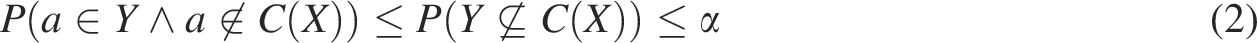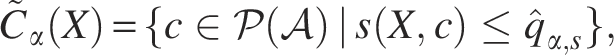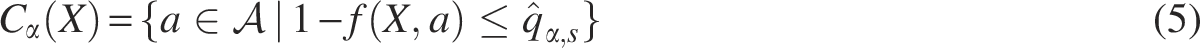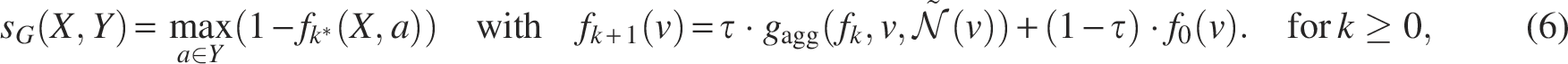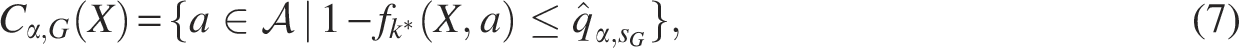Abstract
Bacterial antimicrobial resistance is one of the most pressing global health challenges. Infections with resistant pathogens increase patient morbidity and mortality due to limited treatment options. Rapid and reliable identification of resistance is therefore crucial. However, conventional culture-based diagnostics are slow, typically requiring at least 48 hours from patient sample arrival to result. In contrast, matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry, routinely used for species identification, can provide data 24 hours earlier. Repurposing this technique for antimicrobial resistance prediction has shown promise, but limited predictive performance and a lack of statistically grounded uncertainty estimates have hindered clinical integration. To address these issues, we propose an antimicrobial resistance detection framework using a knowledge-graph-enhanced conformal predictor. Conformal prediction outputs sets of likely effective antibiotics with statistical guarantees, ensuring that resistance detection meets a predefined error rate. Our approach improves upon standard conformal prediction by integrating domain knowledge through a drug- and species-specific knowledge graph that captures interdependencies between antibiotics, such as inferable resistance patterns between broad- and narrow-spectrum agents, as well as co-resistance patterns within antibiotic classes. This predictor is layered on top of a novel classifier that surpasses state-of-the-art models and overcomes key technical limitations of earlier approaches. We evaluated our framework on two clinically relevant species, Klebsiella pneumoniae and Escherichia coli, using the DRIAMS dataset. Our results demonstrate that our conformal predictor consistently achieved the expected coverage guarantees and that the knowledge-graph enhancement significantly reduced false discovery rates compared to standard conformal approaches. By adding statistically grounded uncertainty estimates and improving predictive performance, our framework strengthens the reliability of early antimicrobial resistance predictions from MALDI-TOF data. This could support the clinical integration of such rapid diagnostics by increasing trust in their results and enabling better-informed early treatment decisions.1
Keywords
INTRODUCTION
Antimicrobial resistance is a global health challenge. Every year, almost five million people worldwide die from infections with multidrug-resistant bacteria (Murray et al., 2022). One way to tackle antimicrobial resistance is fast detection in the diagnostic laboratory; the earlier the treating clinician is aware of the resistant bacteria, the earlier the treatment regimen can be adapted. This is particularly important in the case of severe, life-threatening infections, such as sepsis. However, current techniques to detect antimicrobial resistance rely on initial overnight culturing of the patient sample and subsequent culture-based antibiotic susceptibility testing. The resulting turnaround time of at least 48 h poses a substantial challenge in the care of critically ill patients, where immediate diagnostic information is crucial for treatment decisions. In contrast, species identification is available 24 h earlier than resistance testing. This is possible through matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS), which only takes a few minutes to generate characteristic protein mass spectra from bacterial isolates. Recent findings show that these data can be repurposed for detecting antimicrobial resistance (Weis et al., 2022a). In particular, machine learning models have been used to identify patterns in MALDI-TOF spectra that correlate with specific resistance phenotypes.
MALDI-TOF MS-based antimicrobial resistance prediction has therefore shown the potential to deliver results 24 hours earlier than conventional culture-based approaches. Yet, the clinical applicability of this approach has largely been examined retrospectively, and prospective evaluation and clinical deployment face several challenges, including missing uncertainty quantification. The clinical application of a model would require rigorous, statistically valid uncertainty estimates to allow for an informed application of the method. Such uncertainty estimates are particularly important, as models’ performance can vary highly between different health centres, species, and antibiotics (Park et al., 2024; Weis et al., 2022a). However, the models available so far do not provide this kind of uncertainty estimation with statistically valid guarantees, limiting the clinicians’ trust in the models’ outputs.
To close this gap, we propose a conformal predictor for the task of detecting all antibiotics (from a set of clinically relevant antibiotics) to which a bacterial isolate exhibits resistance from its MALDI-TOF spectrum. This predictor provides statistically valid guarantees on the rate of predictions where an antibiotic was not flagged, even though the bacterial isolate does exhibit resistance, also called coverage guarantee. Crucially, this guarantee is valid simultaneously for all relevant antibiotics without the need for multiple comparison corrections.
Our contribution
We start by formalizing the task of detecting antimicrobial resistance by formulating the problem as a multilabel classification task. Then, we enhance a generic conformal predictor for multilabel classification by using a knowledge graph capturing the interdependencies in antibiotic resistance patterns. We further introduce a novel classifier architecture, based on a residual multi-layer perceptron with a graph neural network drug encoder (ResMLP-GNN), for predicting resistance of single antibiotics, which we use as a base classifier for our knowledge-graph-enhanced conformal predictor. See Figure 1 for an illustration of our prediction pipeline.
