Abstract
Dental enamel forms as a progressively thickening extracellular layer by the action of proteins secreted by ameloblasts. The most abundant enamel protein is amelogenin, which is expressed primarily from a gene on the X-chromosome (AMELX). The two most abundant non-amelogenin enamel proteins are ameloblastin and enamelin, which are expressed from the AMBN and ENAM genes, respectively. The human AMBN and ENAM genes are located on chromosome 4q13.2. The major secretory products of the human AMELX, AMBN, and ENAM genes have 175, 421, and 1103 amino acids, respectively, and are all post-translationally modified, secreted, and processed by proteases. Mutations in AMELX have been shown to cause X-linked amelogenesis imperfecta (AI), which accounts for 5% of AI cases. Mutations in ENAM cause a severe form of autosomal-dominant smooth hypoplastic AI that represents 1.5%, and a mild form of autosomal-dominant local hypoplastic AI that accounts for 27% of AI cases in Sweden. The discovery of mutations in the ENAM gene in AI kindreds proved that enamelin is critical for proper dental enamel formation and that it plays a role in human disease. Here we review how enamelin was discovered, what is known about enamelin protein structure, post-translational modifications, processing by proteases, and its potentially important functional properties such as its affinity for hydroxyapatite and influence on crystal growth in vitro. The primary structures of human, porcine, mouse, and rat enamelin are compared, and the human enamelin gene, its structure, chromosomal localization, temporal and spatial patterns of expression, and its role in the etiology of amelogenesis imperfecta are discussed.
Introduction
“No idea is so modern that it will not some day be antiquated.”
— Ellen Glasgow (Urist, 1966)
There has been considerable confusion about what is meant by the term enamelin. This is because the definition of enamelin has changed over time. The classic perspective of amelogenesis held that the enamel matrix of developing mammalian teeth was comprised of two classes of proteins: amelogenins and enamelins (Termine et al., 1980; Deutsch, 1989). The amelogenins did not bind to enamel crystals, and therefore could be extracted from secretory-stage enamel scrapings with guanidine HCl. The enamelins, which consisted of proteins tightly bound to the crystallites, were extracted after the amelogenins by dissolving the mineral, typically with guanidine HCl/EDTA. The classic theory was conceived over 20 years ago, prior to the availability of any significant amino acid, cDNA, or genomic sequence data for the proteins in the enamel matrix. This framework proved useful for over a decade, but became antiquated as the nature of enamel proteins and the genetic mechanisms that produced them were more fully understood (Fincham et al., 1999).
In the early 1980s, partial amino acid sequences for amelogenin were reported from pigs (Fukae et al., 1979, 1980; Fukae and Shimizu, 1983), cattle (Zalut et al., 1980; Fincham et al., 1981; Takagi et al., 1984), and humans (Fincham et al., 1983). The protein data were followed by cDNA sequences, with mouse amelogenin being the first to be cloned (Snead et al., 1983) and characterized (Snead et al., 1985). These advances gave amelogenin a specific identity, and the term amelogenin became restricted to proteins expressed from amelogenin genes, which were found to localize on the sex chromosomes (Lau et al., 1989; Sasaki and Shimokawa, 1995). Alternative RNA splicing (Sasaki, 1984; Gibson et al., 1991; Simmer, 1995) and proteolytic processing (Suga, 1970; Bartlett and Simmer, 1999; Simmer and Hu, 2002) were shown to expand greatly the number of amelogenins present in the enamel matrix. Eventually, the amelogenin gene on the X-chromosome (AMELX) was shown to cause X-linked amelogenesis imperfecta (Lagerström et al., 1991; Hart et al., 2002). Characterizing the non-amelogenin enamel proteins, or “enamelins”, lagged ten to 15 years behind amelogenin, but followed the same course of starting as a class of proteins and ending as a specific gene product.
Non-amelogenin enamel proteins combined to account for only 10% of the protein in immature enamel. For a while, it appeared as though there might not be any true enamelins, but that this class of proteins consisted largely of serum albumin (Limeback and Simic, 1989; Limeback et al., 1989; Strawich and Glimcher, 1989, 1990; Strawich et al., 1993). Albumin is not synthesized by ameloblasts (Couwenhoven et al., 1989; Yuan et al., 1996) and is not considered to be a true enamel protein, although it is commonly found in enamel protein preparations. It is still not certain how it gets there. Radiolabeled serum albumin injected into rabbits did not incorporate into the enamel layer, suggesting a physiological barrier between the extravascular fluid and the enamel matrix (Kinoshita, 1979; Kinoshita and Ogura, 1979). Albumin detected in normal enamel is likely to be an artifact of tissue preparation (Chen et al., 1995; Fincham et al., 1999).
A cDNA clone encoding a protein called tuftelin was characterized in the late 1980s and was thought to encode the major enamelin component in developing enamel (Deutsch et al., 1989, 1991). Although the tuftelin clone was isolated from a cDNA expression library using antibodies raised against the major 66-kDa protein in the guanidine/EDTA fraction of developing bovine enamel (Deutsch et al., 1987), the protein encoded by the cDNA was not present in the enamel matrix in appreciable quantities, and has never been isolated or characterized from immature enamel. Immunohistochemistry with tuftelin-specific antibodies showed positive signal at the dentino-enamel junction (DEJ), suggesting a role in enamel crystal nucleation (Deutsch et al., 1997). The term “enamelin” should no longer be used to refer to tuftelin (Brookes et al., 2002).
Today, we know that the two predominant non-amelogenin proteins in immature enamel are ameloblastin (also called amelin and sheathlin) and enamelin, which are the expression products of the AMBN and ENAM genes, respectively. These proteins were discovered by biochemists using a novel way of separating amelogenins from non-amelogenin proteins, which differed from the sequential dissociative extraction scheme used widely elsewhere. They separated porcine enamel fractions on SDS-polyacrylamide gels (SDS-PAGE), and soaked the gels in 25% isopropanol (Fukae and Tanabe, 1985, 1987b). The non-amelogenins remained in the gel, while the amelogenins went into solution. They were able to characterize two distinct parts of the protein we now call ameloblastin. Polypeptides from the N-terminal region of the protein had apparent molecular weights in the range of 13–17 kDa; they were concentrated in the sheath space that partially separates rod and interrod enamel and were designated as “sheath proteins” (Uchida et al., 1991b, 1995). The C-terminus of ameloblastin was characterized, shown to bind calcium, and was described as the 27- and 29-kDa calcium-binding proteins (Fukae and Tanabe, 1987a; Murakami et al., 1997; Yamakoshi et al., 2001). A second group of non-amelogenin enamel proteins was also characterized. A 32-kDa non-amelogenin was isolated from developing enamel that comprised about 1% of total enamel protein and showed many enamelin-like biochemical properties. It was acidic, and separated on 2-D gels into seven constituents having isoelectric points ranging from 3 to 4.5. It also bound strongly to hydroxyapatite (HA) crystals and inhibited HA crystal growth in vitro (Tanabe et al., 1990). Immunohistochemistry with antibodies against the 32-kDa (Uchida et al., 1991a) and the related 89-kDa (Uchida et al., 1991b) non-amelogenin proteins showed a concentration of signal in the crystallite-containing rod and interrod areas of the enamel matrix, suggesting that the proteins were bound to mineral in vivo. It was therefore concluded that this protein represented the enamel matrix component that constituted the enamelin class of proteins, and was specifically designated as enamelin.
