Abstract
High-dose ionizing radiation (IR) alters the expression levels of non-coding RNAs (ncRNAs). However, the roles of ncRNAs and mRNAs in mediating radiation protection by radioprotectants remain unknown. Microarrays were used to determine microRNA (miRNA), long ncRNA (lncRNA), and mRNA expression profiles in the bone marrow of irradiated mice pretreated with amifostine, CBLB502, and nilestriol. Differentially expressed mRNAs were functionally annotated by Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway analyses. Some histone cluster genes were validated by real-time PCR, and the effects of radioprotectant combinations were monitored by survival analysis. We found that these radioprotectants increased the induction of lncRNAs and mRNAs. miRNA, lncRNA, and mRNA expression patterns were similar with amifostine and CBLB502, but not nilestriol. The radioprotectants exhibited mostly opposite effects against IR-induced miRNAs, lncRNAs, and mRNAs while inducing a common histone gene downregulation following IR, mainly via nucleosome assembly and related signaling pathways. Notably, the effects of nilestriol significantly complemented those of amisfostine or CBLB502; low-dose drug combinations resulted in better radioprotective effects in pretreated mice. Thus, we present histone gene downregulation by radioprotectants, together with the biological functions of miRNA, lncRNA, and mRNA, to explain the mechanism underlying radioprotection.
Introduction
Exposure to high-dose ionizing radiation (IR) could lead to life-threatening injuries, primarily to the more radiosensitive, self-renewing tissues. IR primarily affects hematopoietic and gastrointestinal systems, resulting in acute radiation syndrome. The pre-administration of radioprotectants and radiomitigators, such as amifostine, CBLB502, and nilestriol reportedly protect against IR damage and promote recovery. 1,2
Although the underlying protective mechanism of these radioprotectants against high-dose IR remains unclear, several biological and molecular mechanisms have been suggested. 2,3 Amifostine (S-2[3-aminopropylamino]-ethylphosphorothioic acid; WR-2721) is a thiol compound whose active metabolite is a free radical scavenger, which regulates cell cycle checkpoints and gene expression while facilitating DNA repair. 4 CBLB502, a truncated derivative of the Salmonella flagellin protein, shows high radioprotective efficacy by binding to Toll-like receptor 5 of target cells and activating nuclear factor-κB signaling, which modulates the expression of numerous genes, including apoptosis inhibitors, scavengers of reactive oxygen species, and a spectrum of cytokines. 5 -7 Nilestriol, an estrogen derivative, is a long-acting radiomitigator that acts systemically by accelerating the post-irradiation restoration of the radiosensitive tissues by activating pro-inflammatory signaling pathways and stimulating hematopoietic system regeneration. 3,8,9 The estrogenic activity of nilestriol is 230-fold and 510-fold greater than that of estriol when used intravenously and orally, respectively. 10
Nevertheless, the complex mechanisms underlying the altered biological processes after irradiation and regulation by radioprotectants are not completely understood. In addition to the mRNAs, increasing evidence has pointed to significant changes in the expression levels of non-coding RNAs, such as microRNAs (miRNAs) or long non-coding RNAs (lncRNAs) upon IR and their critical roles in the cellular response to IR.
MiRNAs are endogenous small non-coding RNAs (21–23 nucleotides) that act as post-transcriptional modulators of gene expression by either inhibiting mRNA translation or inducing mRNA degradation. 11 A single miRNA can influence the expression of up to 1,000 genes and influence various cellular processes, such as cell differentiation, proliferation, and apoptosis. 12 -15 IR can alter miRNA expression both in vivo and in vitro, depending on cell type, radiation dose, and exposure time. 16 -21 Although several studies have suggested that miRNAs regulate the DNA damage response (DDR) at the transcriptional level 22,23 in a p53-dependent manner, 24,25 little is known about how miRNAs respond to IR.
Recently, lncRNAs have gained attention and been related to multiple cellular processes, including DDR. 26,27 Several lncRNAs play key roles in p53-dependent DDR and immune regulation. 28 -30 Besides other lncRNAs reportedly regulate apoptosis and the cell cycle after DNA damage induction, DNA repair pathways, and DDR protein modification. 31 Despite significant progress in understanding the roles of lncRNAs in the last few years, their specific functions and regulation networks are unclear.
