Abstract
Due to the complex environments in real fields, it is challenging to conduct identification modeling and diagnosis of plant leaf diseases by directly utilizing in-situ images from the system of agricultural Internet of things. To overcome this shortcoming, one approach, based on small sample size and deep convolutional neural network, was proposed for conducting the recognition of cucumber leaf diseases under field conditions. One two-stage segmentation method was presented to acquire the lesion images by extracting the disease spots from cucumber leaves. Subsequently, after implementing rotation and translation, the lesion images were fed into the activation reconstruction generative adversarial networks for data augmentation to generate new training samples. Finally, to improve the identification accuracy of cucumber leaf diseases, we proposed dilated and inception convolutional neural network that was trained using the generated training samples. Experimental results showed that the proposed approach achieved the average identification accuracy of 96.11% and 90.67% when implemented on the data sets of lesion and raw field diseased leaf images with three different diseases of anthracnose, downy mildew, and powdery mildew, significantly outperforming those existing counterparts, indicating that it offered good potential of serving field application of agricultural Internet of things.
Keywords
Introduction
Plant diseases are one of the main factors to decrease the yield and quality of global agriculture production. 1 Cucumber is one of the most popular vegetables on our table. However, cucumber is severely affected by more than 30 diseases during its growth cycle, the most common ones of which are anthracnose, downy mildew, powdery mildew, and so on. The diseases have been posing threats to the cucumber production and bringing huge losses to farmers. Therefore, it is of great significance to timely identify the real types of diseases and take corresponding preventive measures to improve the quality and yield of cucumber so as to reduce the economic loss of farmers. At present, the identification of cucumber diseases mainly depends on the field investigation of farmers and plant pathologists to discriminate the specific type of cucumber diseases according to their experience.2,3 However, manual diagnosis is often laborious, time-consuming and subjective, inevitably resulting in the low-efficiency and misdiagnosis. Meanwhile, the techniques based on laboratory testing and biosensors are difficult to be implemented for farmers due to the lack of expertise and high cost.
During recent years, the agricultural Internet of things (IoTs) has been one important infrastructure for timely acquiring the field videos and environmental data. 2 In modern precision agriculture, it is expected to offer online collection and identification of plant disease images in a timely and accurate way through the system of IoTs. For example, the plant leaf disease images are captured and uploaded using mobile devices and in-situ video surveillance, subsequently the diagnosis result of diseases is automatically provided. 3 To this end, it is in great demand to conduct timely and accurate identification of cucumber leaf diseases in automatic way, in addition to the support of IoTs. During the previous decade, deep learning has gained significant attention in the task of automating image classification and recognition.1,4–6 To meet with the requirement of huge number of annotated training samples, the modification operations of the images—such as random crop, rotation, translation, flip, and scale—have been popularly employed for enlarging data sets.7,8 Furthermore, for the purpose of enriching the versatility of training images, generative adversarial networks (GANs) and its variations have been introduced in producing new samples on various image data sets. 9 As highlighted in Mai et al. 10 and Hu et al., 11 the introduction of GAN network was really helpful in improving the diversity and variation of training images, thus making contribution to high identification accuracy. In addition, it has been claimed that feeding lesion images was effective for achieving better identification in comparison to diseased leaf images, which demonstrated that, even in the era of deep learning, one sophisticated region extraction still helps in improving the performance of practical image-based disease diagnosis.1,11,12
Regrettably, it is challenging to achieve good identification accuracy under field conditions through the system of IoTs. This reason is that the existing methods might perform poorly under real field conditions when using IoTs, resulting from the fact that they did not fully take into account the existence and disturbance of uneven illumination and clutter backgrounds in the diseased leaf images. Actually, one cannot expect to always obtain the satisfactory images from the system of IoTs because they may be taken by remote cameras and farmers without enough expertise. As a result, the background of the images might account for a large part of the whole image area. The disease spots might have the similar color as ground soil in the background. Therefore, the introduction of an efficient preprocessing step to segment out the region of disease spots is still required. 12 Moreover, it is required to manually annotate them one by one when preparing large number of training samples, which is time-consuming and labor-intensive in addition to the requirements of plant expertise. Therefore, the training samples may be still limited even when applying the IoTs to agriculture. In addition, it is worthy of further exploring powerful, but light-weight, identification models of plant diseases because there are too many parameters in the considered deep convolutional neural networks.
To address these aforementioned issues, the proposed approach aimed to implement the recognition of cucumber leaf diseases under the condition of small sample size and IoTs, including three major procedures. To acquire good lesion images, a two-stage method, combining GrabCut with support vector machine (SVM), was used to segment the disease spots from cucumber leaves by extracting the color, texture, and border features of cucumber leaves. Subsequently, to enlarge training data sets, the new training samples were generated by the activation reconstruction generative adversarial networks (AR-GAN) for data augment. Finally, the dilated and inception convolutional neural network (DICNN) was developed for implementing the identification of cucumber leaf diseases. The performance comparisons were made on the data sets of diseased images with three different leaf diseases of anthracnose, downy mildew, and powdery mildew, demonstrating that the proposed approach effectively overcame the aforementioned shortcomings.