The term enamelin is now restricted to the enamelin gene (ENAM) and its protein products. The classic theory, based upon the division of enamel proteins into two groups according to their mineral-binding properties, poorly accommodated the growing molecular data. It turned out that intact amelogenin binds to hydroxyapatite (Aoba et al., 1987; Ryu et al., 1998), as do some enamelin (Tanabe et al., 1990; Brookes et al., 2002) and ameloblastin cleavage products (Brookes et al., 2001). Furthermore, mineral binding is only one biochemical property. Solubility, ion binding, protein-protein interactions, susceptibility to proteolysis, and effects on crystal growth are other important properties potentially related to the function of enamel proteins. As a consequence, enamel matrix components are currently grouped according to the gene that expressed them (AMELX, EMBN, ENAM), so that all of the protein products of the enamelin gene are designated as enamelin, with the different proteolytic cleavage products being distinguished by their apparent molecular weight on SDS-PAGE.
This is the story of enamelin, how it was discovered, where it is expressed, its structure and post-translational modifications, its processing by proteases, its genetics, and how investigations into the composition of the enamel matrix of developing teeth improved our understanding of enamel formation and led to the discovery that defects in the enamelin gene are a major factor in the etiology of autosomal-dominant amelogenesis imperfecta.
Enamelin Protein Sequences and Immunohistochemistry
Enamelin is not abundant in the immature enamel matrix, so efforts to isolate and characterize enamelin components used animals with large developing teeth (cattle and pig) as a source of enamel proteins. Results using the bovine model never advanced to the point of obtaining protein sequence data (Menanteau et al., 1988; Ogata et al., 1988). Persistence made the porcine animal model the standard system for the study of enamel proteins, and brought about the discovery of sheathlin (ameloblastin) (Uchida et al., 1995; Hu et al., 1997a), enamelin (Fukae et al., 1996), enamelysin (Bartlett et al., 1996), and enamel matrix serine proteinase 1 (EMSP1) (Tanabe, 1984; Simmer et al., 1998). The first enamelin protein sequence in the literature (89-kDa enamelin, MPMNMPRMPGFSPKREPM) was published in a discussion of the Proceedings of the Fifth International Symposium on the Composition, Properties and Fundamental Structure of Tooth Enamel (Fukae, 1989). The first peer-reviewed publication of enamelin sequence (Tanabe et al., 1990) was from the N-terminus of the most abundant enamelin cleavage product in developing enamel: the 32-kDa enamelin (LXHVPGRIPPGYGRPPTP).
Antibodies were raised against the 32-kDa enamelin N-terminal sequence and affinity-purified (Uchida et al., 1991a). These antibodies proved to be highly specific. Western blots clearly demonstrated that the 32-kDa enamelin was a proteolytic cleavage product of a much larger protein. The newly formed surface enamel contained enamelin proteins of 140- and 89-kDa apparent molecular weight. The enamelin cleavage products decreased in size with increasing depth toward the DEJ, including molecules with apparent molecular weights of 56, 45, and 32 kDa. Light and electron micrographs of immunostained sections of pig incisors were startling in their clarity and detail. The earliest signal for enamelin protein was observed in differentiating ameloblasts as the basal lamina disappeared, and in predentin at the future DEJ. During the secretory stage, the signal extended from the DEJ to the Tomes’ process of the secretory ameloblast, being especially strong beneath the secretory face of the Tomes’ process, and scarce along the non-secretory face and the sheath space separating rod from interrod enamel (Fig. 1). The enamelin signal disappeared abruptly during the early maturation stage.
Antibodies were also raised against the 89-kDa enamelin (Uchida et al., 1991b), which is now known to be the N-terminal half of the secreted protein, and contains the entire sequence of the 32-kDa enamelin (Fukae et al., 1996). Immunohistochemistry of secretory-stage enamel sections using this antibody showed that enamelin cleavage products concentrate in the rod and interrod enamel (among the crystallites) and are scarce in the sheath space (Uchida et al., 1991b). This “reverse honeycomb pattern” suggested that enamelin cleavage products, which have acidic isoelectric points and bind HA crystals in vitro, are apparently bound to the enamel crystals in vivo. If one compares the immunolocalization patterns for amelogenin, the amelogenin C-terminus, the 13- to 17-kDa ameloblastin cleavage products, and the 89-kDa enamelin (Uchida et al., 1991b; Fukae et al., 1993), it becomes apparent that enamel proteins and their cleavage products generally segregate into different compartments (rod and interrod vs. sheath space; outer vs. inner enamel layers) (Fig. 2). These discoveries strengthened the interpretation that the proteolytic cleavage products of enamel proteins are not simply degradation products, but are functional polypeptides, and that their functions are potentially different from those of the intact proteins and other cleavage products (Bartlett and Simmer, 1999; Simmer and Hu, 2002).
Extensive isolation and characterization of enamelin cleavage products and the direct sequencing of PCR (polymerase chain-reaction) products provided a wealth of information concerning the proteolytic processing of enamelin (Fukae et al., 1996), and led directly to the cloning of the full-length porcine enamelin cDNA (Hu et al., 1997b). Knowing the complete enamelin protein sequence permitted investigators to generate anti-peptide antibodies to selected regions of the enamelin protein, including the C-terminal region, which had never been isolated or characterized at the protein level. The enamelin signal from four different regions of the enamelin protein—(a) the enamelin N-terminus, (b) the 32-kDa N-terminus, (c) the 34-kDa N-terminus, and (d) the enamelin C-terminus—provides a vivid picture of the effect of proteolytic processing on the distribution and abundance of enamelin cleavage products in developing enamel (Fig. 3). The antibody against the enamelin N-terminus stained the entire enamel matrix, with no particular concentration at the secretory face of the Tomes’ process (Fig. 3a). The signal decreased from the surface to the DEJ, and electron micrographs detected a greater concentration of the enamelin N-terminus in the sheath space relative to the rod and interrod enamel (Dohi et al., 1998). The 32-kDa antibody showed a slight concentration at the secretory face and a scarcity of signal along the non-secretory face of the Tomes’ process (Fig. 3b). The signal decreased going from the surface to the DEJ, although at the DEJ itself, it was particularly intense (Uchida et al., 1991a). Throughout the entire enamel layer, the 32-kDa enamelin showed a “reverse honeycomb” distribution pattern, being particularly scarce in the sheath space (Uchida et al., 1991a). The anti-peptide antibody specific for the N-terminus of the 34-kDa enamelin cleavage product, which is in the center of the protein, shows a concentration of signal beneath the secretory face of the Tomes’ process (Fig. 3c). The signal diminishes rapidly with depth, indicating that this portion of the enamelin protein is rapidly degraded or re-absorbed into the ameloblast and does not accumulate in the matrix (Hu et al., 1997b). The most restricted localization pattern was observed with the antibody specific for the enamelin C-terminus (Fig. 3d). Signal was observed only beneath the secretory face of the Tomes’ process, at the mineralization front. This highly restricted localization supports the interpretation that intact (uncleaved) enamelin participates in the elongation of enamel crystals, which occurs at the mineralization front (Hu et al., 1997b).
Immunohistochemistry combined with Western blot analyses shows that intact enamelin (186 kDa) and the large enamelin cleavage products (155 kDa, 142 kDa, 89 kDa) are present only near the enamel surface and do not accumulate in the matrix (Hu et al., 1997b). All of these proteins contain the original enamelin N-terminus, since enamelin, like amelogenin, is processed by successive cleavages from its C-terminus. The smaller polypeptides from the enamelin C-terminal half appear to undergo successive cleavages or are re-absorbed into ameloblasts, which prevents their accumulation in the deeper enamel layers. The 32-kDa enamelin is resistant to further proteolytic digestion and accumulates in the rod and interrod enamel probably bound to the mineral, while the extreme N-terminus does not bind mineral and concentrates in the sheath space, along with N-terminal polypeptides from ameloblastin.
