Abstract
Background. Housing animals in an enriched environment (EE) enhances behavioral function. However, the mechanism underlying this EE-mediated functional improvement and the resultant changes in gene expression have yet to be elucidated. Objectives. We attempted to investigate the underlying mechanisms associated with long-term exposure to an EE by evaluating gene expression patterns. Methods. We housed 6-week-old CD-1 (ICR) mice in standard cages or an EE comprising a running wheel, novel objects, and social interaction for 2 months. Motor and cognitive performances were evaluated using the rotarod test and passive avoidance test, and gene expression profile was investigated in the cerebral hemispheres using microarray and gene set enrichment analysis (GSEA). Results. In behavioral assessment, an EE significantly enhanced rotarod performance and short-term working memory. Microarray analysis revealed that genes associated with neuronal activity were significantly altered by an EE. GSEA showed that genes involved in synaptic transmission and postsynaptic signal transduction were globally upregulated, whereas those associated with reuptake by presynaptic neurotransmitter transporters were downregulated. In particular, both microarray and GSEA demonstrated that EE exposure increased opioid signaling, acetylcholine release cycle, and postsynaptic neurotransmitter receptors but decreased Na+/Cl−-dependent neurotransmitter transporters, including dopamine transporter Slc6a3 in the brain. Western blotting confirmed that SLC6A3, DARPP32 (PPP1R1B), and P2RY12 were largely altered in a region-specific manner. Conclusion. An EE enhanced motor and cognitive function through the alteration of synaptic activity–regulating genes, improving the efficient use of neurotransmitters and synaptic plasticity by the upregulation of genes associated with postsynaptic receptor activity and downregulation of presynaptic reuptake by neurotransmitter transporters.
Introduction
Enriched environment (EE) is a classical paradigm that has been used extensively to study the effect of rearing animals with a complex combination of physical, cognitive, and social stimulations. 1 Exposure to an EE can facilitate biochemical and histologic changes even in the adult brain, consequently promoting brain functions. 2 An EE elicits various plastic responses, ranging from biochemical changes to neurogenesis, axonal sprouting, and dendritic arborization.3,4 Studies conducted in rodents have convincingly demonstrated that EE exposure increases the production of neurotrophins such as brain-derived neurotrophic factor and nerve growth factor in the cerebral cortex and hippocampus.5,6 Enhanced expression of neurotrophins and immediate early genes, changes in neurotransmitter receptors, and the activation of cAMP response element–binding protein (CREB) after exposure to an EE can affect neural plasticity.7-10 Many reports have shown that EE exposure improves performance in various behavioral tasks, including motor function, learning, and memory, suggesting positive effects for EE exposure as a potential therapeutic strategy for those recovering from brain damage.11-13
Likewise, an EE is known to have beneficial effects on brain functions, including the promotion of neural plasticity and the enhancement of motor and cognitive performance. However, the neurobiological mechanisms underlying this behavioral improvement have not yet been fully elucidated, although a few researchers have focused on hippocampal plasticity following physical exercise. 14 Although changes in gene expression after short-term exposure to an EE have been reported,15,16 little has been investigated regarding the effect of long-term exposure to an EE, which is more compatible with the rehabilitation and related therapies for human patients. Therefore, the purpose of this study was to investigate the underlying changes in the global gene expression patterns associated with EE-mediated improvement of motor and cognitive function. For this purpose, we performed gene set enrichment analysis (GSEA), a powerful and objective method to identify pathways that are affected by specific stimuli, because many biological events are mediated by a battery of proteins rather than a single protein.
Here we demonstrated that EE exposure enables more efficient use of neurotransmitters and enhances synaptic plasticity through the upregulation of genes involved in postsynaptic neurotransmitter receptors and downregulation of those associated with presynaptic neurotransmitter reuptake. Improvements in motor performance and the learning paradigms acquired after long-term exposure to an EE were presumably affected by these mechanisms. According to these results, it can be hypothesized that EE exposure promotes precise motor abilities as well as memory function through characteristic changes in gene expression patterns in the adult brain.
Methods
Animals and Enriched Environment
A total of 32 CD-1 (ICR) mice were randomly assigned to either an EE (n = 16) or standard condition (SC; n = 16) group at 6 weeks of age. The EE mice were housed in a huge cage (86 × 76 × 31 cm3) containing novel objects, such as tunnels, shelters, toys, and running wheels for voluntary exercise, and allowing for social interaction (12-15 mice/cage) for 2 months (Figure 1A), whereas the SC controls were housed for the same duration in standard cages (27 × 22.5 × 14 cm3) without social interaction (3-4 mice/cage; Figure 1B). All animals were housed in a facility accredited by the Association for Assessment and Accreditation of Laboratory Animal Care and were given food and water ad libitum under alternating 12-hour light/dark cycles, according to animal protection regulations. The experimental procedure was approved by the institutional review board. A schematic timeline of this experiment from birth to 14 weeks of age is provided in Figure 1C.
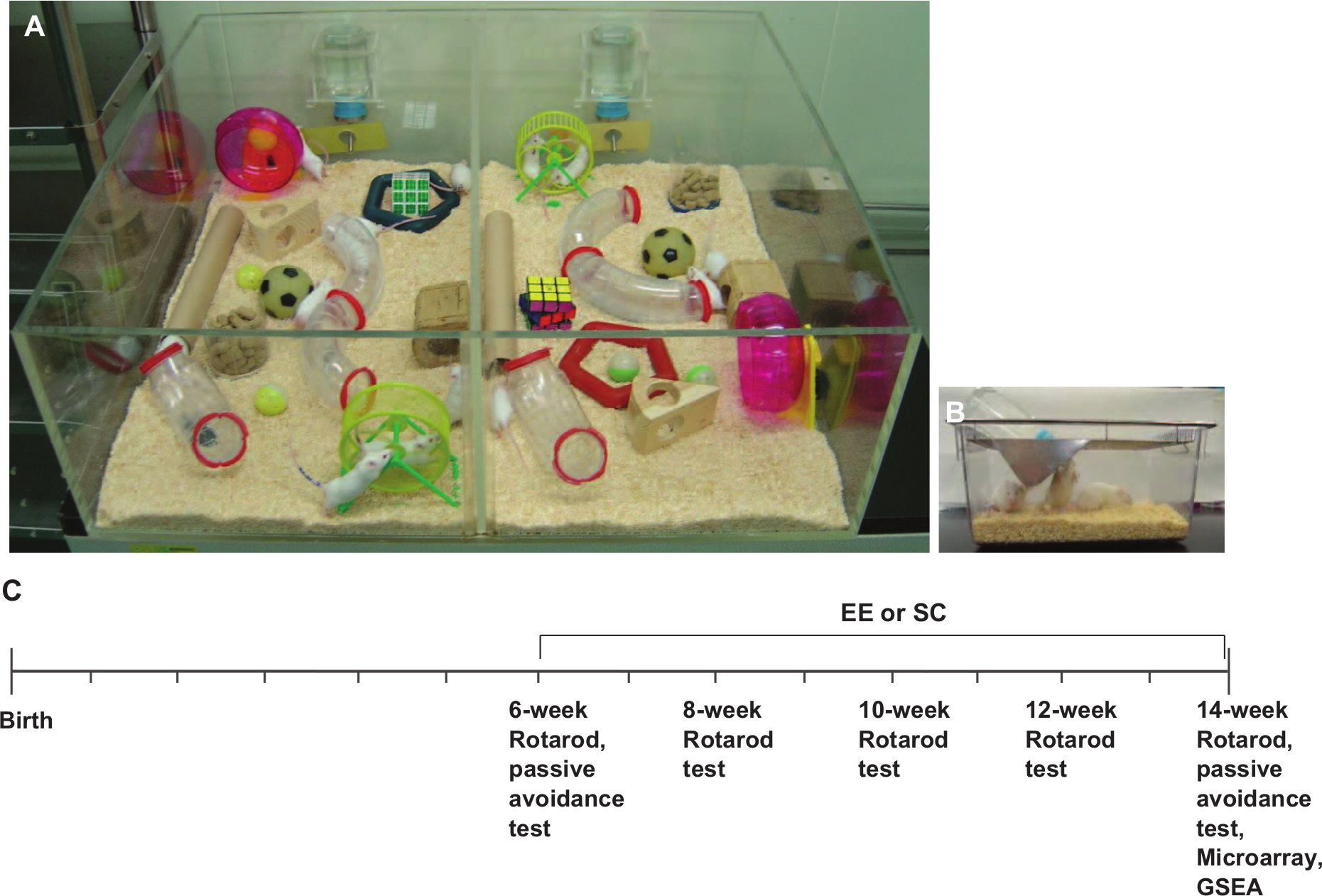

Enriched environment and experimental design: A. CD-1 (ICR) mice were randomly assigned to either an enriched environment (EE; n = 16) or standard conditions (SC; n = 16) starting at 6 weeks of age for 2 months. The EE was provided in a huge cage (86 × 76 × 31 cm3) containing various objects, such as tunnels, shelters, toys, and running wheels for voluntary exercise, and allowed for enriched social interaction (12-15 mice/cage). B. SC comprised a standard cage (27 × 22.5 × 14 cm3) with limited social interaction (3-4 mice/cage). C. A schematic timeline of the experiments.