Related work
With the development of machine learning and computer vision, great efforts have been devoted to developing the new approaches that could automate the process of recognition of diseases using image processing techniques. So far, leaf disease identification and diagnosis of cucumbers and other plants have made great progress based on classical machine learning, such as SVM, AdaBoost, k-nearest neighbor (kNN), and probabilistic neural networks (PNNs).13–16
In recent years, deep learning–based algorithms have increasingly prevailed the area of image classification and recognition because they can offer promising results and larger potential of feature extraction and recognition by passing input data through several non-linearity functions in comparison with traditional shallow methods.4–6 It has been witnessed that deep convolution neural network was remarkably superior to the traditional methods in plant disease recognition. H Yalcin 17 used pre-trained AlexNet to identify and classify the phenological stages of wheat, barley, and other plants. In order to classify plants in the natural environment on a large scale, V Bodhwani et al. 18 designed a 50-layer deep residual learning framework consisting of five stages. M Afonso et al. 19 proposed a quick and non-destructive method based on ResNet18 to identify blackleg disease of potato plants in the field. In order to improve the classification accuracy of cucumber leaf disease spots MA Khan et al. 20 used the Sharif saliency method and pre-trained VGG model to extract deep features which were finally fed to multi-classification SVM for disease recognition. Particularly, in addition to classical deep convolution neural networks, such as AlexNet, VGG, and ResNet, many approaches have been proposed based on the latest deep convolutional neural networks associated with a classifier for the identification of diseases using the images of the plant parts.21,22 To tackle the problems of excessive parameters of AlexNet and single feature scale, a global pooling dilated convolutional neural network was proposed for plant disease identification by combining dilated convolution with global pooling. 1 S Zhang et al. trained the deep convolutional neural network for conducting symptom-wise recognition of cucumber diseases with the aid of lesion image segmentation. 13 UP Singh et al. 21 proposed an approach for identifying mango leaf disease named as anthracnose using multilayer convolutional neural network. In order to detect tomato diseases and pests, Fuentes et al. 22 combined VGG and ResNet with each of these detectors: region-based fully convolutional network, faster region-based convolutional neural network, and single shot multi-box detector. T-T Le et al. 23 used mask region-based convolution neural networks (Mask R-CNNs) to detect complex banana fruits in the image and predict the types of bananas. J Chen et al. 24 skillfully added the Inception module to the VGGNet model, proposed the INC-VGGN model to identify rice diseases. In order to realize the detection of apple at different growth stages, Y Tian et al. proposed an improved YOLO-V3 network processed by DenseNet method. The DenseNet method is used to process feature layers with low resolution in the YOLO-V3 network. This effectively enhances feature propagation, promotes feature reuse, and improves network performance. 25 Aiming at various problems with the rice disease images, such as noise, blurred image edge, large background interference, and low detection accuracy, G Zhou et al. 26 proposed a method for detecting rapid rice disease based on fuzzy c-means and k-means (FCM-KM) FCM-KM and Faster R-CNN. However, it is well-known that the deep learning–based algorithms require large annotated data sets on a per-pixel level in order to successfully train the large number of free parameters of the deep network. 27
Acquisition of plant disease images is a complex and costly endeavor, which often requires the collaboration of professional people from different fields at various growth stages. 1 For the purpose of alleviating over-fitting problem, different research has addressed several automated data augmentation techniques for enlarging the data sets. 7 In particular, GANs and its variations have been introduced in producing new samples on various image data sets in addition to the traditional modification operations of the images.9,10 The main reason to use GANs in generating new images was the effective generation with the same characteristics as the given training distribution, when given a limited number of input images. To increase the resolution and quality of the generated samples, the variations of GANs have been involved in designing new generator and discriminator CNN networks, modifying GAN configurations, and reformulating GAN objective functions.22,28 For the purpose of obtaining large number of plant seedling images, SL Madsen et al. 28 designed WacGAN using a supervised conditioning scheme that enabled the model to produce visually distinct samples for multiple classes. ME Purbaya et al. 29 improved GAN and added Elastic Net or a combination of L1 and L2 regularizer to generate multiple plant leaf images. Radford A et al. and Zhang M et al. used deep convolutional generative adversarial network (DCGAN), with the combination of CNN with GAN, 30 to generate citrus canker images to improve the accuracy of the classification network. 31 G Hu et al. 11 segmented disease spots from tea leaf’s disease images using SVM; subsequently, data augmentation was implemented by means of one improved conditional deep convolutional generative adversarial network (C-DCGAN) for generating new samples. Finally, the VGG16 model was trained for identifying the tea leaf’s diseases. 11 To improve the plant disease recognition performance in data deficient application, Zhu JY et al. and Nazki H et al. proposed a pipeline for synthetic augmentation of plant disease data sets using AR-GAN, where an additional perceptual loss was introduced into CycleGAN for generating more natural images.32,33 For the purpose of reducing latent similarities and enriching the versatility of training images, QH Cap et al. 34 proposed another improved CycleGAN by only transforming leaf areas with a variety of backgrounds, instead of transforming entire image from the source to the target domain. A Förster et al. used a hyperspectral microscope to capture hyperspectral images of barley leaves, and then used CycleGAN to forecast the spread of disease symptoms on barley plants. Because of the complexity of the lesion structure, the image quality of the complete diseased leaf directly generated by GAN might be not satisfactory. 35 In order to solve this problem, R Sun et al. proposed a binarization plant lesion generation method using a binarization generator network. After generating plant lesions with specific shape, the image pyramid and edge smoothing algorithm were used to synthesize a complete lesion leaf image. 36
Aiming at effectively dealing with the disease identification under field conditions in the system of IoTs, this article developed one new identification approach of cucumber leaf diseases using deep learning and small sample size. Compared with the traditional algorithms, our work had the following advantages. The proposed two-stage segmentation method effectively acquired lesion images from the raw images with uneven illumination and clutter background under field conditions in the system of IoTs. Training data set was enlarged using AR-GAN for avoiding the manual selection and annotation of large number of raw images. In addition, the DICNN effectively implemented the accurate identification of cucumber leaf diseases with powerful feature extraction and fast convergence ability.