Enamelin mRNA Expression
Investigators have applied molecular biology techniques to define the temporal and spatial patterns of enamelin expression. These techniques have different levels of sensitivity, resulting in apparent discrepancies relative to the tissue specificity of the gene whose expression is being defined. Some of these techniques are ultra-sensitive, so that extremely low levels of “leaky” mRNA expression can be detected. Trace levels of expression, however, can never be totally excluded from having physiological significance, even when knockout mice or human kindreds with defined mutations show no apparent abnormality in the tissue showing trace expression. Subtle abnormalities might occur under unusual circumstances. On the other hand, there may be little or no selection pressure to develop genetic controls that would totally restrict expression of a particular gene to its intended site when such expression is not harmful, and too much emphasis on the sites of trace levels of expression might obscure the true specificity of a protein’s expression and function.
The technique that generated the most restricted pattern of enamelin expression was in situ hybridization. In situ hybridization in a day 1 mouse developing incisor detected enamelin mRNA expressed by ameloblasts, but not by odontoblasts or other cells in the dental pulp. Extensive in situ hybridization analyses of enamelin expression in mouse molars from post-natal days 1, 2, 3, 7, 9, 14, and 21 have been reported (Hu et al., 2001a). Enamelin mRNA in maxillary first molars was first observed in pre-ameloblasts on the cusp slopes at day 2. The onset of enamelin expression was approximately synchronous with the initial accumulation of predentin matrix. Enamelin was expressed by ameloblasts throughout the secretory, transition, and early-maturation stages and was terminated in maturation-stage ameloblasts on day 9. No enamelin expression was observed in pulp or bone, or along the developing root.
Northern blot analysis of mRNA obtained from enamel organ epithelia (EOE), dental pulp organ, liver, heart, kidney, brain, spleen, skeletal muscle, and lung detected enamelin mRNA only in the EOE and dental pulp, with expression levels being at least an order of magnitude higher in the EOE (Hu et al., 1998).
Reverse-transcription/polymerase chain-reaction (RT-PCR), as well as Western blots, detected low levels of enamelin expression during root formation. Tissue samples were prepared from the apical portion of the forming root in porcine permanent incisor tooth germs. The specific source of enamelin expression was presumed to be cells derived from Hertwig’s epithelial root sheath (Fukae et al., 2001).
As of November, 2003, six human enamelin cDNAs have been detected in non-dental tissues as expressed sequence tags (ESTs). Two were from eye (BU741192 and BM726916), one from kidney (AI627857), one from a kidney tumor (AW466983), one from lymph (BU428532), and one from prostate (AI675060) tissues.
In summary, enamelin is expressed primarily by secretory ameloblasts and is presumed to function only during dental enamel formation. Enamelin expression by ameloblasts is low compared with that by amelogenin and ameloblastin, with enamelin protein accounting for only a few percent of total protein in the forming enamel layer. Much lower levels of enamelin expression have been observed in dental pulp, presumably secreted by odontoblasts, and along the forming root. Enamelin mRNAs have been detected as ESTs from several tissues, with no suggestion that such expression is physiologically significant.
Cloning Enamelin cDNAs and Genes
Extensive protein and cDNA sequencing of enamelin from the pig (Fukae et al., 1996) led directly to the isolation and characterization of full-length enamelin cDNA clones from the pig (Hu et al., 1997b), and subsequently from the mouse (Hu et al., 1998) and humans (Hu et al., 2000). As of November, 2003, no enamelin cDNA sequences have been reported from other organisms; however, a preliminary rat enamelin coding sequence (NW_042906) has been predicted from a National Center for Biotechnology Information genomic contig (NW_042906) using GenomeScan. In addition, the mouse and human enamelin genes (Hu et al., 2001b) have been cloned and characterized. The chromosomal localization of the enamelin gene was determined by fluorescent in situ hybridization (FISH) and by radiation hybrid mapping. The human enamelin gene is located on chromosome 4q13 near the ameloblastin gene (AMBN), in a region previously linked to local hypoplastic AI (Kärrman et al., 1997). These results were confirmed, and it was concluded that the distance between the human ENAM and AMBN genes is less than 15 kilobases, and that the enamelin gene is closer than the ameloblastin gene to the centromere (Dong et al., 2000). This order of the genes relative to the centromere appears to be in error. The enamelin and ameloblastin genes are now available at the NCBI Web site (http://www.ncbi.nlm.nih.gov), which places the AMBN gene closer than the ENAM gene to the centromere (Fig. 4).
Enamelin Genes and cDNA
Enamelin cDNA and genomic clones permitted the derivation of enamelin amino acid sequences from pig, human, mouse, and rat homologues. These sequences are shown aligned in Fig. 5. The first ATG codon starting from the 5′ end of the human, mouse, and pig cDNA sequences are all in the appropriate context for translation, but are followed in four codons by a translation stop signal (TGA), with the true translation initiation codons being located a short distance downstream. This feature has been observed in other genes (Kozak, 1984) and may play a role in the regulation of translation. In every enamelin homologue, translation begins with an amino acid sequence that satisfies the structural criteria for a functional signal peptide (Gierasch, 1989). The appropriate signal peptidase cleavage sites are predicted by computer analyses and, in the case of the pig protein, have been confirmed by Edman sequencing of the enamelin amino-terminus (Fukae, 1989; Fukae et al., 1996).
The mouse enamelin gene has 10 exons. The human enamelin gene has 9 or 10 exons (there is a segment on the human enamelin gene that corresponds to a noncoding exon 2 in the mouse gene, but this sequence was not found on the human enamelin cDNA). In both the mouse and human enamelin genes, eight of the exons are coding (Fig. 4) (Hu et al., 2001b). In contrast, the human ameloblastin gene has 13 coding exons (Toyosawa et al., 2000), while the human amelogenin gene has six (Salido et al., 1992). All of the coding exons in these genes have type O splice junctions, meaning that none of the introns splits a codon. Type O splice junctions allow exons to function as modules, so skipping exons by alternative splicing does not shift the reading frame. Despite this feature, no alternatively spliced enamelin cDNAs have ever been identified.
The primary structures of mammalian enamelin from the four known eutherian species show a high degree of sequence homology (Fig. 5), except in a particular segment between the 32-kDa and 25-kDa enamelin cleavage products. In the coding region for this section, there is a 33-nucleotide segment that is tandemly repeated in the mouse and rat sequences, but not in the pig or human enamelin genes. There are 14 copies of the repeat in the mouse and 16 in the rat. The human and pig enamelin cDNAs encode a pre-protein of 1142 amino acids, while the mouse has 1274 amino acids, and rat enamelin is larger still. Enamel proteins seem to be able to accommodate repeated segments into their structures. Exon 6 (the largest exon) in the amelogenin gene contains short repeat sequences that differ in length and sequence in virtually every amelogenin gene characterized (Hu et al., 1996). In the human ameloblastin gene, exons, 7, 8, and 9 each encode homologous 13 amino acid segments (Toyosawa et al., 2000). The repeat sequences in the rodent enamelin genes occur within exon 10, the largest and last exon. The inserted polypeptides significantly affect the amino acid compositions, calculated isoelectric points, and isotope-averaged molecular masses of the rodent enamelin homologues (Table). In practice, though, these differences may not be significant with respect to the proteins’ functional properties. This is due to the high impact of post-translational modifications on the size, structure, charge, stability, and function of enamelin.
Enamelin Post-translational Modifications
The deduced molecular weight of pig enamelin is 124 kDa, but the protein’s apparent molecular weight on SDS-PAGE is 186 kDa (Hu et al., 1997b). The principal reason for the difference is post-translational modifications. Conserved among the four known enamelin homologues are five phosphorylation sites, five N-linked glycosylation sites, and five cysteines that could potentially form disulfide bonds (Fig. 5). The 32-kDa enamelin is the most abundant enamelin cleavage product in the enamel matrix, and provides a good example of the influence of post-translational modifications on the character of the protein.