Rotarod Behavioral Test
A rotarod test was used to assess motor coordination and locomotor function. All animals received a pretreatment performance evaluation at 5 to 6 weeks of age. For this assessment, mice were placed on a rotarod treadmill (Ugo Basile, Comerio, Italy), and the latency to fall, which is the length of time the animals remained on the rolling rod, was measured. Rotarod tests were then performed at 2-week intervals until 8 weeks after the commencement of the treatment at a constant speed (48 rpm) and an accelerating speed (4-80 rpm). The latency period of when the mice fell from the rod was measured twice for each test, and individual tests were terminated at a maximum latency of 300 s. To avoid any stress related to the tests, we conducted the tests gently.
Passive Avoidance Task
To evaluate memory function based on learning to avoid an aversive stimulus, a 2-compartment step-through passive avoidance task (PAT) was conducted.17,18 A Plexiglas shuttle box (41.5 × 21 × 35 cm3) of the PAT apparatus (Jeung Do Bio & Plant Co, Seoul, South Korea) was divided into bright and dark compartments by a wall with a guillotine door. The floor rods in the dark compartment were connected to an electrical stimulator. The training trial of the PAT was performed first. Mice were placed in the bright compartment and allowed to explore for 30 s, at which point the guillotine door was raised to allow the mice to enter the dark compartment. When the mice entered the dark compartment, an electrical foot shock (0.5 mA) was delivered for 2 s. As a baseline study, test sessions were performed to assess short-term working memory (30 minutes) and long-term memory (24 hours) before the mice were assigned to either the EE or the SC group (n = 10 each). The test sessions were then performed 8 weeks after the treatment. The difference (Δ) in the step-through latency of the retention test relative to the preoperative evaluation was calculated. The step-through latency to enter the dark compartment was recorded for up to 300 s.
RNA Preparation
Total RNA was extracted from mouse brain hemispheres using Trizol (Invitrogen Life Technologies, Carlsbad, CA) and purified using RNeasy columns (Qiagen, Valencia, CA) according to the manufacturers’ protocols. After being processed for DNase digestion and clean-up procedures, the RNA samples were quantified, aliquoted, and stored at −80°C until further use. For quality control, RNA purity and integrity were evaluated by denaturing gel electrophoresis and OD 260/280 ratio and analyzed with an Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, California).
Labeling and Purification
Total RNA was amplified and purified using the Ambion Illumina RNA amplification kit (Ambion, Austin, Texas) to yield biotinylated cRNA according to the manufacturer’s instructions. Briefly, 550 ng of total RNA was reverse- transcribed to cDNA using a T7 oligo (dT) primer. Second-strand cDNA was synthesized, transcribed in vitro, and labeled with biotin-NTP. After purification, the cRNA was quantified using an ND-1000 spectrophotometer (NanoDrop, Wilmington, Delaware).
Hybridization and Data Export
Next, 750 ng of labeled cRNA samples were hybridized to each mouse-8 expression bead array for 16 to 18 hours at 58°C, according to the manufacturer’s instructions (Illumina, Inc, San Diego, California). The detection of array signals was carried out using Amersham Fluorolink streptavidin-Cy3 (GE Healthcare Bio-Sciences, Little Chalfont, UK) following the bead array manual. Arrays were scanned with an Illumina BeadArray Reader confocal scanner according to the manufacturer’s instructions.
Microarray Data Analysis
The quality of the hybridization and overall chip performance were monitored by visual inspection of both internal quality control checks and the raw scanned data. Raw data were extracted using the software provided by the manufacturer (Illumina Genome Studio v2009.2, Gene Expression Module v1.5.4). Array data were filtered by the detection value of P < .05 (similar to signal to noise) in at least 50% of the samples (we applied a filtering criterion for data analysis; a higher signal value was required to obtain a detection value of P < .05). Selected probe signal values were transformed by the logarithm function and normalized by the quantile method. Comparative analyses between the test group and control group were carried out using the local-pooled-error test and fold change. The false discovery rate was controlled by adjusting P values using the Benjamini-Hochberg algorithm. The differentially expressed gene probes were filtered with the adjusted P < .05 of a local-pooled-error test and with |fold change| ≥2. Hierarchical clustering analyses were performed for filtered probes using complete linkage and Euclidean distance as measures of similarity. Biological gene ontology (GO) analysis for the filtered differentially expressed gene probes was performed using PANTHER 6.1 (http://www.pantherdb.org/panther/ontologies.jsp). Fisher exact test was applied to determine the affected pathways.
Gene Set Enrichment Analysis
To explore the differentially regulated pathways, we performed GSEA using the microarray data, as described previously, using a GSEA program (http://www.broad.mit.edu/gsea/msigdb/index.jsp).19-21 We used C2 curated gene sets (http://www.broadinstitute.org/gsea/msigdb/collections.jsp#C2, version 3), the sizes of which ranged from 10 to 1000 genes. A P value <.05 was considered significant.
Quantitative Real-Time Reverse Transcription-Polymerase Chain Reaction (qRT-PCR)
Differentially expressed genes of interest were selected for the validation of the microarray results by qRT-PCR; 1 µg of purified total RNA was used as a template to generate the first strand cDNA using a cDNA kit (Invitrogen Life Technologies, Carlsbad, California). Then, 2 µL of cDNA in a total volume of 20 µL was used in the following reaction. The qRT-PCR was performed in triplicate on a LightCycler 480 (Roche Applied Science, Mannheim, Germany) using the LightCycler 480 SYBR Green master mix (Roche Applied Science, Mannheim, Germany), and the thermocycler conditions were as follows: amplifications were performed starting with a 300-s template preincubation step at 95°C, followed by 45 cycles at 95°C for 10 s, 53°C for 10 s, and 72°C for 10 s. The melting curve analysis began at 95°C for 5 s, followed by 1 minute at 60°C. The specificity of the produced amplification product was confirmed by the examination of a melting curve analysis and showed a distinct single sharp peak with the expected Tm for all samples. A distinct single peak indicates that a single DNA sequence was amplified during qRT-PCR. The primers were as follows: mouse Drd1a, 5′-GCTTTTACA TCCCCGTAGCC-3′ and 5′-AGCCAGCAGCACACGA ATAC-3′; mouse Slc6a3, 5′-CAACAACACCTGGAACA GCC-3′ and 5′-AATGCCACGACTCTGATGGA-3′; mouse Slc6a4, 5′-TCTTCACGGTGCTTGGCTAC-3′ and 5′-CGATGAGCACAAACCATTCC-3′; mouse Ppp1r1b, 5′-AGGCCTCTCCACATCAGAGA-3′ and 5′-CCTGGCT GTCTTCTTCTTCG-3′; mouse P2ry12, 5′-GAAGGAGA GCACCCTATGGC-3′ and 5′-AGCACCTCAGCATGCTT GTC-3′; mouse Pdyn, 5′-CCATCCCAGAATCCAGAGA A-3′ and 5′-CCAGGGTAGGGTGCATAAGA-3′; mouse GAPDH, 5′-AACTTTGGCATTGTGGAAGG-3′ and 5′-ACACATTGGGGGTAGGAACA-3′. GAPDH (glyceraldehyde 3-phosphate dehydrogenase) was used as the internal control. The expression of each gene of interest was obtained using the 2−ΔΔCt method.