Thus, we aimed to compare the effects of radioprotectants (amifostine, CBLB502, and nilestriol) on whole-genome expression profiles of miRNAs, lncRNAs, and mRNAs in the bone marrow of mice after whole-body high-dose irradiation, using microarray analysis. Functional and pathway analyses of the differentially expressed RNAs before and after IR and between treatment groups were performed to determine the potential molecular mechanisms contributing to their radioprotective effects. Our findings provide fundamental insights into the mechanisms of radioprotection while highlighting the roles of non-coding and coding RNAs as potential targets for the treatment and prevention of IR-induced damage.
Material and Methods
Mice and Irradiation
Male C57BL/6 mice (6–8 weeks old) were purchased from SPF (Beijing) Biotechnology Co. Ltd. (Beijing, China) and randomly divided into 8 groups: phosphate-buffered saline (PBS) control, IR-only, radioprotectant-only (amifostine, CBLB502, or nilestriol), and radioprotectant + IR (3 mice per group). Animals were irradiated using a 60Co γ-ray radiation source at the Beijing Institute of Radiation Medicine (Beijing, China) with a total dose of 8.0 Gy at a dose rate of 1.29 Gy/min. All animal experiments were approved by the Institutional Animal Care and Use Committee of Academy of Military Medical Sciences, Beijing, China (permit number: IACUC of AMMS-13-2015-011).
Drug Administration
Nilestriol (Beijing Institute of Radiation Medicine) was dissolved in PBS and administered orally at 10 mg/kg at 48 h pre-irradiation. Amifostine (Dalian Merro Pharmaceutical Factory, Dalian, China) and CBLB502 (Beijing Institute of Radiation Medicine) were dissolved in PBS and administered intraperitoneally at 0.5 h pre-irradiation at 150 and 0.2 mg/kg, respectively. The PBS was administered to the control animals.
RNA Isolation and Microarray Analysis
At 12 h post-irradiation, bone marrow cells were collected and pooled from 3 animals per group and single-cell suspensions were prepared. RNA isolation and microarray analysis were performed as described previously. 32 For miRNA analysis, the miRNA was labeled with the miRNA Complete Labeling and Hyb Kit (Agilent Technologies) and hybridized to the Agilent Mouse miRNA microarray (8 × 60 K) V19.0 (Agilent Technologies). For lncRNA and mRNA analyses, total RNA was amplified and labeled with a Low Input Quick Amp Labeling Kit (Agilent Technologies) and then hybridized to the Agilent Mouse lncRNA 4 × 180 K microarray (Agilent Technologies). Raw data were normalized by the Quantile algorithm Gene Spring Software 11.0 (Agilent Technologies).
Data Processing
Data processing was performed using the SBC Analysis System from Shanghai Biotechnology Corporation (Shanghai, China). miRNAs showing a fold-change (FC) > 2.0 and lncRNAs or mRNAs showing an FC > 3.0 between IR and/or radioprotectant-treated and PBS-treated samples were considered significantly differentially expressed. The intersecting set of miRNAs, lncRNAs, and mRNAs between the different groups was analyzed using Venny 2.1.0 (http://bioinfogp.cnb.csic.es/tools/venny/). The microarray data were deposited in the NCBI Gene Expression Omnibus (GEO) database (accession no. GSE137013).
Functional Annotation
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses of differentially expressed mRNAs were conducted using the Database for Annotation, Visualization, and Integrated Discovery (DAVID) webtool. 33,34 Pathway enrichment and biological process analyses with GO-SLIM annotation for miRNAs were conducted in the radioprotectant treatment groups using DIANA Tools mirPath V3 (http://diana.imis.athena-innovation.gr). 35 The GO terms and KEGG pathways with p values of <0.05 in the amifostine group were considered as significantly enriched compared to all radioprotectant groups.