The 32-kDa enamelin has 106 amino acids (extending from Leu174 to Arg279), but has two phosphorylated serines (Ser191 and Ser216) and 3 glycosylated asparagines (Asn245, Asn252, and Asn264) (Fig. 6). The isotope-averaged molecular weight of the 32-kDa phosphoprotein (without glycosylations) is 11,657.6 Daltons; its predicted isoelectric point is 5.27. Based upon the primary structure of the phosphorylated, but unglycosylated, 32-kDa enamelin, the ProtParam tool (http://us.expasy.org/cgi-bin/protparam) computes its instability index (II) as 48.01, classifying it as unstable. The size, isoelectric point, and stability of the 32-kDa enamelin appear to be affected greatly by its high degree of glycosylation. The structures of the N-linked oligosaccharides on the 32-kDa enamelin were determined by a combination of sequential exoglycosidase digestion and two-dimensional sugar mapping (Yamakoshi, 1995; Yamakoshi et al., 1998). Eight different oligosaccharide structures were observed, five biantennary and three triantennary types. The 32-kDa enamelin was cleaved, and glycopeptides containing each of the three glycosylation sites were isolated (Yamakoshi, 1995; Yamakoshi et al., 1998). It was determined that Asn245 can have any of the five biantennary complexes. Asn252 uses two of the biantennary complexes, while Asn264 uses any of the three triantennary complexes. The variability of the glycosylation pattern for the 32-kDa enamelin resulted from differences in the number, site, and mode of linkage of N-acetylneuraminic acid to the core-sugar chains, and in their degree of sialylation. Extensive and variable glycosylation presumably accounts for the large number of distinct 32-kDa enamelin bands that can be distinguished by two-dimensional gel electrophoresis, its lower-than-predicted isoelectric point, its large apparent molecular weight, and its great stability relative to the other enamelin cleavage products. Polyclonal antibodies raised against recombinant 32-kDa enamelin expressed in bacteria could detect the recombinant protein, but not the 32-kDa protein isolated from in vivo (Ryu et al., in press). Post-translational modifications were the only difference between the recombinant and native proteins. Clearly, post-translational modifications can have a profound effect on the structure and function of enamelin.
Perhaps the most significant post-translational modification is proteolytic processing (Bartlett and Simmer, 1999; Simmer and Hu, 2002). Proteolysis allows enamelin cleavage products to accumulate in different parts of the enamel matrix, and determines their relative abundance. Some enamelin cleavage products appear to be insoluble. The 32-kDa enamelin may be released from the mass of insoluble enamel protein by proteolytic cleavage, freeing it to bind enamel crystallites, potentially regulating their shape or habit (Brookes et al., 2002). Enamelysin (MMP-20) is the predominant proteolytic activity in the secretory-stage enamel matrix (Li et al., 1999; Ryu et al., 1999), and is assumed to catalyze the processing of enamelin. Targeted knockout of the enamelysin gene in mice resulted in profound enamel defects (Caterina et al., 2002). Enamelysin is inactive against the 32-kDa enamelin, but the 32-kDa enamelin can be degraded by KLK4, the main degradative matrix enzyme that is expressed throughout the maturation stage (Fig. 7).
Enamelin Protein-Protein Interactions
Little is known about the interactions of enamelin with other enamel matrix proteins. After it was learned that amelogenin has an affinity for N-acetylglucosamine (Ravindranath et al., 1999), it was believed that enamelin might interact with amelogenin through its oligosaccharide chains. It turned out that the 32-kDa enamelin contains N-acetylglucosamine, but this sugar is buried within the oligosaccharide structure. Far Western analyses showed that native 32-kDa enamelin did not bind to amelogenin (Yamakoshi et al., in press). The 32-kDa enamelin bound to amelogenin only after its N-acetylglucosamine was exposed to the surface by sequential removal of the overlying oligosaccharides, but such enamelin structures are not found in vivo.
Enamelin and Amelogenesis Imperfecta (AI)
Amelogenesis imperfecta (AI) is a heterogeneous group of inherited defects in dental enamel formation. The malformed enamel can be unusually thin, soft, rough, and stained. Clinically, AI presents as a spectrum of enamel malformations that are categorized as hypoplastic, hypocalcified, or hypomaturation types, based primarily upon the thickness and hardness of the dental enamel (Witkop and Sauk, 1976; Witkop, 1989). The different types of AI are thought to reflect differences in the timing, during amelogenesis, when the disruption occurred. The earliest defects affect formation of the dentino-enamel junction, and result in an enamel layer that shears easily from the underlying dentin. A severe defect might block subsequent enamel formation and cause enamel agenesis, where there is almost no clinical or radiographic evidence of enamel. The teeth are yellowish brown, rough in texture, and widely spaced. Defects that occur during the secretory stage of amelogenesis interfere with crystal elongation and leave the enamel layer pathologically thin, or hypoplastic. Defects that occur during the maturation stage of amelogenesis disrupt the removal of the organic matrix and interfere with hardening of the enamel layer, and lead to pathologically soft or hypomaturation forms of AI. The dental crowns are of normal size and come into contact with adjacent teeth, but the mottled, brownish-yellow enamel is soft and has a radiodensity approaching that of dentin. Sometimes there appears to be a failure in mineralization of the organic matrix, resulting in hypocalcified enamel. The enamel layer may or may not be of normal thickness, but is extremely soft and wears away quickly following tooth eruption. When mode of inheritance is included in the classification of AI, 14 subclasses are distinguished (Witkop and Sauk, 1976; Witkop, 1989). This classification system is difficult to apply clinically. It has been proposed that when sufficient information is learned about the genetic causes of AI, a gene-based classification system should be adopted (Aldred and Crawford, 1995).
Alterations in the human amelogenin gene are responsible for X-linked forms of AI (Lagerström et al., 1991; Aldred et al., 1992; Hart et al., 2003), but only 5% of families with AI show an X-linked pattern of inheritance (Bäckman and Holmgren, 1988). The first linkage of an autosomal-dominant form of AI (ADAI) was to chromosome 4q11-q21 (Kärrman et al., 1997). The human ameloblastin gene (AMBN) was shown to localize in this region, and it was concluded that AMBN is linked to ADAI (MacDougall et al., 1997). Subsequent studies, however, localized the enamelin gene (ENAM) within the linked interval (Hu et al., 2000), and mutational studies of families with AI linked to chromosome 4q were determined to have critical defects in the enamelin gene (Rajpar et al., 2001; Mårdh et al., 2002), while no defects in the ameloblastin gene were detected (Mårdh et al., 2001). The ameloblastin gene is still considered to be a candidate gene for AI, but its linkage to the disease is unproven.
At present, there are four published reports of defects in the human enamelin gene that cause autosomal-dominant amelogenesis imperfecta (ADAI) (Rajpar et al., 2001; Kida et al., 2002; Mårdh et al., 2002; Hart et al., 2003). The mutation sites are shown in Fig. 8. The first reported human enamelin mutation was a splice donor site mutation after enamelin codon 178, which resulted in a severe form of thin and smooth hypoplastic AI that is believed to affect only 1.5% of all AI cases (Rajpar et al., 2001). This mutation occurs at the beginning of the intron following the sixth coding exon. It is difficult to predict the effect of this mutation on the enamelin protein structure. It was proposed that skipping coding exon 6 would be the most likely scenario and would delete amino acids 158–178; but failure to delete the intron following this exon seems equally probable and would cause the following extraneous amino acids to be added after Gln178: EKFFFLYTVSEK*, along with the deletion of amino acids 179–1142. It was also proposed that the mutation could interfere with the proteolytic activity of enamelin, but this can be dismissed, since the original data showing that enamelin had proteolytic activity (Moradian-Oldak et al., 1996) were erroneous, the mistaken conclusion being caused by the co-purification of enamel matrix serine proteinase 1 (now known as kallikrein-4, KLK4) with the 32-kDa enamelin cleavage product (Simmer et al., 1998).