Western Blot Analysis
To confirm the expression of P2ry12, Drd1a, Slc6a3, Pdyn, and Ppp1r1b levels by long-term exposure to an EE or SC, 30 µg of extracted proteins were dissolved in a sample buffer, boiled for 5 minutes, and separated on a 10% sodium dodecyl sulfate (SDS) reducing gel. The separated proteins were then equally loaded and transferred onto polyvinylidene difluoride membranes (Amersham Pharmacia Biotech, Little Chalfont, UK) using a transblot system (Bio-Rad, Hercules, California). Blots were blocked for 1 hour in Tris-buffered saline (TBS) containing 5% nonfat dry milk (Bio-Rad, Hercules, California) at room temperature, washed 3 times with TBS, and incubated overnight at 4°C with the following antibodies: antidopamine receptor D1 antibody (DRD1A, 1:1000; Abcam, Cambridge, UK), antidopamine transporter antibody (SLC6A3, 1:1000; Abcam, Cambridge, UK), anti-DARPP-32 antibody (PPP1R1B, 1:1000; Cell Signaling Technology, Beverly, MA), anti-P2Y12 antibody (P2RY12, 1:1000; Abcam, Cambridge, UK), antiprodynorphin (anti-PDYN) antibody (PDYN, 1:1000; Abcam, Cambridge, UK), and anti-GAPDH antibody (GAPDH, 1:1000; Cell Signaling Technology, Beverly, Massachusetts). The next day, the blots were washed 3 times with TBS plus Tween 20 and incubated for 1 hour with horseradish peroxidase–conjugated secondary antibodies (1:2000; Santa Cruz, California) at room temperature. After being washed 3 times with TBS plus Tween 20, the protein was visualized with an enhanced chemiluminescence detection system (Amersham Pharmacia Biotech, Little Chalfont, UK).
Statistical Analysis
All data were expressed as means ± standard error of the mean, and statistical analyses were conducted using the premier vendor for Statistical Package for Social Sciences (SPSS), Predictive Analytics Software (PASW) Statistics version 18.0. Student t test was used for the comparison of continuous variables between 2 groups, and analysis of variance (ANOVA) followed by multiple comparisons with Bonferroni or Tukey methods was used for the statistical analysis of variables among more than 2 groups. A P value <.05 was considered statistically significant.
Results
Rotarod Test Showed That EE Exposure Improved Motor Function
To determine whether EE exposure improves motor function, rotarod tests were performed with constant (48 rpm) and accelerating (4-80 rpm) paradigms every 2 weeks after exposure to either an EE or SC (n = 16 per group). In the pretreatment evaluation, no statistical differences were seen between the groups. EE-induced improvement of rotarod performance relative to SC was evident at 2 weeks posttreatment. A significant improvement was observed starting at 4 weeks posttreatment for the constant-speed condition (t = 2.527, P = .014; Figure 2A) and at 2 weeks posttreatment for the accelerating speed condition (t = 3.906, P < .001; Figure 2B). This improved motor function was maintained throughout the study period. Finally, at 8 weeks after treatment, the mean rotarod latencies of the EE mice increased significantly to 162.0 ± 23.1 s for the constant-speed condition (t = 2.448, P = .017; Figure 2A) and to 179.4 ± 17.6 s for the accelerating speed condition (t = 2.974, P = .004; Figure 2B), compared with the rotarod latencies of the SC mice (84.1 ± 21.8 and 113.7 ± 13.4 s, respectively).
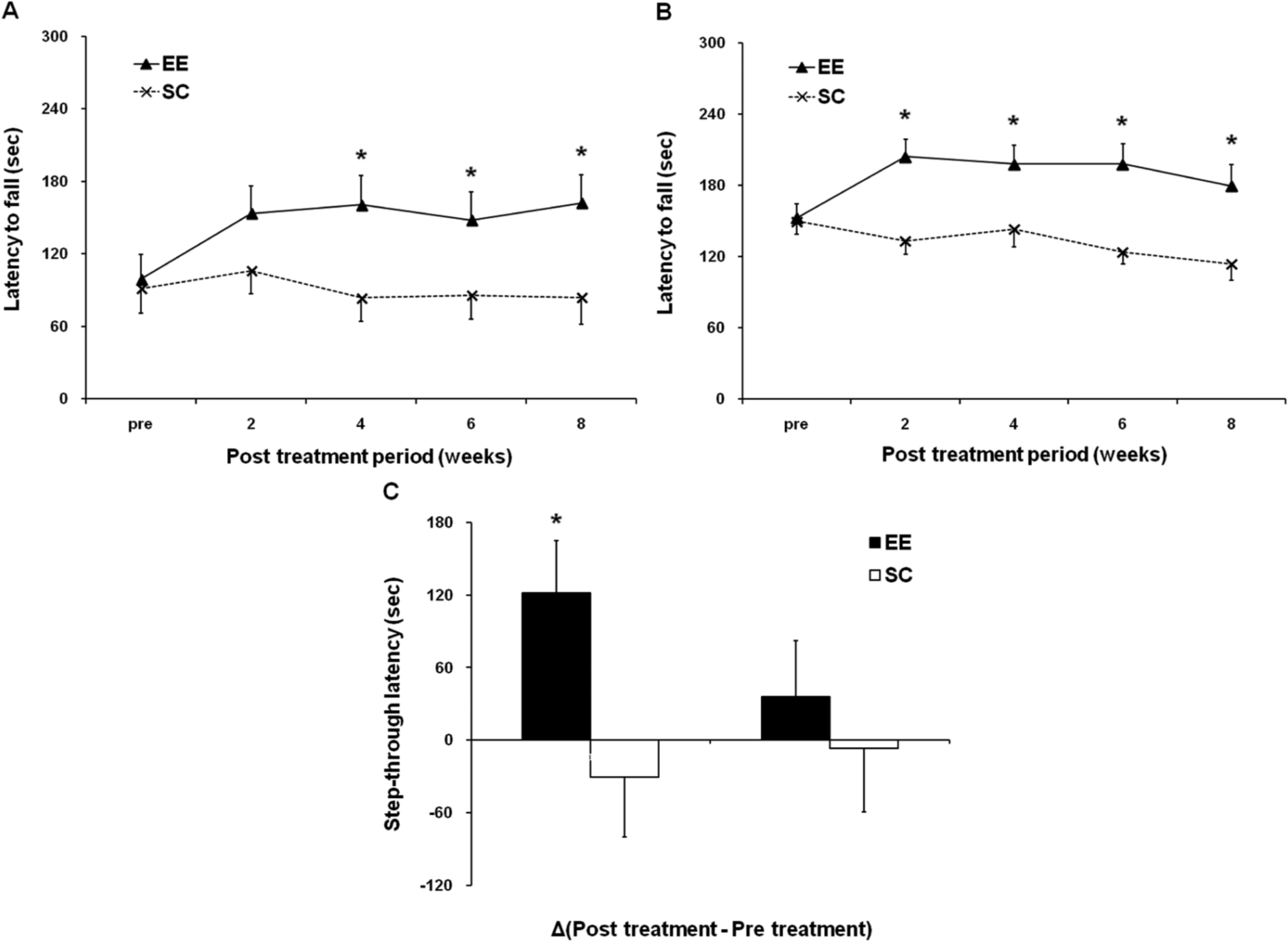
Rotarod test and passive avoidance test show that an enriched environment (EE) improves neurobehavioral function. To determine whether an EE can improve motor function, a rotarod test was performed at constant (48 rpm) and accelerating (4-80 rpm) paradigms every 2 weeks after exposure to either an EE or standard conditions (SC; n = 16 per group). A. For the constant speed rotarod test, a significant improvement was observed starting 4 weeks after exposure to an EE (P < .05). After an 8-week exposure to an EE, the mean rotarod latency of the EE mice was significantly higher (EE vs SC: 162.0 ± 23.1 vs 84.1 ± 21.8 s; P < .05). B. For the accelerating speed rotarod test, a significant improvement was observed starting 2 weeks after the exposure to an EE (P < .05). After an 8-week exposure to an EE, the mean rotarod latency of the EE mice was significantly higher (EE vs SC: 179.4 ± 17.6 vs 113.7 ± 13.4 s; P < .05). C. To determine whether an EE improves cognitive function, passive avoidance tests were performed before and at 8 weeks after exposure to either an EE or SC (n = 10 per group). EE-induced improvement of the short-term working memory test 30 minutes after an aversive stimulus relative to the pretreatment evaluation baseline (Δ = 121.4 ± 43.3 s) was significantly evident at 8 weeks posttreatment when compared with the SC group (Δ = 30.4 ± 46.5 s; P < .05). P < .05.