Construction of the Protein-Protein Interaction Network
The protein-protein interaction (PPI) network for the treatment groups was constructed using STRING (http://string-db.org). 36 The representative subnetwork containing nodes with high levels of interconnection was further derived from the PPI network at a confidence score of 0.900. The network graphs were produced using Cytoscape 3.7.1 (The Cytoscape Consortium, San Diego, CA, USA). 37
Real-Time PCR Analysis
Real-time PCR for histone cluster genes was performed on the Stratagene Mx3000P (Agilent) using the TB Green Premix Ex TaqTM kit (Takara, Shiga, Japan) as previously described. 21 The primers sequences are listed in Supplementary Table 1. The endogenous glyceraldehyde 3-phosphate dehydrogenase (Gapdh) gene was used as an internal control. Fold differences for each of the genes were calculated using the 2−ΔΔCT method. 38
Survival and Body Weight Analysis Following Radioprotectant Combination
Mice (30 per group) were administered 75 mg/kg amifostine and 0.02 mg/kg CBLB502 5 intraperitoneally at 0.5 h pre-irradiation, or 5 mg/kg nilestriol orally at 48 h pre-irradiation, or a combination of amifostine + nilestriol or CBLB502 + nilestriol. Mice were irradiated and survival time was recorded daily for 30 days, and body weights of surviving mice were recorded on day 30.
Statistical Analysis
Data are presented as means ± SEM. The statistical analyses were performed using Prism 7 software (GraphPad, San Diego, CA, USA). Real-time PCR results were analyzed by 1-way analysis of variance (ANOVA) followed by the Dunnett post hoc test. The Kaplan–Meier survival curves were compared using the log-rank test. The body weights were compared using 1-tailed Student’s t-test. p < 0.05 was considered significant.
Results
Radioprotectants Induce Numerous Differentially Expressed RNAs After Irradiation
As shown in Table 1, the numbers of differentially expressed lncRNAs and mRNAs increased by 2.42–2.77- and 2.37–3.66-folds, respectively, in irradiated mice pre-administered amifostine, CBLB502, and nilestriol, compared to the numbers in radioprotectant-only groups. However, there were no differences in the numbers of differentially expressed miRNAs between the radioprotectant-only groups and radioprotectant + IR groups (Table 1). To understand the mechanism of radioprotectant-induced gene expression, we focused on RNAs commonly regulated by the drugs. We found that the intersecting sets of RNAs regulated by all 3 radioprotectants were increased by 20.66- and 40.00-fold in the numbers of altered lncRNAs and mRNAs in the radioprotectant + IR groups compared to those in the radioprotectant-only groups, respectively. However, there was only a 1.08-fold increase in the numbers of miRNAs altered in the radioprotectant + IR groups (Figure 1). These results support the essential roles of these commonly regulated radioprotectant-induced RNAs after irradiation.
Numbers of Differentially Expressed miRNAs, lncRNAs, and mRNAs in Mice Pretreated With Radioprotectant-Only (amifostine, CBLB502, and nilestriol), or Pretreated With Radioprotectant Before Ionizing Radiation (radioprotectant + IR).

Fold-changes were set to 2.0 (miRNAs) or 3.0 (lncRNAs and mRNAs).

Venn diagram showing the common miRNAs, lncRNAs, and mRNAs induced by radioprotectant-only and pre-treatment with radioprotectants before ionizing radiation (radioprotectant + IR). The fold-change of miRNAs was set to 2.0; the fold-changes of lncRNAs and mRNAs were set to 3.0.
Amifostine and CBLB502 Treatments Result in Similar RNA Profiles
Overall, there was a similar expression pattern of altered RNAs in mice pretreated with amifostine and CBLB502, regardless of irradiation exposure, which differed from the pattern of change in RNAs induced by nilestriol (Figure 2). Specifically, most altered miRNAs, lncRNAs, and mRNAs were either co-upregulated or co-downregulated (71, 3606, and 1051, respectively) between the CBLB502 and amifostine groups, with only a few showing opposite trends of change (1.39%, 0.36%, and 0.76%, respectively) (Table 2). In contrast, although there were several co-regulated miRNAs, lncRNAs, and mRNAs between the amifostine and nilestriol groups (46, 1107, and 591, respectively), higher percentages of altered RNAs were found in the opposite regulatory direction (9.80%, 16.45%, and 4.68%, respectively), indicating the different regulatory patterns between these 2 radioprotectants. Similarly, 14.89%, 17.44%, and 5.42% of the respective RNAs in the opposite regulatory direction showed different patterns between the CBLB502 and nilestriol groups (Table 2). Taken together, these results suggested that amifostine and CBLB502 have a more similar regulation mechanism, which is distinct from that of nilestriol, during radioprotection.