A milder form of AI was shown to be caused by translation termination at enamelin codon 53 (Mårdh et al., 2002). This mutation deleted amino acids 53 through 1142. Given that the first 39 amino acids comprise the signal peptide, only 13 amino acids would be synthesized, essentially resulting in a null mutation. This milder form of AI, known clinically as autosomal-dominant local hypoplastic AI, accounts for 27% of the autosomally inherited cases in Northern Sweden. It was proposed that more severely hypoplastic enamel was observed in the kindred with the splice donor site mutation, because the secretion of an abnormal protein acted in a dominant-negative way to disturb enamel formation.
Most recently, a splice donor site mutation after enamelin codon 196 was shown to cause autosomal-dominant hypoplastic AI (Kida et al., 2002). There are normally 6 Gs at the end of coding exon 7, which are followed by a 7th G at the beginning of the adjacent intron. One of these Gs was deleted in a Japanese family with AI. If the splice donor site function is preserved in the mutant condition, the deletion would shift the reading frame at the start of the last coding exon, resulting in the inclusion of the following 80 amino acids after Gly196: ILTLDILDIMALGVALLIIQKKCLNKILKNPKKKILLKQKVQAQNPQLIQQSLRRILPNQILKGVREEMTPAPQETVPQD*. The affected members of the family showed hypoplastic enamel in both their deciduous and permanent teeth that resulted in a yellowish appearance and hypersensitivity to cold stimuli. This same mutation has recently been characterized in a family from Australia (Hart et al., 2003).
Current Research
Enamel biomineralization is a fascinating but highly complex process. Advances in our understanding of amelogenesis are coming at a steady pace, and enamelin is destined to be a prominent part of the molecular mechanism. Enamelin was only recently defined as a specific protein, and collective research efforts are under way to better understand the structure of enamelin, its post-translational modifications, and proteolytic processing. Functional studies are under way to define the ability of enamelin cleavage products to bind other matrix proteins and enamelin crystals, and to influence the growth of hydroxyapatite crystals in vitro. The ultimate test of our understanding of enamelin’s role in dental enamel formation will be to reconstitute the enamel matrix and grow enamel-like crystals in vitro. Because enamelin is processed by MMP-20 almost immediately following its secretion, it is likely that recombinant enamelin will be needed to assay enamelin function in vitro. Collective research efforts are under way to express enamelin in a variety of prokaryotic and eukaryotic systems, as well as to better define the extent of its post-translational modifications, including its processing by proteinases. Experiments are under way to generate targeted knockouts of the enamelin gene in mice, which would provide a means of assaying its function in vivo. The temporal and spatial patterns of enamelin expression are being refined, and promoter analyses are under way to better understand the transcriptional regulation that gives the enamelin gene its tissue-specific pattern of expression.
Perhaps the most exciting new developments involve the characterization of mutations in the human enamelin gene that cause amelogenesis imperfecta. We anticipate that such studies will allow the diagnosis of AI to become gene-based. Whenever the gene and mutation that cause AI in a given family are learned, careful descriptions of the associated clinical phenotypes will allow genotype-phenotype correlations to be identified. By studying the outcomes of different restoration procedures (i.e., bonding and veneers vs. crowns) for each genotype/phenotype condition, practicing dentists will use gene-based diagnoses to choose among various treatment options, and thereby restore the dentition in a way that achieves the best results.
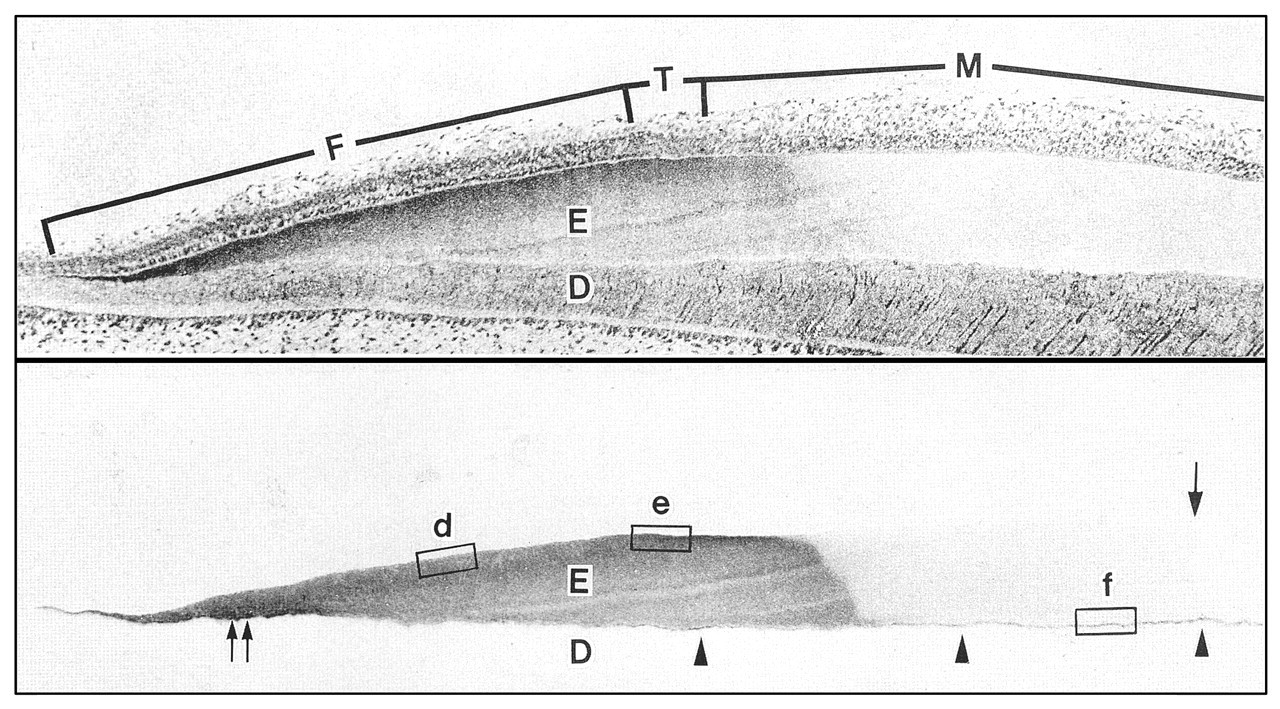
Consecutive sections of a developing porcine incisor showing the stages of enamel formation. The top section is stained with toluidine blue, and the bottom section is immunostained with the 32-kDa enamelin antibody. Enamelin signal is observed in the enamel matrix throughout the matrix formation (secretory) stage and persists at the DEJ (arrowheads) during enamel maturation. Key: enamel, E; dentin, D; matrix formation stage, F; transition stage, T; maturation stage, M; arrowheads mark the DEJ. Adapted from reference (Uchida et al., 1991a).
