditions. 12 Compared with conventional techniques for detection of specific proteins, proteomics technology separates and identifies proteins with high throughput and high sensitivity. It has low detection limits and aids discovery of new disease-related proteins. 13 Thus, in the present study, we evaluated the changes in a diverse panel of potential biomarkers in the serums of former uranium miners after long-term radon exposure using iTRAQ technology.
The uranium miners were divided into 2 groups according to the cumulative radon exposure dose, and the mixed serum samples from the 2 groups were detected using iTRAQ. A total of 25 co-differentially expressed proteins were screened, including CIP2A, GCN1, and ANKK1 among others. Biological process analysis of these proteins using Metascape showed that 5 GO terms were enriched, namely, negative regulation of very-low-density lipoprotein particle clearance, endocytosis, regulated exocytosis, tissue homeostasis, and cell morphogenesis involved in differentiation. Many proteins may be biomarkers of some diseases, and most of them are closely related to lipoprotein metabolism. For example, CIP2A functions as an oncoprotein by directly binding to the tumor suppressor PP2AB56a, which is overexpressed in a variety of human cancers.14-16 Anthrax toxin receptor 1 (ANTXR1) is highly upregulated in the tumor endothelium and expressed in diverse types of cancer. Tissue overexpression and elevated serum ANTXR1 levels play pivotal roles in the development and progression of colorectal cancer (CRC). Thus, detection of serum ANTXR1 may serve as a diagnostic biomarker and clinical predictor of CRC. 17 Meanwhile, high ANTXR1 expression was strongly correlated with stromal and immune infiltration in gastric cancer. 18 APOC1, APOC2, APOC3, Orm1, and Orm2 are associated with lipoprotein metabolism. APOC1, APOC2, and APOC3 have distinct effects on the major metabolic pathways involved in lipoprotein metabolism and play an important role in the etiology of human hyperlipidemia. 19 Orm1 and Orm2 are yeast members of a conserved family of endoplasmic reticulum membrane proteins whose physiologic significance is emphasized by the linkage of a human family member to asthma susceptibility, which plays a central role in lipid homeostasis and protein quality control.20,21
Lipids are widely distributed in cellular organelles and are critical components of all membranes. In addition to their role as structural components, lipids in membranes also serve important functions in different organelles. Dysregulation of lipid metabolism contributes to the progression of various metabolic diseases, including cardiovascular diseases, obesity, hepatic steatosis, and diabetes. 22 Apolipoproteins are structural components of lipoproteins and play a critical role in lipid transport. Genetic mutations in apolipoproteins are widely associated with metabolic and cardiovascular diseases. Increasing evidence suggests that apolipoproteins may also be related to the development and outcomes of certain cancers, such as ovarian, renal, and bladder cancer. 23 For example, the expression of apolipoprotein ApoA-1 and ApoC-III in small cell lung cancer (SCLC) was significantly different from that in normal lung tissue, suggesting that ApoA-1 and ApoC-III may be used as differentiating and predictive markers for SCLC. Based on the results of the proteomic tests, differentially expressed serum proteins related to lipid metabolism, such as APOC1, APOC2, APOC3, Orm1, and Orm2, may reflect the harmful effects of long-term radon exposure.
Among the 25 co-differentially expressed proteins, we selected GCN1, CIP2A, and IGHV1-24 for further verification. We selected these 3 proteins because they had a high degree of differential expression in the serum of uranium miners. The results showed that GCN1 and CIP2A levels were consistent with the iTRAQ results, indicating that the expression levels of GCN1 and CIP2A in serums were related to radon exposure, and the expression of IGHV1-24 in serum was unstable, which may be affected by many factors.
Conclusions
In conclusion, ANTXR1, GCN1, CIP2A, and differentially expressed proteins related to lipid metabolism (APOC1, APOC2, APOC3, ORM1, and ORM2) may be used as early markers of the harmful effects after long-term radon exposure.
Footnotes
Acknowledgments
The authors thank all the researchers for this study.
Author Contributions
Conceptualization, D.X. and L.H.; Methodology, Y.Y. and Y.B.; Sample Collection, Z.Z., D.J., and L.X.; Formal Analysis, Y.Y. and Y.B.; Investigation, Y.Y. and Y.B.; Writing – Original Draft Preparation, D.X. and L.H.; Writing – Review & Editing, L. Y and C.D.; Supervision, Z.Y.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by China Institute for Radiation Protection.