Passive Avoidance Test Also Showed That an EE Improved Cognitive Function
To determine whether an EE improves cognitive function, passive avoidance tests were performed before and 8 weeks after exposure to either an EE or SC (n = 10 per group). In the pretreatment evaluation, no statistical differences were seen between the groups. However, the EE-induced improvement of retention test performance at 30 minutes after an aversive stimulus relative to that in the pretreatment evaluation (Δ = 121.4 ± 43.3 s) was significantly evident at 8 weeks posttreatment as compared with the difference for the SC group (Δ = 30.4 ± 46.5 s; t = −2.388; P = .028). However, the retention test 24 hours after an aversive stimulus did not exhibit a significant improvement after EE exposure relative to that in the pretreatment evaluation (Δ = 35.9 ± 49.5 s) as compared with the difference in the SC group (Δ = 6.9 ± 52.2 s), although the result suggested a modest tendency toward improved cognitive function after EE exposure. This suggests that an EE improves short-term working memory rather than long-term memory (Figure 2C).
Microarray Data Revealed Global Changes in Gene Expression Patterns in the Brain of Mice Exposed to an EE
To investigate the underlying mechanisms associated with EE-induced behavioral improvement, we performed microarray analyses of total RNA extracts from mouse brain hemispheres. Hierarchical clustering using the significantly regulated genes showed that the gene expression patterns were similar among samples within the same group but were more than 2-fold different between the groups (Figure 3A). Hierarchical clustering analysis showed that the gene expressions in the EE group were distinct from those of the control group in that 17 genes were significantly upregulated and 31 genes were significantly downregulated by more than 2-fold in the EE group compared with the control group (P < .05; Figure 3B). To analyze further the differences in gene expression between the groups, we next categorized genes that exhibited significantly altered expression levels after exposure to an EE according to their biological functions. The analysis using the GO database revealed that the expression levels of 11 genes involved in signal transduction; 9 genes in nucleoside, nucleotide, and nucleic acid metabolism; 9 genes in developmental processes; 8 genes in neuronal activities; and 6 genes in transport pathways were significantly altered after exposure to an EE (Figure 3C). To determine the statistical significance of these changes, we performed Fisher’s exact test. “Neuronal activities” was the only category shown to be statistically regulated by an EE (P < .05; Table 1), suggesting that an EE may preferentially regulate neuronal activity genes in brain tissue. Furthermore, 8 genes involved in neuronal activities were found to be differentially regulated. Among these genes, there were significant increases in Drd1 (dopamine receptor D1A, 2.3-fold), Pdyn (PDYN, 2.6-fold), Ppp1r1b (protein phosphatase 1, regulatory subunit 1B, 2.1-fold), and P2ry12 (purinergic receptor P2Y, G protein-coupled 12, 2.9-fold) after EE exposure (Supplementary Table). In contrast, a drastic decrease was observed in Slc6a3 (dopamine transporter, 7.4-fold) as well as Slc6a4 (serotonin transporter, 3.8-fold) after EE exposure, raising the possibility that the presynaptic reuptake of these neurotransmitters might be reduced by long-term exposure to an EE (Supplementary Table).
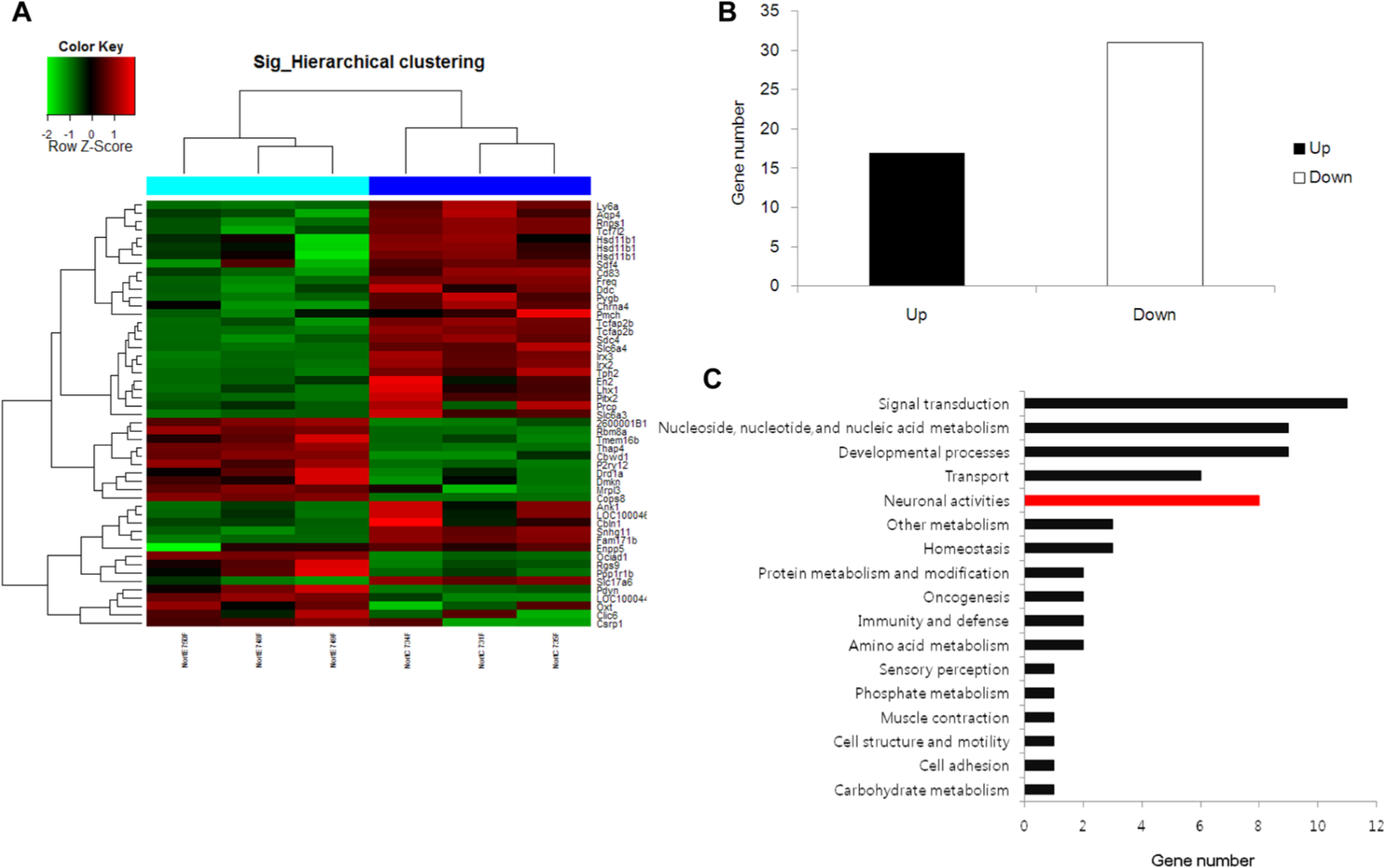
Microarray data reveal the global change of gene expression patterns in the brains of mice exposed to an enriched environment (EE). After an 8-week exposure to either an EE or standard conditions (SC), CD-1 (ICR) mice were euthanized. Total RNA was isolated from the whole brains and subjected to microarray analysis. Genes that demonstrated significantly different expression levels between the EE and SC groups by more than 2-fold were regarded as significantly regulated genes and used for further analysis (P < .05). A. Hierarchical clustering using the significantly regulated genes shows that gene expression patterns were similar among samples within the same group but different between the groups (more than 2-fold). B. The number of up- or downregulated genes differed in the intact brain hemispheres following EE exposure. C. The significantly regulated genes were categorized into gene sets, which were predetermined based on the functions of the genes. The absolute numbers are shown. D. The values normalized to the sizes of gene sets are shown. The categories that had a significantly higher number of significantly regulated genes than the other categories are marked in red.