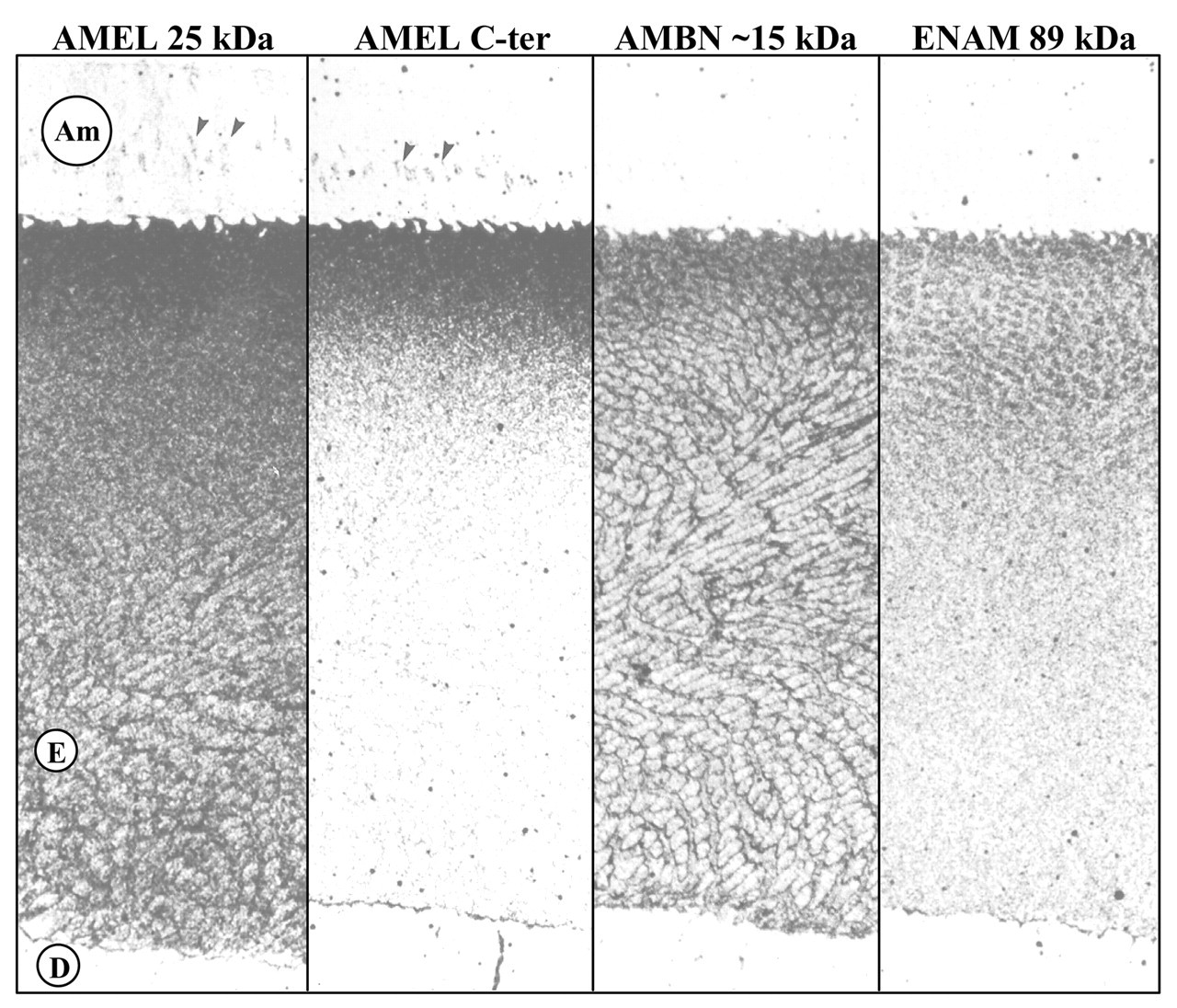
Immunohistochemistry of the porcine secretory-stage enamel showing immunolocalization of enamel proteins using light microscopy (Uchida et al., 1991b). The four antibodies used (from left to right) were a polyclonal antibody raised against intact (25 kDa) amelogenin, an anti-peptide antibody specific for the amelogenin C-terminus, polyclonal antibodies raised against 13- to 17-kDa ameloblastin cleavage products, and polyclonal antibodies raised against the 89-kDa enamelin. The amelogenin C-terminal antibody, indicative of the intact protein, was restricted to the surface enamel. The ameloblastin antibody produced a honeycomb pattern over the entire thickness of the immature enamel. The enamelin antibody generated a reverse honeycomb pattern. Key: secretory ameloblasts (Am), immature enamel (E), dentin (D), Golgi (arrowheads). Approximate magnification, 520x.
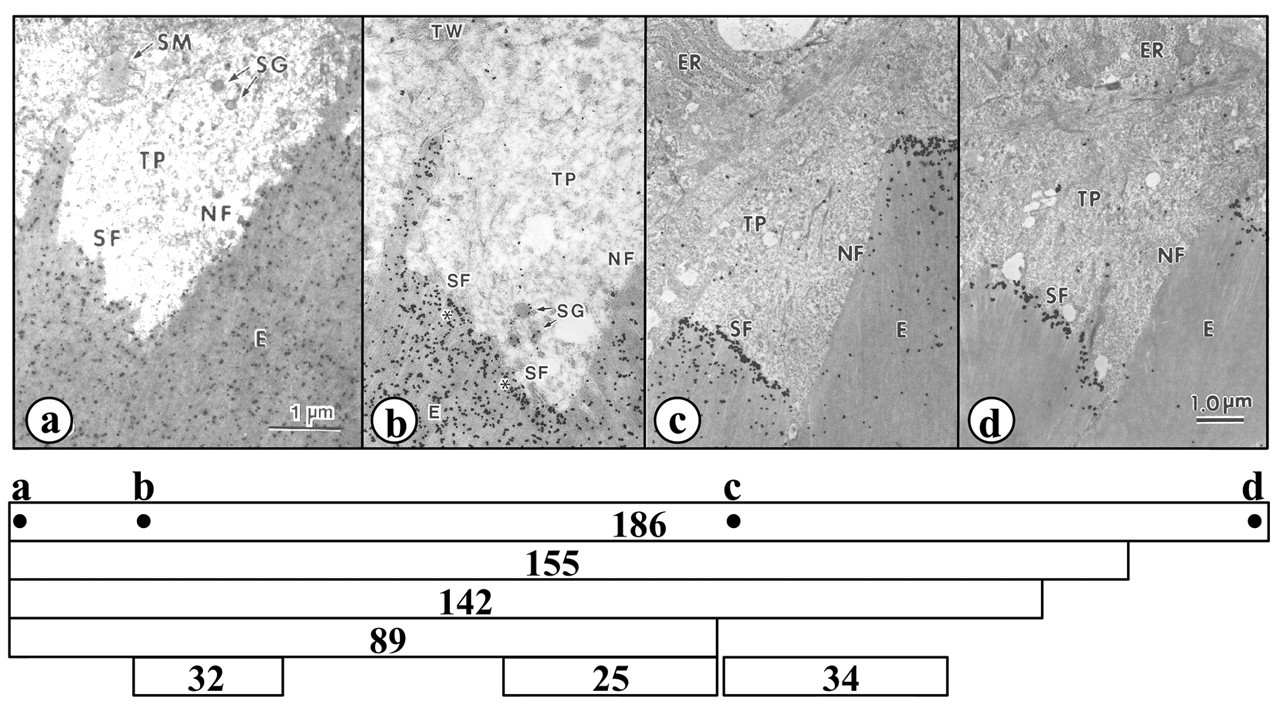

Electron micrographs of porcine secretory-stage enamel showing immunolocalization of enamelin at the enamel surface near the ameloblast Tomes’ process. The results from four affinity-purified anti-peptide antibodies are shown: (a) enamelin N-terminus (Dohi et al., 1998); (b) 32-kDa N-terminus (Uchida et al., 1991a); (c) 34-kDa N-terminus; and (d) enamelin C-terminus (Hu et al., 1997b). Below the histology is a diagram showing the different enamelin cleavage products and the positions of the sequences used to make antibodies. Pig enamelin has 1142 amino acids. The first 38 amino acids constitute the signal peptide. The secreted protein has an apparent molecular weight of 186 kDa (amino acids 39–1142); partially characterized enamelin cleavage products are the 155 kDa (39-unknown), 142 kDa (39 to unknown), 89 kDa (39 to 665), 32 kDa (174–279), 25 kDa (515–665), and the 34 kDa (670-unknown) (Fukae et al., 1996). Key: Tomes’ processes (TP), enamel matrix (E), secretory granules (SG), secretory face (SF), non-secretory face (NF) of Tomes’ process endoplasmic reticulum (ER), terminal web (TW), stippled material (SM). Bar = 1.0 μm.