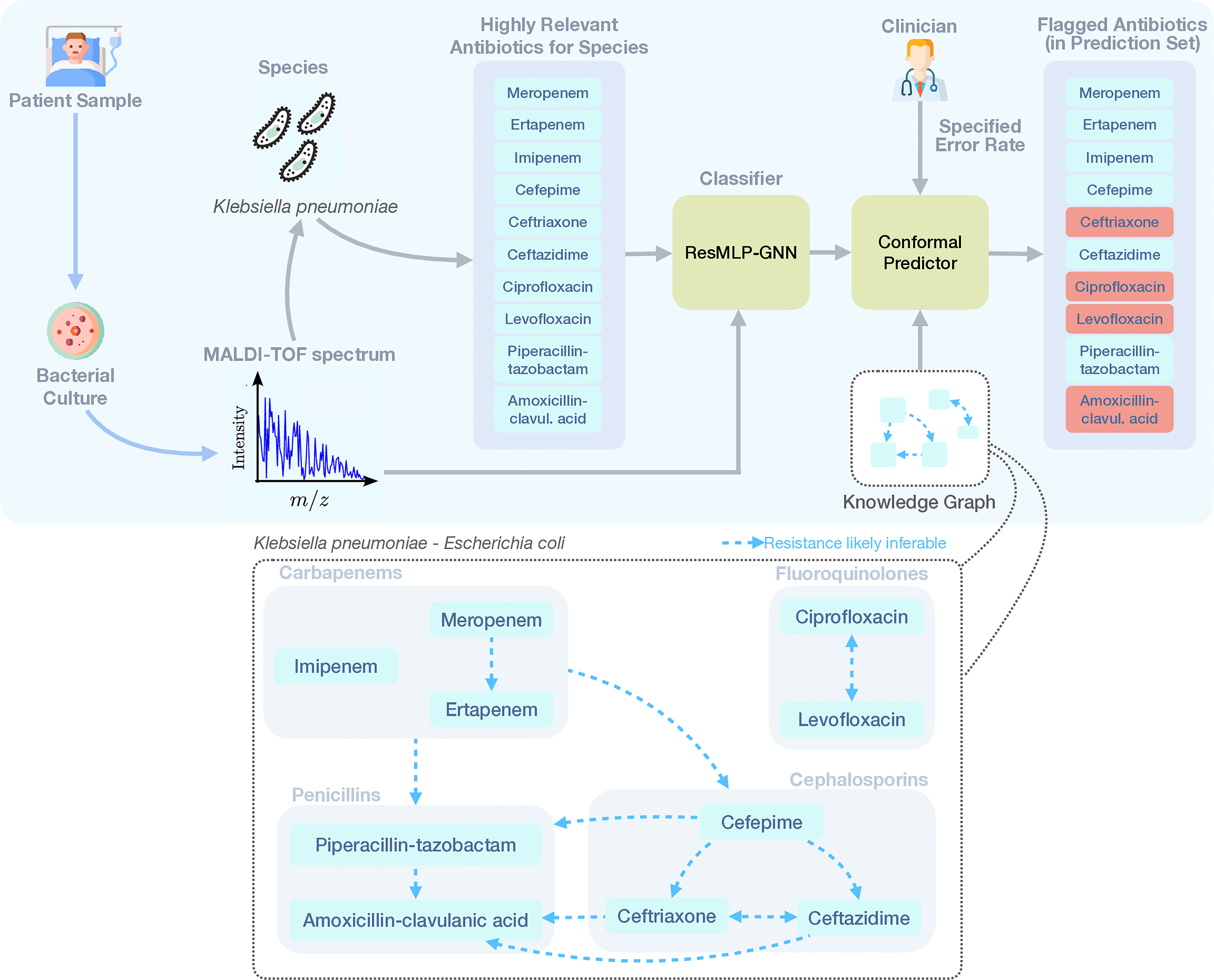
Illustration of our prediction pipeline. Our novel binary classifier, ResMLP-GNN, takes as input the matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) spectrum X and an antibiotic a and returns a score
Using the DRIAMS-A dataset (Weis et al., 2021), we compare the performance of our method with relevant baselines. ResMLP-GNN shows improved performance compared to previous architectures and supports the prediction of highly relevant drug combinations such as amoxicillin-clavulanic acid and piperacillin-tazobactam. Compared to the generic conformal predictor, our knowledge-graph-enhanced counterpart shows superior performance in reducing the false discovery rate associated with a chosen coverage guarantee.
Various subsequent works addressing antimicrobial resistance prediction from MALDI-TOF spectra have emerged following Weis et al. (2022a). Weis et al. (2022b) extended their prediction models by adding hierarchical stratification to the original dataset, which led to significant improvements. De Waele et al. (2023) also used the DRIAMS dataset to create a drug recommender system that showed stable transfer results between different centres. Visonà et al. (2023) proposed incorporating representations of chemical antibiotic structures in the training of a neural network, which could significantly improve the performance of antimicrobial resistance prediction. However, relevant limitations remained as these state-of-the-art multimodal models rely on molecular fingerprints to represent antibiotics, and lack support to date for commonly used and highly relevant antibiotic combinations.
Conformal prediction (Angelopoulos and Bates, 2023; Vovk et al., 2005) is a statistical method that generates prediction sets with a specified level of coverage, ensuring that predictions remain valid for new data under minimal assumptions. By leveraging calibration data, it provides rigorous, adaptable confidence measures that can be recalibrated to maintain validity under distribution shifts (Gibbs and Candes, 2021; Tibshirani et al., 2019). Conformal predictors have been studied in other medical applications, such as drug discovery (Fisch et al., 2022), cancer treatment (Nolte et al., 2024), or sepsis prediction (Yang et al., 2024). They have also been previously applied to antimicrobial resistance prediction using epidemiological patient data for Escherichia coli (Inda-Díaz et al., 2023). However, while previous work only guarantees individual coverage for each drug, we extend this coverage guarantee for all drugs simultaneously by applying conformal prediction on a set of predictions for all species-relevant drugs. To our knowledge, our work is the first to apply conformal prediction in the context of antimicrobial resistance prediction from MALDI-TOF spectra.
METHODS
Machine learning approaches offer promising solutions for predicting antimicrobial resistance patterns, yet their clinical application demands rigorous statistical guarantees of prediction reliability. In this section, we develop a framework that provides such guarantees through conformal prediction, while addressing the challenges of heterogeneous resistance patterns across different antibiotics. Specifically, Section 2.1 introduces antimicrobial resistance prediction as a multilabel classification task and presents a generic conformal prediction framework providing coverage guarantees in this setting. To address the challenge of heterogeneous resistance patterns, Section 2.2 introduces a novel enhancement that leverages domain knowledge encoded in a graph structure. Section 2.3 details the underlying classifier architecture employed within the conformal prediction framework. Finally, Section 2.4 provides a thorough description of experimental procedures and implementation details.
Prediction of antimicrobial resistance with statistical coverage guarantees
Given the MALDI-TOF mass spectrum X of a bacterial isolate, the species identification determines a set of clinically relevant antibiotics
This guarantee extends beyond the true resistant subset Y as a whole. Notably, it implies individual coverage guarantees simultaneous for each antibiotic
To construct such a set, we employ the conformal prediction framework, which utilizes a pre-trained classifier f to provide prediction sets with statistical coverage guarantees. Classifier f outputs a score
The prediction set
The effectiveness of the conformal predictor and the quality of the resulting prediction sets
This choice of score function enables a more computationally efficient construction of the prediction set
While the conformity score function in Eq. (4) offers computational efficiency, it might yield overly large prediction sets. This limitation arises from the significant variation in antimicrobial resistance rates, which leads to the heterogeneous performance of classifier f across different antibiotics: the classifier typically achieves better performance for antibiotics with high resistance rates compared with those with more imbalanced data distributions. This performance heterogeneity affects the conformal predictor in Eq. (5), as the inclusion of antibiotics in
We propose enhancing the confidence scores generated by the above-mentioned classifier by incorporating domain knowledge through a graph structure that captures inherent interdependencies in antimicrobial resistance patterns. These interdependencies arise from well-established microbiological principles: resistance to certain antibiotics can be predicted with high confidence based on resistance to functionally related antibiotics. These can be classified into two key patterns: 1. Hierarchical relationships between narrow- and broad-spectrum antibiotics. Within these hierarchies, resistance to narrow-spectrum antibiotics (e.g., amoxicillin-clavulanic acid) can be inferred from resistance to broad-spectrum antibiotics (e.g., meropenem). 2. Co-resistance patterns, where functionally highly similar antibiotics will exhibit similar resistance phenotypes, e.g., the 3rd generation cephalosporins ceftriaxone and ceftazidime.