Materials and methods
Disease identification approach
The proposed approach comprised three components. The first one was to acquire the lesion image by segmenting the disease spots from the whole image taken under field condition. The second component was to implement the data augmentation. The third one trained and tested the identification neural network. Figure 1 illustrates the proposed identification scheme of cucumber leaf diseases.

Schematic diagram of identifying cucumber leaf diseases.
Lesion image segmentation
Through the agricultural IoTs deployed in field, the images of cucumber leaves were often taken with the existence of complex environmental background. The environmental backgrounds, also known as noise, would severely interfere with the modeling of leaf disease identification. Naturally, it is definitely beneficial for obtaining good identification model of leaf diseases if eliminating clutter background before conducting model training.
There have been lots of image segmentation algorithms that could be classified into unsupervised and supervised methods. For example, fuzzy cluster,K-means, and SVM were widely used to develop the image segmentation algorithms.11,37,38 Due to the uneven illumination and clutter background in diseased leaf images, it is very difficult to get good segmentation of disease spots using the unsupervised methods. G Hu et al. 11 attempted to use SVM with few shot learning for segmenting the disease spots from tea leaf images with clutter background. Based on graph frame theory, GrabCut could divide the image into the foreground pixel set and the background pixel set of two disjoint subsets of the image. 39 It has been claimed that GrabCut outperformed many traditional image segmentation methods with the aid of a little bit human interference.
In order to acquire lesion images, one two-stage image segmentation method, with the combination of GrabCut and SVM algorithms, was proposed for extracting the diseased spots from cucumber leaf images. As shown in Figure 1, in the first stage, the GrabCut took charge of extracting the cucumber leaves from the raw image, by taking into account texture and boundary information. In the following stage, SVM was employed for extracting diseased spots from leaves according to the texture and color features. Specifically, the diseased spots of cucumber leaves, color, and texture feature vectors of healthy cucumber leaves were used as training samples for SVM. Color feature vector was obtained by color feature extraction from color histogram in RGB space, and texture feature was extracted by the Gabor filter instead of the gray-level co-occurrence matrix. Similarly to the visual stimulus response of simple cells in human vision system, the Gabor wavelet is sensitive to the edge of image and has good characteristics in extracting local space and frequency domain information of objects.
Figure 2 illustrates one typical segmentation procedure and results of cucumber leaves and disease spots. It can be seen, from Figure 2, that the GrabCut was able to overcome the noises and showed strong capacity of segmenting leaves from the rest in raw images, which lay a solid foundation for the following segmentation of lesion images. It should be pointed out that a little bit human intervention was required because the diseased leaf segmentation from clutter background was carried out by interactively selecting the region of interest disease spots on raw diseased leaf images. However, the required action was to only frame the leaf area containing disease spots. In addition, the human intervention could be easily completed during photo-taking and annotation. Compared with the requirements of professional photography skill and image preprocessing in those traditional methods, the interactive segmentation offered an appropriate option to achieve good tradeoff between the accuracy and usability of segmentation method under the condition of IoTs.

Segmentation results of cucumber leaf and disease spots.
Data set augmentation
Compared with many existing GANs and its variation models, such as, DCGAN and SinGAN, the recently proposed AR-GAN was found successful at generating a more realistic and attractive composite image visually by preserving the semantic identity and features of the objects in the input mask. 33 The generator and discriminator in AR-GAN formed a game process and finally reached the Nash equilibrium. AR-GAN normalized the original objective function and cyclic consistency optimization on the basis of GANs. In addition, through improving the activation reconstruction (AR) loss function, the feature activation for natural images was optimized. Figure 3 presents the AR-GAN framework.
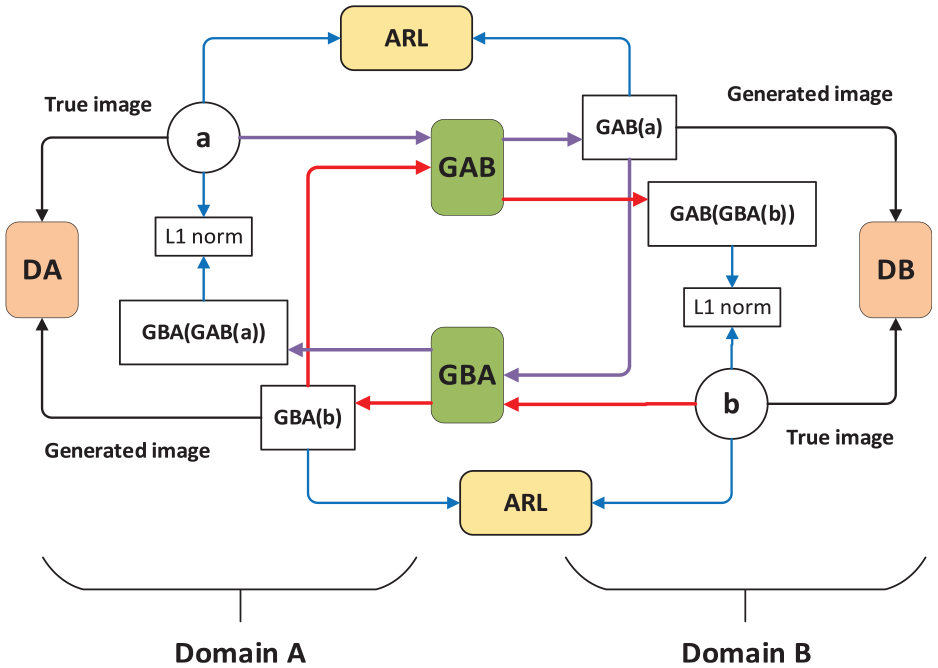
The framework AR-GAN.