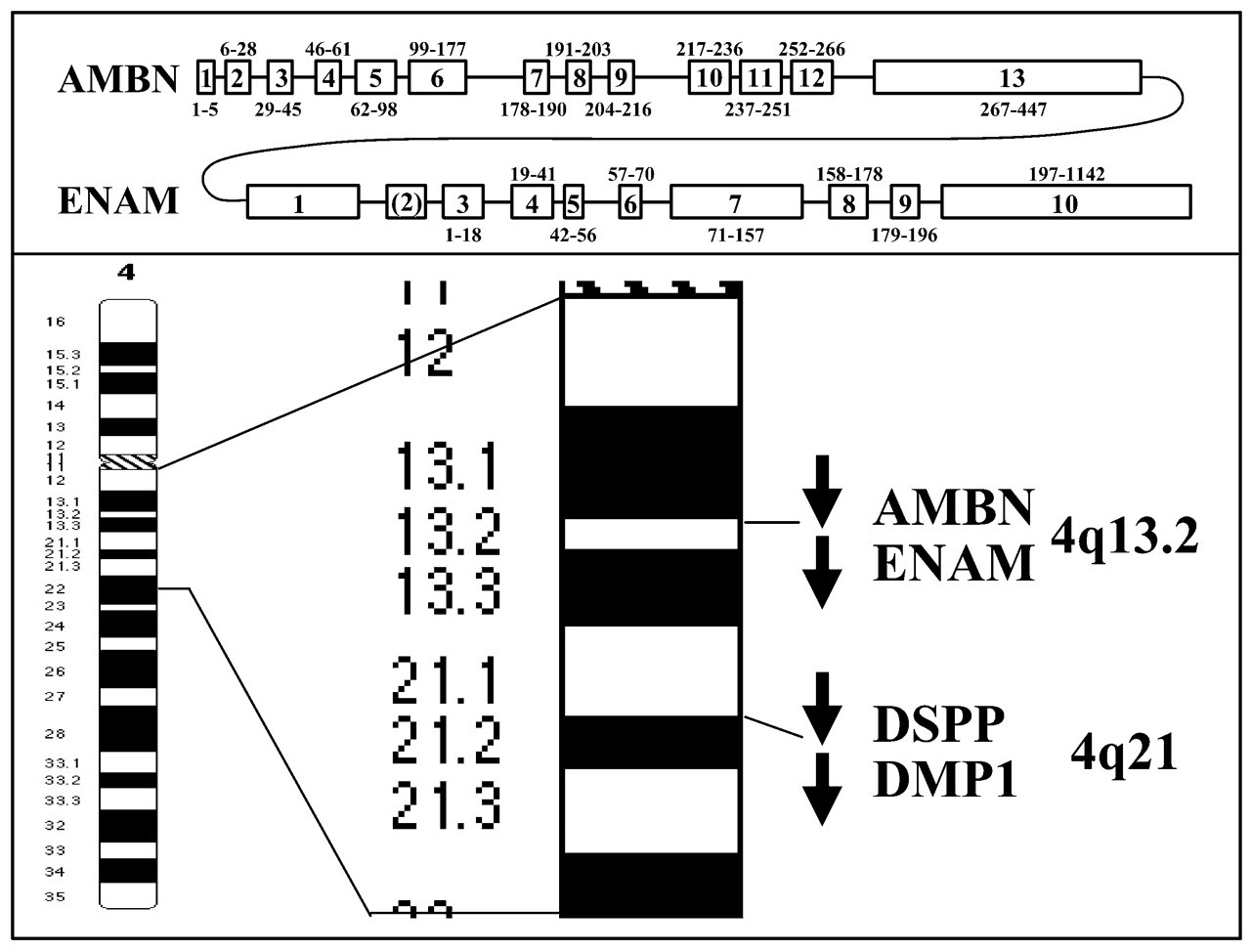
Structures of the human ameloblastin (AMBN) and enamelin (ENAM) genes and their chromosomal localizations. Exons are indicated by numbered boxes, introns by a line. The numbers below each exon show the range of amino acids encoded by that exon. The AMBN gene has 13 exons, all coding. AMBN exons 7 through 9 are repeat sequences (Toyosawa et al., 2000). The human ENAM gene is shown with 10 exons for consistency with the mouse enamelin gene (Hu et al., 2001b). It is not known if exon 2, which is noncoding, is used in humans. The enamelin gene has 8 coding exons. The AMBN and ENAM genes are located together on the long arm of chromosome 4. The order of the genes is centromere, AMBN, ENAM, DSPP, DMP1, teleomere.

Alignment of the derived protein sequences of pig, human, mouse, and rat enamelin. The number of the last amino acid in each row is indicated on the right. Amino acid residues at the amino- or carboxyl-termini of known porcine enamelin cleavage products are labeled above the pig sequence. Potentially modified amino acids are in bold. An asterisk indicates an amino acid that is known to be modified in pig enamelin. Known polymorphisms in the human enamelin protein are underlined: (Arg/Gln)286, (Ile/Thr)648, (Arg/Gln)763, and (Gly/Asp)948.