To test our framework, we chose Klebsiella pneumoniae as a model organism. Klebsiella pneumoniae is a clinically highly relevant bacterial species with a high propensity to acquire resistance and virulence factors (Sattler et al., 2024). At the same time, it exhibits a high number of the aforementioned antimicrobial resistance interdependencies in the antibiotics relevant for its treatment. We further tested our framework on Escherichia coli, which shares the same resistance interdependencies and is another highly relevant clinical pathogen of the same bacterial order. To create the knowledge graph on the antimicrobial drug interdependencies, we focused on a set of antibiotics that are, in principle, considered in the treatment of serious systemic infections, as testing of antibiotics used in mild forms of infections can be deferred to conventional culture-based methods. See Supplementary Table S1 for details on the construction of the knowledge graph (Gniadkowski, 2001; Drawz and Bonomo, 2010; Kiffer et al., 2006; Poole, 2004) and the lower panel of Figure 1 for an illustration of the knowledge graph.
Knowledge-graph-based conformity score refinement
Given the knowledge graph described above, let
We explore several choices for the aggregation function
For
The message passing mechanism allows the conformity scores to be refined based on the graph structure while maintaining a connection to the original classifier output through
In the following, we present the classifier architecture to serve as the base classifier for our conformal predictor. We compare the performance of our proposed classifier to various state-of-the-art classifiers from related work.
Antibiotic feature extraction
One of the key limitations of previous best-performing approaches was that the antibiotic representations employed to enhance prediction performance did not support combination antibiotics, such as ß-lactam/ß-lactamase inhibitors (Visonà et al., 2023). As some antibiotics included in our knowledge graph consist of multiple drugs, we modified the existing architecture to support learned representations of small molecules represented as attributed graphs, where nodes are atoms, edges are bonds, and attributes correspond to atom and bond types, respectively. To this end, we used a state-of-the-art self-supervised method for small molecule representation learning, called Mole-BERT (Xia et al., 2023), that relies on atom masking and node type prediction. Instead of relying on raw atom types for the masked prediction task, the model learns new class labels for atom types in context, using a VQ-VAE (van den Oord et al., 2018). Here, multiple drugs in the same input are represented as disconnected components in the same graph.
Graph-based representations for all antibiotics were obtained by pre-training Mole-BERT from scratch using a backbone of 5 graph isomorphism network layers (GIN, Xu et al., 2018) and default hyperparameters for 100 epochs on the ZINC-250k dataset (Irwin et al., 2012).
Antimicrobial resistance prediction model architectures
Antimicrobial resistance prediction scores were obtained for all drugs in the defined knowledge graph for both Klebsiella pneumoniae and Escherichia coli, using a set of different models. For starters, we included classical baselines trained on single species-drug combinations without chemical information, as originally presented by Weis et al. (2022a), including logistic regression (LR), gradient boosting machine (GBM), and multi-layer perceptron (MLP). As an additional baseline, we included a multimodal residual multi-layer perceptron that uses chemical structure by means of Morgan fingerprints (ResMLP-FP), as presented by Visonà et al. (2023).
Our novel model architecture, termed ResMLP-GNN, is a modification of ResMLP-FP that learns multigraph-based antibiotic representations using the Mole-BERT method described in the previous section. Here, the 6000-dimensional binned MALDI-TOF spectra are passed through a 1D convolutional and a fully connected layer. Their representations are concatenated with either Morgan fingerprints (ResMLP-FP) or Mole-BERT embeddings (ResMLP-GNN). For ResMLP-GNN, antibiotic representations are fed to a series of GIN and fully connected layers. The concatenated representations are fed to a series of residual blocks, and an antimicrobial resistance score is returned in the end. The model is trained to minimize the binary cross-entropy loss between predictions and ground truth experimental labels. A schematic representation of the model is available in Supplementary Figure S2.
Experimental details
Dataset
For all analyses, we used the publicly available Database of Resistance Information on Antimicrobials and MALDI-TOF Mass Spectra (DRIAMS, Weis et al., 2021), which contains MALDI-TOF mass spectra of clinical bacterial isolates from four Swiss diagnostic centers between 2015 and 2018. Experimental details are given in the original paper (Weis et al., 2022a). Briefly, the spectra were acquired with the Microflex Biotyper System by Bruker Daltonics with standard settings for species identification, which measures the m/z range between 2,000 and 20,000. After standard preprocessing, including baseline correction and normalization, the spectra are binned into intervals of 3 m/z units each, resulting in a consistent representation across all samples. Therefore, each data point includes a 6000-dimensional binned vector representation of a MALDI-TOF mass spectrum from a bacterial isolate, along with annotations for susceptibility or resistance to all tested antibiotics. In our analysis, we used the Klebsiella pneumoniae and Escherichia coli isolates from the DRIAMS-A subset, see Supplementary Tables S2 and S3 for details.
Data splits
Each spectrum in the dataset maps to a single isolate, which is tested in the clinic against a set of several antibiotics, with antimicrobial resistance correlated across multiple drugs. To prevent the model from learning associations between antibiotics that would not be realistic in a clinical setting, we split the data by sample, ensuring that all labels corresponding to the same spectrum are kept together in the same split. Following this logic, 60% of the samples were kept as a global set for classifier training and validation, and 40% were used for performance held-out testing. Due to the highly variable resistance prevalence per drug and species, all splits were stratified on the global resistance labels per species (see Supplementary Table S3 for reference).
Training the classifiers
Hyperparameter tuning on the logistic regression, gradient boosting machine, and multi-layer perceptron baselines was carried out following the same approach as Weis et al. (2022a), using a grid search over a 10-fold cross-validation on the training set, and reporting performance on the test set. The best-performing of all three models per drug was reported as a classical baseline to compare our ResMLP architectures to (see Supplementary Table S4 and S5 for details).
For both ResMLP-FP and ResMLP-GNN, a base model was trained for 500 epochs with early stopping on all available species and drug combinations pooled. The base models were subsequently fine-tuned to each unique species-drug combination present in the knowledge graph for 150 epochs with early stopping. Model weights were averaged over training using stochastic weight averaging (Izmailov et al., 2019). Drug representation heads were frozen during fine-tuning of ResMLP-GNN (as all examples use the same drug).