As shown in Figure 3, domain A and domain B were two unaligned domains, the generators (GAB and GBA) translated an image from one domain to another, and the discriminator (DA and DB) of each domain was responsible for judging whether the image belongs to the domain. As shown by the purple and red arrows in the figure, image data flows between the cycle a to GBA(GAB(a)) and b to GAB(GBA(b)). The AR-GAN applied L1 loss between the input a (or b) and the AR input GBA(GAB(a)) (or GAB(GBA(b))) to enforce self-consistency. The AR module containing feature extraction network was introduced to calculate the AR loss, cycle-consistency loss, and antagonism loss in the network. The definition of the countermeasure loss function in AR-GAN was as follows
Another parallel adversarial loss
AR loss as a perception loss trained along with the other loss functions aiming to enforce perceptual realism between the real and the generated images and increasing the stability of the model. Let
where
where
For more details about the AR-GAN, readers are referred to the literature. 33 As mentioned before, there were 200 samples for each type of cucumber leaf diseases in this article. Such scale of training sample was far from enough when implementing the modeling of deep learning–based disease identification. To improve the diversity and variation, that is, quality of training samples, AR-GAN was employed for generating more training samples in addition to traditional data augmentation methods. More specifically, our data augmentation was implemented by integrating image rotation, image translation with AR-GAN.
Disease identification model
Deep convolutional neural network plays an important role in the identification of cucumber leaf diseases in this article. It is crucial for achieving accurate recognition to design an effective convolution neural network. Inception V3 is the third of four versions of the inception model proposed by Google on the basis of GoogLeNet. 40
As the task becomes more and more complicated, the performance of many deep learning models previously used on a large scale has reached a bottleneck, such as AlexNet, VGG, and ResNet. It is common sense that the easiest way to obtain a high-quality deep network model is to increase its depth or width, but this design idea usually brings the problems of over-fitting and gradient disappearance. The emergence of the Inception model makes it possible to maintain the sparsity of the network structure and take advantage of the high computing performance of dense matrices. 41 Compared with other deep learning networks, Inception V3 had two significant improvements. First, Inception V3 used parallel small-size filters to decompose large-size filters in convolution, which saved more computing resources and improved the speed of training. Second, it was the multi-scale convolution kernel in Inception V3 in charge of extracting multi-scale features of the input image. Through the fusion of these features, the recognition accuracy and robustness of the model were improved.
This article designed DICNN based on Inception V3 for identifying cucumber leaf diseases. Figure 4 shows the structure of the DICNN network, which had 52 layers, including 6 complete convolution layers (Conv1 to Conv6), 3 complete pooling layers (Pooling1 to Pooling3), and one Softmax classifier layer. The remaining 42 layers were composed of 3 Inception blocks. Among them, Block 1 contained 3 modules with a total of 9 layers, Block 2 contained 5 modules with a total of 23 layers, and Block 3 contained 3 modules with a total of 10 layers. Different from canonical Inception networks, in DICNN, the first convolution layer Conv1 was replaced by dilated convolution neural network, abbreviated as Dilated Conv1 in Figure 4. Convolution layer Conv3 used the convolution of no padding instead of the original convolution. And the full connection layer used for regression operation was also replaced by convolution layer Conv6. And there was a non-linear activation function ReLU and Batch Normalization behind all convolution layers, which was able to not only restrain the over-fitting problem to some extent but also to improve the training efficiency. The number of the output channels of each layer, the size of the convolution kernel, and the number of the extracted maps are denoted in Figure 4, respectively. The convolutional kernel numbers of Conv1, Conv2, …, Conv6 were 32, 32, 64, 80, 192, and 1000, respectively. The kernel sizes of Dilated Conv1, Conv2, Conv3, and Conv5 were all 3 × 3 dpi, and the kernel sizes of Conv4 and Conv6 were both 1 × 1 dpi. The pooling sizes of Pooling1 and Pooling2 were all 3 × 3 dpi, and the pooling sizes of Pooling3 were 7 × 7 dpi, respectively.

The structure of DICNN.
There were some key issues about the proposed DICNN. First, the first convolution layer in DICNN employed dilated convolution for reducing the loss of image spatial hierarchy information and internal data structure in the pooling process. 14 Second, no padding convolution in DICNN effectively contributed to the reduction in the position deviation during the convolution process, as addressed in Zhang and Peng. 42 Finally, DICNN replaced the full connection layer of the logic regression part of Inception V3 with convolution layer so as to become a real full convolution neural network, which greatly reduced the number of parameters of the whole network and saved computing resources. When feeding into DICNN, it was required to resize the image resolution of the training sample to 299 × 299 and use the RGB color channel. After the treatment of multiple convolution layers and pooling layers, the probability of each disease was regressed by the last Softmax layer. The output layer had three neurons representing the three types of cucumber leaf diseases, that is, anthracnose, downy mildew, and spot target. Given the output layer, Softmax function was used to calculate the estimated probability of three cucumber leaf diseases.