Hierarchical clustering of differentially expressed miRNAs, lncRNAs, and mRNAs in mouse bone marrow. Mice were pretreated with amifostine, CBLB502, and nilestriol or PBS, followed by whole-body irradiation at 8.0 Gy or non-irradiated sham treatment. Bone marrow was removed from the mice 12 h after radiation for miRNA and RNA microarray analysis.
Comparison of Regulation of Differentially Expressed miRNAs, lncRNAs, and mRNAs Between Any 2 Radioprotectant Pretreatments Before IR.

Opposite regulation means upregulation in 1 group and downregulation in another group or vice versa. Fold-changes were set to 2.0 (miRNAs) or 3.0 (lncRNAs and mRNAs).
RNA Protection Against Irradiation Occurs During Radioprotection But Not Before Irradiation
To further investigate the role of the radioprotectants in IR-induced damage, we comparatively analyzed the differentially expressed RNAs between radioprotectant + IR and the IR-only groups (radioprotectant + IR/IR, i.e. FC of RNA expression in the radioprotectant + IR group vs. that in the IR-only group > 2). Consequently, we identified the effects on RNA expression that were triggered by radioprotectant treatments but excluded those effects induced by IR. Then we analyzed the regulatory direction of the differentially expressed RNAs in radioprotectant + IR/IR and radioprotectant/PBS groups against that of differentially expressed RNAs in the IR-only group to reveal the effects of pre-treatment with radioprotectants on IR-induced RNAs in non-irradiated mice and irradiated mice without IR-induced RNA changes. As shown in Figure 3 and Supplementary Table 2, most differentially expressed RNAs in the radioprotectant/PBS groups showed the same regulatory direction in the IR-only group (i.e., IR-induced RNAs and radioprotectant-induced RNAs were either co-upregulated or co-downregulated), but few RNAs showed the opposite regulatory direction (i.e., upregulated in IR-only group but downregulated in radioprotectant-only group or vice versa). Specifically, there were 21/6, 21/7, and 23/9 miRNAs; 830/24, 868/18, and 146/79 lncRNAs; and 165/15, 206/10, and 55/28 mRNAs showing the same/opposite direction of regulation upon induction by amifostine, CBLB502 and nilestriol alone, respectively (Supplementary Table 2). However, many more differentially expressed RNAs showed opposite regulatory direction against IR-only treatment in the radioprotectant + IR/IR groups as compared to those in radioprotectant/PBS groups. Specifically, 1/35, 2/37, and 5/7 miRNAs; 12/2187, 28/2279, and 60/257 lncRNAs; and 26/356, 42/367, and 39/97 mRNAs showed the same/opposite regulatory direction by amifostine, CBLB502 and nilestriol, respectively, in irradiated mice (Supplementary Table 2). These results suggested that miRNAs, lncRNAs, and mRNAs might protect against IR-induced damage by inverting the expression profile of IR-induced RNAs during radioprotection but not before irradiation.

Differential regulation of IR-induced RNAs by radioprotectant-only and radioprotectant + IR treatment. First, the differentially expressed RNAs in the radioprotectant-only group were compared to those in the PBS group. The overlapping RNAs that were differentially expressed in the same or the opposite regulatory directions were assessed between the radioprotectant-only/PBS and IR-only groups. Similarly, the differentially expressed RNAs in the radioprotectant + IR group were compared to those in the IR-only group. The numbers of RNAs for each differential regulatory direction were assessed by overlapping the differentially expressed RNAs between the radioprotectant + IR/IR and IR-only groups. Fold-changes were set to 2.0 (miRNAs) and 3.0 (lncRNAs and mRNAs). Bars represent the sum of the gene count values shown in Supplementary Table 2.
Radioprotectants Commonly Downregulate the Expression of Histone Genes
To investigate the global mechanism during radioprotection, we next focused on the common RNAs regulated by the 3 radioprotectants. The intersecting set of differentially expressed miRNAs in radioprotectant + IR groups was functionally annotated and shown in Supplementary Table 3 and Table 4. The radioprotectants induced miRNAs related to similar KEGG pathways and GO functions with variable gene counts and p-values. Meanwhile, the GO and KEGG pathway analysis of downregulated mRNAs (differentially expressed genes, DEGs) in the radioprotectant + IR groups, demonstrated the involvement of similar biological processes, including nucleosome assembly, DNA methylation of cytosine, DNA replication-dependent and DNA replication-independent nucleosome assembly, and DNA-templated initiation of transcription (Figure 4; see the full list in the Supplementary Files).