Genes That Were Significantly Different Between EE and SC Groups as Classified by Biological Function. a
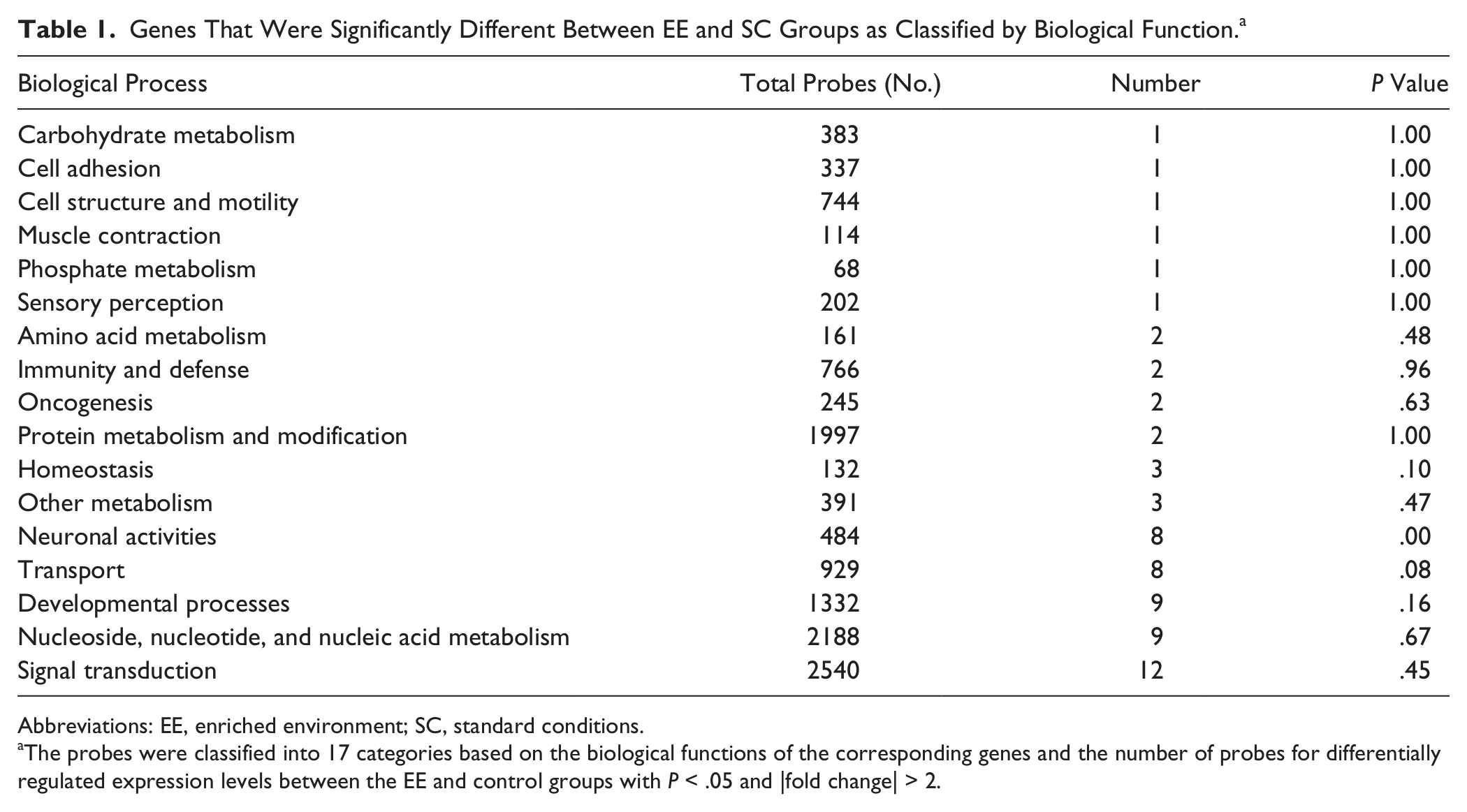
Abbreviations: EE, enriched environment; SC, standard conditions.
The probes were classified into 17 categories based on the biological functions of the corresponding genes and the number of probes for differentially regulated expression levels between the EE and control groups with P < .05 and |fold change| > 2.
GSEA Showed Genes Involved in Synaptic Plasticity Are Regulated by Long-term Exposure to an EE
To determine more thoroughly the pathways selectively regulated by exposure to an EE compared with the control cages, we adopted GSEA as previously described.19-21 GSEA allowed us to determine the selectively regulated pathways among 3272 gene sets, whereas the GO analysis we performed above used only 17 categories. Compatible with the GO analysis results, GSEA revealed that several pathways associated with synaptic plasticity were significantly altered by EE exposure (Figure 4). GSEA showed that the gene sets involved in synaptic plasticity were regulated by exposure to an EE. Namely, genes associated with transmission across chemical synapses, opioid signaling, neurotransmitter receptor binding and downstream transmission in the postsynaptic cells, G protein activation, kainate receptor activation, and depolarization of the presynaptic terminal triggering the opening of calcium channels and the acetylcholine release cycle were upregulated, whereas those linked with dopamine neurotransmitter release and Na+/Cl−-dependent transporters were downregulated.