Experiments
Data set
In the experiments, the data set of cucumber raw diseased leaf images used in this article consisted of three types of disease images: anthracnose, downy mildew, and spot target diseases. The image data set was from the cucumber leaf disease image repository collected through the system of IoTs developed by the Anhui Province Key Laboratory of Smart Agricultural Technology and Equipment, Anhui Agricultural University. It was much convenient to collect in-situ leaf images through our IoT system than traditional ways. However, as discussed before, it was not feasible to directly utilize these stored images as our training samples because they were often taken by remote cameras and farmers without enough expertise and there existed several diseases on the same leaf.
Hence, the selection of raw diseased leaf images needed conducting in addition to annotation, whereas both image selection and annotation processes were often difficult, laborious, and time-consuming, as well as requiring a high level of expertise to avoid misclassification or other errors. Therefore, although there were lots of cucumber leaf images stored in the image repository of IoT system, 200 raw diseased leaf samples for each type of cucumber leaf diseases were chosen from the stored image repository, and the size of each sample was adjusted to 1024 × 682 pixels in order to improve processing efficiency before conducting image analysis.
Experimental settings
The deep learning models, including AR-GAN and DICNN, were trained on two Nvidia Tesla P100 (16 GB memory) graphics cards using the TensorFlow framework. These training strategy and hyper-parameters of deep learning models were determined based on preliminary experiments. More specifically, the batch gradient descent algorithm was used to optimize the weights of the network. The batch sizes of AR-GAN and DICNN were set to 8 and 24, respectively. The initial learning rate was set to 0.001 and dropped out every 20 epochs with a drop factor of 0.1. The maximum number of epochs used for training was set to 20,000.
Lesion image segmentation
The lesion images from segmenting cucumber leaf disease spots under uneven illumination and clutter backgrounds are presented in Figure 5. In order to show the validity of the proposed segmentation algorithm, the comparisons were made among K-means, fuzzy clustering, SVM, and the proposed two-stage lesion image segmentation methods on the basis of cucumber leaf disease samples with downy mildew, anthracnose, and spot target.
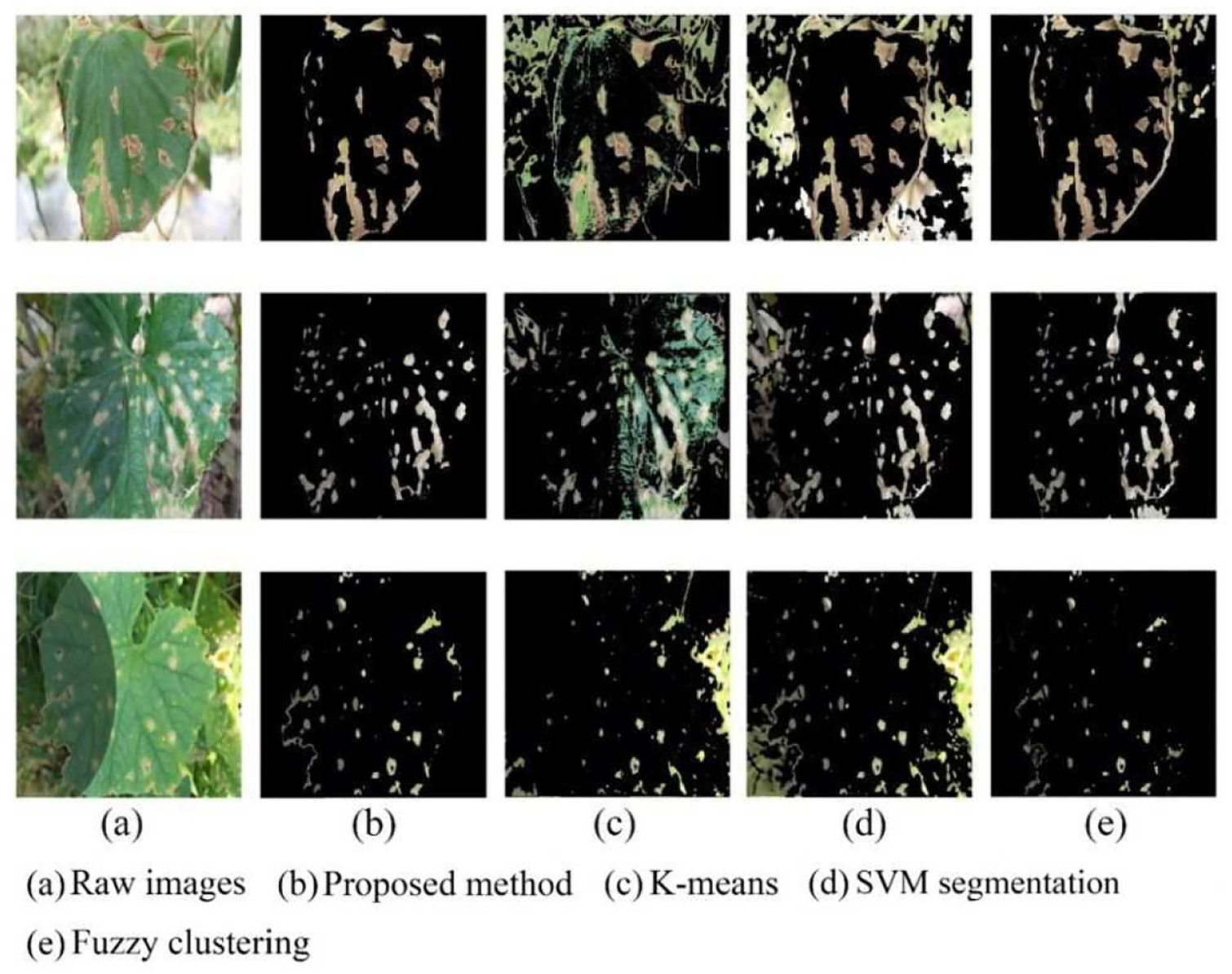

Lesion images from different segmentation methods: (a) raw images, (b) proposed method, (c) K-means, (d) SVM segmentation, and (e) fuzzy clustering.