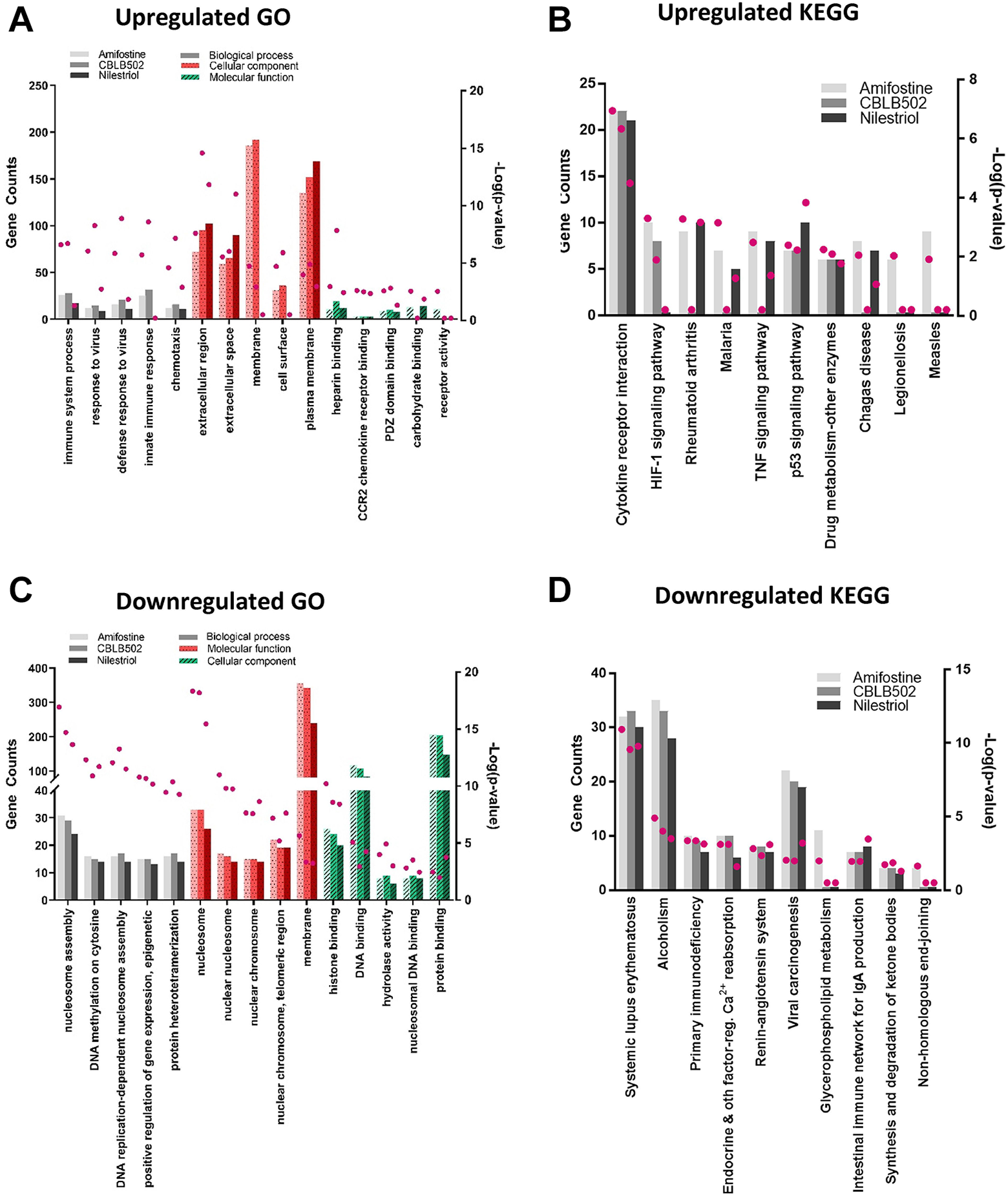
GO enrichment and KEGG pathway analysis of DEGs in radioprotectant treatment. Gene Ontology (GO) enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses of upregulated DEGs (A, B) and downregulated DEGs (C, D) associated with pretreatment with radioprotectants in irradiated mice. The gene counts (histogram) and –log (p-value) (pink dot) were set to 0 when the GO terms or KEGG pathways were not in the list of GO or KEGG enrichment results. Light color: amifostine, medium color: CBLB502, dark color: nilestriol.