For the base ResMLP-FP and ResMLP-GNN models, learning rate, dropout, drug embedding size, and the number of residual blocks were tuned to minimize the loss on a randomly selected validation set containing 10% of the training samples (6% of all samples in DRIAMS-A). The best-performing model was selected for each case using a combination of the hyperband algorithm (Li et al., 2018) and Bayesian optimization sampling (Snoek et al., 2012). Hyperparameter tuning for fine-tuned models followed the same approach, albeit only for dropout and learning rate, since the model backbone was fixed by the base architecture.
Evaluating conformal predictors
We used the 40% test split of the data to validate our graph conformal predictor presented in Eq. (7) against the baseline model presented in Eq. (5). Using Monte Carlo cross-validation for 30 iterations, where we randomly split the test data into noisy splits with an average of 25% test and 75% tuning data, we performed hyperparameter search for
RESULTS
Incorporating multigraph-based antibiotic representations improves resistance detection performance for highly relevant species-drug combinations
Our work advances the state-of-the-art in antimicrobial resistance prediction by implementing multigraph-based antibiotic representations instead of traditional molecular fingerprints. This novel approach addresses a critical limitation of previous models, which were restricted to handling single-drug antibiotics. By leveraging multigraph-based representations, we successfully incorporated clinically relevant antibiotic combinations from our knowledge graph, including amoxicillin-clavulanic acid and piperacillin-tazobactam—combinations routinely used to treat infections caused by Klebsiella pneumoniae and Escherichia coli. Notably, the chemical representations learned through Mole-BERT demonstrated sufficient expressiveness to classify antibiotics by functional class in the latent space, maintaining this discriminative power even when processing antibiotic combinations (see Supplementary Fig. S1).
Moreover, our newly developed ResMLP-GNN demonstrated better median performance (in terms of both AUROC and AUPRC) than both classical and multimodal baselines on 8 out of 10 tested drugs for Klebsiella pneumoniae, and 5 out of 8 tested drugs for Escherichia coli. In all cases in which median performance was not the highest, the reported 95% confidence intervals (obtained via stratified bootstrapping) overlapped with the best-performing model. See Figure 2 for the full ROC and PR curves with ResMLP-GNN, and Supplementary Table S4 and S5 for a detailed performance comparison with the appropriate baselines. It is also noteworthy that performance positively correlates with resistance prevalence, which is not uniform across the tested antibiotics (see Supplementary Table S3), further motivating the need for regularization using prior knowledge.
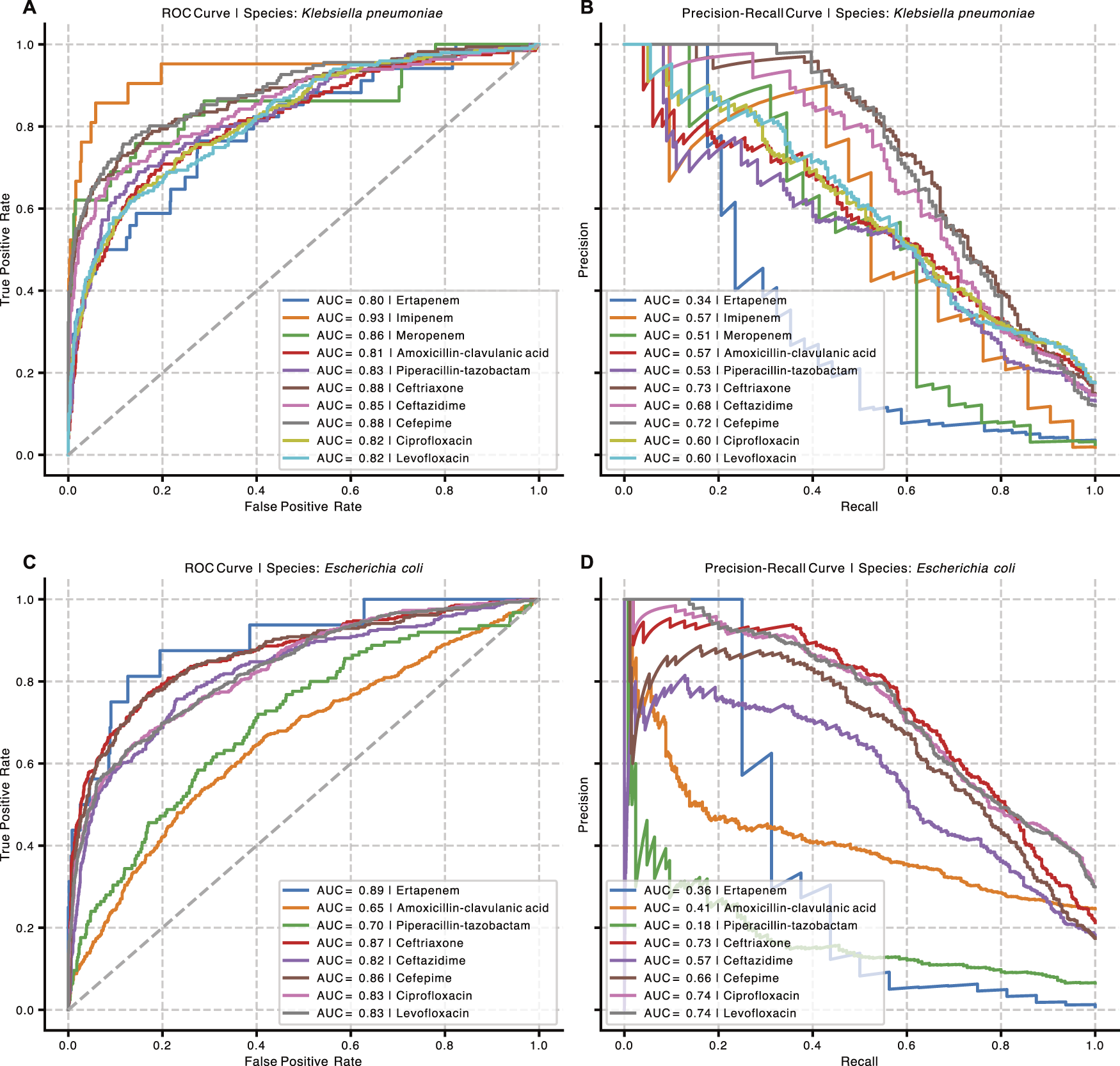
Receiver-Operating Characteristic and Precision-Recall curves showcasing test performance for fine-tuned ResMLP-GNN models on DRIAMS-A, for all selected highly relevant drugs on Klebsiella pneumoniae