As shown in Figure 5, K-means segmentation, fuzzy clustering algorithm, and SVM segmentation methods had some deficiencies in segmenting disease spots. In contrast, the proposed method was significantly superior to other segmentation methods. As for K-means segmentation, too many healthy areas of leaves were retained on the lesion images. Due to the uneven local illumination, K-means failed in discriminating the junction of the shaded area from the normal area for the boundary of the leaf. In the results of SVM segmentation proposed in Hu et al., 11 a large area of backgrounds was not removed. This was because SVM had deficiency in dealing with uneven illumination. Moreover, it was observed that the segmentation result would get worse and worse as the ratio of the leaf area to the whole image decreased. Compared with K-means and SVM, the fuzzy clustering method had more powerful ability of handling uneven illumination and clutter background with the expense of high computation cost; however, it still suffered from the problem of missing disease spots. Contrastively, the proposed combination of GrabCut and SVM could achieve satisfactory segmentation results on cucumber diseased leaf images taken under real field condition, which implied that the good training samples could be offered for further implementing disease recognition.
Disease identification accuracy
The 600 samples in the data set of raw diseased leaf images were divided into training and test data sets in a ratio of 8:2 by randomly selecting samples from each type of diseased cucumber leaf images. Then, all of 600 images were segmented to acquire lesion images. Of total raw diseased leaf and lesion images, 80% was used as training samples and 20% was used as test samples in the following experiments.
Identification using various data set augmentation methods
In this experiment, we compare our scheme of data set augmentation against several traditional methods, such as rotation, translation, and classical GAN variation. Totally, there were five data augmentation methods taken into consideration for making comparisons. More specifically, the notation “RT + AR-GAN” denoted that before the training samples were augmented by AR-GAN, their rotation and translation were required. AR-GAN denoted that the training samples were augmented directly by AR-GAN. RT + C-DCGAN and C-DCGAN followed the same naming principle. The rest one method of data augmentation was rotation and translation (abbreviated as RT). The 300 lesion samples for each disease were generated through the rotation and translation operation. Both AR-GAN and C-DCGAN generated 6000 new lesion samples for each disease. For a fair comparison, these five groups of training samples, as well as the training samples in the original lesion image data set, were fed into the proposed DICNN for training. The recognition results are shown in Figure 6.

Identification accuracy using different data set augmentation methods.
As can be seen in Figure 6, the worst identification accuracy was observed if without any implementation of data augmentation. This phenomenon indicated that, in the case of small sample size, enlarging the scale of sample data set did result in the improvement of the accuracy of disease recognition by effectively alleviating the over-fitting problem in the deep convolutional neural networks to a certain extent. Apparently, compared with no augmentation, the identification accuracy increased by a certain amount if employing image rotation and translation. Meanwhile, the introduction of GAN network led to more gain in the recognition accuracy because it brought the increase in the sample scale and diversity. Moreover, the operation of image rotation and translation could contribute to the increase of the identification accuracy even in the case that GAN network was employed. Under scrutiny, it was observed that the rotation and translation effectively reduced the probability that GAN network suffered from over-fitting problem, resulting in the quality improvement of training samples. However, in comparison with C-DCGAN, AR-GAN had better potential of synthesizing high-quality training samples from a finite number of original lesion images. Figure 7 presents some lesion images of three cucumber leaf diseases generated by AR-GAN. From Figure 6, it can be observed that AR-GAN achieved the improvement in the average recognition accuracy of about 8% and 6% over C-DCGAN, respectively, when with and without rotation and translation operations. Therefore, it can be drawn that the proposed integration of rotation, translation, and AR-GAN was as an effective solution for implementing the sample augmentation.