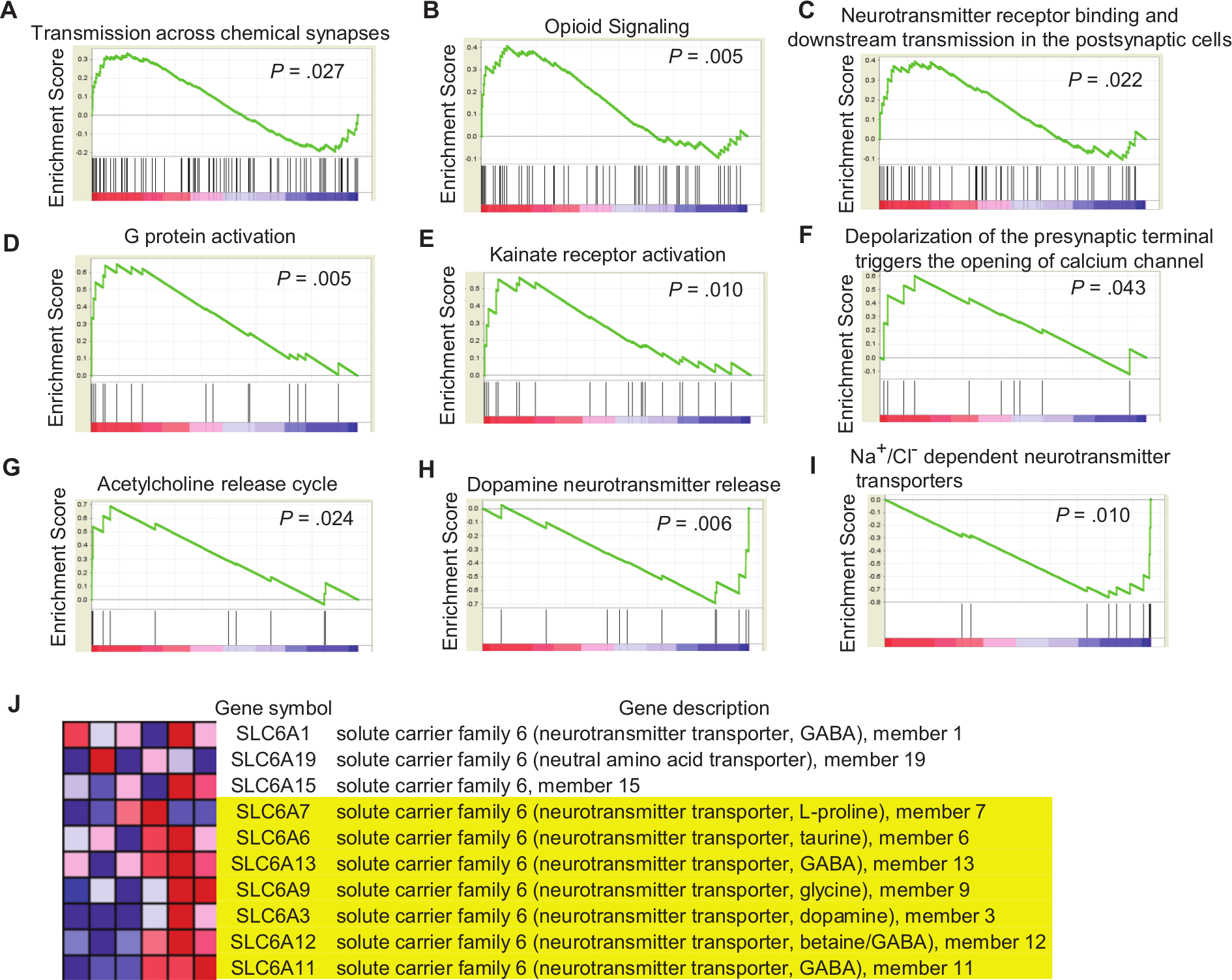
Gene set enrichment analysis (GSEA) shows that genes involved in synaptic plasticity are regulated by long-term exposure to an enriched environment (EE). After mice had an 8-week exposure to either an EE or standard conditions (SC), total RNA isolated from the whole brains were subjected to microarray analysis. GSEA was performed using the microarray data. A-G. GSEA shows significantly upregulated gene sets in the EE group compared with the SC group. H-I. GSEA shows significantly downregulated gene sets in the EE group compared with the SC group. J. Leading-edge analysis of “Na+/Cl−-dependent neurotransmitter transporters” gene sets shows that 7 genes (highlighted in yellow), including Slc6a3, comprise the leading edge responsible for the global downregulation of the gene set of Na+/Cl−-dependent neurotransmitter transporters.
The global expression levels of genes associated with the transmission of neurotransmitters across chemical synapses were significantly higher in the EE group compared with the control group, suggesting that an EE may activate the transfer of information across chemical synapses (Figure 4A; P = .027). In agreement with this, genes involved in opioid signaling were globally upregulated in the EE group, implying that an EE can activate opioid signaling (Figure 4B; P = .005). This GSEA result of opioid signaling is compatible with previous reports that showed, based on biochemical and functional analyses, that exercise activates opioid production and signaling.22-24 In line with this enhanced transmission across chemical synapses, genes involved in neurotransmitter receptor binding and downstream transmission in postsynaptic cells as well as G protein activation were globally upregulated (Figures 4C and 4D; P = .022 and .005, respectively), suggesting that EE exposure may increase postsynaptic sensitivity. Compatible with this general postsynaptic receptor activation by an EE, the global expression levels of genes involved in kainate receptor activation were significantly higher in the EE group than in the controls (Figure 4E; P = .010), suggesting that an EE may activate non-NMDA (N-methyl D-aspartate) glutamate receptors as well.
Although EE exposure consistently upregulated genes related to transmission across synapses and postsynaptic receptor activation, genes involved in neurotransmitter release did not show uniform change after long-term exposure to an EE. Genes involved in the depolarization of presynaptic terminal triggering, for example, the opening of the calcium channel, were significantly, but only marginally, upregulated in the EE group compared with the controls (Figure 4F; P = .043), suggesting that EE exposure may activate the release of some neurotransmitters from presynaptic terminals. Genes associated with acetylcholine and dopamine release were significantly upregulated and downregulated, respectively (Figure 4G, P = .024; Figure 4H, P = .006), suggesting that an EE can increase acetylcholine release and reduce dopamine release after long-term exposure to it.
It is interesting to note that genes associated with Na+/Cl−-dependent transporters were globally downregulated in the EE group compared with the controls (Figure 4I; P = .010). Given that the activity of a neurotransmitter is terminated by reuptake at presynaptic neurons, the downregulation of genes associated with Na+/Cl−-dependent transporters can enhance the activity of neurotransmitters at the synaptic level. Leading-edge analysis showed that this global downregulation of Na+/Cl−-dependent transporters was associated with the selective downregulation of 7 genes, including Slc6a3, which is responsible for the presynaptic reuptake of dopamine (Figure 4J). The Slc6a3 gene was also downregulated in the microarray results. Taken together, the GSEA results suggested that an EE may enhance synaptic activity by increasing postsynaptic sensitivity and reducing presynaptic neurotransmitter reuptake.
Intersection of Microarray Data and GSEA Suggest That Neurotransmitter Bioactivity Is Increased in the Synaptic Level by Long-term Exposure to an EE
To investigate the novel mechanisms that were regulated by exposure to an EE, we interpreted both microarray data and GSEA results. Among the biological functions of interest in the microarray data, PDYN, Ppp1r1b, and Slc6a3 in the neuronal activity category were statistically regulated by EE exposure. These genes contributed to the effects on opioid signaling and Na+/Cl−-dependent transporters in the GSEA results. The PDYN and Ppp1r1b genes involved in opioid signaling were globally upregulated in the EE group.22-24
Among the genes involved in Na+/Cl−-dependent neurotransmitter transporters, the Slc6a3 gene is a dopamine transporter that is a member of the Na+/Cl−-dependent neurotransmitter transporter family. Slc6a3 is an integral membrane protein that removes dopamine from the synaptic cleft and deposits it into surrounding cells, thus terminating the signal of the neurotransmitter. The Slc6a3 gene involved in Na+/Cl−-dependent neurotransmitter transporters was globally downregulated in the EE group; downregulating the expression of Slc6a3 leads to the increased synaptic occupancy of dopamine. It is interesting to note that the GSEA results demonstrated that dopamine neurotransmitter release was significantly downregulated after long-term exposure to an EE. Among the genes related to dopamine release, tyrosine hydroxylase (TH) and dopa decarboxylase (DDC) genes were downregulated by an EE. In the intersection of microarray and GSEA results, we suggest that dopamine biosynthesis was reduced and the bioactivity of neurotransmitters in the synaptic cleft was increased.
Validation of Microarray Data Using qRT-PCR and Western Blotting
We next validated the microarray data using qRT-PCR and Western blotting. We first performed qRT-PCR on the genes that had a significant change of more than 2-fold in the microarray and found that the RT-PCR data were compatible with those from the microarray (Figure 5A). The qRT-PCR confirmed that dopamine receptor Drd1a, protein phosphatase 1 regulatory subunit 1B Ppp1r1b, and endogenous opioid peptide Pdyn significantly increased after long-term exposure to an EE, whereas Na+/Cl−-dependent neurotransmitter transporters Slc6a3 and Slc6a4 significantly decreased in a pattern similar to the microarray data (Figure 5A).