To evaluate our knowledge-graph-enhanced conformal predictor against the baseline conformal predictor defined in Eq. (5), we analyzed three key metrics: Empirical Error Rate, False Discovery Rate (FDR), and False Negative Rate (FNR) (detailed definitions available in the Appendix in the section ADDITIONAL METRICS). The results presented in Figure 3 demonstrate that incorporating the knowledge graph substantially reduces the FDR for Klebsiella pneumoniae, with the most pronounced improvement observed at
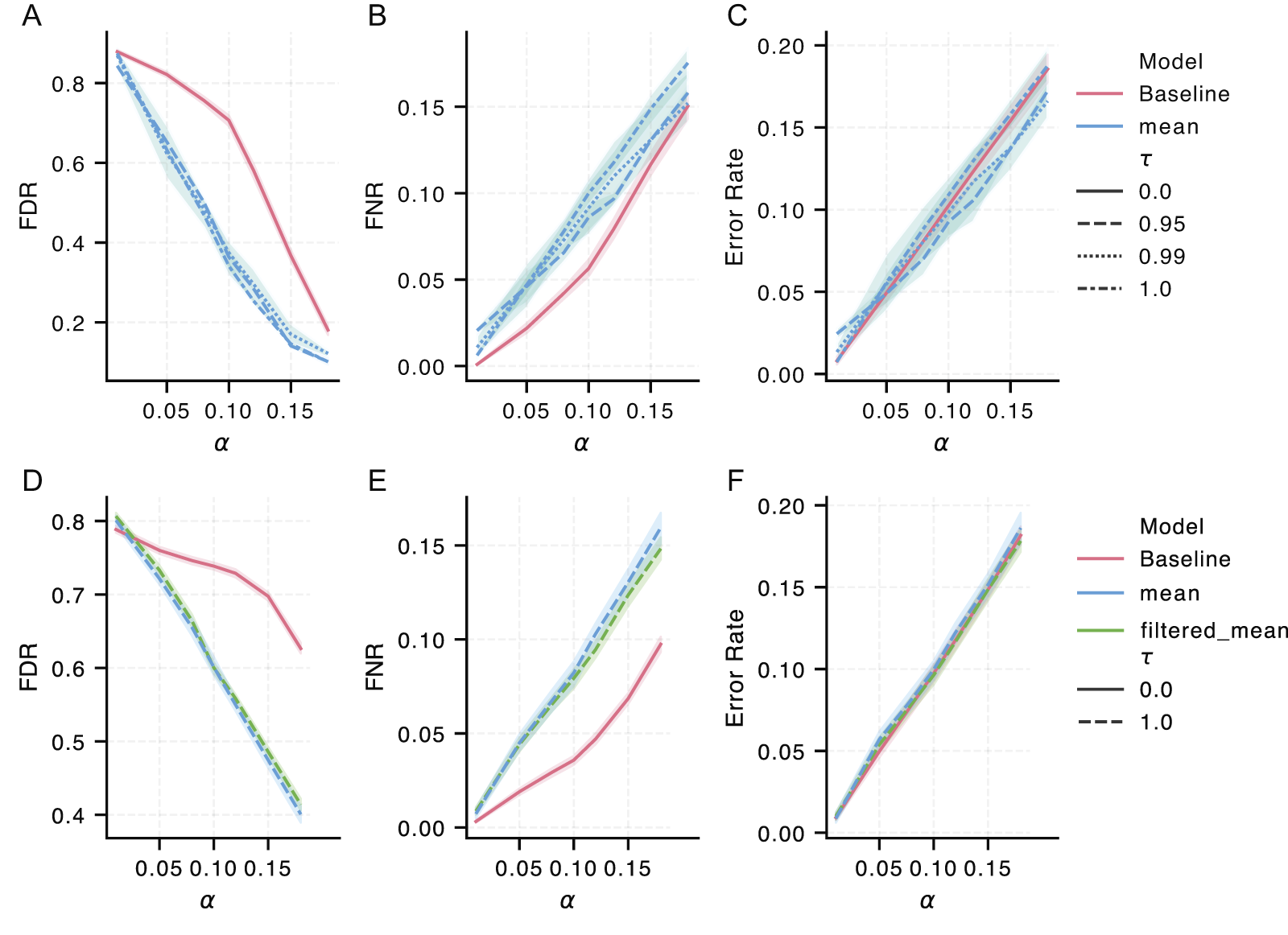
For our hyperparameter optimization, we evaluated performance using the area under the curve of the FDR, calculated across a range of specified error rates
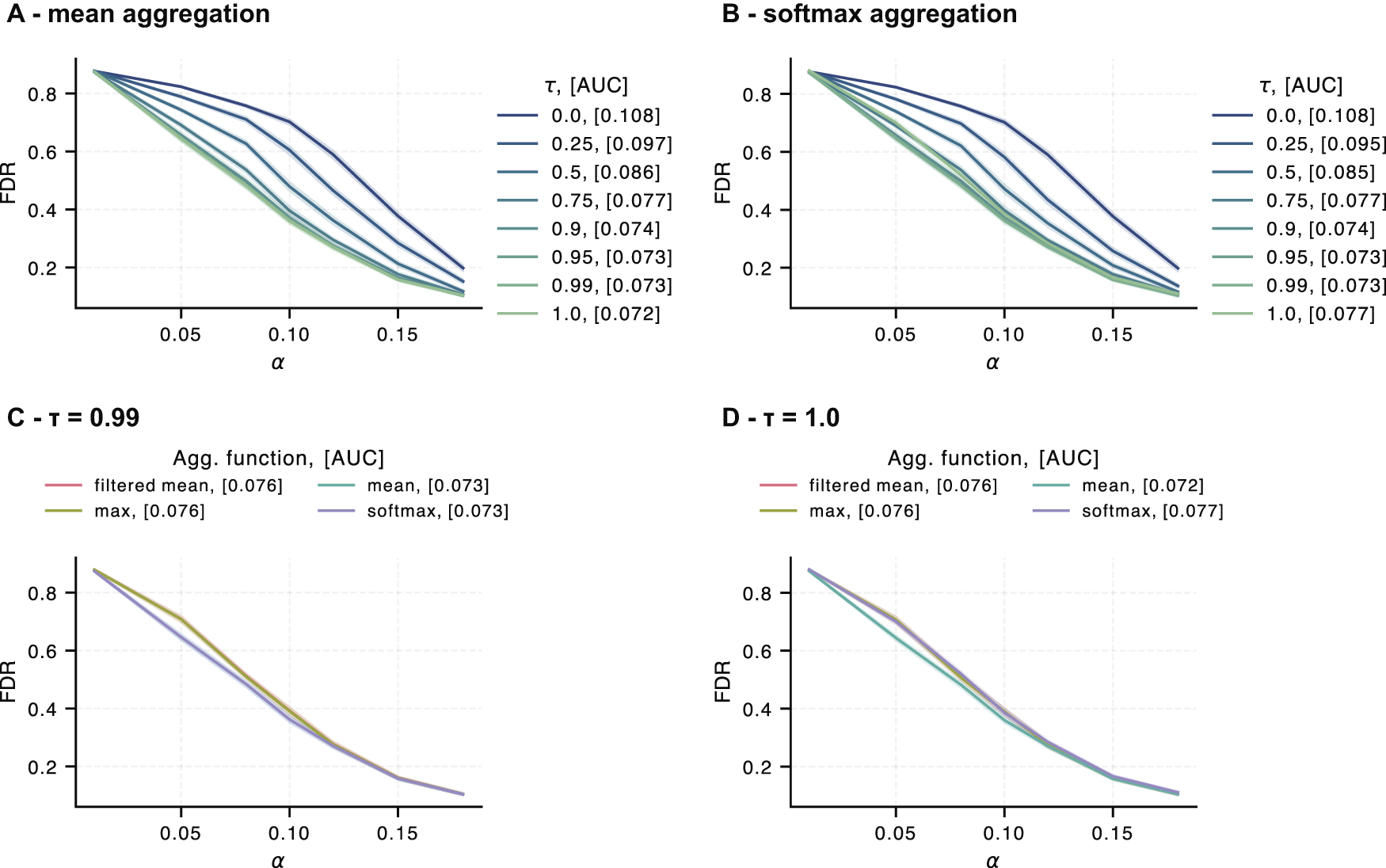
Hyperparameter study for our conformal predictor with knowledge-graph inclusion for Klebsiella pneumoniae: Subplots
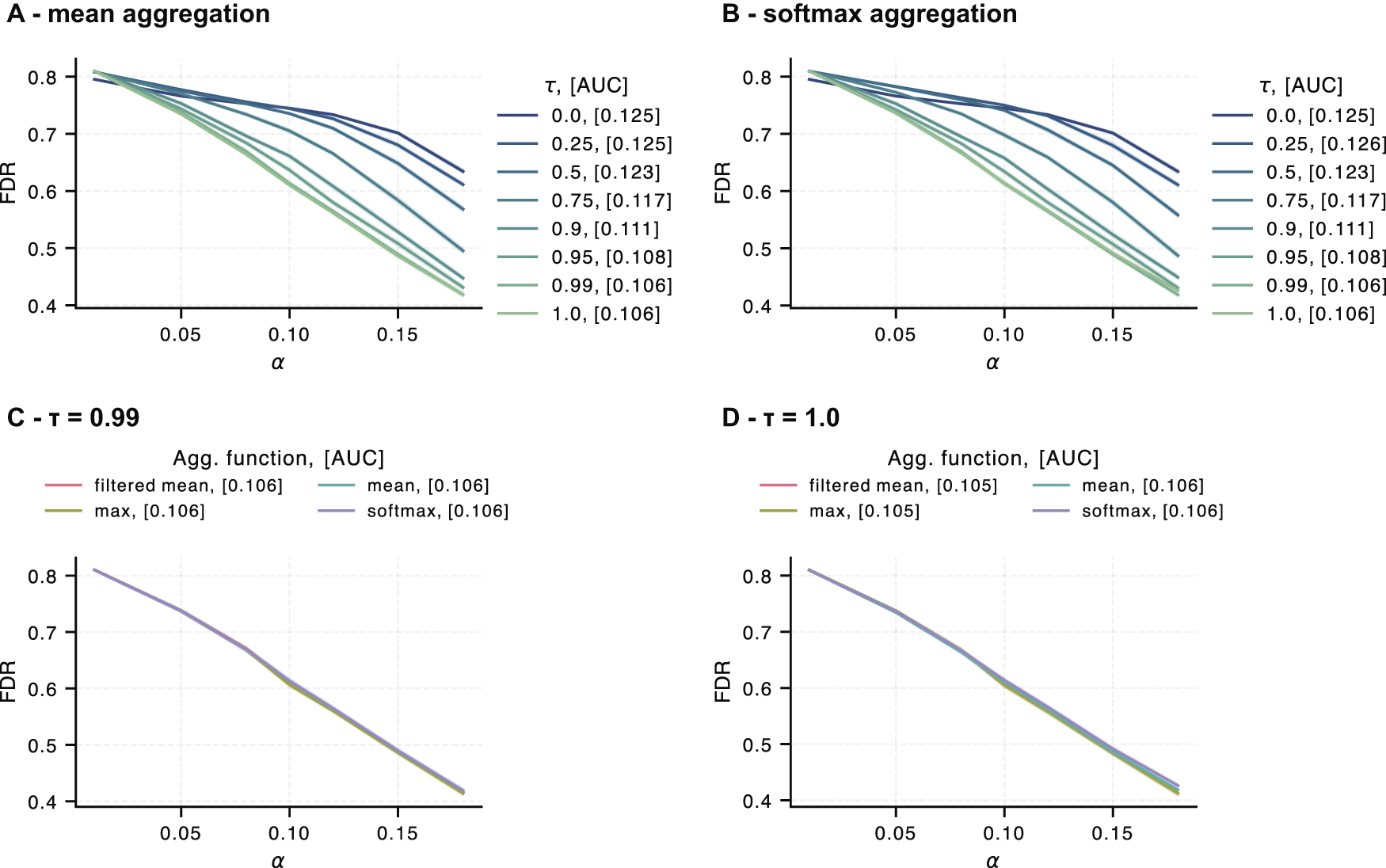
Hyperparameter study for our conformal predictor with knowledge-graph inclusion for Escherichia coli: Subplots
We evaluated the performance of the best conformal predictors on a per-drug basis using individual empirical error rate and antibiotic-specific FDR metrics (detailed definitions in the Appendix in the section ADDITIONAL METRICS). Figure 6 shows the results for the grouped drug classes fluoroquinolones, cephalosporins, penicillins, and carbapenems for Klebsiella pneumoniae. Figure 7 shows the equivalent results for Escherichia coli.