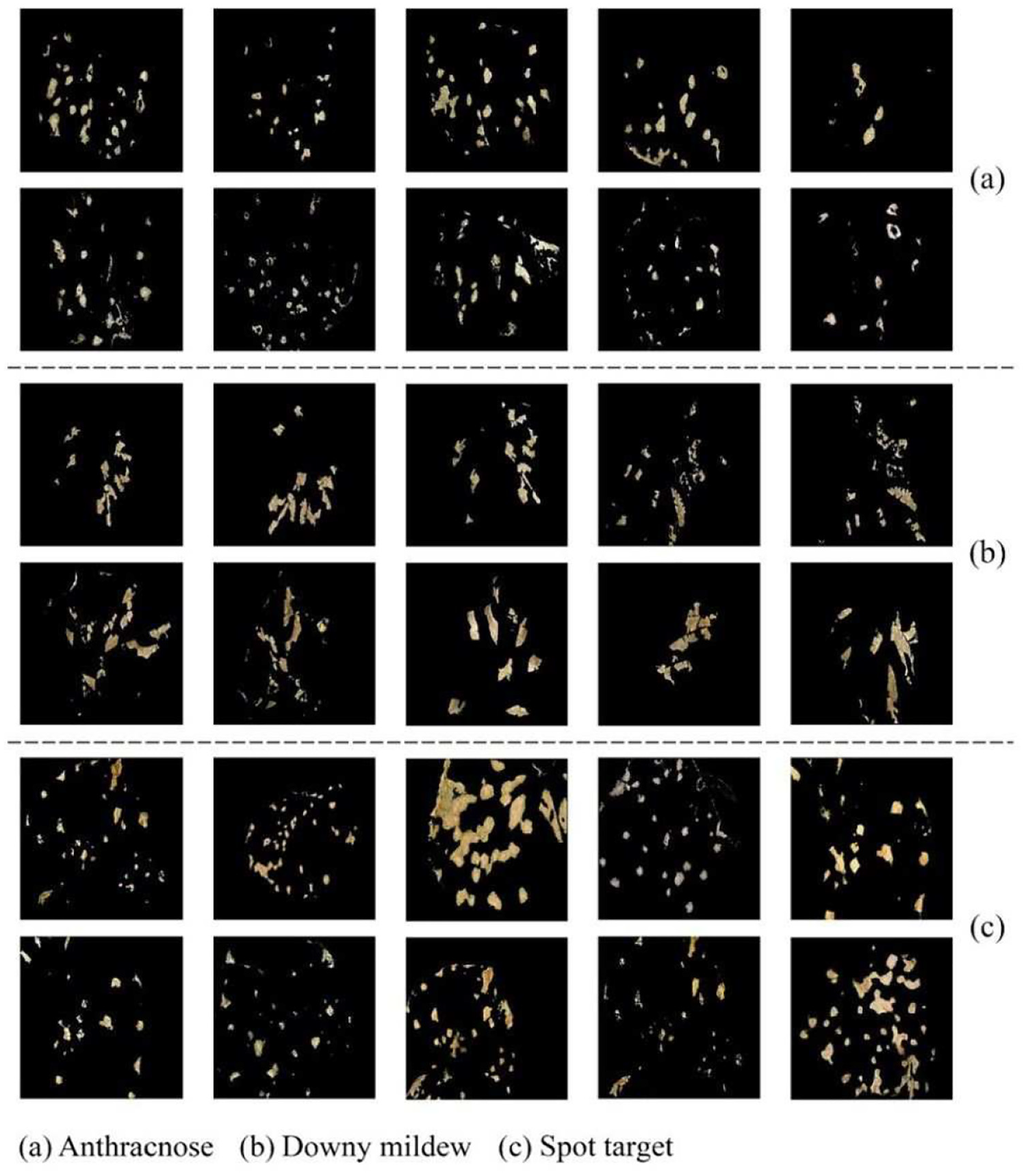
Lesion images generated by AR-GAN: (a) anthracnose, (b) downy mildew, and (c) spot target.
Identification using various machine learning methods
We compared our DICNN against VGG16 used in Hu et al., 11 as well as several traditional machine learning algorithms, such as SVM, kNN, and AdaBoost. When evaluating DICNN and VGG16, the integration of rotation, translation, and AR-GAN was used for generating 6000 new lesion samples for each disease as the training samples. To evaluate SVM, kNN, and AdaBoost, the 160 lesion training samples for each disease were randomly selected from the original lesion image data set.
Figure 8 shows the comparison of traditional machine learning and deep learning methods in cucumber leaf disease identification accuracy. It can be seen from Figure 8 that the deep learning methods, both DICNN and VGG16, were significantly superior to traditional machine learning algorithms in the identification accuracy of cucumber leaf diseases. At the same time, DICNN offered higher average identification accuracies, with the increase by 10%, than VGG16. Moreover, it was found out that it took DICNN fewer iterations, as well as lower time overhead, to achieve the stable recognition results than VGG16. All the above advantages of DICNN were from its inherent characteristic of network structure. The first convolution layer in DICNN employed dilated convolution, which reduced the loss of image spatial hierarchy information and internal data structure in the pooling process by means of enlarging the local receptive field and enhancing the feature extraction ability of the convolution layer. Using convolution layer instead of full connection layer greatly reduced the number of parameters of the whole network and saved computing resources. The employment of Inception network structure enhanced the feature extraction ability of the convolution layer from the input images. Due to the use of fully connected layer, VGG16 might take too much computation and convergence time to calculate its large number of weight parameters. In addition, the ability of VGG16 extracting features was limited resulting from dependence on the resolution of feature maps. Therefore, the proposed DICNN offered the advantage of network structure over VGG16.