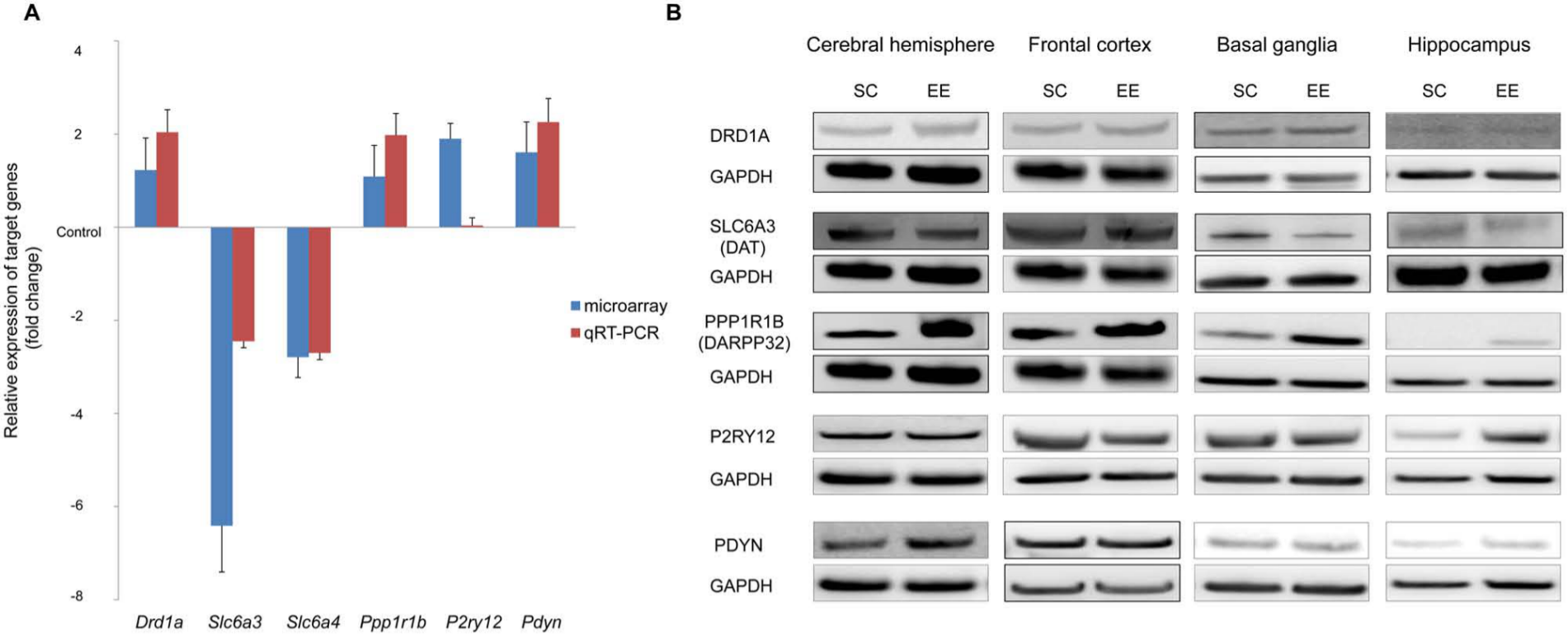
Validation of microarray results by quantitative real-time reverse transcription-polymerase chain reaction (qRT-PCR) and Western blotting: Total RNA (A) and protein (B) were isolated from the brains of mice after an 8-week exposure to either an enriched environment (EE) or standard conditions (SC) and were subjected to qRT-PCR (A) or Western blotting on those genes with a significant change of more than 2-fold in the microarray. A. The qRT-PCR confirmed that dopamine receptor Drd1a, protein phosphatase 1 regulatory subunit 1B Ppp1r1b, and endogenous opioid peptide Pdyn significantly increased after long-term exposure to an EE, whereas Na+/Cl−-dependent neurotransmitter transporters Slc6a3 and Slc6a4 significantly decreased in a similar pattern to the microarray data. B. Western blotting showed that dopamine receptor DRD1A protein increased in the basal ganglia, whereas dopamine transporter SLC6A3 protein decreased in the basal ganglia after exposure to an EE. In addition, cAMP-regulated neuronal phosphoprotein, DARPP32 (PPP1R1B) increased in all analyzed regions. The protein level of G protein-coupled purinergic receptor P2RY12 increased in the hippocampus after long-term exposure to an EE.
We next determined the protein levels of these genes (Drd1a, Slc6a3, Ppp1r1b, P2ry12, and Pdyn) in specific regions of the brain such as the frontal cortex, basal ganglia, and hippocampus (Figure 5B). Western blotting showed that dopamine receptor DRD1A protein increased in the basal ganglia, whereas dopamine transporter SLC6A3 protein decreased in the basal ganglia after exposure to an EE. It is important to note that cAMP-regulated neuronal phosphoprotein DARPP32, encoded by the Ppp1r1b gene, drastically increased in all analyzed regions, including the frontal cortex, basal ganglia, and hippocampus by sustained exposure to an EE. The protein level of G protein-coupled purinergic receptor P2RY12 increased in the hippocampus after long-term exposure to an EE. The endogenous opioid peptide PDYN increased in the cerebral hemisphere, although this increase was not significant in the regional analysis.
Discussion
To explore the molecular mechanisms of the therapeutic effects of an EE, we used microarray analysis and GSEA-containing probe sets representing approximately 10 000 genes to analyze the levels of gene transcripts in mouse brain hemispheres after 2 months of exposure to an EE in comparison with controls kept in standard cages. To our knowledge, this is the first study to investigate such mechanisms by using GSEA to explore global gene expressions and novel pathways that were differentially regulated after long-term exposure to an EE.
Rampon et al 15 previously described the effects of an EE on the gene expression in the brain using oligonucleotide microarrays. They found that the expressions of genes associated with neuronal structure, synaptic plasticity, and transmission were significantly changed by EE exposure. Unlike the short-term intermittent exposure to an EE in their study, we focused on the long-term chronic exposure to an EE, which is more usual in the field of rehabilitation and related therapies. When we compared our results with those of Rampon et al, most of the genes that were changed by EE exposure in the Rampon group’s study were not significantly regulated by the EE in our study, suggesting that chronic exposure to an EE shows a differential change in gene expression or desensitizes the upregulation of gene expression. This difference may be attributable to the difference in EE exposure time. In the Rampon et al study, the EE was provided for a relatively short period such as 3 hours, 6 hours, 2 days, or 14 days, whereas we provided the long-term exposure to an EE for 2 months. In addition, the EE was provided for only 6 h/d in the Rampon study, whereas we provided the EE for 24 h/d. Thus, the longest EE exposure time in their study was 3.5 days (14 days × 6 h/d), whereas we provided 2 full months. Given that the gene expression patterns are significantly different between early changes (3 and 6 hours) and late changes (2 and 14 days) after exposure to an EE, it is not surprising that the genes significantly regulated by the EE in the study by Rampon et al were not altered by the EE in our study.
Here we adopted GSEA for the first time to find specific pathways that are affected by an EE. The microarray-based determination of gene expression patterns has been used to find pathways that are affected by EE exposure.15,16 However, because a post hoc examination of top-ranking genes could be subjective and prone to bias, finding pathways altered by an EE can be properly conducted only when the analysis is performed by unbiased scientists who know the function of the genes. In our GSEA analysis, we used a priori defined gene sets to analyze the data to minimize this analyzer-dependent potential bias. Although GSEA is a powerful method to find pathways activated or inactivated by a specific factor, it also has limitations; GSEA is an in silico analysis and is not a definitive study to determine the activation or inactivation of pathways.
In the present study, GSEA revealed that several pathways associated with synaptic plasticity were significantly altered by the EE. As we already described in the results, gene sets expression associated with chemical synaptic transmission, opioid signaling, postsynaptic neurotransmitter receptor binding and G protein activation, kainate receptor activation, the opening of the presynaptic calcium channel, and acetylcholine release were upregulated, whereas those associated with dopamine release and neurotransmitter transporters were downregulated.
The expression of genes related to transmission across chemical synapses was upregulated. The chemical synapses enable cell-to-cell communication via the secretion of neurotransmitters. These chemical agents, released by the presynaptic neurons, produce secondary current flow in postsynaptic neurons by activating specific receptor molecules. Neurotransmitter receptors are proteins that are embedded in the plasma membranes of postsynaptic cells and have an extracellular neurotransmitter binding site that detects the presence of neurotransmitters in the synaptic cleft. G protein activation is important for transducing transmitter binding into postsynaptic responses. The neurotransmitter receptors have an intracellular domain that indirectly affects channels through the activation of intermediate molecules called G proteins. Neurotransmitter binding to these receptors activates G proteins that make intracellular messengers. Thus, G proteins can be thought of as transducers for signal transmission that couple neurotransmitter binding to the regulation of postsynaptic ion channels and receptors. 25 Activation of kainate receptor, a non-NMDA glutamate receptor, also has a role in synaptic plasticity, affecting the postsynaptic cell firing and the presynaptic neurotransmitter release underlying modulation of excitatory synaptic transmission in the hippocampus. 26 Transmitter agents are released presynaptically in units, reflecting their storage within synaptic vesicles. These vesicles discharge their contents into the synaptic cleft when the presynaptic depolarization generated by the invasion of an action potential opens voltage-gated calcium channels, allowing Ca2+ to enter the presynaptic terminal.