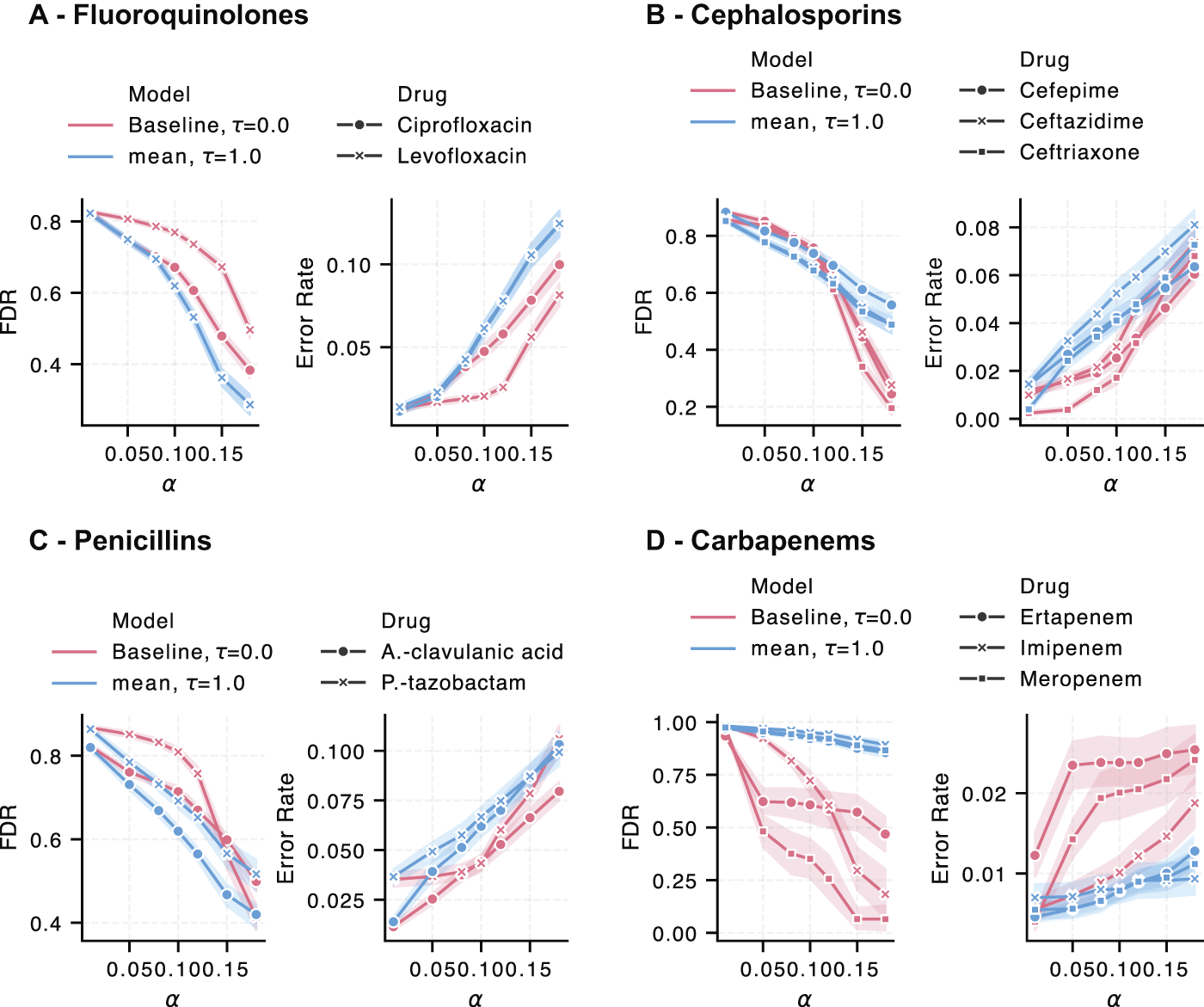
Individual drug false discovery rates (FDR) and empirical error rates (Error Rate) of our conformal predictor with knowledge-graph inclusion (blue) vs. the baseline conformal predictor in Eq. 5 (red) for Klebsiella pneumoniae. Drugs are grouped by their drug class, with subplots
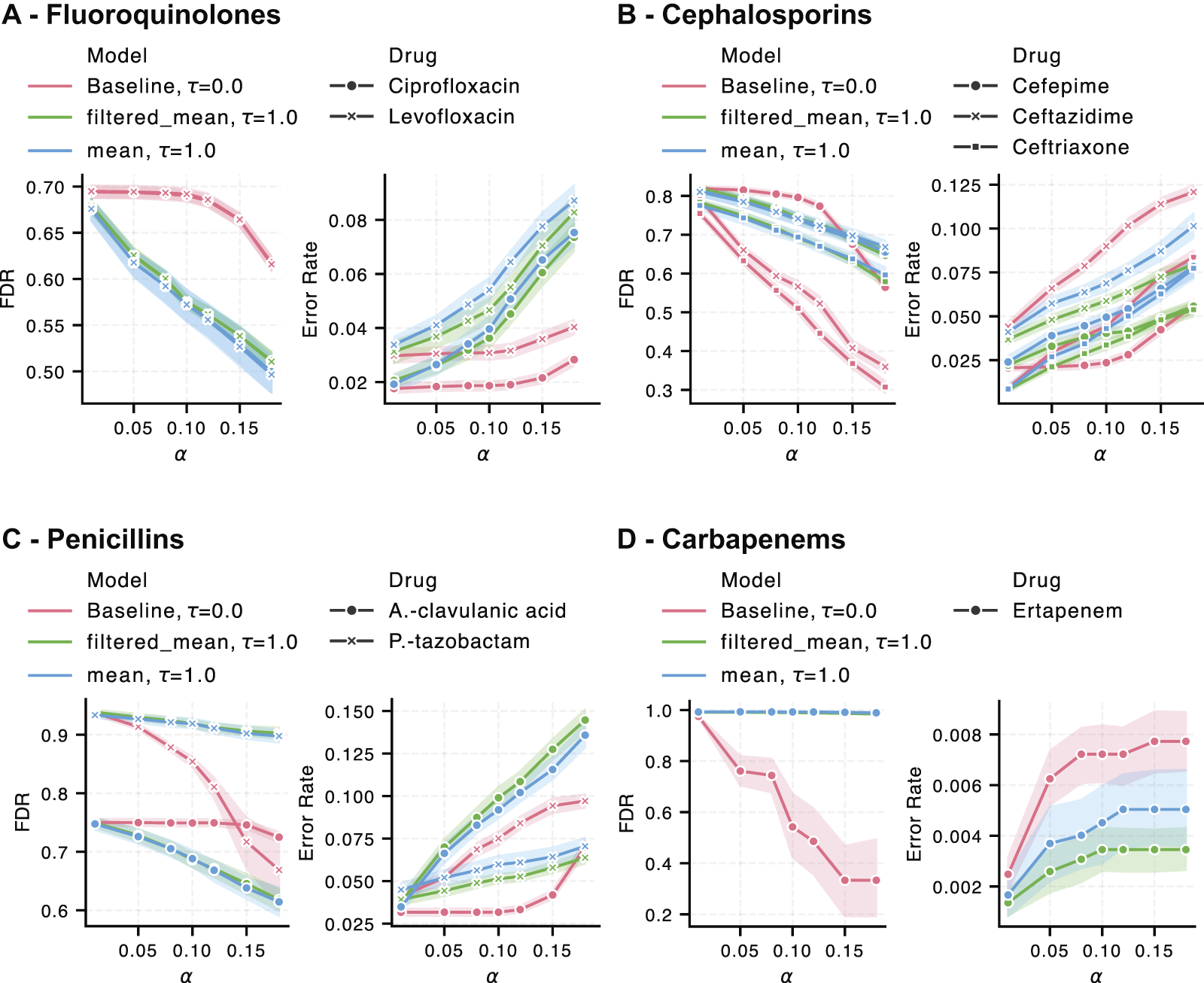
Individual drug false discovery rates (FDR) and empirical error rates (Error Rate) of our conformal predictor with knowledge-graph inclusion (blue/green) vs. the baseline conformal predictor in Eq. 5 (red) for Escherichia coli. Drugs are grouped by their drug class, with subplots
We observed that individual error rates consistently fell below the target threshold
The results demonstrate improved FDR performance compared to the baseline conformal predictor for three major antibiotic classes: fluoroquinolones, penicillins, and cephalosporins. The only exception is the carbapenem class, which shows increased FDR rates. However, this trade-off for carbapenems (the least represented class in our dataset and the most challenging for our predictors) should be considered in context: While the baseline conformal predictor only offers coverage guarantees near each carbapenem’s resistance prevalence rate, our model delivers stronger empirical error rates, although at the cost of higher FDR rates for these specific drugs.