PPI network analysis revealed a prominent subnetwork mainly consisting of histone and related genes in the intersecting set of DEGs downregulated by the radioprotectants, but not in radioprotectant-upregulated DEGs (Supplementary Figure 1 and Figure 2). To confirm this, we selected some GO terms enriched from the intersecting set of downregulated DEGs by radioprotectants to examine their relationship with histones related genes. However, for IR-only-induced DEGs, only 9 histones and related genes were identified among the downregulated DEGs related to of nucleosome assembly, which was fewer than that observed for the radioprotectant + IR-induced downregulated DEGs (Table 3, Supplementary Tables 5 and 6), and there were no histone genes among the upregulated IR-only or radioprotectant + IR-induced DEGs (Supplementary Table 5 and 7). Interestingly, no DEGs in the radioprotectant-only groups were related to the regulation of histone genes (Supplementary Figures 3 -5, Supplementary Table 8). We further verified the downregulation of the expression of selected histone cluster genes for histones H1–H4 in irradiated mice by RT-PCR, whereas some were generally reversed by amifostine and nilestriol but not CBLB502. However, they were downregulated in non-irradiated mice treated with amifostine and CBLB502 but not nilestriol (Figure 5).

Validation of expression of selected histone cluster genes. Total RNA was extracted from the bone marrow of irradiated or non-irradiated mice with or without radioprotectant pretreatment, followed by real-time PCR analysis of Hist1h1a, Hist1h1d, Hist1h2bb, Hist1h2bg, Hist3h2ba, Hist1h3a, Hist2h3b, and Hist2h4 genes. Gapdh was used as an internal control. Con: Control, IR: irradiation. The experiments were performed using 3 mice, and data are presented as the mean ± SEM. * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Selected GO Enrichment Analysis of Histone Cluster Genes in Downregulated DEGs of IR-Only or Radioprotectant + IR Groups.

IR: ionizing radiation.
Fold-changes of DEGs were set to 3.0.
Moreover, all radioprotectant + IR groups showed the activation of commonly-modulated, specific regulatory pathways, such as immune response, cellular response to IL-1, p53-mediated intrinsic apoptotic signaling pathway, and positive regulation of the ERK1 and ERK2 cascade. Other processes and pathways involved in either 1 or 2 of the radioprotectant treatments include IFN signaling, TNF, cAMP biosynthesis process, and SMAD phosphorylation (Table 4).
Selected GO Enrichment Analysis of Radioprotectant + IR Groups.

Fold-changes of DEGs were set to 3.0.
Nilestriol Demonstrates Complementation of Amifostine and CBLB502 Effects
Either amifostine or CBLB502 and nilestriol showed lower overlap for differentially expressed RNAs having the same regulatory direction and an increase in the overlap for differentially expressed RNAs having opposite regulatory directions, indicating that these radioprotectants induced different radioprotective mechanisms involving some unique pathways (Figure 6A, Supplementary Table 9). Therefore, we examined the potential complementary effects of combining radioprotectants. Although pre-treatment of irradiated mice with a low-dose combination of radioprotectants offered limited survival benefits, the combination of low-dose amifostine and nilestriol, or CBLB502 and nilestriol showed higher radioprotective effects than a single administration of corresponding radioprotectants (all p < 0.05) (Figure 6B). Moreover, on day 30, the body weights of irradiated mice pretreated with combinations of radioprotectants showed an increasing trend. Most differences were not significant, except the comparison of groups of CBLB502 vs. CBLB502 + nilestriol (p = 0.026) (Figure 6C). Thus, the lower overlap of differentially expressed RNA profiles suggested a complementary effect between radioprotectants.