In addition, the results of our study demonstrated the EE-driven changes in gene expression patterns and neural plasticity in the neurotransmitter system, including monoamine transporters. These results may provide potentially novel and interesting insights into the effects of environmental stimulation such as voluntary exercise and social interaction. Noradrenaline, dopamine, and serotonin were suggested in previous studies to be implicated in brain plasticity and neurobehavioral functions.27-29 Thus, the monoamine neurotransmitter system could be a reasonable candidate for mediating the effects of an EE. For example, the inhibition of the Na+/Cl−-dependent transporters, such as the norepinephrine, dopamine, and serotonin transporters, is known to be the major therapeutic mechanism for the downregulation of monoamine reuptake. Consequently, the downregulation of reuptake transporters can lead to enhanced activity of monoamine neurotransmitters at the synaptic level because their actions are not terminated by reuptake into the presynaptic nerve terminal after the release of neurotransmitters into the synaptic cleft. Among the genes involved in Na+/Cl−-dependent neurotransmitter transporters, the Slc6a3 gene was significantly regulated by EE exposure according to both microarray and GSEA results. Slc6a3, a dopamine transporter, is an integral membrane protein that removes dopamine from the synaptic cleft and deposits it into the surrounding cells, thus terminating the signal of the neurotransmitter. 30 Alterations in Slc6a3 activity can influence the synaptic occupancy of dopamine. The interventions such as physical exercise and EE exposure that downregulate Slc6a3 expression may, therefore, lead to behavioral improvement by increasing the synaptic occupancy of dopamine in a manner similar to pharmacological treatment using reuptake inhibitors. In the same manner as dopamine transporters, we suggest that other Na+/Cl−-dependent neurotransmitter transporters of solute carrier family 6, which are serotonin, GABA, glycine, L-proline, and taurine transporters, are downregulated after long-term exposure to an EE, thereby enhancing the synaptic plasticity of the neurotransmitters.
It is interesting to note that the GSEA results showed that genes associated with dopamine neurotransmitter release were significantly downregulated after long-term exposure to an EE. TH and DDC are related to dopamine neurotransmitter release. TH as an enzyme is responsible for the rate-limiting step in dopamine biosynthesis. In addition, the expressions of TH and Slc6a3 proteins have been shown to be closely related. For example, Slc6a3 knockout mice have been shown to express reduced striatal TH protein. 31 Although the precise link between TH and Slc6a3 is unclear, it has been speculated that the increased synaptic bioavailability of dopamine, through the downregulation of Slc6a3, may lead to increased activation of the dopamine autoreceptor, leading to the downregulation of TH. 32 Similar to the action of selective serotonin reuptake inhibitors and other monoamine reuptake transporter blockers used as a first-line choice of antidepressive agents, an EE combining physical, cognitive, and social stimulation might be a potential neurotherapeutic strategy for behavioral improvement and functional recovery.
The GSEA results suggest that opioid signaling is significantly upregulated by EE exposure. The expression level of PDYN, which is involved in opioid signaling, was significantly higher in the EE group than in controls. Opioids are thought to play an important role as neuromodulators and neurotransmitters. Endogenously produced opioids are released during exercise and contribute to the feeling of well-being following exercise. 24 In addition, one of the major precursor molecules for the endogenous opioid peptides is coded by PDYN. Dynorphin is widely expressed in the brain. After running, increased dynorphin mRNA in the medial caudate putamen was found. 22 Dynorphin transcription is upregulated by phosphorylated CREB. 33 The phosphorylation state of CREB in the hippocampus also plays an important role in learning and memory processes, which parallel CREB expression. The activation of transcription factor CREB can be mediated through an EE. 34 Ppp1r1b, also known as dopamine- and cAMP-regulated neuronal phosphoprotein (DARPP-32), is related to opioid signaling as well.
Several factors may be involved in the EE-mediated improvement of neurobehavioral functions. An EE can lead to increased exercise, motor skill training and learning, and social interactions. Given that exercise alone can activate opioid signaling and improve synaptic plasticity,22-24,35 we can assume that the neurobehavioral improvement is at least in part a result of exercise. However, learning and social interactions also play an important role in the recovery of neurobehavioral function, as suggested by a previous report that the recovery of motor function in rats with cerebral infarction was improved in the following order: rats in an EE, those in the same size of cage without activity-stimulating equipment, and those in individual cages with access to a running wheel.2,36 The EE, social interaction, and physical activity were determinants of functional outcome after cerebral infarction.2,36 Thus, the improvement of neurobehavioral functions in our current study could be attributable to the combination of physical exercise, learning, and social interactions.
When the protein levels of the defined genes regulated by the EE were determined in specific regions of the brain using Western blotting, dopamine transporter SLC6A3 protein characteristically decreased in the basal ganglia where the dopamine transporter is largely distributed, whereas DARPP32, a key factor in the integration of dopamine and glutamate, increased in all analyzed regions, including the frontal cortex and basal ganglia where the DARPP32 was abundantly found. 37 This result suggests that EE-induced downregulation of presynaptic dopamine transporter leads to increase in the neurotransmitter in the synaptic level, and upregulation of DARPP32 as a major target for the dopamine plays a pivotal role in motor and cognitive functions. 38 In addition, dopamine receptor DRD1 increased in the basal ganglia, and G protein-coupled purinergic receptor P2RY12 increased in the hippocampus, where the proteins were largely distributed, mediating the signaling functions in neurotransmission and regeneration.39,40 It is possible that the upregulation of P2RY12 might be associated with enhanced memory function as suggested by the result of the passive avoidance test, although this possibility needs further investigation. The endogenous opioid peptide PDYN also increased in the cerebral hemisphere, although this increase was not significant in the regional analysis that included the frontal cortex, basal ganglia, or hippocampus. Given that the level of PDYN is often regulated in the hypothalamus,41-43 it is possible that the exposure to an EE increased PDYN mainly in the hypothalamus. This EE-induced upregulation of dynorphin is compatible with previous reports that exercise or an EE increases the level of endogenous opioids,22-24 and the loss of an EE increases vulnerability to cocaine addiction.44,45
In conclusion, behavioral tests showed that an EE enhanced motor and cognitive functions in mice. The analysis of the gene expression pattern changes in their brains revealed that this EE-directed improvement of motor and cognitive functions is associated with the alteration of synaptic activity–regulating genes. In other words, the genes associated with postsynaptic receptor activity were upregulated and the presynaptic neurotransmitter transporters such as dopamine transporters were downregulated, potentially leading to the more efficient use of neurotransmitters and enhanced synaptic plasticity. However, the exact mechanisms underlying this EE-mediated improvement await further investigations, including the ultrastructural examination of the synaptic structural changes and the postsynaptic and glial transporters. Further studies should also be performed to elucidate whether EE exposure enhances the bioactivity of neurotransmitters at the synaptic level, increases acetylcholine release, and reduces dopamine release.
Footnotes
Authors’ Note
M-YL and JHY contributed equally to this work: M-YL, conception and design, collection and/or assembly of data, and manuscript writing; JHY, conception and design, and collection and/or assembly of data. The other authors contributed as follows: JYK and JHS, collection and/or assembly of data; ESP and CHK, administrative support, data analysis, and interpretation; HK, data analysis and interpretation, manuscript writing, and final approval of manuscript; S-RC, conception and design, data analysis and interpretation, manuscript writing, and final approval of manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants from the 2011 Chyung Ki Lee Research Fund and the National Research Foundation (NRF-2010-0020408; 2010-0024334; 2011-0019357) funded by the Ministry of Education, Science and Technology, Republic of Korea.