Clinical motivation
In this study, we demonstrate how a combination of conformal prediction and domain-specific knowledge on top of cutting-edge machine learning models can pave the way to clinically applicable antimicrobial resistance prediction from MALDI-TOF mass spectra. MALDI-TOF MS-based resistance prediction has shown the potential to provide results 24 hours earlier than conventional culture-based approaches. However, despite promising results, their practical applicability has mainly been demonstrated retrospectively (Weis et al., 2022a; Yu et al., 2023). Examples of successful prospective clinical evaluations are rare and would only focus on a particular resistance mechanism (Wang et al., 2022). Therefore, widespread application in clinical diagnostics is still a long way off. The lack of broader clinical adoption comes from several remaining challenges. These include uncertain model reliability across different clinical settings (Park et al., 2024) and the absence of rigorous uncertainty metrics—a critical requirement for clinical decision-making where lives are at stake. The urgency of addressing this gap, and, therefore, enabling faster prediction of antimicrobial resistance in clinical diagnostics, is underscored by established research showing that inadequate initial antimicrobial treatment leads to dramatically increased mortality rates (Tumbarello et al., 2007). Particularly in severe systemic infections such as sepsis, this 24-hour advantage of MALDI-TOF MS-based antimicrobial resistance prediction can be crucial for patient survival.
Technical novelty
Our approach of incorporating conformal prediction provides statistically guaranteed error rates for antimicrobial resistance predictions from MALDI-TOF spectra, enabling confident clinical decision-making based on these rapid predictions. While conformal prediction has been previously applied to antimicrobial resistance prediction using epidemiological patient data for Escherichia coli (Inda-Díaz et al., 2023), which represents a completely different clinical setting, our study is the first to apply this technique to MALDI-TOF spectra. Our approach allows clinicians to specify guaranteed error levels based on their assessment of clinical risk. For instance, in severe infections, they can bound the error rate—the rate at which the predictor would give a susceptible result for an antibiotic, which is indeed resistant—to a conservative threshold, receiving predictions with higher certainty at the cost of potentially flagging more antibiotics as resistant. This will reduce the occurrence of very major errors in antimicrobial resistance prediction.
We further enhanced our conformal predictor by integrating domain-specific knowledge through the incorporation of a knowledge graph in the conformity score function. Using Klebsiella pneumoniae and Escherichia coli as strategic model organisms—chosen for both their clinical relevance in nosocomial infections and their well-documented hierarchical resistance patterns—we demonstrated how antibiotic hierarchies and co-resistance patterns can be leveraged to improve the conformal prediction sets. This improvement via integration of domain expertise with machine learning aligns with findings from other medical fields, such as radiology, where adding human knowledge to algorithmic approaches has shown superior performance (Deng et al., 2020). Our approach could be particularly powerful when adapted to incorporate local resistance patterns and epidemiology in the construction of the knowledge graph, allowing hospitals to customize the system based on their specific resistance profiles while maintaining rigorous error guarantees.
Looking beyond antimicrobial resistance prediction, our approach can be applied to improve conformal predictors in other multilabel classification tasks where simultaneous error guarantees across labels are needed and a knowledge graph capturing interdependencies across labels is available. A typical use case could be the field of sepsis prediction from intensive care unit patient data using machine learning models (Moor et al., 2023). Furthermore, since our pooling approach only updates the classifier’s output, our inclusion of the knowledge graph is agnostic to the type of conformity score function used. In future work, we will explore whether similar improvements of the prediction set can be seen with other conformity score functions.
The conformal prediction framework adapts and improves as the underlying machine learning model evolves. Building upon baseline techniques (Weis et al., 2022a), we have made significant technical improvements through the integration of learnable embeddings for antibiotic representation. While recent architectures have demonstrated enhanced performance through transfer learning between antibiotic classes (Visonà et al., 2023), these frameworks were limited by their inability to handle clinically highly relevant antibiotic drug combinations such as piperacillin-tazobactam. By incorporating MoleBERT embeddings, we overcame this limitation while maintaining or improving performance for most clinically relevant antibiotics for both Klebsiella pneumoniae and Escherichia coli. This advancement is particularly important given that many newly approved antibiotics are
To put our results into a future perspective, the implementation of this approach can build upon existing MALDI-TOF MS infrastructure, which is already established in clinical microbiology laboratories for species identification. Therefore, technical integration within existing or new software frameworks would be straightforward. However, clinical validation through trials will be necessary to demonstrate real-world utility. Since this work applies to a health care setting, rigorous quality control practices will be required to meet health care product standards, following regulations such as those from the IVDR in the European Union. Once implemented, regular recalibration would be required to maintain performance, aligning with standard clinical laboratory quality control practices.
Furthermore, we acknowledge several limitations of our study. Firstly, we have only evaluated the usefulness of the knowledge graph on species of the order Enterobacterales, where numerous antibiotic resistance interdependencies exist, and future studies must assess whether the demonstrated improved performance holds for other clinically relevant species and antibiotics. Additionally, while simultaneous and individual-level statistical guarantees represent a significant advance in clinical applicability, refinement of the individual-level guarantees remains an important future direction. For instance, as the coverage guarantee is not conditional on the drug being resistant, we do not cover the FNR of the individual drugs. This will be tackled in future studies to possibly further meet the clinical needs for such an algorithm.
CONCLUSION
Our work addresses a critical gap between the theoretical potential and practical implementation of MALDI-TOF MS-based antimicrobial resistance prediction. By combining conformal prediction with domain knowledge and advanced antibiotic representations, we enable resistance predictions with statistical guarantees while extending capabilities to highly relevant drug combinations. As underlying machine learning approaches continue to advance through various complementary innovations, this framework provides a robust foundation for translating rapid antimicrobial resistance prediction into clinical practice, potentially improving patient outcomes in cases where timely appropriate therapy is critical.
AUTHORS’ CONTRIBUTIONS
N.C.B. led the development and implementation of the conformal prediction methodology. L.M. led the development and implementation of the machine learning models and data processing pipelines. D.C. contributed ideas and technical development for the graph enhancements to the machine learning and conformal inference frameworks. J.S. provided medical domain expertise and designed the knowledge graph structure. K.B. supervised the project and provided strategic guidance. All authors contributed to the writing and revision of the article.
Footnotes
ACKNOWLEDGMENT
The authors would like to thank the Research in Computational Molecular Biology conference (RECOMB) for coordinating the review process and providing valuable feedback that improved this article.
AUTHOR DISCLOSURE STATEMENT
K.B. is co-founder and scientific advisor of Computomics GmbH, Tübingen, Germany.
FUNDING INFORMATION
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Supplemental Material
Supplemental Material
Supplemental Material
Supplemental Material
Supplemental Material
Supplemental Material
Supplemental Material
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.