Complementary effects of combined radioprotectants during radioprotection. (A) KEGG analysis of DEGs in irradiated mice exclusively induced by radioprotectant-only in the overlap of amifostine + CBLB502 (left), amifostine + nilestriol (middle), and CBLB502 + nilestriol (right). Top 5 KEGG items are shown. (B) Kaplan–Meier survival curve of C57BL/6 irradiated mice pretreated with 75 mg/kg amifostine, 0.02 mg/kg CBLB502, 5 mg/kg nilestriol, combination of amifostine + nilestriol, or combination of CBLB502 + nilestriol. Statistical analysis of survival was performed by the log-rank test (n = 30 per group, # p < 0.05 for amifostine vs. amifostine + nilestriol; * p < 0.05 for nilestriol vs. amifostine + nilestriol; *** p < 0.001 for nilestriol vs. CBLB502 + nilestriol, † p < 0.05 for CBLB502 vs. CBLB502 + nilestriol). (C) Body weights of surviving mice on day 30. Statistical analysis was performed by 1-tailed Student’s t-test; p values are indicated.
Discussion
The expression profiles of non-coding RNAs and coding RNAs have been described in irradiating human cells at low and high doses of IR 20,29 ; however, the changes in the RNA levels response to radioprotectant treatment upon IR were unclear. In this study, we showed that pre-treatment with different radioprotectants induced the differential expression of a more extensive set of miRNAs, lncRNAs, and mRNAs in the mouse bone marrow after exposure to high-dose γ-rays. The amifostine and CBLB502 pre-treatments resulted in similar patterns of differentially expressed miRNAs, lncRNAs, and mRNAs, which differed from those induced with nilestriol, in both IR and non-IR exposed mice. Further analysis of the intersecting set of differentially expressed mRNAs suggested that radioprotectants downregulate histone gene expression and the nucleosome assembly process under IR stimulation. Importantly, RNAs differentially expressed in irradiated mice pretreated with radioprotectants showed opposite regulatory direction to those of RNAs differentially expressed in mice in the IR-only group. Pre-treatment of irradiated mice with a low-dose of either amifostine and nilestroil or CBLB502 and nilestriol produced a better radioprotective effects.
Although the mechanisms of amifostine and CBLB502-mediated radioprotection are poorly understood, 39 they are both likely to have a cytoprotective effect and rapid response against IR-induced stress within several hours, 5,40 with achieving complex radiation-cytoprotective effects by interfering with genes involved in scavenging free radicals, cell cycle regulation, apoptosis, and inflammation. 6,7,41 -46 However, nilestriol requires a longer time, usually hours or days, to exert its protective effects, and its post-irradiation mechanism does not involve primary radiation-induced cellular processes. 3,9 These differences in reaction time and mechanism may explain the similarities between amifostine and CBLB502 in terms of the influence on IR-induced RNAs, which was distinct from that of nilestriol. Nevertheless, the high similarity in the RNA profile of amifostine and CBLB502 groups provide insights into possible shared regulatory mechanisms of radioprotection.
Interestingly, our results also showed that radioprotectants mostly exert their radioprotective effects after irradiation but not at the pre-irradiation stage. The radioprotectants, particularly amifostine and CBLB502, induced opposite expression of RNAs to that induced by IR mostly in irradiated mice, but rarely in non-irradiated mice. Several reports have shown conflicting results regarding the effects of radioprotectants at the pre-irradiation stage. For instance, CBLB502 pre-treatment in non-irradiated mice increased IL-6 and G-CSF levels, which are pivotal to radioprotection 5,6 ; however, pre-administration of amifostine significantly reduced radiation-induced increases in the excretion rates of prostaglandins and thromboxane in irradiated rats, but not in non-irradiated rats. 47 Thus, our results provide evidence at the genome-wide RNA expression level that some radioprotectants might not show prominent effects if administered before irradiation.