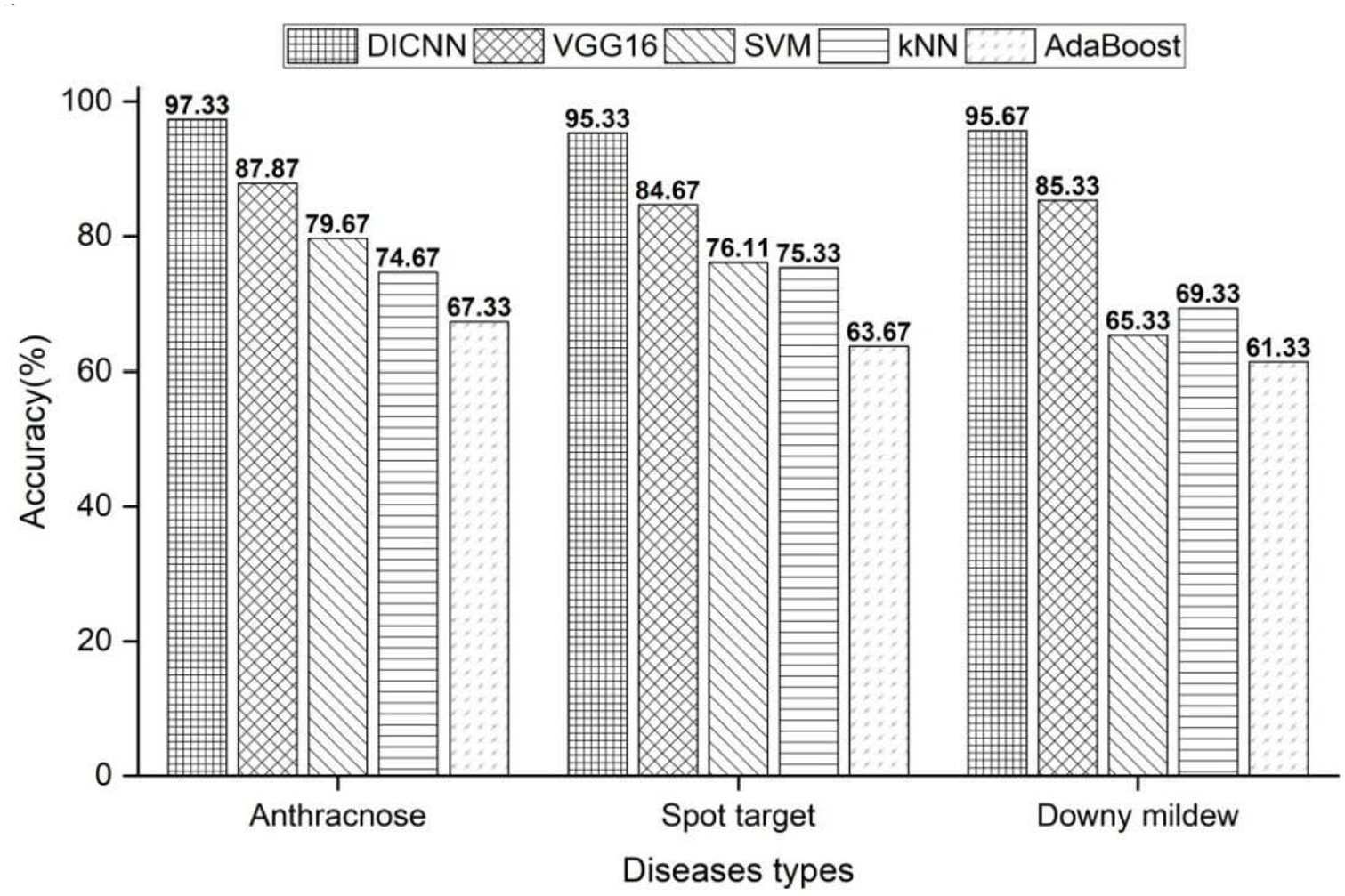
Identification accuracy using different machine learning methods.
Identification on raw diseased leaf and lesion images
In order to provide one good reference for practical application of the proposed approach, we performed the disease identification experiments by the aforementioned deep learning and traditional shallow machine learning methods, on the augmented training samples of raw diseased leaf image and segmented lesion image data set. Figure 9 depicts the average identification accuracies of three diseases when using raw diseased leaf and lesion images. The proposed approach in this article was denoted as “AR-GAN + DICNN.” The symbol “C-DCGAN + VGG16” represented the identification method of cucumber leaf diseases proposed in Hu et al. 11

Identification accuracy on raw diseased leaf and lesion images.
From Figure 9, it can be seen that the improvement in the identification accuracy was acquired when using lesion image data set than using raw diseased leaf image data set, which was consistent with the conclusion in the existing literature. However, the traditional shallow machine learning methods received significant increase in identification accuracy on the segmented lesion image data set. In contrast, deep learning method obtained less improvement from the lesion image data set. This can be explained by the fact that deep learning methods needed a large number of training samples to train and can automatically learn the high-level abstract features from the original images, while the identification accuracies of shallow machine learning methods heavily depended on image segmentation and feature extraction algorithms, which led to the limitation of directly recognizing disease type from the raw diseased leaf image. It should be mentioned that the proposed approach acquired less gain in the average identification accuracies than the existing C-DCGAN + VGG16 between two experiments of lesion and raw diseased leaf image data sets, which should owe to the combination of our data set augmentation and inherent advantage of network structure. Note that the proposed approach achieved excellent ability of disease identification with the accuracy of 96.11% and 90.67% on the data sets of lesion and raw field images, respectively, which indicated that it had a good potential of providing reliable service of leaf disease recognition for both plant pathologists and farmers by integrating with agricultural IoTs, which would help in saving human efforts and reducing pesticide usage.
Conclusion
This research developed a novel identification approach of cucumber leaf diseases based on small sample size and deep convolutional neural network. The lesion images were acquired by one two-stage segmentation method that offered strong discrimination ability to extract disease spots from cucumber leaves with little human intervention. The high-quality training samples were generated under the operation of rotation, translation, and AR-GAN. With the improvement in convolutional layers, the proposed DICNN exhibited powerful feature extraction and fast convergence ability. Experimental results demonstrated that the proposed approach could effectively identify cucumber leaf diseases. The research explored a feasible way for field agricultural IoTs to timely implement the identification of plant leaf diseases, which is of great practical significance. The future work will focus on identifying more types of cucumber leaf diseases, as well as extending our approaches into the disease recognition of more plants.
Footnotes
Handling Editor: Lyudmila Mihaylova
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The subject is, in part, sponsored by the Key Research and Development Plan of Anhui Province (nos 1804a07020108 and 201904a06020056) and the Natural Science Major Project for Anhui Provincial University (no. KJ2019ZD20).