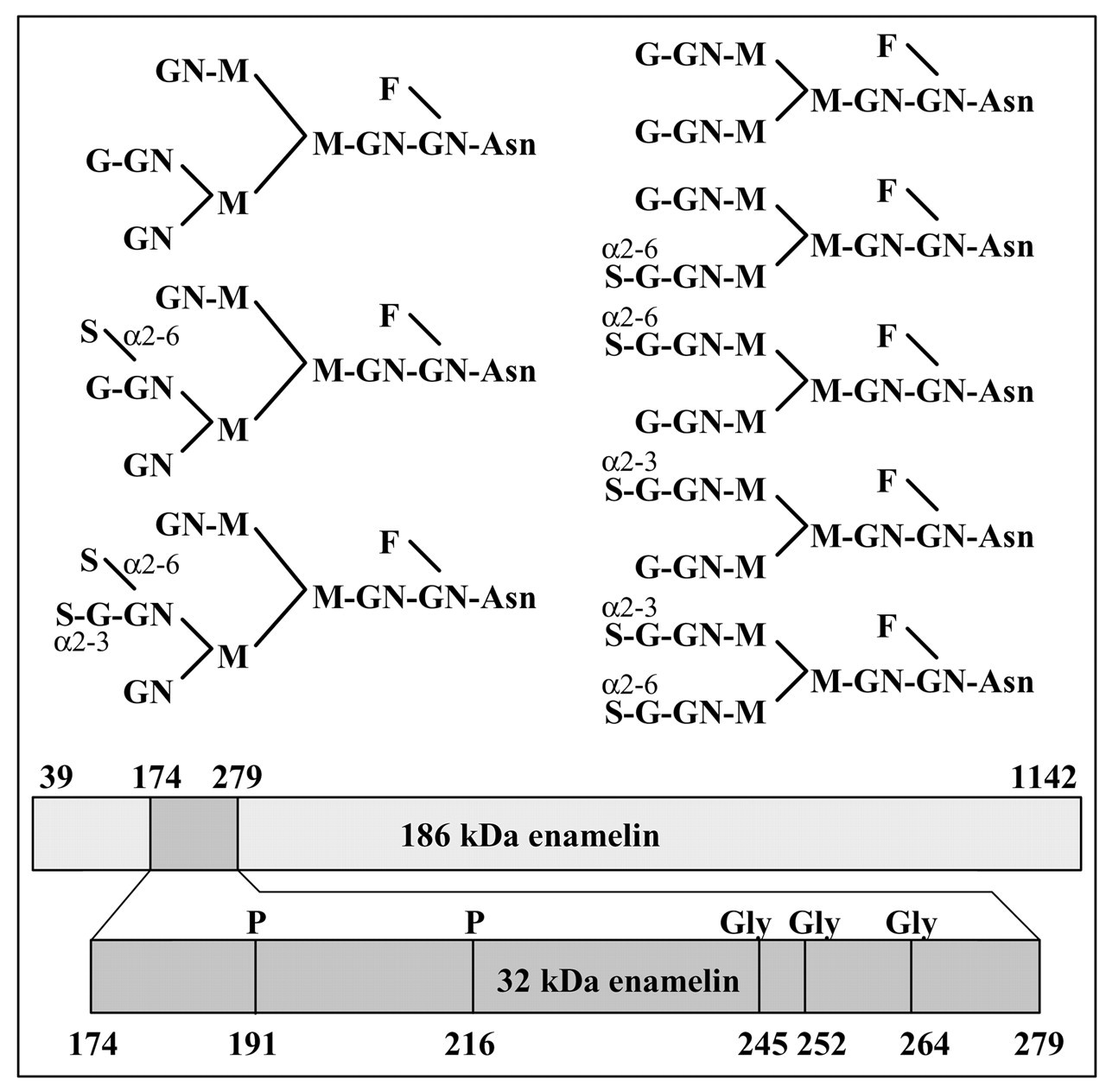
Structures of the pyridylaminated-oligosaccharides liberated from the 32-kDa enamelin by glycopeptidase F digestion (Yamakoshi, 1995). Asn245 uses all five of the biantennary types shown on the right. Asn252 uses the top two biantennary types. Asn264 uses the three triantennary types shown on the left (Yamakoshi et al., 1998). Key: fucose, F; galactose, G; N-acetylglucosamine, GN; mannose, M; N-acetylneuraminic acid, S. Diagram showing the position of the 32-kDa enamelin cleavage product in the 186-kDa secreted enamelin protein and the positions of the two phosphorylations (P) and glycosylations (Gly).
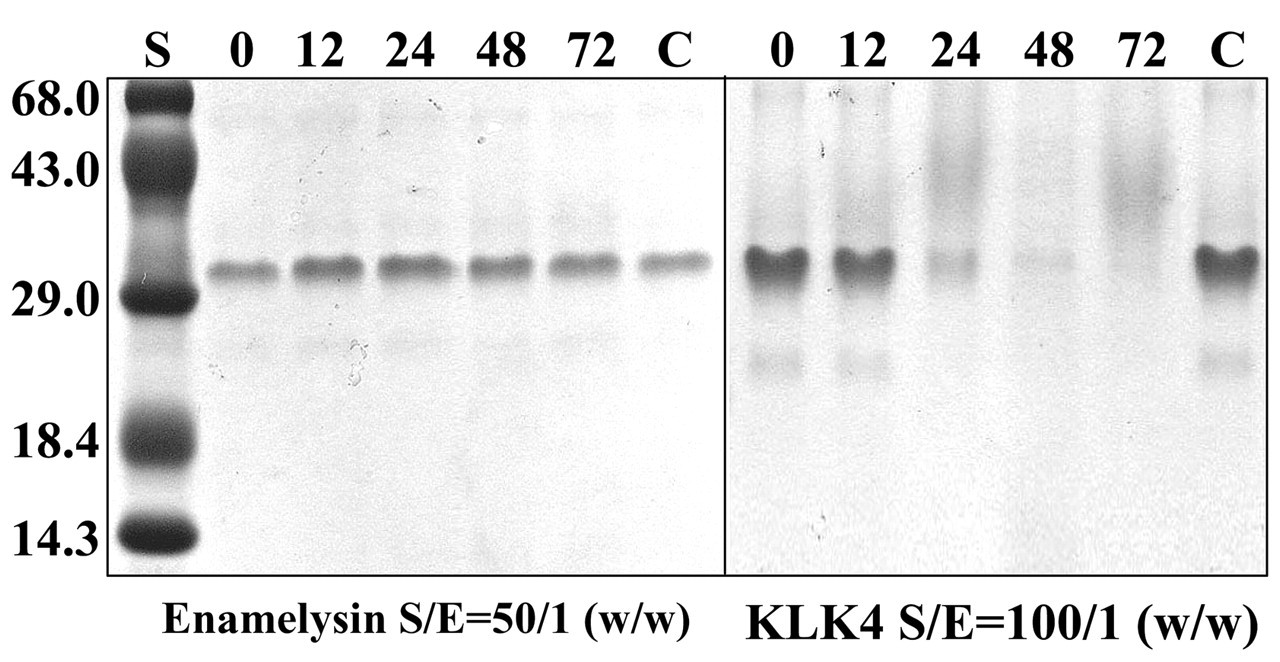
Digestion of the 32-kDa enamelin by enamelysin (MMP-20) and kallikrein 4 (KLK4). The 32-kDa enamelin, the 25-kDa amelogenin, MMP-20, and KLK4 were isolated from developing pig teeth. Cleavage of the 32-kDa enamelin by MMP-20 was not detected even after 72 hrs of incubation at a substrate:enzyme ratio of 50:1 (w/w). MMP-20 did digest amelogenin (data not shown). KLK4 fully degraded the 32-kDa enamelin after 24 hrs of incubation at a substrate:enzyme ratio of 100:1 (w/w).
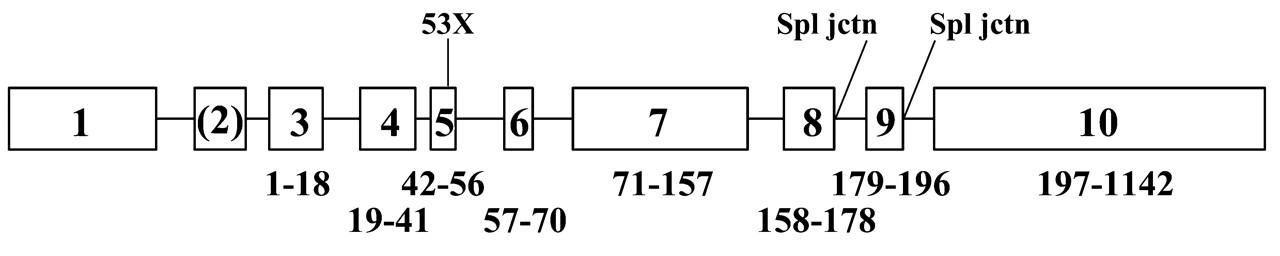
Sites of human enamelin gene mutations identified in families with AI. Exons are indicated by numbered boxes, introns by a line. The numbers below each exon show the range of amino acids encoded by that exon. Lines indicate the location of defined mutation sites in the enamelin gene. An “X” indicates that the mutation created a stop codon. Mutations affecting splice junctions are indicated by “spl jctn”.
Footnotes
Acknowledgements
This investigation was supported in part by USPHS Program project grants DE13221 and DE13237, and by R01 DE12769 from the National Institute of Dental and Craniofacial Research, National Institutes of Health, Bethesda, MD 29892. We thank Dr. Thomas Hart for his analysis of the order of enamelin and ameloblastin genes with respect to the centromere and Dr. Tim Wright for information concerning human enamelin polymorphisms I648T and R763Q.