Notably, we found that IR reduces the expression levels of several histone cluster genes involved in nucleosome assembly, which is consistent with previous reports showing that IR downregulated histone expression in parallel with inhibition of DNA synthesis. 48,49 IR exposure induces highly lethal DNA damage resulting in local and global decondensation of chromatin, 50,51 which resulted in chromatin remodeling, including nucleosome disruption and histone eviction at double-strand break sites, which may facilitate DDR. 52,53 However, based on our results, pre-treatment with radioprotectants before IR may further downregulate histone cluster genes, including most histone proteins involved in nucleosome assembly, DNA methylation, and a variety of similar biological processes. This finding indicates that a rapid response of radioprotectants upon IR facilitates subsequent nucleosome remodeling inducing DDR and DNA repair. Our findings provide novel evidence that the downregulation of histone cluster genes is the primary biological response to IR-induced damage and that radioprotectants regulate this response following exposure regardless of their specific radioprotective mechanisms.
Finally, our data showed that differentially expressed RNA profiles of amifostine and CBLB502 had a lower overlap with those of nilestriol, indicating that they have different radioprotective mechanisms, which potentially exert complementary effects. Therefore, the unique RNAs and pathways induced by amifostine or CBLB502 might be complementary to those induced by nilestriol and vice versa. Nonetheless, extensive investigations are required to clarify the differences in their mechanisms.
Conclusion
Our comparison of the common miRNAs, lncRNAs, and mRNAs with altered expression in radioprotectant-pretreated mice with or without IR revealed different patterns between amifostine, CBLB502, and nilestriol. Pathway enrichment and biological process analyses of the genes commonly altered by all radioprotectants confirmed the involvement of these genes in nucleosome assembly and their antagonistic regulation against IR-induced RNAs while demonstrating a complementary radioprotection effect of radioprotectants as a combination. These findings will facilitate future research on the functional genes and regulatory mechanisms that mediate radioprotection.
Supplemental Material
Supplemental Material, Supplementary_figure_1 - RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation
Supplemental Material, Supplementary_figure_1 for RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation by Changhui Ge, Fei Su, Hanjiang Fu, Yuan Wang, Baolei Tian, Bin Liu, Jie Zhu, Yong Ding and Xiaofei Zheng in Dose-Response
Supplemental Material
Supplemental Material, Supplementary_Figure_2 - RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation
Supplemental Material, Supplementary_Figure_2 for RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation by Changhui Ge, Fei Su, Hanjiang Fu, Yuan Wang, Baolei Tian, Bin Liu, Jie Zhu, Yong Ding and Xiaofei Zheng in Dose-Response
Supplemental Material
Supplemental Material, Supplementary_figure_3 - RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation
Supplemental Material, Supplementary_figure_3 for RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation by Changhui Ge, Fei Su, Hanjiang Fu, Yuan Wang, Baolei Tian, Bin Liu, Jie Zhu, Yong Ding and Xiaofei Zheng in Dose-Response
Supplemental Material
Supplemental Material, Supplementary_figure_4 - RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation
Supplemental Material, Supplementary_figure_4 for RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation by Changhui Ge, Fei Su, Hanjiang Fu, Yuan Wang, Baolei Tian, Bin Liu, Jie Zhu, Yong Ding and Xiaofei Zheng in Dose-Response
Supplemental Material
Supplemental Material, Supplementary_Figure_5 - RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation
Supplemental Material, Supplementary_Figure_5 for RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation by Changhui Ge, Fei Su, Hanjiang Fu, Yuan Wang, Baolei Tian, Bin Liu, Jie Zhu, Yong Ding and Xiaofei Zheng in Dose-Response
Supplemental Material
Supplemental Material, Supplementary_Table_1-9 - RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation
Supplemental Material, Supplementary_Table_1-9 for RNA Profiling Reveals a Common Mechanism of Histone Gene Downregulation and Complementary Effects for Radioprotectants in Response to Ionizing Radiation by Changhui Ge, Fei Su, Hanjiang Fu, Yuan Wang, Baolei Tian, Bin Liu, Jie Zhu, Yong Ding and Xiaofei Zheng in Dose-Response
Footnotes
Authors’ Note
CG, XZ and YD conceived and designed the experiments. CG, FS, HF, YW, BT, BL, and JZ performed the experiments and data analysis. CG wrote the manuscript with comments provided by XZ and YD.
Acknowledgments
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Chinese National Natural Science Foundation projects under Grant numbers 91540202 and 81773038, and Wu Jie Ping Medical Fund under Grant number 320.6750.15230, and Key Scientific Research Projects under Grant number BWS18J008.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
